Bradi 1 g 25180 The Gene in Question

Bradi 1 g 25180: The Gene in Question Group ECR: Olivia Carneiro & David Cellucci http: //www. free-powerpoint-templates-design. com
Goals for Research Develop an understanding of the gene’s function Identify the mutation and the possible impacts on the plant physiology Form a basis of genome analysis techniques in lab Gain exposure to various bioinformatic platforms for genetic studies

Brachypodium distachyon Model Organism ● Purple False Brome Image Courtesy of Wikipedia

Hypothesis of ECR Mutation You can simply impress your audience and add a unique zing and appeal to your Presentations. Easy to change colors, photos and Text. Get a modern Power. Point Presentation that is beautifully designed. We hypothesize that our gene encodes for plant growth regulation, as we noticed in the mutants that there was not as much visible growth.
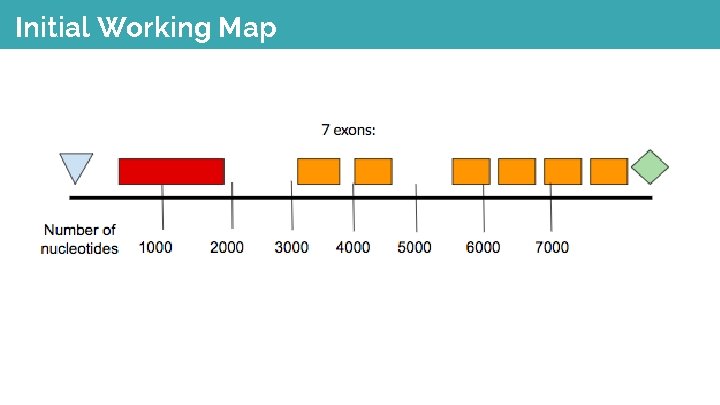
Initial Working Map
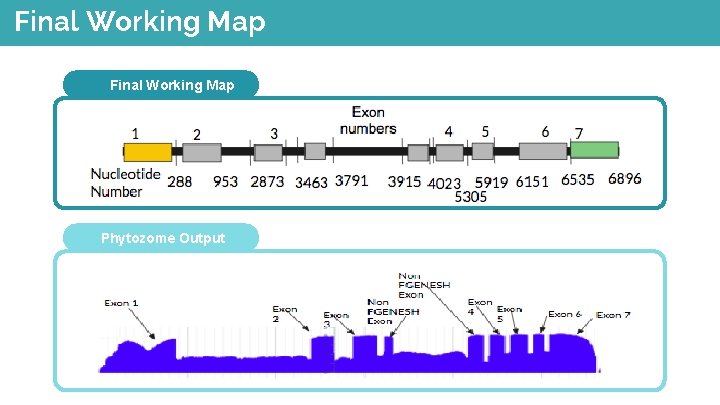
Final Working Map Phytozome Output

Protein Domains Phytozome BLAST domains ● Mutation Located: S-Locus glycoprotein ● Overall name: G-type lectin s-receptor-like serine/threonine-protein kinase
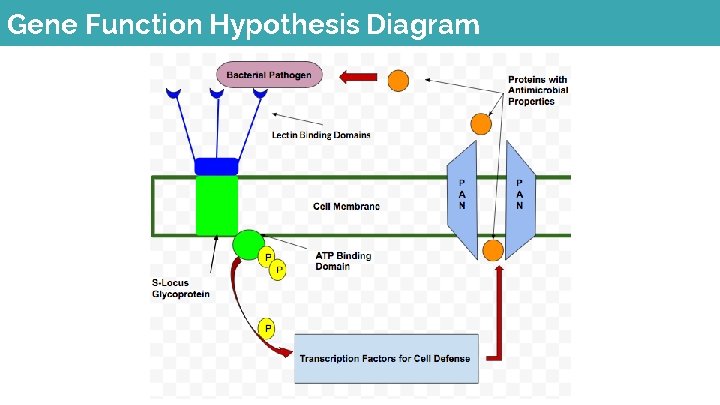
Gene Function Hypothesis Diagram
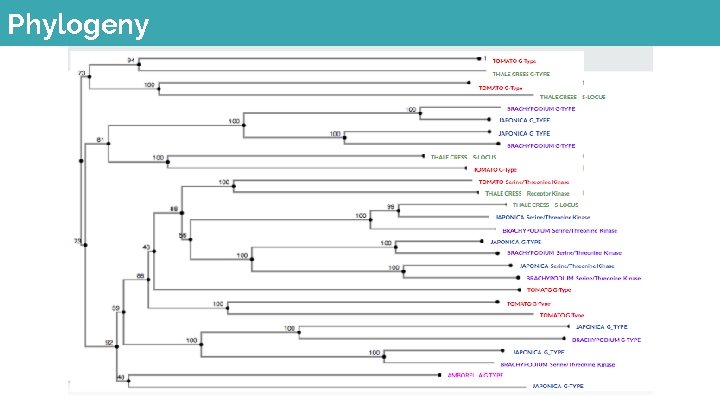
Phylogeny
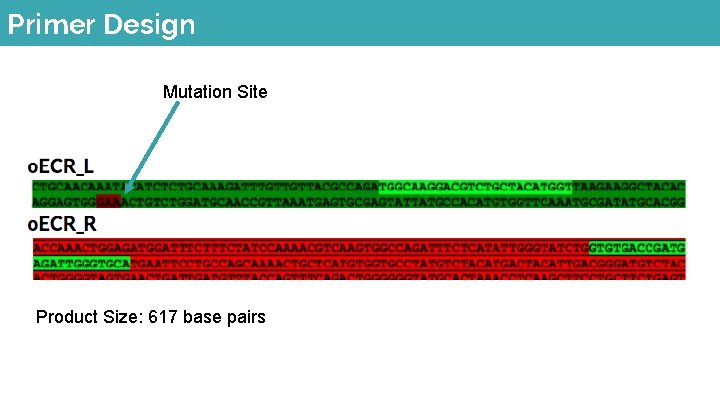
Primer Design Mutation Site Product Size: 617 base pairs
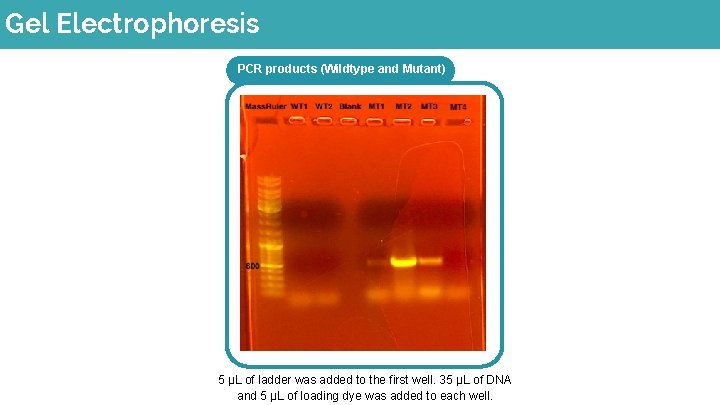
Gel Electrophoresis PCR products (Wildtype and Mutant) 5 μL of ladder was added to the first well. 35 μL of DNA and 5 μL of loading dye was added to each well.
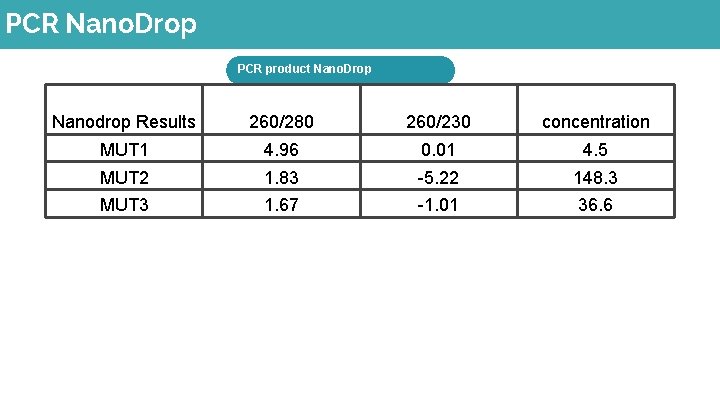
PCR Nano. Drop PCR product Nano. Drop Nanodrop Results 260/280 260/230 concentration MUT 1 4. 96 0. 01 4. 5 MUT 2 1. 83 -5. 22 148. 3 MUT 3 1. 67 -1. 01 36. 6

ECR Sequencing Results ECR Mutant 2 ECR Mutant 3
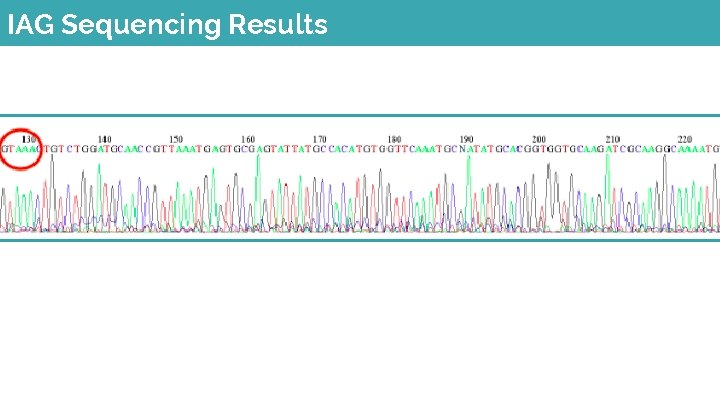
IAG Sequencing Results
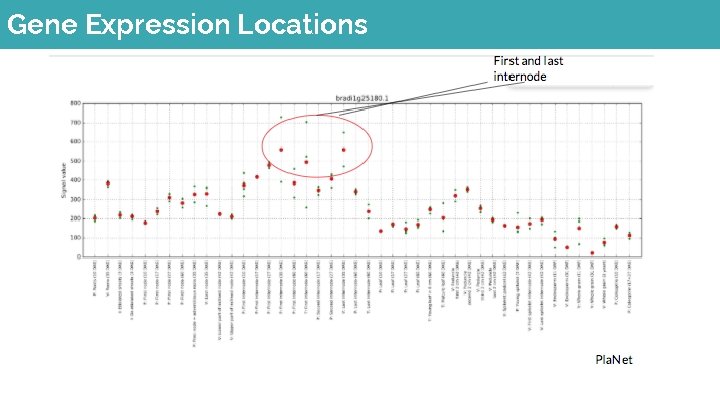
Gene Expression Locations
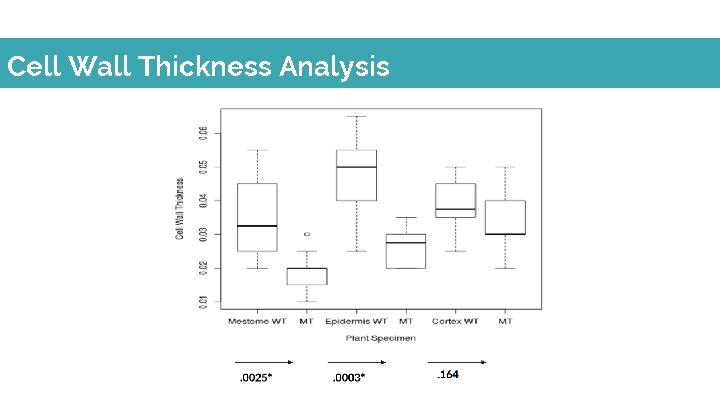
Cell Wall Thickness Analysis
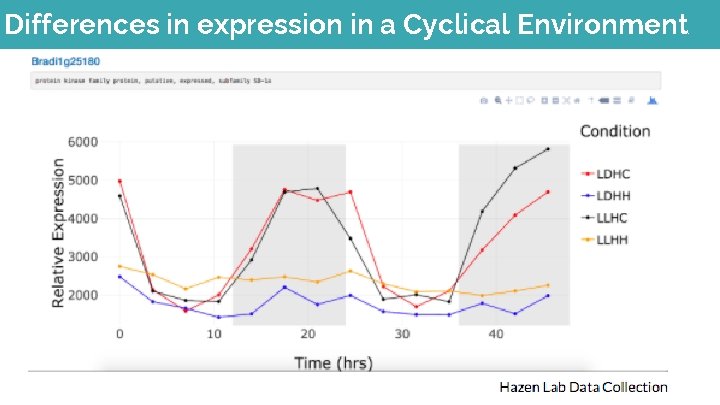
Differences in expression in a Cyclical Environment
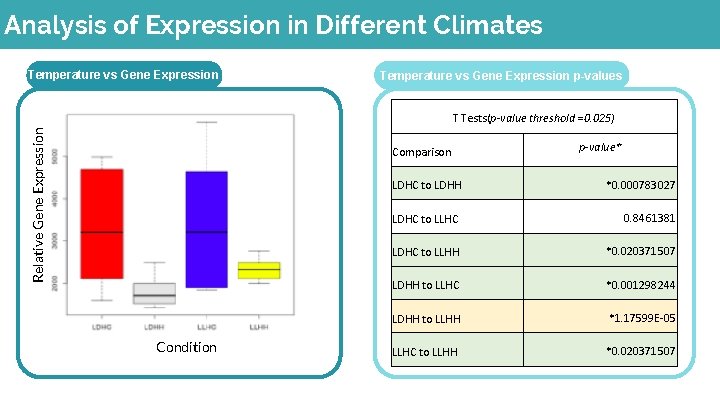
Analysis of Expression in Different Climates Temperature vs Gene Expression p-values Relative Gene Expression T Tests(p-value threshold =0. 025) Comparison Condition p-value* LDHC to LDHH *0. 000783027 LDHC to LLHC 0. 8461381 LDHC to LLHH *0. 020371507 LDHH to LLHC *0. 001298244 LDHH to LLHH *1. 17599 E-05 LLHC to LLHH *0. 020371507
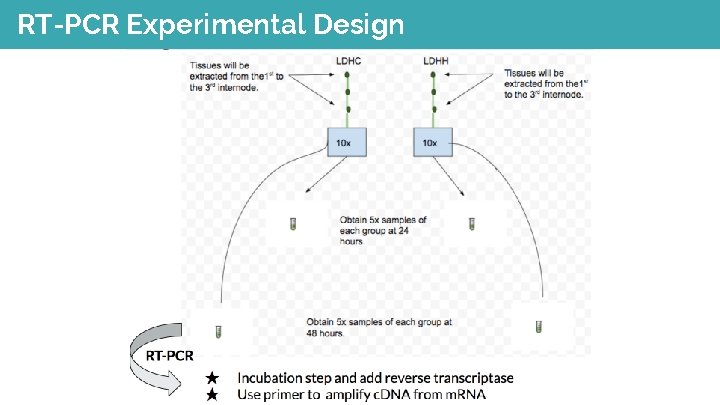
RT-PCR Experimental Design
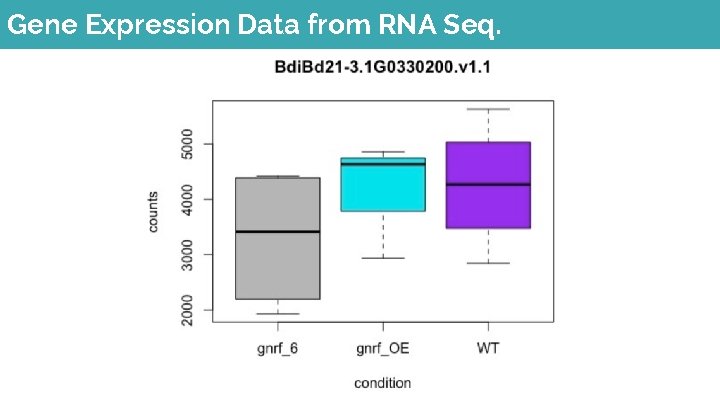
Gene Expression Data from RNA Seq.
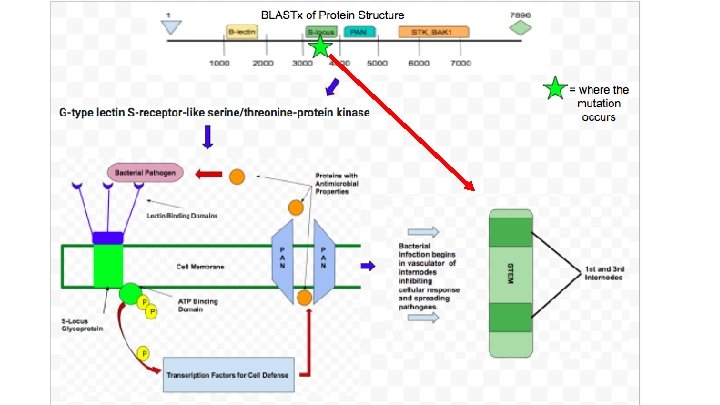
Gene Function Diagram

Summary of Results ● 4 Protein domains are encoded for by 9 exons. D-Mannose binding lectin, SLocus Glycoprotein Domain, Pan-like Domain, and a Protein tyrosine kinase ● The mutation is a Nonsense GAA→ TAA mutation caused by Na. N 3 ● Our Mutants did not contain the mutation but the MW section confirmed the mutation’s presence ● The gene in question is vital to the vasculature of the plant. Specifically, the 1 st and 3 rd(last) internodes

Conclusion ● Our gene encodes for the formation of cellular defense to external pathogens ● The mutation inhibits the formation of the gene (stop codon) which leaves the cell defenseless to pathogens that enter through the vasculature ● The 1 st and 3 rd internodes act as checkpoints for these pathogens and they no longer have this defense so the plant is susceptible to harmful pathogens

References Maimbo, Milimo, et al. “S-Glycoprotein-Like Protein Regulates Defense Responses in Nicotiana Plants against Ralstonia Solanacearum. ” Plant Physiology, American Society of Plant Biologists, 1 Apr. 2010, www. plantphysiol. org/content/152/4/2023. Reece, Richard J. Analysis of Genes and Genomes. John Wiley & Sons, 2003. Sanabria, Natasha M. , et al. “Molecular Characterisation and Regulation of a Nicotiana Tabacum S-Domain Receptor-like Kinase Gene Induced during an Early Rapid Response to Lipopolysaccharides. ” Gene, vol. 501, no. 1, 2012, pp. 39– 48. , doi: 10. 1016/j. gene. 2012. 03. 073. Uemura, Kazuhide, et al. “Preparation of Recombinant Mannan-Binding Protein with a Native Oligomeric Structure. ” Recognition of Carbohydrates in Biological Systems, Part B: Specific Applications Methods in Enzymology, vol. 363, 2003, pp. 16– 26. , doi: 10. 1016/s 0076 -6879(03)01040 -1. Yan, Liming, et al. “Structural Basis for the Impact of Phosphorylation on the Activation of Plant Receptor-like Kinase BAK 1. ” Nature News, Nature Publishing Group, 1 May 2012, www. nature. com/articles/cr 201274. Resources: https: //phytozome. jgi. doe. gov/pz/portal. html# http: //www. bioline. com/media/calculator/01_13. html https: //blast. ncbi. nlm. nih. gov/Blast. cgi http: //www. softberry. com/berry. phtml? topic=fgenesh&group=programs&subgroup=gfind http: //primer 3. ut. ee/ http: //doua. prabi. fr/software/cap 3 https: //mafft. cbrc. jp/alignment/server/ http: //bar. utoronto. ca/efp_brachypodium/cgi-bin/efp. Web. cgi

Acknowledgements Professor Hamid Razifard, Ph. D Amber De. Neve, Ph. D Student, Teaching Assistant Professor Samuel Hazen, MS, Ph. D Kate Dorfman, Biology Lab Coordinator Entire Tues. /Thurs. Section IAG Group M/W Section Thank You!
- Slides: 25