BNC BIOPHYSICS NANOSCIENCE CENTRE CNISM Tuscia University Viterbo

BNC BIOPHYSICS & NANOSCIENCE CENTRE CNISM & Tuscia University Viterbo, Italy Salvatore Cannistraro www. unitus. it/biophysics/ AFM COST MEETING – Paris – May 2011 1
BNC BIOPHYSICS & NANOSCIENCE CENTRE CNISM & Tuscia University Viterbo, Italy Research activities Scanning Probe Nanoscopies & Spectroscopies Surface-Enhanced Raman Spectroscopy Surface Plasmons Resonance Modelling & Molecular Dynamics Biosensing & Bioelectronics AFM COST MEETING – Paris – May 2011 2
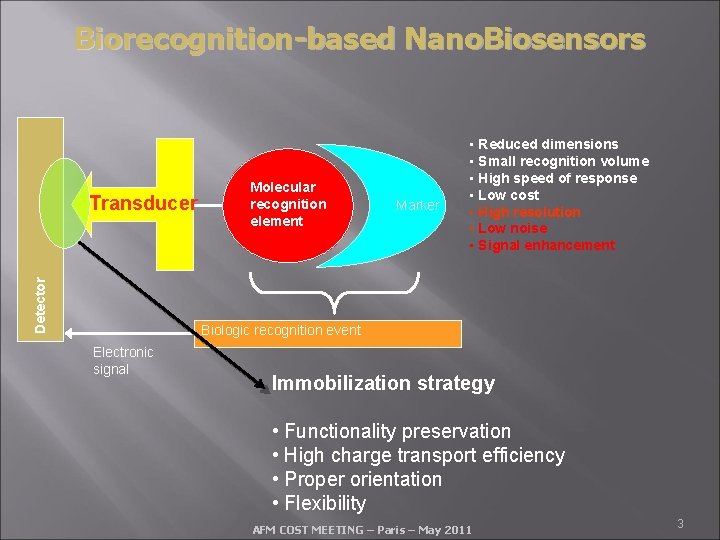
Biorecognition-based Nano. Biosensors Detector Transducer Molecular recognition element Marker • Reduced dimensions • Small recognition volume • High speed of response • Low cost • High resolution • Low noise • Signal enhancement Biologic recognition event Electronic signal Immobilization strategy • Functionality preservation • High charge transport efficiency • Proper orientation • Flexibility AFM COST MEETING – Paris – May 2011 3

Biosensing Current resolution limit (ELISA): 10 -10 M, 1013 molecules Ideal goal for early diagnosis: detection of a single molecule Main limit: NOISE Concrete goal: resolution limit to 10 -18 M, 105 molecules Such result can be achieved by improving biosensors with: Molecular biorecognition exploitation: one signal per event Nanostructures conjugation: noise lowering and signal enhancement AFM COST MEETING – Paris – May 2011 4
Single molecule techniques for early diagnostics Single molecule techniques: • • Atomic Force Microscopy and Spectroscopy (AFM & AFS) Scanning Tunnelling Microscopy (STM) Conductive-AFM Surface Enhanced Raman Spectroscopy (SERS) Recognition capability of biomolecules (1: 1) Nanotechnological structures: nanoparticles, nanotubes etc. AFM COST MEETING – Paris – May 2011 5
Biorecognition refers to highly specific interactions between two biological molecules, exhibiting unambiguous one-to-one complementarity. Biorecognition is involved in many important biological processes, including genome replication and transcription, enzymatic activity, immune response, cellular signalling, . . . AFM COST MEETING – Paris – May 2011 6

Biorecognition with Atomic Force Spectroscopy Bizarri AR, Cannistraro S (2009) J. Phys. Chem. B 113: 16449 -16464 Bizzarri AR, Cannistraro S. (2010) Chem. Soc. Rev. 39: 734 -749 AFM COST MEETING – Paris – May 2011 7

Protein – proten interaction 1. Azurin-Cytochrome c 551: a transient complex, with high k Azurin-Cytochrome c 551: off , where the linking spacer may play a crucial role. 2. p 53 -Mdm 2: a stable complex, with low k p 53 -Mdm 2: off, formed by a human tumor suppressor and its down-regulator. 3. p 53 -Azurin: a bacterial protein showing an anti-cancer action a forms a heterogeneous complex with p 53. 4. p 28 -p 53: an azurin-derived peptide fragment displaying the same anticancer potentiality of the whole protein. AFM COST MEETING – Paris – May 2011 8

Azurin-Cythocrome c 551: a Transient Complex Biological interest: electron transfer interaction involved in electron transfer interaction the nitrate respiration of bacterium Pseudomonas aeruginosa. First AFS study on an electron transfer complex Docking simulations: best complex from close contact between the hydrophobic regions of the two proteins Optim. ET [Bizzarri et al. , JMR 20, 122 (2007)] Immobilization strategies: • PEG molecules for flexible linking of cytochrome to the tip, targeting –NH 2; • oriented azurin bonding to the Au (electr sens)substrate via disulphide bridge, with or without spacers. AFS results: • Single energy barrier; • koff values consistent with a transient complex: 7 and 14 s-1; • immobilization via organic spacers increases the binding efficiency Bonanni et al. , BJ 89, 2783 (2005) and JPCB 110, 14574 (2006) AFM COST MEETING – Paris – May 2011 9
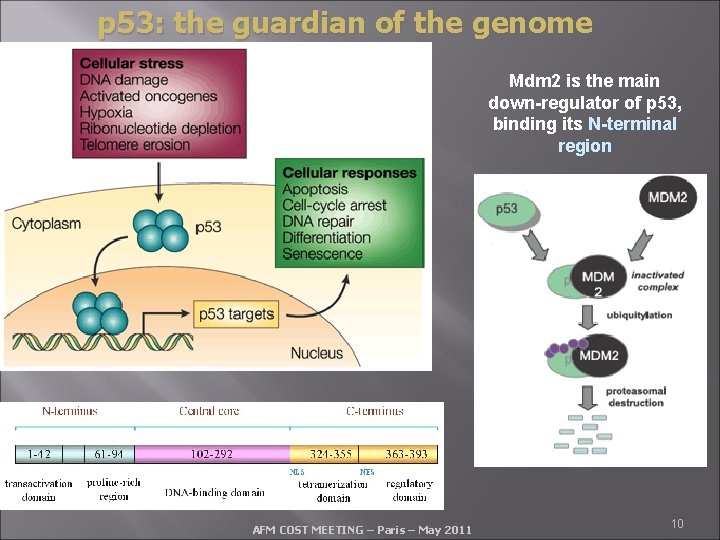
p 53: the guardian of the genome Mdm 2 is the main down-regulator of p 53, binding its N-terminal region AFM COST MEETING – Paris – May 2011 10
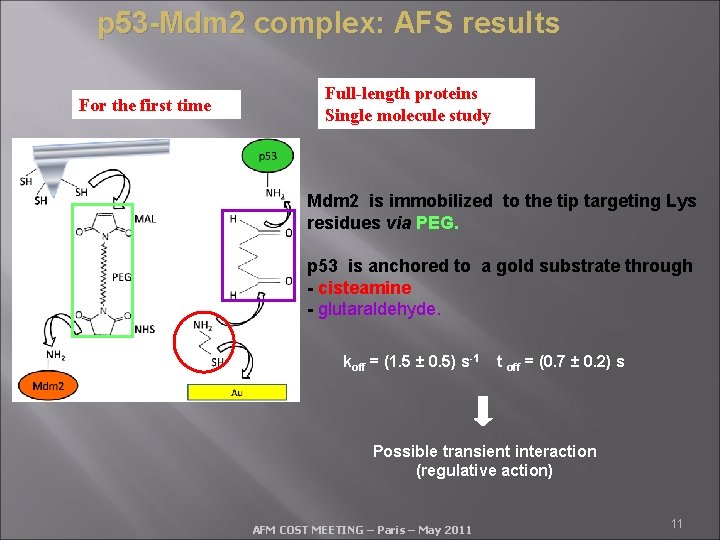
p 53 -Mdm 2 complex: AFS results For the first time Full-length proteins Single molecule study Mdm 2 is immobilized to the tip targeting Lys residues via PEG. p 53 is anchored to a gold substrate through - cisteamine - glutaraldehyde. koff = (1. 5 ± 0. 5) s-1 t off = (0. 7 ± 0. 2) s Possible transient interaction (regulative action) AFM COST MEETING – Paris – May 2011 11
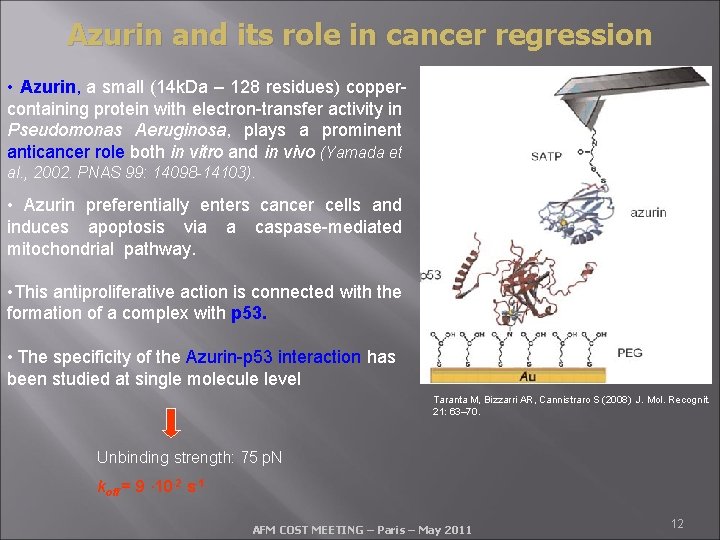
Azurin and its role in cancer regression • Azurin, a small (14 k. Da – 128 residues) coppercontaining protein with electron-transfer activity in Pseudomonas Aeruginosa, plays a prominent anticancer role both in vitro and in vivo (Yamada et al. , 2002. PNAS 99: 14098 -14103). • Azurin preferentially enters cancer cells and induces apoptosis via a caspase-mediated mitochondrial pathway. • This antiproliferative action is connected with the formation of a complex with p 53. • The specificity of the Azurin-p 53 interaction has been studied at single molecule level Taranta M, Bizzarri AR, Cannistraro S (2008) J. Mol. Recognit. 21: 63– 70. Unbinding strength: 75 p. N koff = 9 · 10 -2 s-1 AFM COST MEETING – Paris – May 2011 12
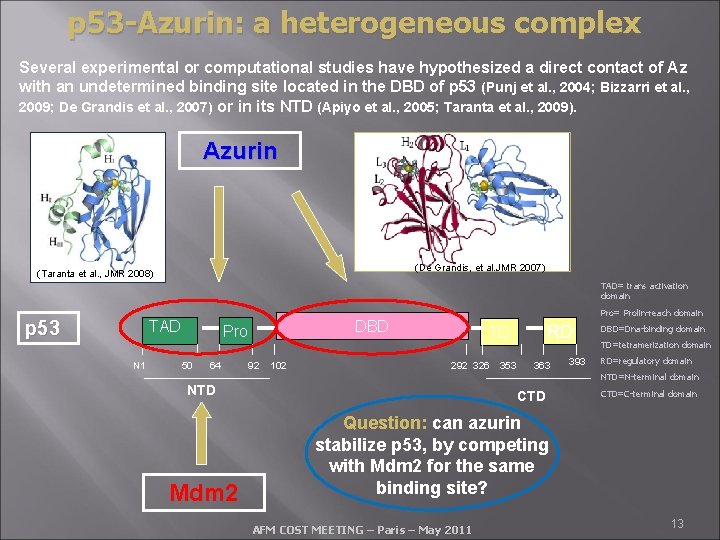
p 53 -Azurin: a heterogeneous complex Several experimental or computational studies have hypothesized a direct contact of Az with an undetermined binding site located in the DBD of p 53 (Punj et al. , 2004; Bizzarri et al. , 2009; De Grandis et al. , 2007) or in its NTD (Apiyo et al. , 2005; Taranta et al. , 2009). Azurin (De Grandis, et al. JMR 2007) (Taranta et al. , JMR 2008) TAD= trans activation domain p 53 TAD N 1 DBD Pro 50 64 Pro= Prolin-reach domain 92 102 RD TD 292 326 353 363 393 DBD=Dna-binding domain TD=tetramerization domain RD=regulatory domain NTD=N-terminal domain NTD Mdm 2 CTD=C-terminal domain Question: can azurin stabilize p 53, by competing with Mdm 2 for the same binding site? AFM COST MEETING – Paris – May 2011 13
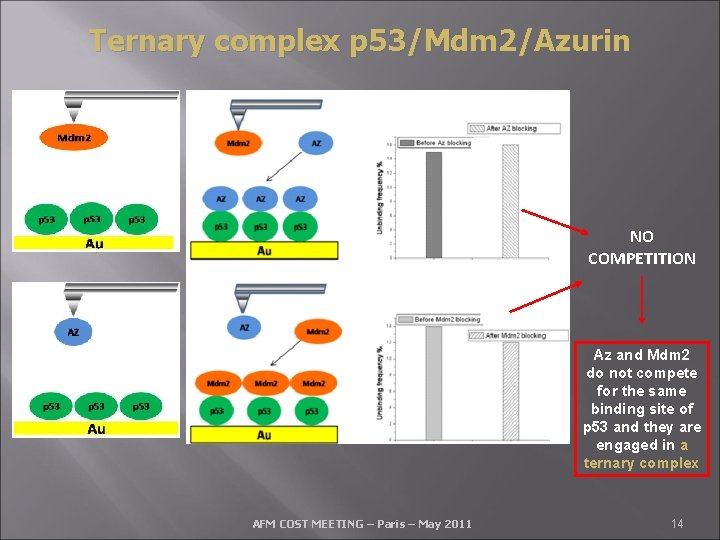
Ternary complex p 53/Mdm 2/Azurin NO COMPETITION Az and Mdm 2 do not compete for the same binding site of p 53 and they are engaged in a ternary complex AFM COST MEETING – Paris – May 2011 14

Surface Plasmon Resonance studies have also shown that Azurin is able to induce a weakening of the Mdm 2 -p 53 interaction by a non competitive inhibition mechanism, which should be figured out as a long range binding regulation. NTD 1 DBD 94 CD studies CTD 292 393 Possible allosteric regulation of this azurin-induced inhibition Az The contribution of our centre has been crucial in disclosing the molecular and kinetic details of the Azurin-p 53 interaction. It has also suggest to search for the azurin smallest peptide fragment retaining both the azurin cellular penetration ability and anti-proliferative activity a. a)Yamada T. et al. , 2009. Mol. Cancer Ther. (2009) 8: 2947 -2958 AFM COST MEETING – Paris – May 2011 15
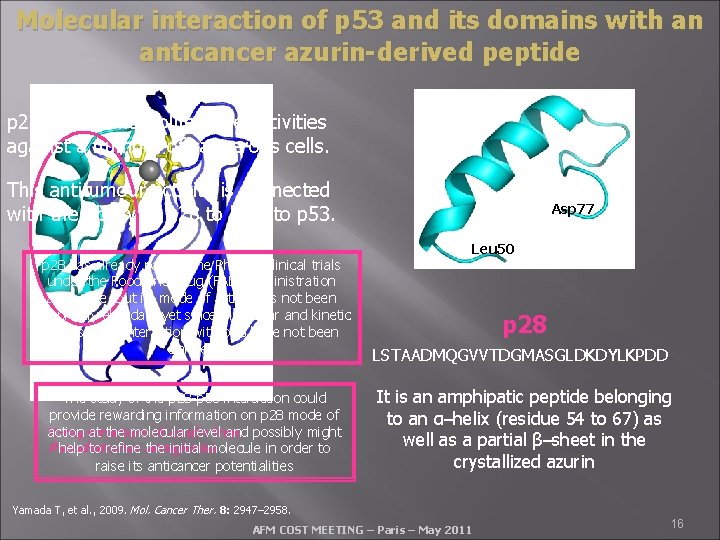
Molecular interaction of p 53 and its domains with an anticancer azurin-derived peptide p 28 shows antiproliferative activities against a number of cancerous cells. This antitumour activity is connected with the ability of p 28 to bind to p 53. p 28 has already passed the Phase I clinical trials under the Food and Drug (FAD) administration allowance, but its mode of action has not been completely elucidate yet since molecular and kinetic details of its interaction with p 53 have not been clarified The study of the p 28 -p 53 interaction could provide rewarding information on p 28 mode of X-ray structure of azurin from action at the molecular level and possibly might Pseudomonas help to refine aeruginosa the initial molecule in order to raise its anticancer potentialities Asp 77 Leu 50 p 28 LSTAADMQGVVTDGMASGLDKDYLKPDD It is an amphipatic peptide belonging to an α–helix (residue 54 to 67) as well as a partial β–sheet in the crystallized azurin Yamada T, et al. , 2009. Mol. Cancer Ther. 8: 2947– 2958. AFM COST MEETING – Paris – May 2011 16
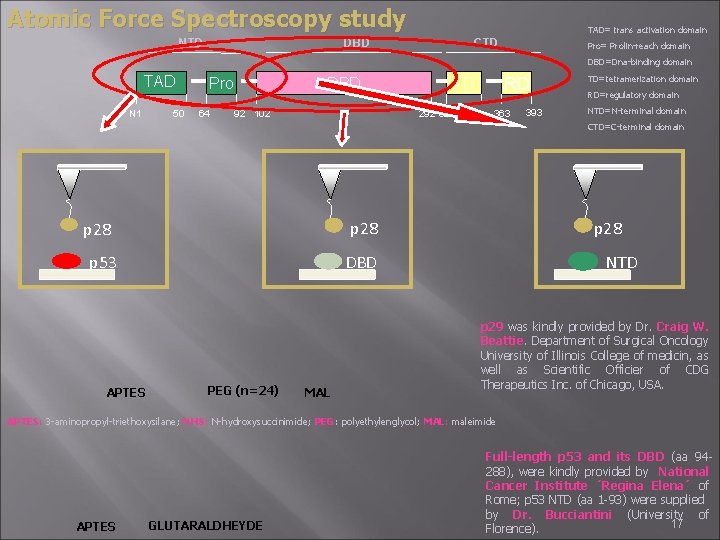
Atomic Force Spectroscopy study DBD NTD TAD= trans activation domain CTD Pro= Prolin-reach domain DBD=Dna-binding domain TAD N 1 50 Pro 64 DBD 92 102 RD TD 292 326 353 363 393 TD=tetramerization domain RD=regulatory domain NTD=N-terminal domain CTD=C-terminal domain p 28 p 53 APTES p 28 DBD PEG (n=24) MAL NTD p 29 was kindly provided by Dr. Craig W. Beattie. Department of Surgical Oncology University of Illinois College of medicin, as well as Scientific Officier of CDG Therapeutics Inc. of Chicago, USA. APTES: 3 -aminopropyl-triethoxysilane; NHS: N-hydroxysuccinimide; PEG: polyethylenglycol; MAL: maleimide APTES GLUTARALDHEYDE Full-length p 53 and its DBD (aa 94288), were kindly provided by National Cancer Institute ´Regina Elena´ of Rome; p 53 NTD (aa 1 -93) were supplied by Dr. Bucciantini (University of 17 Florence).
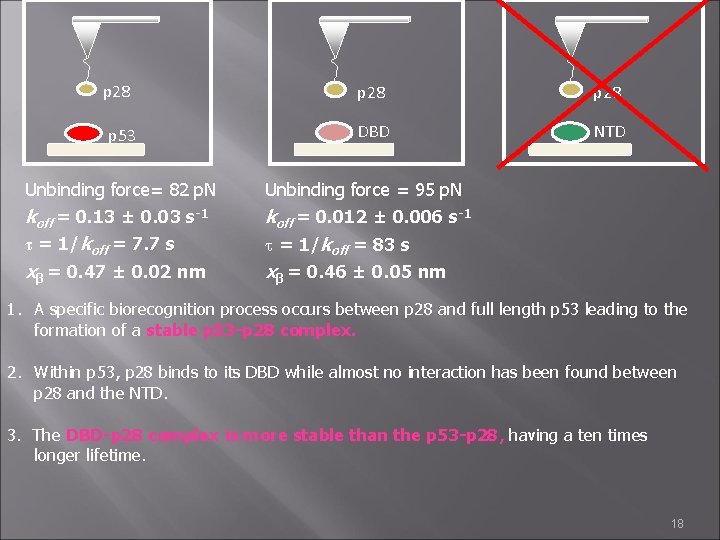
p 28 p 53 p 28 DBD NTD Unbinding force= 82 p. N Unbinding force = 95 p. N koff = 0. 13 ± 0. 03 s-1 τ = 1/koff = 7. 7 s xβ = 0. 47 ± 0. 02 nm koff = 0. 012 ± 0. 006 s-1 t = 1/koff = 83 s xβ = 0. 46 ± 0. 05 nm 1. A specific biorecognition process occurs between p 28 and full length p 53 leading to the formation of a stable p 53 -p 28 complex. 2. Within p 53, p 28 binds to its DBD while almost no interaction has been found between p 28 and the NTD. 3. The DBD-p 28 complex is more stable than the p 53 -p 28, having a ten times longer lifetime. 18
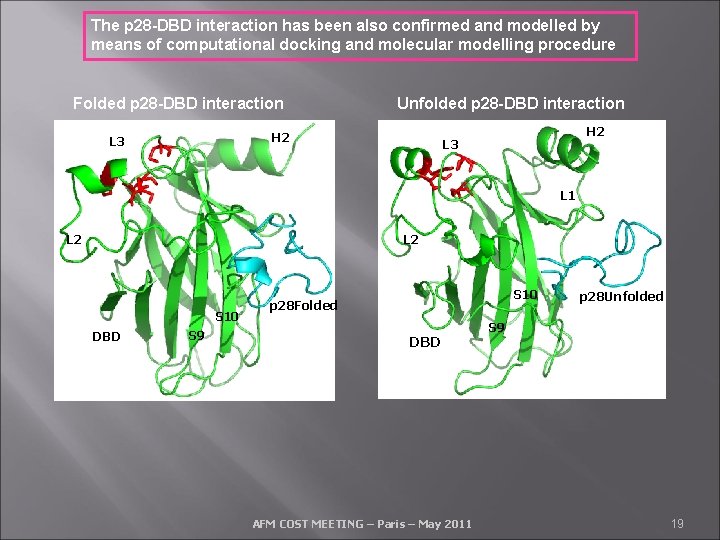
The p 28 -DBD interaction has been also confirmed and modelled by means of computational docking and molecular modelling procedure Folded p 28 -DBD interaction Unfolded p 28 -DBD interaction H 2 L 3 L 1 L 2 S 10 DBD S 9 S 10 p 28 Folded DBD AFM COST MEETING – Paris – May 2011 p 28 Unfolded S 9 19

Conclusions AFS represents one of the most rewarding tools for studying biological processes, allowing to measure forces acting between individual biomolecules with sensitivity of p. N, in near-physiological conditions and without labelling. AFS data permit to obtain information on the dissociation rate constant koff and on the activation free energy for a single ligand-receptor pair, complementing traditional biochemical approaches. In our case, it allowed us to reach unprecedented kinetic results on the p 53/mdm 2 complex formed by the two full-length proteins. Moreover , it evidenced that the p 53 stabilization induced by Azurin cannot be attributed to competitive binding with respect to mdm 2. The occurrence of the ternary complex opens new perspective for the anticancer action of azurin AFM COST MEETING – Paris – May 2011 20

Atomic Force Spectroscopy and Biomolecular Recognition Editor(s): Salvatore Cannistraro; Anna Rita Bizzarri http: //www. crcpress. com/product/isbn/9781439862377 Table of contents Introduction Biorecognition Processes, Dr. Bongrand, France Atomic Force Microscopy and Spectroscopy, Dr. Hoelscher, Germany Theoretical Models, Dr. Friddle, Usa Immobilization Strategies, Dr. Akhremitchev, Usa Data Analysis, Drs. Pellequer and Parot, France Outline of the Most Relevant Applications, Drs. Bizzarri and Cannistraro, Italy Conclusions and Perspectives Price: $129. 95 Cat. #: K 12890 ISBN: 9781439862377 ISBN 10: 1439862370 Publication Date: December 26, 2011 Number of Pages: 240 forthcoming book 21

Biophysics & Nanoscience Centre Anna Rita Bizzarri (Professor of Physics) Ines Delfino (Research Assistant Professor) Chiara Baldacchini (CNR-CNISM Research Fellow) Samuele Raccosta (Post. Doc) Fabio Domenici (Ph. D Student) Xian Jin Xu (Phd Student) Simona Santini (Graduated Student) www. unitus. it/biophysics/ AFM COST MEETING – Paris – May 2011 22
- Slides: 22