BMS 631 LECTURE 10 x Flow Cytometry Theory

BMS 631 - LECTURE 10 x Flow Cytometry: Theory J. Paul Robinson Professor of Immunopharmacology Professor of Biomedical Engineering Purdue University Multiparameter Data Analysis 3 rd Ed. Shapiro p 207 -214 Bindley Bioscience Center Purdue University Office: 494 0757 Fax 494 0517 email; robinson@flowcyt. cyto. purdue. edu WEB http: //www. cyto. purdue. edu Page 1 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Data Analysis • Gating • Data displays – – – histogram dot plot isometric display contour plot chromatic (color) plots 3 D projection Page 2 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Gating • Real-time gating vs. software gating • Establishing regions • Gating strategies • Quadrant analysis • Complex or Boolean gates • Back gating Page 3 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Real-Time vs. Software Gating Real-time or live gating: -restrict the data that will be accepted by a computer (some characteristic must be met before data is stored) Software or analysis gating: -excludes certain stored data from a particular analysis procedure Page 4 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
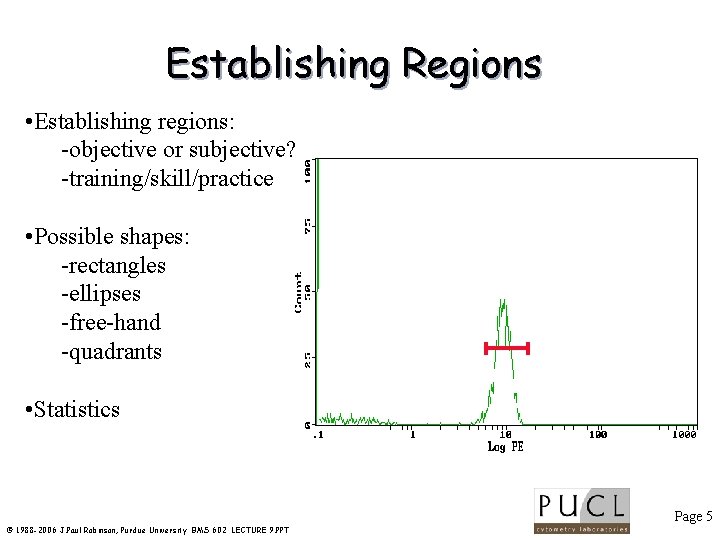
Establishing Regions • Establishing regions: -objective or subjective? -training/skill/practice • Possible shapes: -rectangles -ellipses -free-hand -quadrants • Statistics Page 5 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
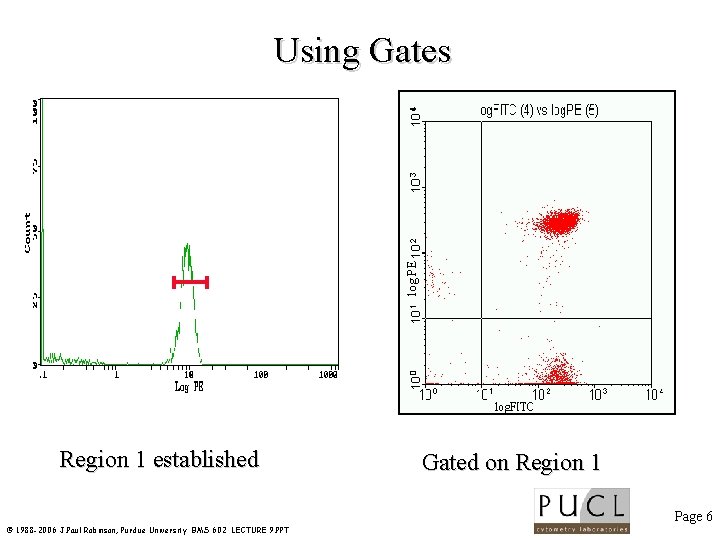
log PE Using Gates Region 1 established Gated on Region 1 Page 6 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
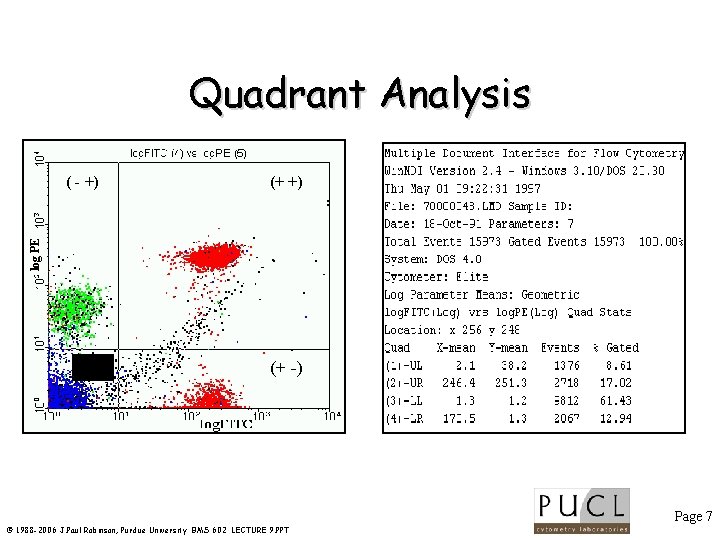
Quadrant Analysis (+ +) log PE ( - +) (- -) (+ -) Page 7 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
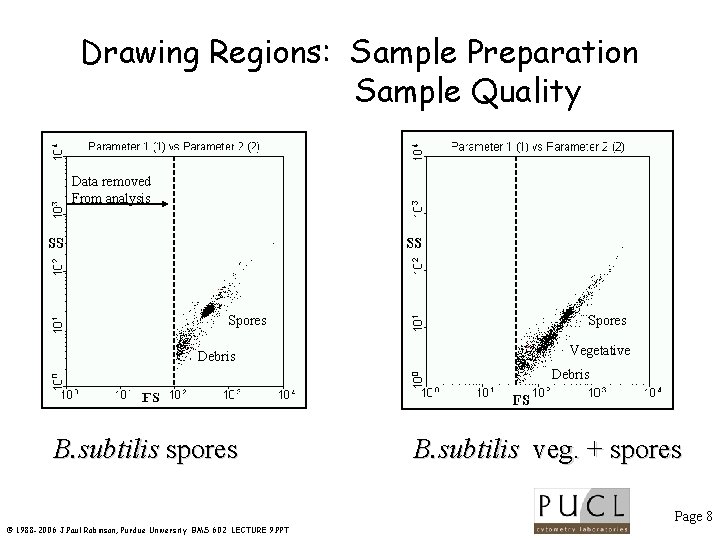
Drawing Regions: Sample Preparation Sample Quality Data removed From analysis Spores Debris Spores Vegetative Debris B. subtilis spores B. subtilis veg. + spores Page 8 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
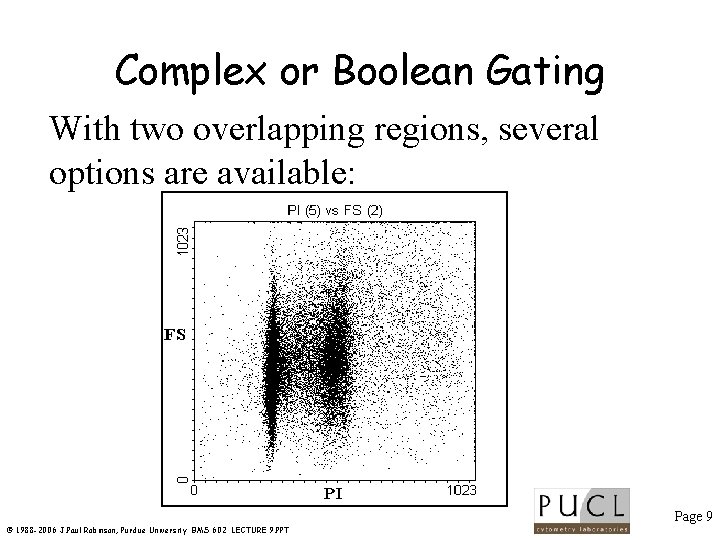
Complex or Boolean Gating With two overlapping regions, several options are available: Page 9 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
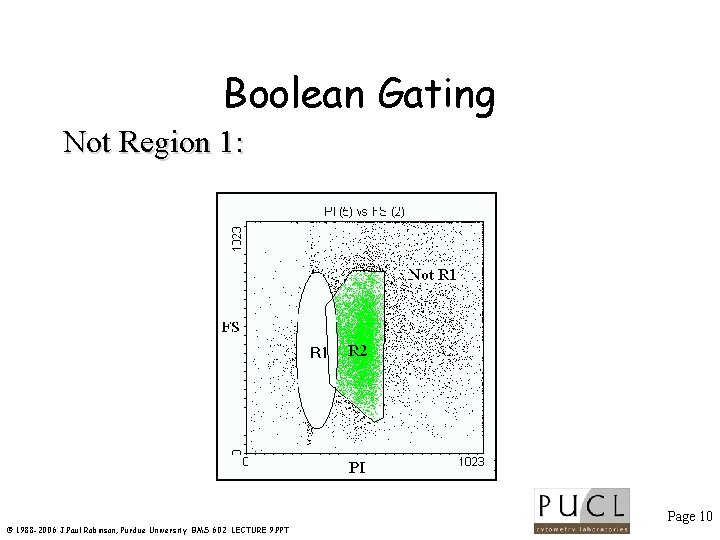
Boolean Gating Not Region 1: Page 10 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
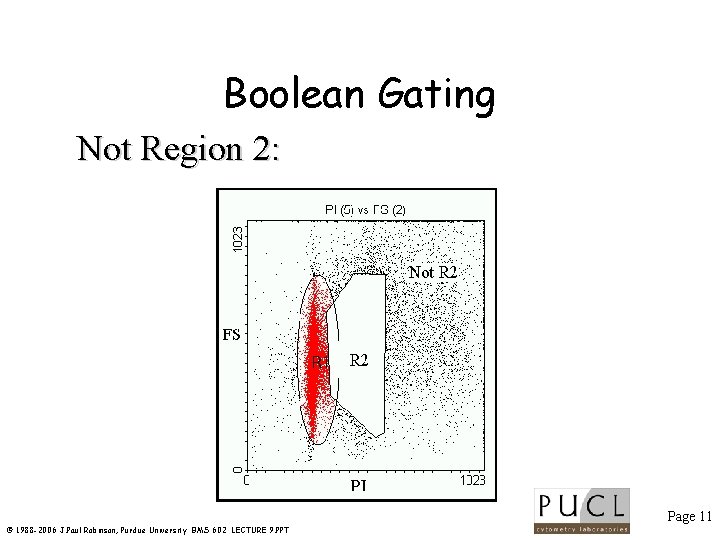
Boolean Gating Not Region 2: Page 11 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
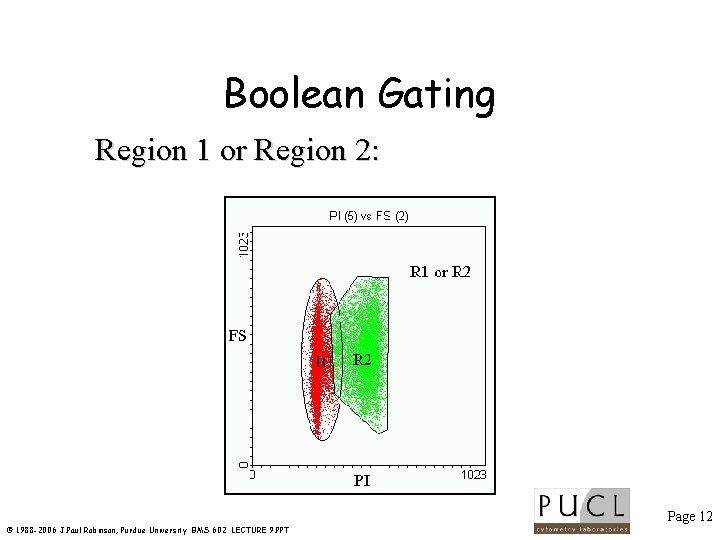
Boolean Gating Region 1 or Region 2: Page 12 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Boolean Gating Region 1 and Region 2: Page 13 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
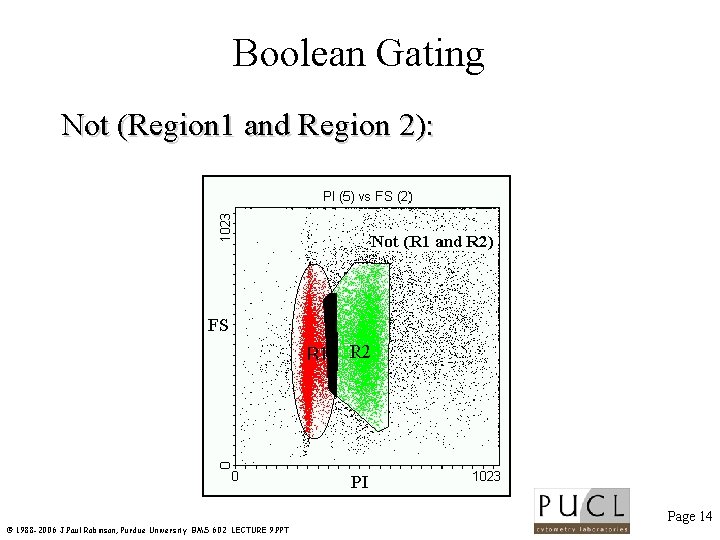
Boolean Gating Not (Region 1 and Region 2): Page 14 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
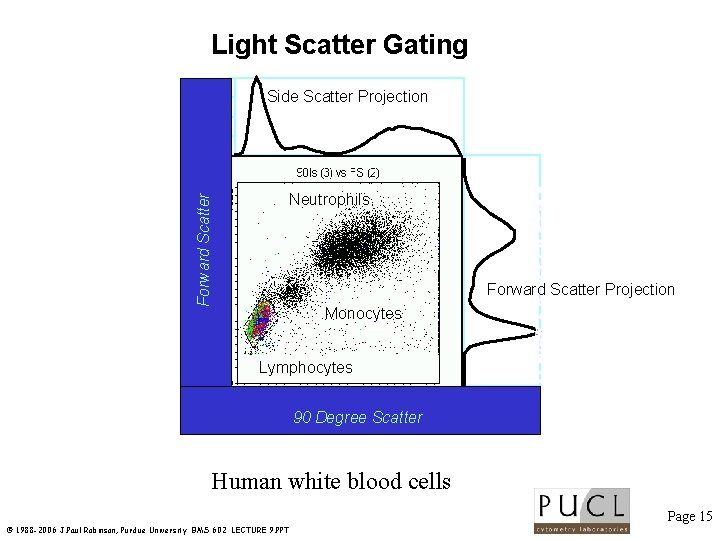
Light Scatter Gating Side Scatter Projection Forward Scatter Neutrophils Forward Scatter Projection Monocytes Lymphocytes 0 200 400 600 800 1000 90 Degree Scatter Human white blood cells Page 15 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
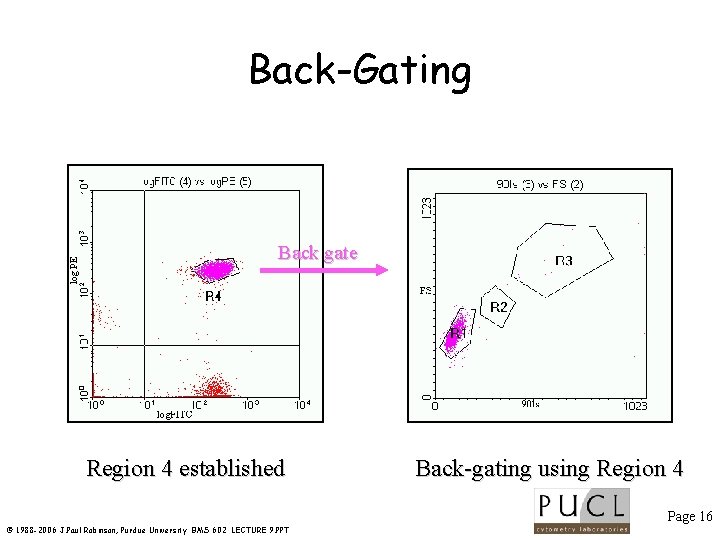
log PE Back-Gating Back gate Region 4 established Back-gating using Region 4 Page 16 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
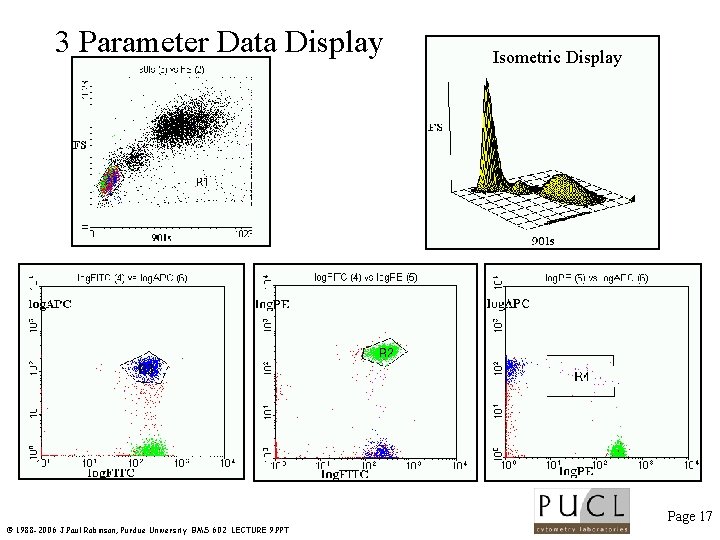
3 Parameter Data Display Isometric Display Page 17 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Methods that can change results: 1. Doublet discrimination 2. Time as a quality control parameter Example: DNA content -need to eliminate debris & clumps -need to gate out doublets -maintain constant flow rate Page 18 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
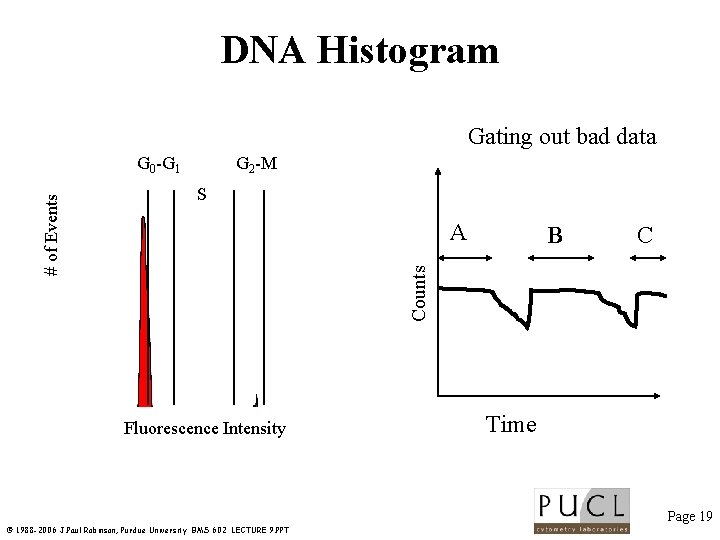
DNA Histogram Gating out bad data G 2 -M S A B C Counts # of Events G 0 -G 1 Fluorescence Intensity Time Page 19 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
Multi-color studies generate a lot of data ++ -- +- QUADSTATS -+ ++ -- +- Log Fluorescence QUADSTATS -+ ++ -- +- 10 QUADSTATS -+ ++ -- +- QUADSTATS Log Fluorescence ++ Log Fluorescence +- -+ Log Fluorescence -- QUADSTATS Log Fluorescence QUADSTATS -+ ++ 9 Log Fluorescence QUADSTATS -+ ++ -- +- Log Fluorescence QUADSTATS Log Fluorescence +- +- Log Fluorescence -+ 8 Log Fluorescence -- -- QUADSTATS 7 Log Fluorescence ++ ++ 6 Log Fluorescence -+ -+ 5 Log Fluorescence +- Log Fluorescence -- +- QUADSTATS -+ ++ -- +- Log Fluorescence QUADSTATS QUADSTATS QUADSTATS -+ ++ -- +- Log Fluorescence -+ ++ -- +- Log Fluorescence -+ ++ -- +- Log Fluorescence Log Fluorescence Log Fluorescence Log Fluorescence Log Fluorescence ++ -- QUADSTATS Log Fluorescence QUADSTATS -+ ++ 4 Log Fluorescence +- Log Fluorescence -+ Log Fluorescence -- QUADSTATS Log Fluorescence ++ Log Fluorescence QUADSTATS 3 Log Fluorescence 2 Log Fluorescence 1 Log Fluorescence 3 color -+ 5 color 4 color -+ ++ -- +- Log Fluorescence Page 20 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
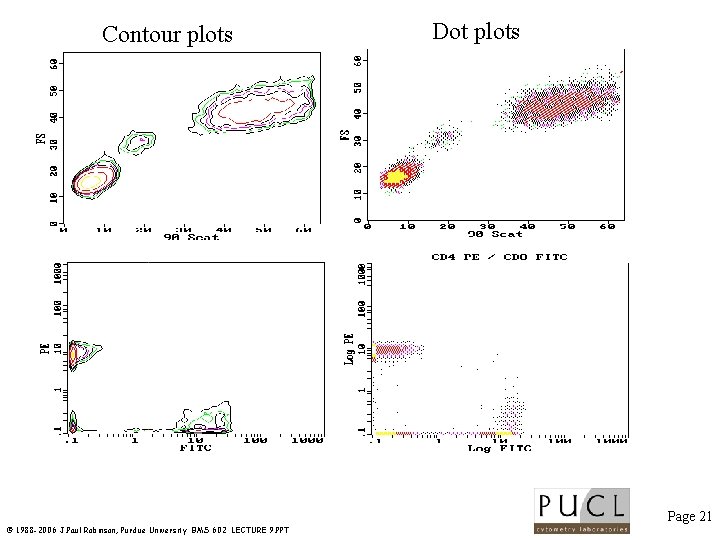
Contour plots Dot plots Page 21 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
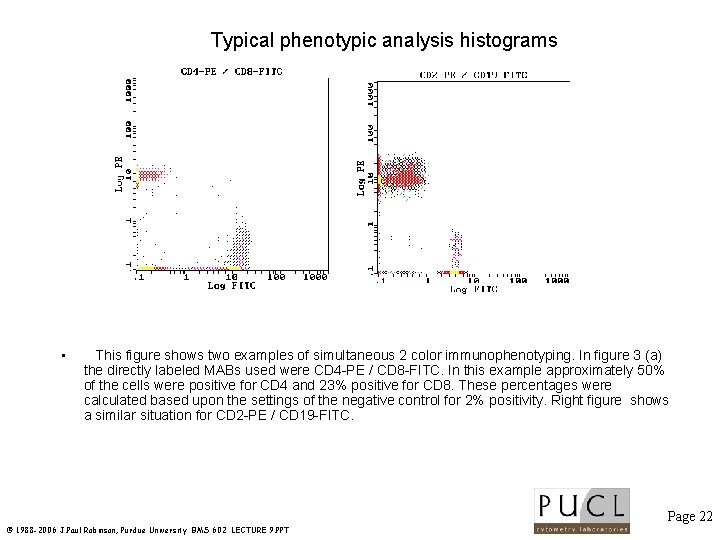
Typical phenotypic analysis histograms • This figure shows two examples of simultaneous 2 color immunophenotyping. In figure 3 (a) the directly labeled MABs used were CD 4 -PE / CD 8 -FITC. In this example approximately 50% of the cells were positive for CD 4 and 23% positive for CD 8. These percentages were calculated based upon the settings of the negative control for 2% positivity. Right figure shows a similar situation for CD 2 -PE / CD 19 -FITC. Page 22 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
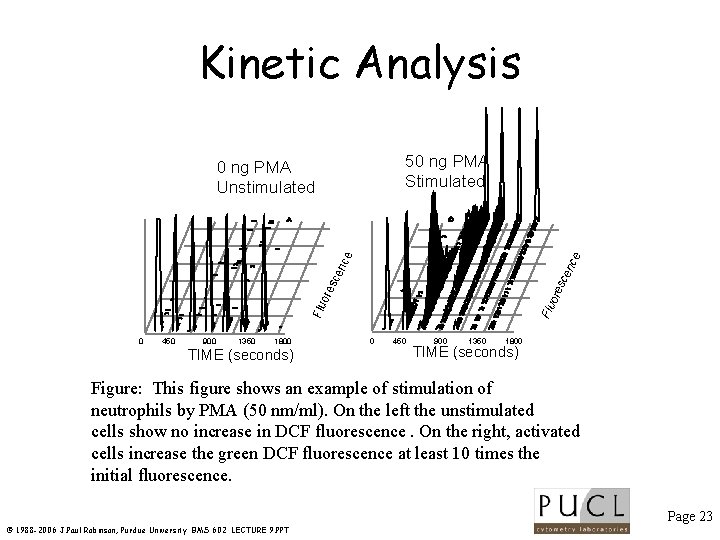
Kinetic Analysis 50 ng PMA Stimulated 0 450 900 1350 1800 TIME (seconds) Flu o res cen ce ce 0 ng PMA Unstimulated 0 450 900 1350 1800 TIME (seconds) Figure: This figure shows an example of stimulation of neutrophils by PMA (50 nm/ml). On the left the unstimulated cells show no increase in DCF fluorescence. On the right, activated cells increase the green DCF fluorescence at least 10 times the initial fluorescence. Page 23 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Color Coded Dot Plots Key to understanding this figure, is the notion that different populations can be identified by colors and the relationship of these populations to one another can be monitored. Page 24 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

50 40 30 0 10 20 Forward Scatter 60 Contour Plot with Projection 0 10 20 30 60 90 Light Scatter 40 50 Page 25 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

4+4+4=12 A PC 3 Color Combinations PE Positives 4+4=8 FITC Negatives Page 26 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
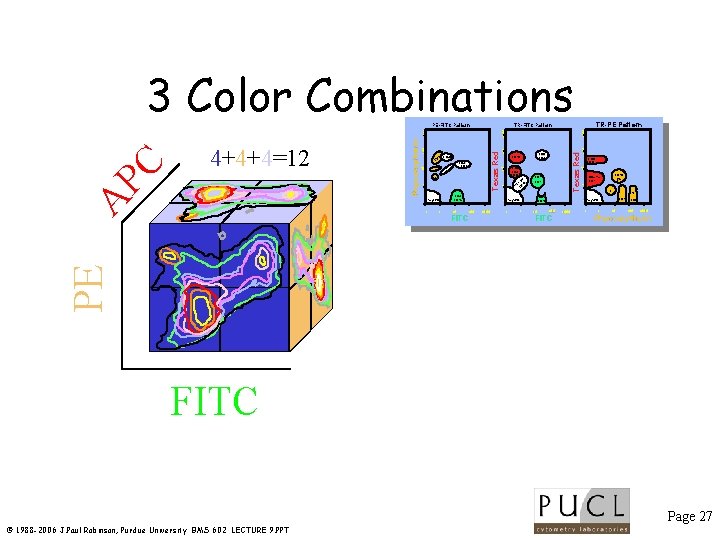
3 Color Combinations. 1 1 CD 3 CD 45 10 . 1 Negative 100 FITC 1000 . 1 1000 100 CD 8 CD 20 10 CD 8 CD 3 CD 20 Texas Red CD 8 CD 3 CD 20 CD 7 CD 2 CD 3 CD 45 Negative 1 10 CD 20 Ig. G CD 8 (dim) CD 2 1 DR A- 0 HL CD 1 L . 1 1 Ig. G 10 K/L 1 Texas Red Ig. M CD 3 CD 4 1000 100 10 TR-PE Pattern TR-FITC Pattern 100 FITC 1000 . 1 1 K CD 4 Negative 10 CD 5 1000 Phycoerytherin PE Phycoerytherin 5 CD 38 CD . 1 4+4+4=12 A PC PE-FITC Pattern FITC Page 27 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Page 28 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
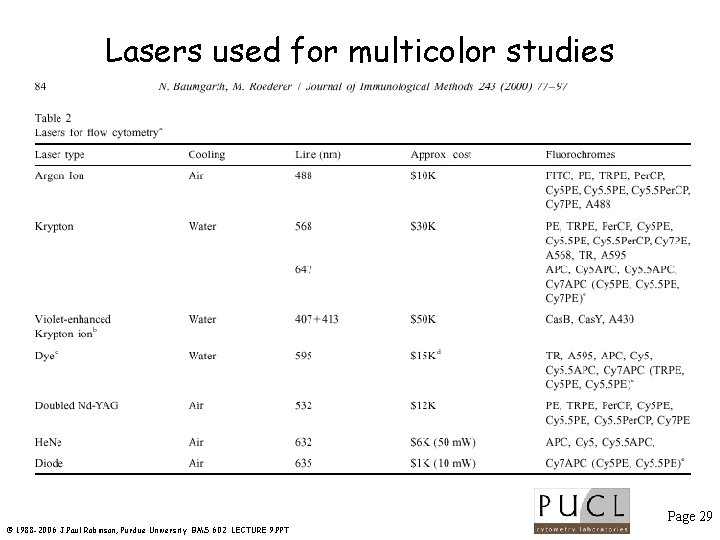
Lasers used for multicolor studies Page 29 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
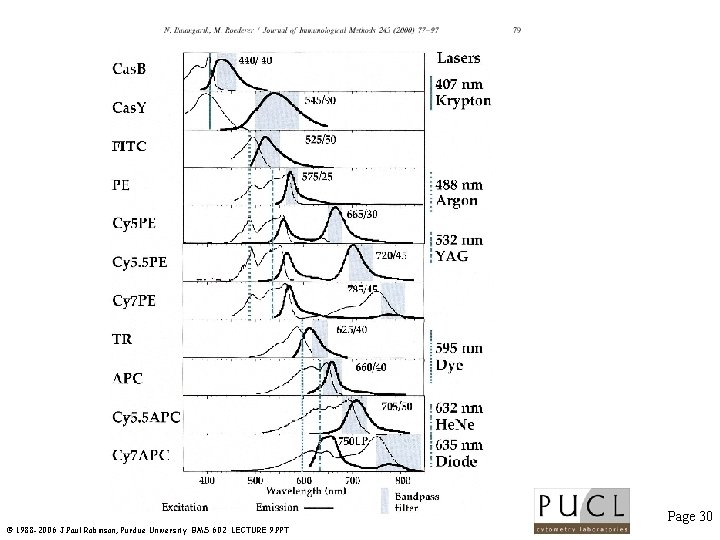
Page 30 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
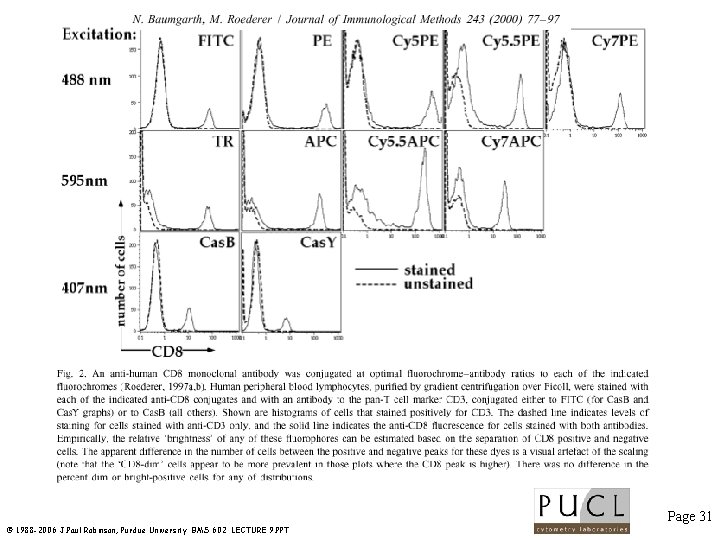
Page 31 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Innovative Data Analysis Page 32 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
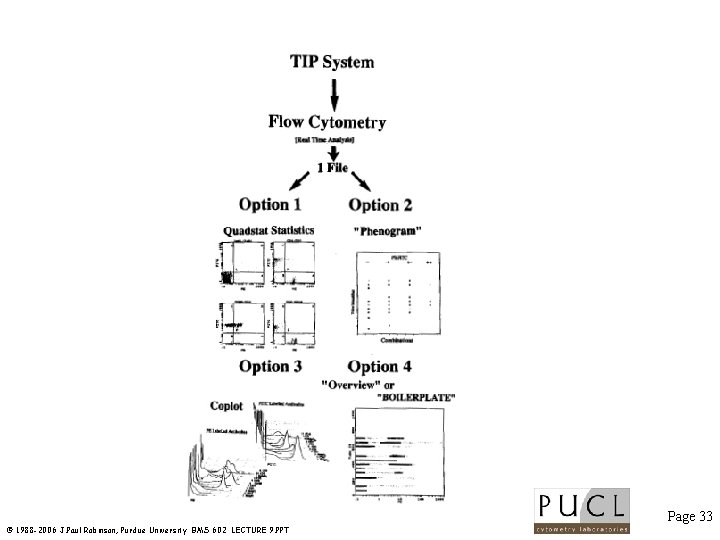
Page 33 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
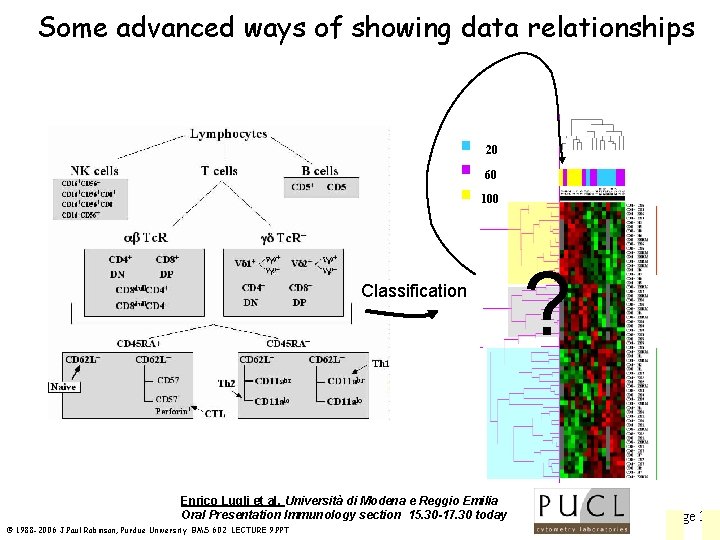
Some advanced ways of showing data relationships 20 60 100 Classification Enrico Lugli et al, Università di Modena e Reggio Emilia Oral Presentation Immunology section 15. 30 -17. 30 today © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT ? Page 34
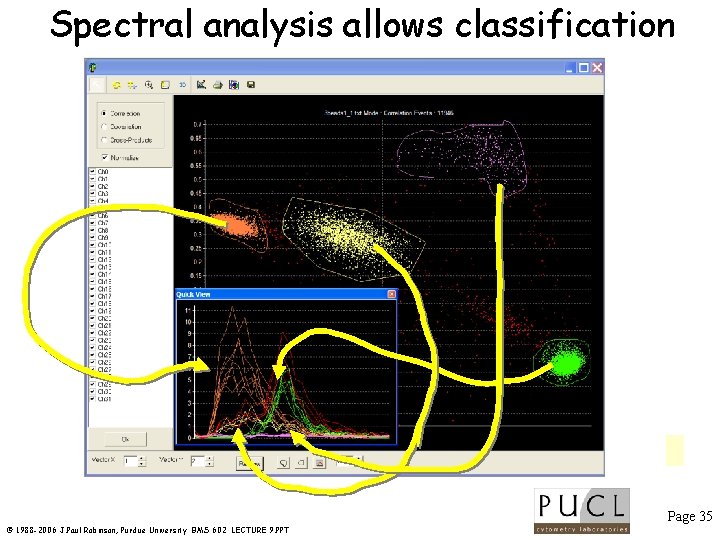
Spectral analysis allows classification Page 35 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT

Conclusions • The more parameters you have, the more complex the analysis will be • But…when you have more parameters (variables) you have more opportunities for population discrimination • Display of data in histogram and dotplot formats assists the analysis process • Displays in 3 D are nice but not particularly useful for analysis. • Multiple parameter displays such as PCA or LDA are more useful for high content data sets • Page 36 © 1988 -2006 J. Paul Robinson, Purdue University BMS 602 LECTURE 9. PPT
- Slides: 36