Blotting Blotting Your nylon filter should now have

Blotting

Blotting • Your nylon filter should now have the DNA from your gel immobilized on the surface • We now want to determine which bands from each restriction digest contain plasmid DNA • To do this, we will hybridize labeled plasmid DNA to your filters, and visualize where it finds complementary plasmid DNA

Blotting: Making Labeled DNA 5’ linear, denatured plasmid DNA random hexanucleotides DNA polymerase (Klenow) d. ATP, d. CTP, d. GTP, d. TTP digoxigenin-11 -d. UTP (DIG-d. UTP) 3’
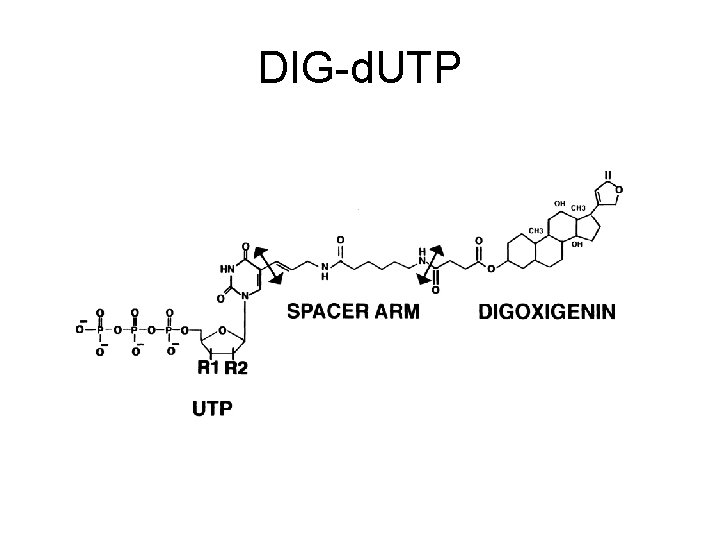
DIG-d. UTP
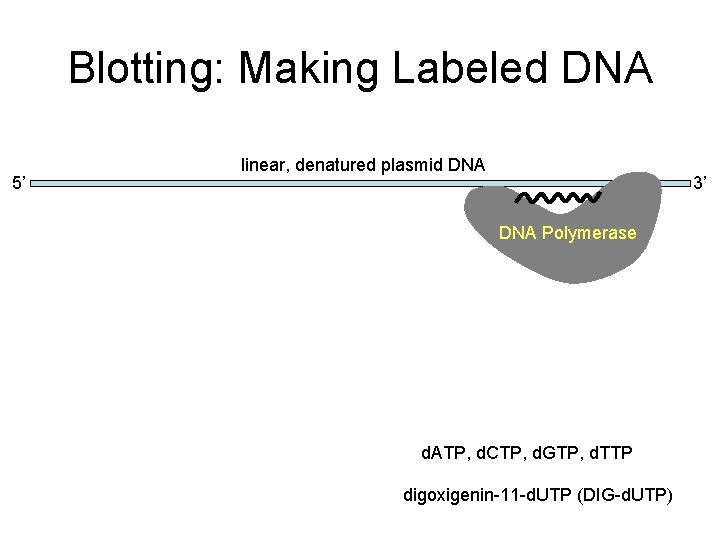
Blotting: Making Labeled DNA 5’ linear, denatured plasmid DNA ATGTTCGTUTGAGUTTGAGGCTA DNA Polymerase d. ATP, d. CTP, d. GTP, d. TTP digoxigenin-11 -d. UTP (DIG-d. UTP) 3’
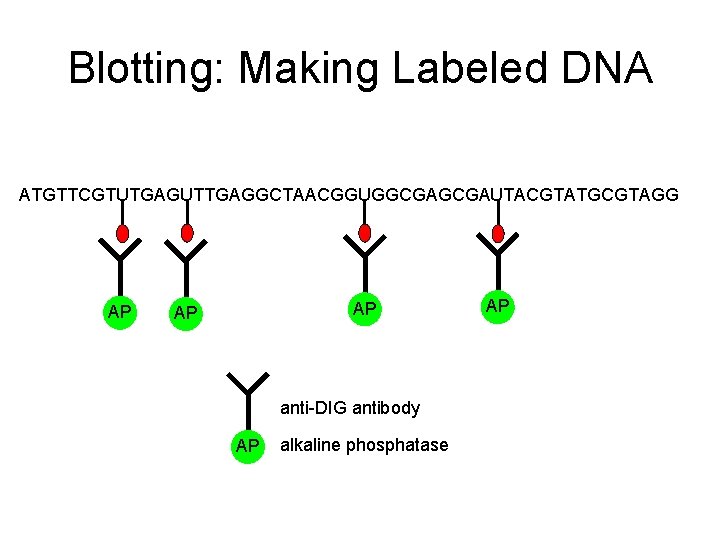
Blotting: Making Labeled DNA ATGTTCGTUTGAGUTTGAGGCTAACGGUGGCGAUTACGTATGCGTAGG AP AP AP anti-DIG antibody AP alkaline phosphatase AP


Blotting: Alkaline Phosphate Detection AP X-phosphate (5 -bromo-4 -chloro-3 -indolyl phosphate) clear and soluble enzyme independent 5, 5’-dibromo-4, 4’-dichloro-indigo blue and insolube NBT reduced NBT clear and soluble purple and insoluble
Blotting

Blotting: What’s Already Happened • The filters were rinsed briefly in 5 X SSC and then incubated in about 20 ml of hybridization solution (5 X SSC, 0. 5% blocking reagent, 0. 1% N-laurylsarcosine (sodium salt), 0. 02% SDS) for 1 -2 hours at 68°C. • The hybridization solution was replaced with 5 ml hybridization solution containing 5 ng of DIG-labeled denatured plasmid DNA and the incubation was continued at 68°C for 6 -24 hours. • Filters were washed, shaking at room temperature, in a large pyrex dish or roller bottles with 2 X SSC, 0. 1% SDS. This was repeated 5 times with fresh 2 X SSC. • Filters were washed at high stringency for 15 minutes at 68°C with 0. 1 X SSC, 0. 1% SDS. This was repeated once.

Blotting: Notes on Protocol • DIG/AP labeling replaces radioactivity • There a lot of washing steps in this protocol. These are important to avoid non -specific background • What is blocking agent for? • The final color reaction is light-sensitive, so do the reaction in a drawer

Plasmid Miniprep

Miniprep – Qiagen Kit • Step 1: Alkaline lysis of cells • Step 2: Bind plasmid DNA to silica membrane • Step 3: Elute plasmid

Miniprep: Alkaline Lysis Start by resuspending cells in a buffer containing: –glucose (to maintain osmotic balance and not burst cells prematurely) –EDTA (to destabilize membranes and inhibit DNAses)

Miniprep: Alkaline Lysis Next add Na. OH and SDS: –SDS (a detergent) disrupts the cell membrane and creates holes in it –Na. OH loosens the cell wall and further breaks down integrity of cell –Na. OH denatures DNA - note that, while chromosomal DNA tends to have breaks in it and separates into single strands, while the two strands of plasmid DNA remain together and can renature readily

Miniprep: Alkaline Lysis Next add potassium acetate. This: –neutralizes the p. H and allows circular DNA to renature –precipitates single-stranded DNA which is insolube in high salt –precipitates the detergent as KDS Spinning the solution removes cell debris and the insoluble ss. DNA and KDS

Miniprep: Alkaline Lysis The solution now contains circular plasmid DNA and soluble small molecules, and some proteins The plasmid DNA can be recovered by: –precipitation in salt (which neutralizes the DNA) and ethanol/isopropanol –binding to a positively charge substrate (such as the membrane in the Qiagen kit) in low salt (where the DNA is negatively charged) and eluting in high salt
- Slides: 17