Bloodbased Biomarkers for Lung Cancer Tony Mok MD

Blood-based Biomarkers for Lung Cancer Tony Mok MD Li Shu Fan Medical Foundation Professor of Clinical Oncology The Chinese University of Hong Kong

How solid is liquid biopsy? ? =
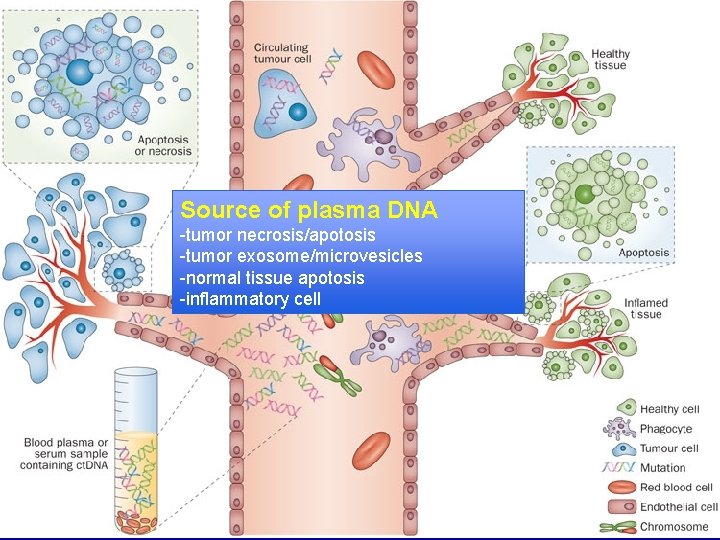
Source of plasma DNA -tumor necrosis/apotosis -tumor exosome/microvesicles -normal tissue apotosis -inflammatory cell

Technologies for tissue samples are potentially applicable to plasma cf DNA Mc. Larty et al MOJ Cell Science & Report 2015

Potential application Pathogenesis Early detection Symptoms treatment selection Diagnosis and treatment selection Resistance to treatment Monitoring Mechanism of resistance and treatment death
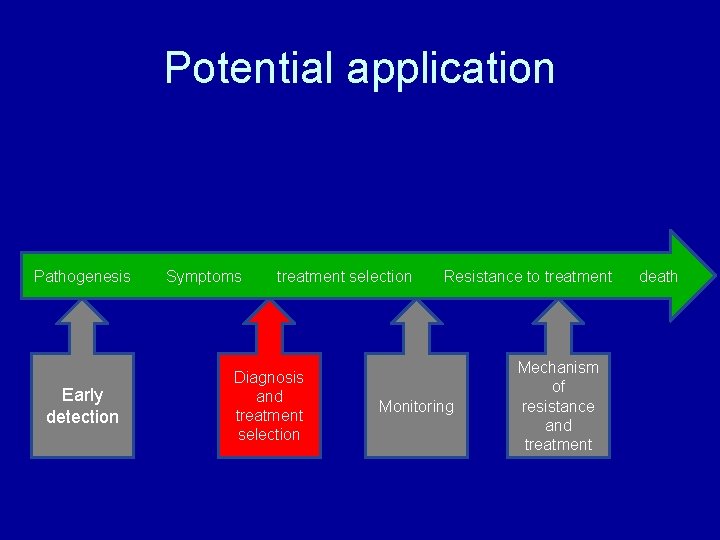
Potential application Pathogenesis Early detection Symptoms treatment selection Diagnosis and treatment selection Resistance to treatment Monitoring Mechanism of resistance and treatment death

Confirming the superior efficacy of EGFR TKI in patients with EGFR mutations Author Study N (EGFR mut +) RR Median PFS Mok et al IPASS 132 71. 2% vs 47. 3 9. 8 vs 6. 4 months Lee et al First-SIGNAL 27 84. 6% vs 37. 5% 8. 4 vs 6. 7 months Mitsudomi et al WJTOG 3405 Maemondo et al We need to find EGFR 86 62. 1% vs 32. 2% mutation at time of diagnosis of adenocarcinoma NEJGSG 002 114 73. 7% vs 30. 7% Zhou et al OPTIMAL 154 83% vs 36% 13. 1 vs 4. 6 months Rosell et al EURTAC 135 56% vs 18% 9. 2 vs 4. 8 months Wu et al LUX Lung 6 364 67% vs 23% 11. 0 vs 5. 6 months 9. 2 vs 6. 3 months 10. 8 vs 5. 4 months Mok et al NEJM 2009, Lee et al WCLC 2009, Mitsudomi et al Lancet Oncology 2010, Maemondo NEJM 2010 Zhou et al Lancet Oncol 2010
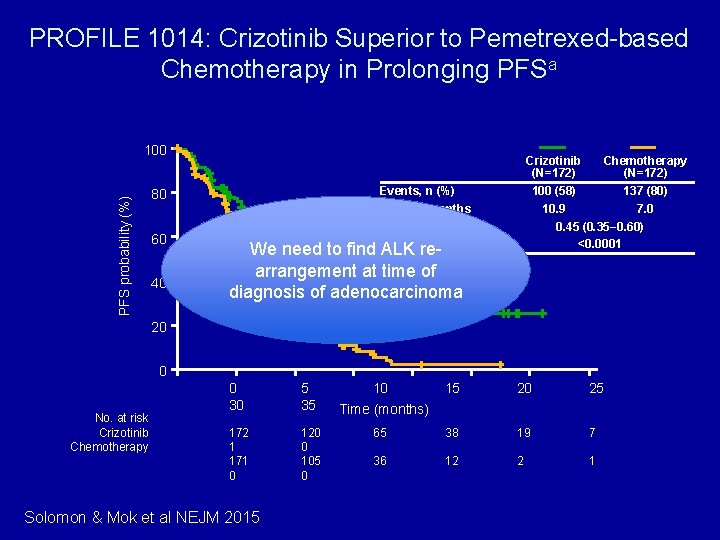
PROFILE 1014: Crizotinib Superior to Pemetrexed-based Chemotherapy in Prolonging PFSa PFS probability (%) 100 Events, n (%) Median, months HR (95% CI) Pb 80 60 40 Crizotinib Chemotherapy (N=172) 100 (58) 137 (80) 10. 9 7. 0 0. 45 (0. 35− 0. 60) <0. 0001 We need to find ALK rearrangement at time of diagnosis of adenocarcinoma 20 0 No. at risk Crizotinib Chemotherapy 0 30 5 35 172 1 171 0 120 0 105 0 Solomon & Mok et al NEJM 2015 10 Time (months) 15 20 25 65 38 19 7 36 12 2 1
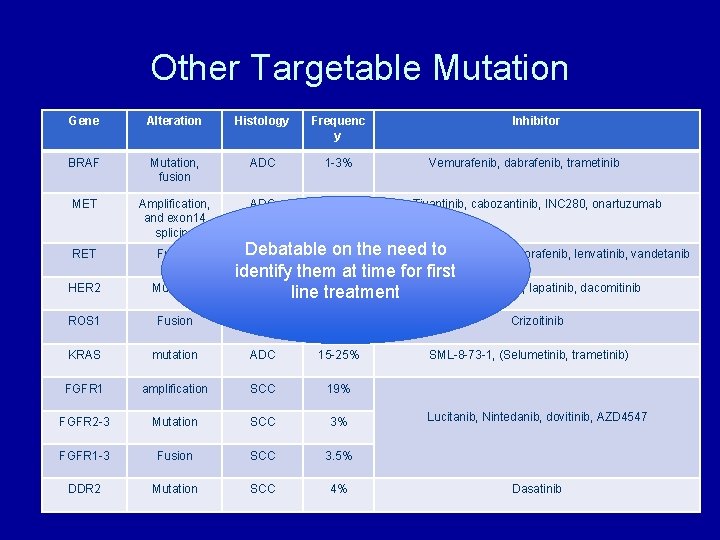
Other Targetable Mutation Gene Alteration Histology Frequenc y BRAF Mutation, fusion ADC 1 -3% MET Amplification, and exon 14 splicing ADC 2 -4% RET Fusion HER 2 Mutation ROS 1 Fusion ADC <1% KRAS mutation ADC 15 -25% FGFR 1 amplification SCC 19% FGFR 2 -3 Mutation SCC 3% FGFR 1 -3 Fusion SCC 3. 5% DDR 2 Mutation SCC 4% Inhibitor Vemurafenib, dabrafenib, trametinib Tivantinib, cabozantinib, INC 280, onartuzumab Debatable on the Carbozantinib need to , sunitinib, sorafenib, lenvatinib, vandetanib ADC 1% identify them at time for first ADC 2 -4% Neratinib, afatinib, lapatinib, dacomitinib line treatment Crizoitinib SML-8 -73 -1, (Selumetinib, trametinib) Lucitanib, Nintedanib, dovitinib, AZD 4547 Dasatinib
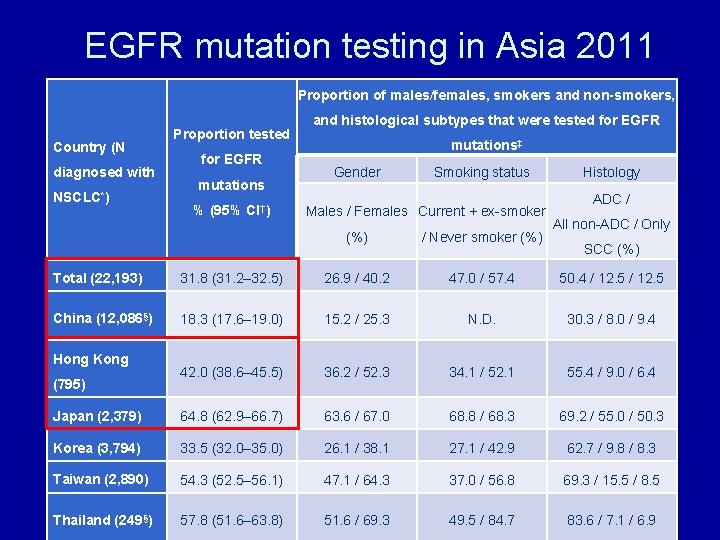
EGFR mutation testing in Asia 2011 Proportion of males/females, smokers and non-smokers, Country (N diagnosed with NSCLC*) Proportion tested for EGFR mutations % (95% CI†) and histological subtypes that were tested for EGFR mutations‡ Gender Smoking status Males / Females Current + ex-smoker (%) / Never smoker (%) Histology ADC / All non-ADC / Only SCC (%) Total (22, 193) 31. 8 (31. 2– 32. 5) 26. 9 / 40. 2 47. 0 / 57. 4 50. 4 / 12. 5 China (12, 086§) 18. 3 (17. 6– 19. 0) 15. 2 / 25. 3 N. D. 30. 3 / 8. 0 / 9. 4 42. 0 (38. 6– 45. 5) 36. 2 / 52. 3 34. 1 / 52. 1 55. 4 / 9. 0 / 6. 4 Japan (2, 379) 64. 8 (62. 9– 66. 7) 63. 6 / 67. 0 68. 8 / 68. 3 69. 2 / 55. 0 / 50. 3 Korea (3, 794) 33. 5 (32. 0– 35. 0) 26. 1 / 38. 1 27. 1 / 42. 9 62. 7 / 9. 8 / 8. 3 Taiwan (2, 890) 54. 3 (52. 5– 56. 1) 47. 1 / 64. 3 37. 0 / 56. 8 69. 3 / 15. 5 / 8. 5 Thailand (249§) 57. 8 (51. 6– 63. 8) 51. 6 / 69. 3 49. 5 / 84. 7 83. 6 / 7. 1 / 6. 9 Hong Kong (795)

cobas® EGFR _ blood Test _ Kit Components Utilizing most of the reagents in the cobas EGFR_FFPET test and requiring additional reagents and the blood-specific data analysis software Blood ct. DNA Preparation Kit Cobas EGFR_Blood ct. DNA SP 2 ml Plasma cobas cell-free DNA Preparation Kit (To be used for other blood based assays) HPEA x 25 PK x 2 PBB x 6 Additional reagents added to cobas DNA preparation Kit cobas EGFR _ blood Test cobas 4800 v 2. 0 HEX cobas EGFR _ Blood Tube FAM 1 EX 19 Del S 768 I 2 L 858 R T 790 M 3 G 719 X L 861 Q* JA 270 Blood-specific cutoffs; Blood-specific data analysis software EX 20 Ins *New primer and probe for L 861 Q Mok et al ASCO 2013
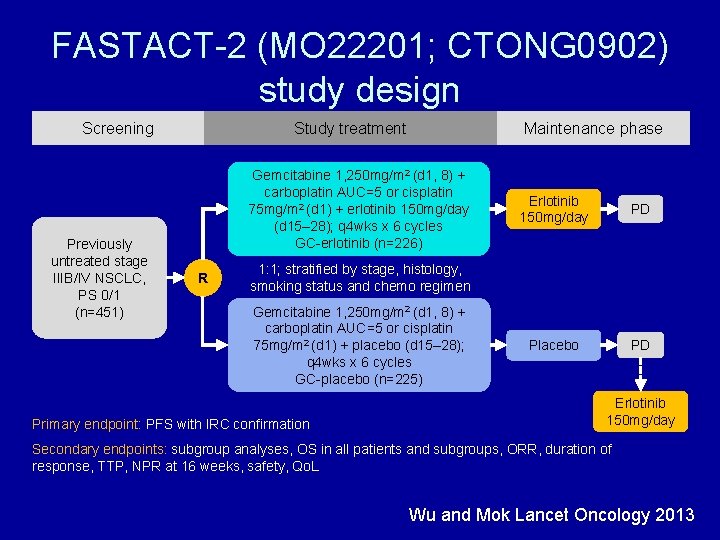
FASTACT-2 (MO 22201; CTONG 0902) study design Screening Previously untreated stage IIIB/IV NSCLC, PS 0/1 (n=451) Study treatment Maintenance phase Gemcitabine 1, 250 mg/m 2 (d 1, 8) + carboplatin AUC=5 or cisplatin 75 mg/m 2 (d 1) + erlotinib 150 mg/day (d 15– 28); q 4 wks x 6 cycles GC-erlotinib (n=226) R Erlotinib 150 mg/day PD Placebo PD 1: 1; stratified by stage, histology, smoking status and chemo regimen Gemcitabine 1, 250 mg/m 2 (d 1, 8) + carboplatin AUC=5 or cisplatin 75 mg/m 2 (d 1) + placebo (d 15– 28); q 4 wks x 6 cycles GC-placebo (n=225) Primary endpoint: PFS with IRC confirmation Erlotinib 150 mg/day Secondary endpoints: subgroup analyses, OS in all patients and subgroups, ORR, duration of response, TTP, NPR at 16 weeks, safety, Qo. L Wu and Mok Lancet Oncology 2013

EGFR Mutation Analysis using cobas 4800_blood test TISSUE SAMPLES 397 (88%) patients consented 301 (66. 7%) samples available 241 (53. 4%) samples analyzable 224 patients with matched tumor and plasma samples PLASMA SAMPLES 451 (100%) patients consented 427 (94. 6%) samples available 427 (94. 6%) samples analyzable

Concordance between tumor and plasma samples • Total of 224 patients had both tumor and baseline plasma samples with available EGFR mutation analysis results (Table 3) – Sensitivity: 77% (69/90) – Specificity: 96% (129/134) – Positive predictive value: 93% (69/74) – Negative predictive value: 86% (129/150) – Overall concordance: 88% (198/224) EGFR Activating Mutations p-EGFR Mut+ (Plasma) p-EGFR Mut(Plasma) Total t-EGFR Mut+ (Tumor) 69 21 90 t-EGFR Mut- (Tumor) 5 129 134 Total 74 150 224
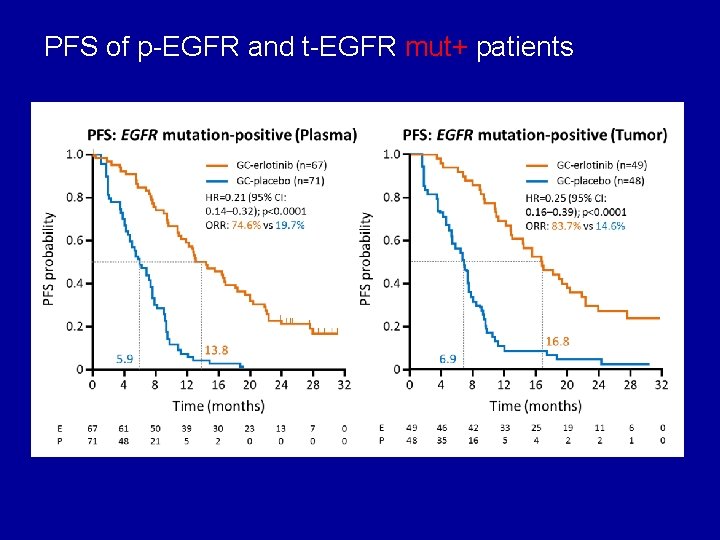
PFS of p-EGFR and t-EGFR mut+ patients
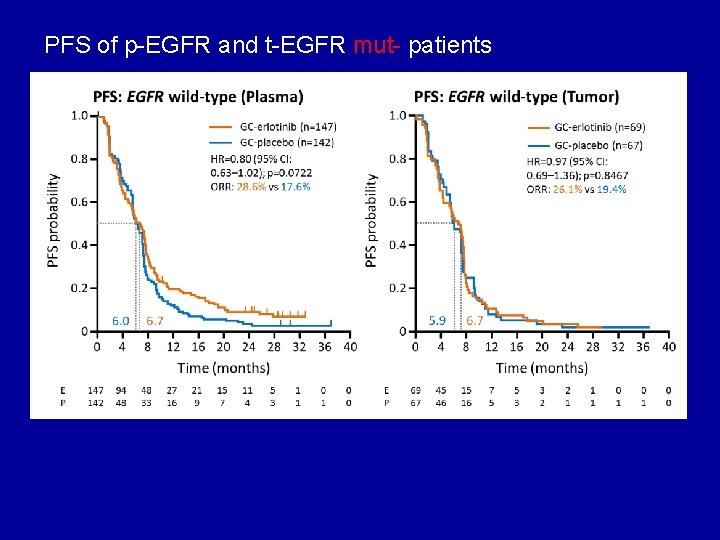
PFS of p-EGFR and t-EGFR mut- patients
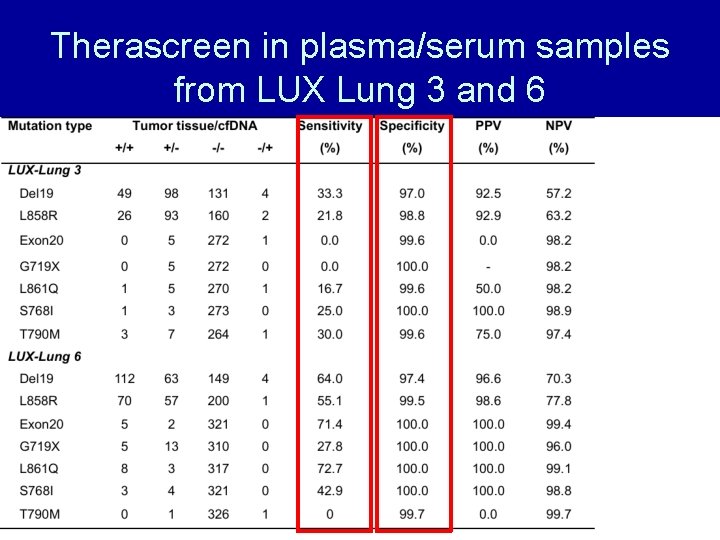
Therascreen in plasma/serum samples from LUX Lung 3 and 6
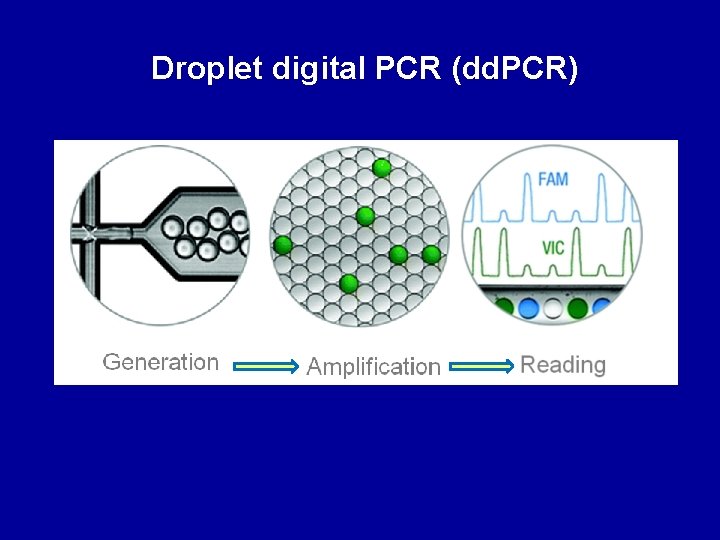
Droplet digital PCR (dd. PCR) Hindson et al. Analytical chemistry 2011
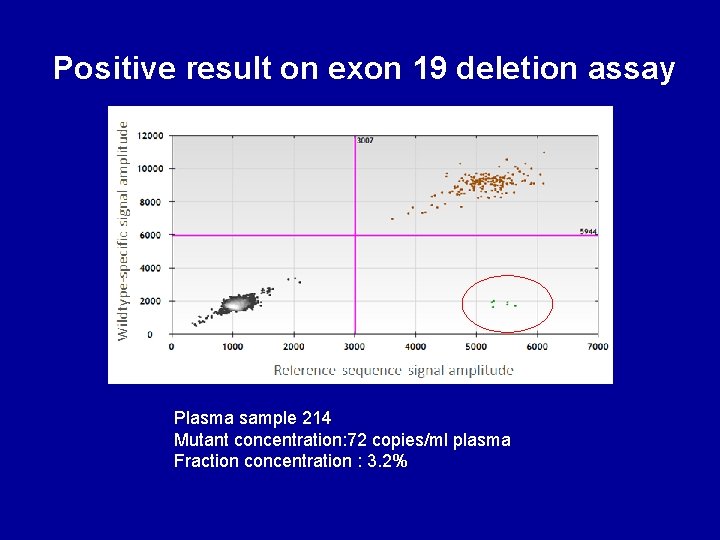
Positive result on exon 19 deletion assay Plasma sample 214 Mutant concentration: 72 copies/ml plasma Fraction concentration : 3. 2%
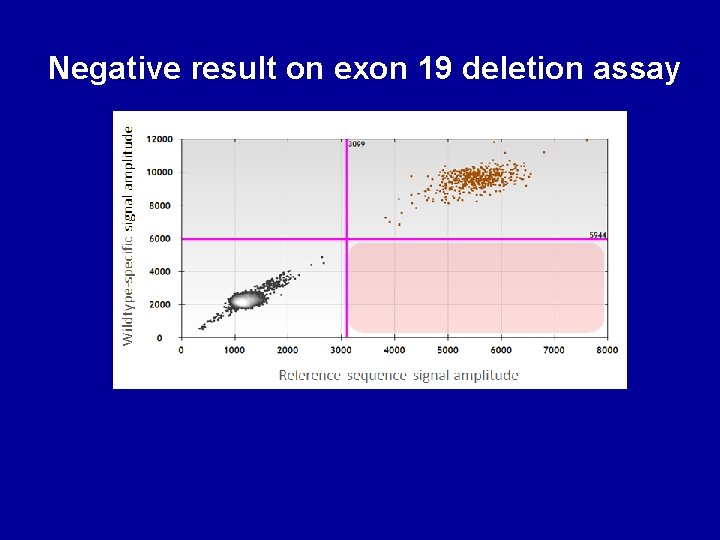
Negative result on exon 19 deletion assay

Analyzing plasma and tumor sample from ASPIRATION study and matched control (n=197) • Tumor sample: COBAS EGFR Mutation Test • Plasma sample: Droplet digital PCR Lee et al WCLC 2013
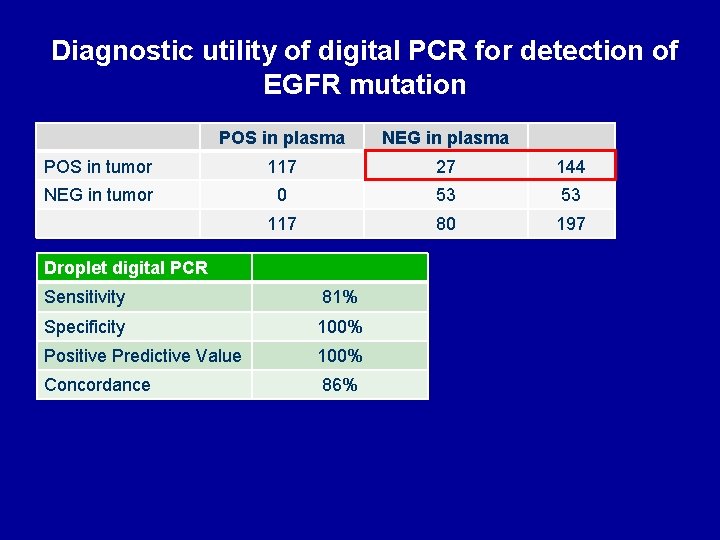
Diagnostic utility of digital PCR for detection of EGFR mutation POS in plasma NEG in plasma POS in tumor 117 27 144 NEG in tumor 0 53 53 117 80 197 Droplet digital PCR Sensitivity 81% Specificity 100% Positive Predictive Value 100% Concordance 86%

Meta-analysis on cf DNA for EGFR mutation • 3110 subjects (27 studies) • Tissue EGFR mutation status as gold standard Sensitivity is • Pooled Analysis technology dependent, eg BEAM vs Sanger – Sensitivity 62% – Specificity 96% – Diagnostic odd ratio – Area under summary ROC 0. 91 High specificity implies clinical applicability for selection of first line EGFR TKI Qiu et al Can Epi Biomarker Prev 2015 Jan; 24(1): 206 -12.
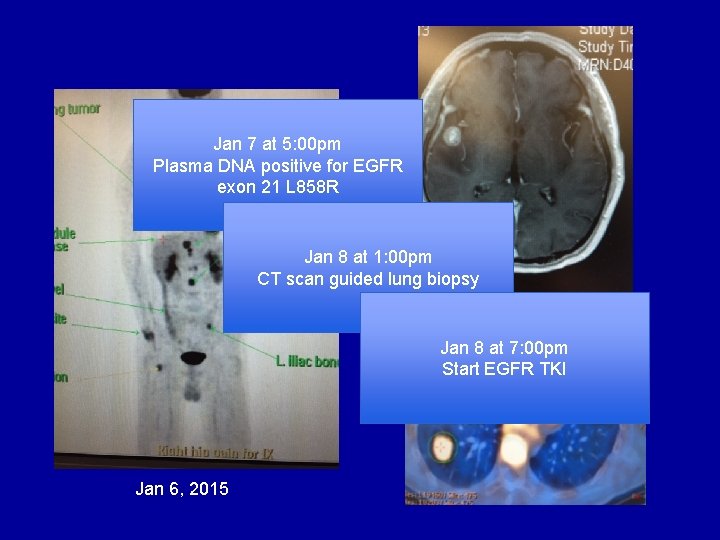
Jan 7 at 5: 00 pm Plasma DNA positive for EGFR exon 21 L 858 R Jan 8 at 1: 00 pm CT scan guided lung biopsy Jan 8 at 7: 00 pm Start EGFR TKI Jan 6, 2015
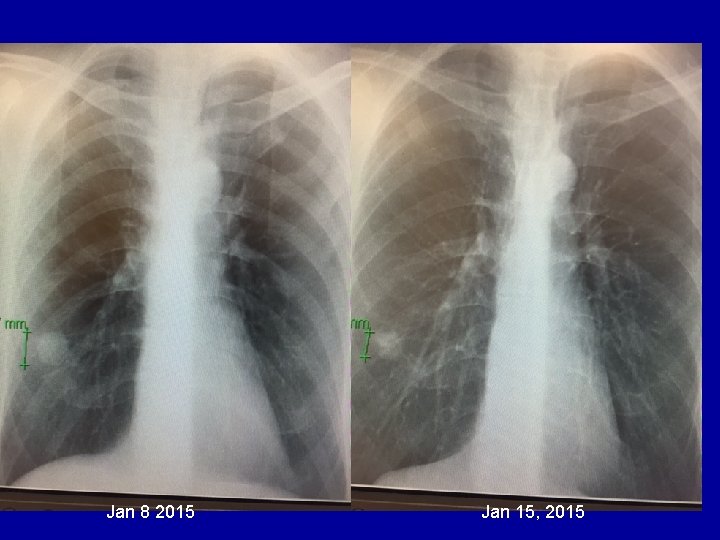
Jan 8 2015 Jan 15, 2015
Potential application Pathogenesis Early detection Symptoms treatment selection Diagnosis and treatment selection Resistance to treatment Monitoring Mechanism of resistance and treatment death

Serial plasma samples at baseline, C 3 and PD BASELINE TISSUE SAMPLES 397 (88%) patients consented 268 (59. 4%) samples available 241 (53. 4%) samples analysable 447 (99. 1%) baseline samples available 447 (99. 1%) baseline samples analysable 238 (52. 8%) patients with matched tumour and plasma results PLASMA SAMPLES 451 (100%) patients consented 362 (80. 3%) C 3 samples available 362 (80. 3%) C 3 samples analysable 376 (83. 4%) PD samples available 376 (83. 4%) PD samples analysable 305 (67. 6%) Patients with plasma results at all three time points
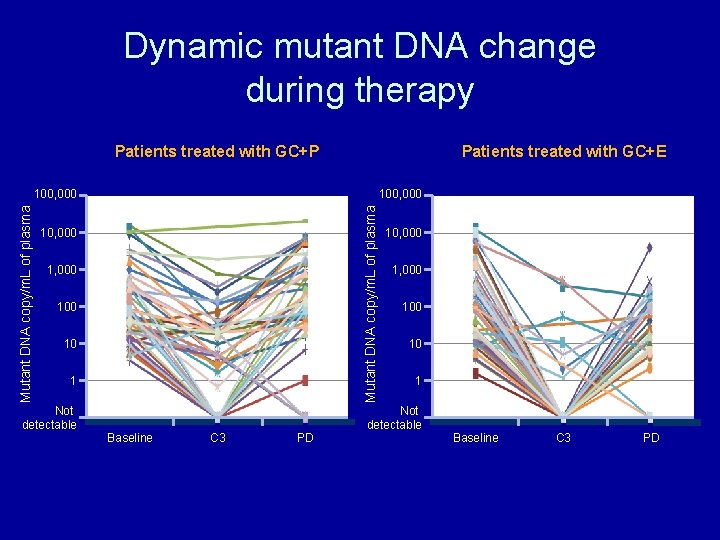
Dynamic mutant DNA change during therapy Patients treated with GC+P Patients treated with GC+E 100, 000 Mutant DNA copy/m. L of plasma 100, 000 1, 000 10 1 Not detectable Baseline C 3 PD 10, 000 100 10 1 Not detectable Baseline C 3 PD
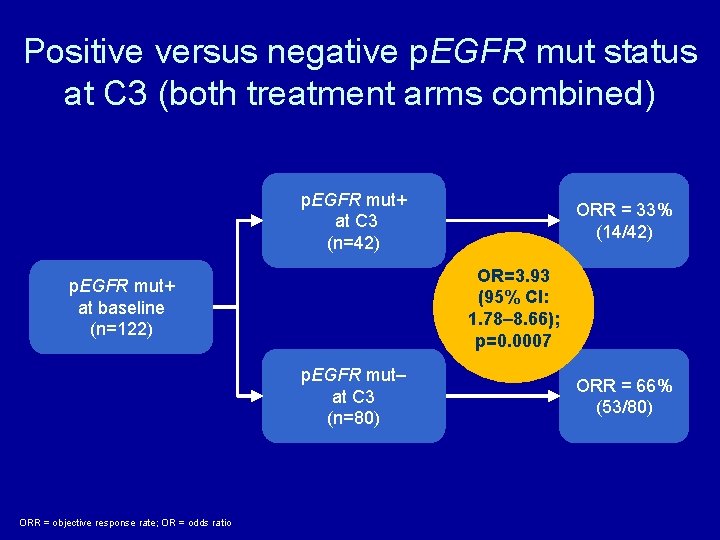
Positive versus negative p. EGFR mut status at C 3 (both treatment arms combined) p. EGFR mut+ at C 3 (n=42) OR=3. 93 (95% CI: 1. 78– 8. 66); p=0. 0007 p. EGFR mut+ at baseline (n=122) p. EGFR mut– at C 3 (n=80) ORR = objective response rate; OR = odds ratio ORR = 33% (14/42) ORR = 66% (53/80)
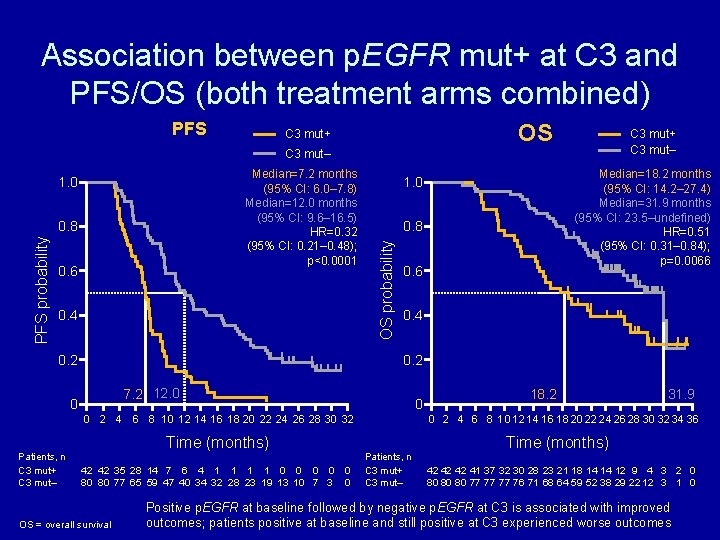
Association between p. EGFR mut+ at C 3 and PFS/OS (both treatment arms combined) PFS OS C 3 mut+ C 3 mut– PFS probability 0. 8 0. 6 0. 4 0. 2 7. 2 12. 0 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 32 Patients, n C 3 mut+ C 3 mut– Median=18. 2 months (95% CI: 14. 2– 27. 4) Median=31. 9 months (95% CI: 23. 5–undefined) HR=0. 51 (95% CI: 0. 31– 0. 84); p=0. 0066 1. 0 OS probability Median=7. 2 months (95% CI: 6. 0– 7. 8) Median=12. 0 months (95% CI: 9. 6– 16. 5) HR=0. 32 (95% CI: 0. 21– 0. 48); p<0. 0001 1. 0 Time (months) 42 42 35 28 14 7 6 4 1 1 0 0 80 80 77 65 59 47 40 34 32 28 23 19 13 10 7 3 OS = overall survival 0 0 18. 2 31. 9 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 32 34 36 Patients, n C 3 mut+ C 3 mut– Time (months) 42 42 42 41 37 32 30 28 23 21 18 14 14 12 9 4 3 2 0 80 80 80 77 77 77 76 71 68 64 59 52 38 29 22 12 3 1 0 Positive p. EGFR at baseline followed by negative p. EGFR at C 3 is associated with improved outcomes; patients positive at baseline and still positive at C 3 experienced worse outcomes
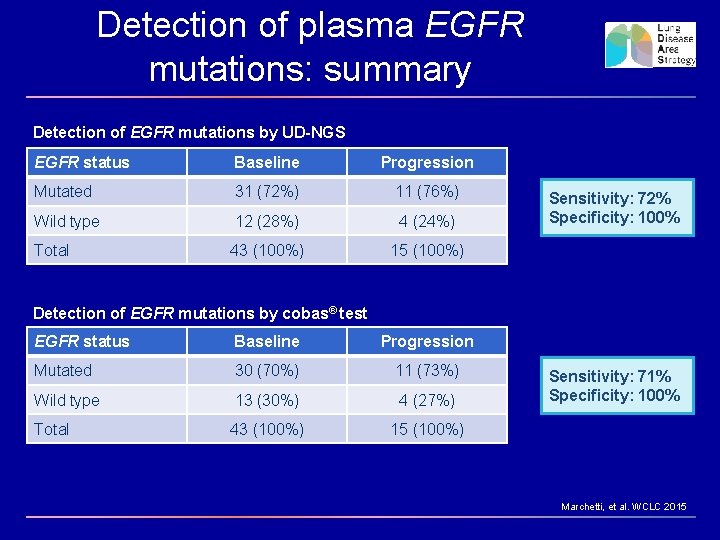
Detection of plasma EGFR mutations: summary Detection of EGFR mutations by UD-NGS EGFR status Baseline Progression Mutated 31 (72%) 11 (76%) Wild type 12 (28%) 4 (24%) Total 43 (100%) 15 (100%) Sensitivity: 72% Specificity: 100% Detection of EGFR mutations by cobas® test EGFR status Baseline Progression Mutated 30 (70%) 11 (73%) Wild type 13 (30%) 4 (27%) Total 43 (100%) 15 (100%) Sensitivity: 71% Specificity: 100% Marchetti, et al. WCLC 2015
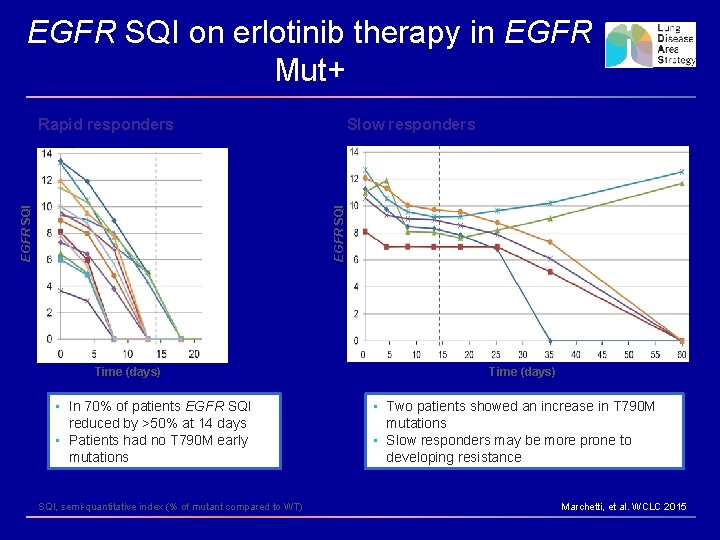
EGFR SQI on erlotinib therapy in EGFR Mut+ Rapid responders EGFR SQI Slow responders Time (days) • In 70% of patients EGFR SQI reduced by >50% at 14 days • Patients had no T 790 M early mutations SQI, semi-quantitative index (% of mutant compared to WT) Time (days) • Two patients showed an increase in T 790 M mutations • Slow responders may be more prone to developing resistance Marchetti, et al. WCLC 2015
Potential application Pathogenesis Early detection Symptoms treatment selection Diagnosis and treatment selection Resistance to treatment Monitoring Mechanism of resistance and treatment death
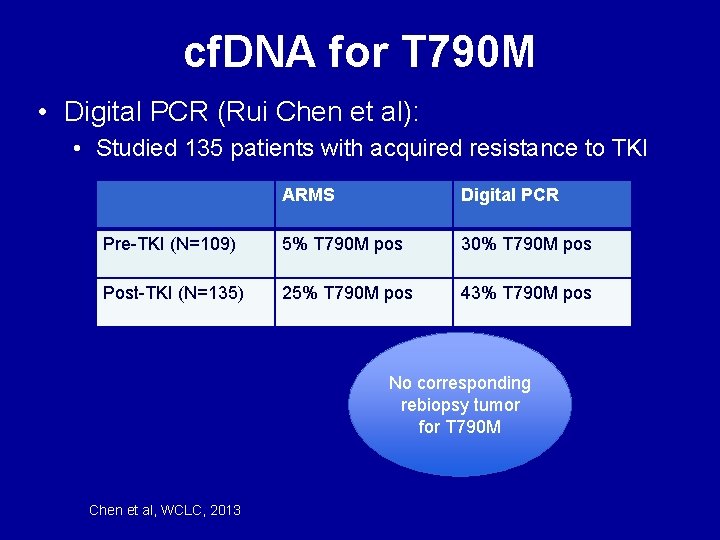
cf. DNA for T 790 M • Digital PCR (Rui Chen et al): • Studied 135 patients with acquired resistance to TKI ARMS Digital PCR Pre-TKI (N=109) 5% T 790 M pos 30% T 790 M pos Post-TKI (N=135) 25% T 790 M pos 43% T 790 M pos No corresponding rebiopsy tumor for T 790 M Chen et al, WCLC, 2013

Serial monitoring of T 790 M from plasma DNA 47% found to Droplet have. PCR Digital T 790 M Zhang et al ASCO 2014
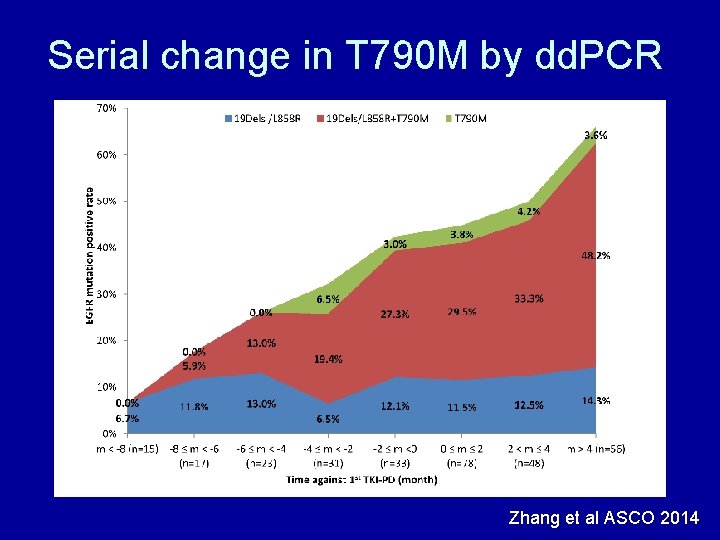
Serial change in T 790 M by dd. PCR Zhang et al ASCO 2014
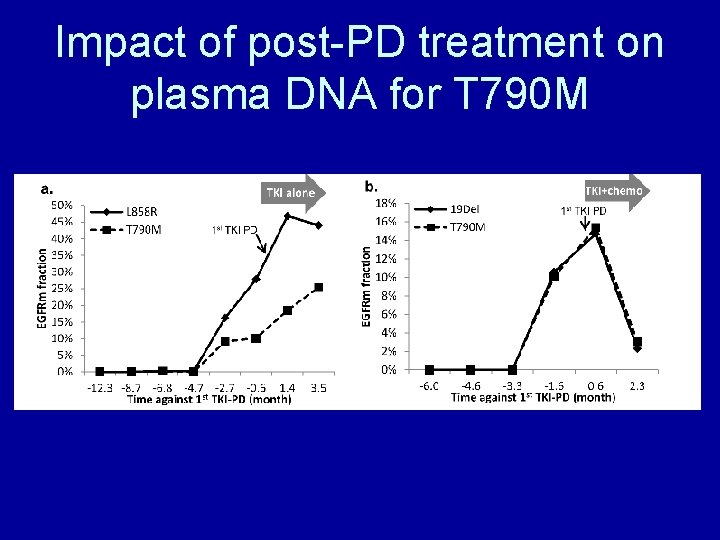
Impact of post-PD treatment on plasma DNA for T 790 M
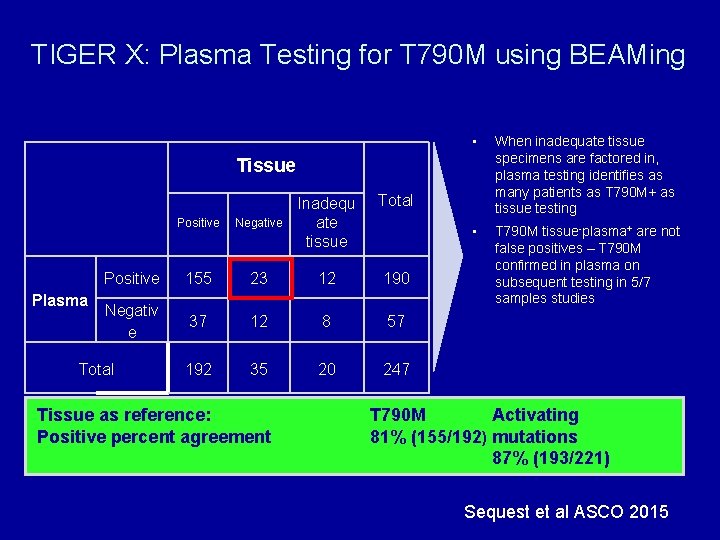
TIGER X: Plasma Testing for T 790 M using BEAMing • When inadequate tissue specimens are factored in, plasma testing identifies as many patients as T 790 M+ as tissue testing • T 790 M tissue-plasma+ are not false positives – T 790 M confirmed in plasma on subsequent testing in 5/7 samples studies Tissue Plasma Total Positive Negative Inadequ ate tissue Positive 155 23 12 190 Negativ e 37 12 8 57 192 35 20 247 Total Tissue as reference: Positive percent agreement T 790 M Activating 81% (155/192) mutations 87% (193/221) Sequest et al ASCO 2015

T 790 M Plasma Testing is a Viable Alternative to Tissue Testing Objective response rate for 188 evaluable patients with both central T 790 M tissue test result and plasma T 790 M result Plasma T 790 M + 55% + (72/130) Tissue T 790 M − 43% (13/30) 53% (85/16 0) 35% (6/17) 27% (3/11) 32% (9/28) 53% (78/147) 39% (16/41) • Similar ORR observed when detecting T 790 M in either tissue or plasma • Not all patients with progression on first-line TKI are candidates for tissue re-biopsy

Targeting EGFR mutation versus NGS
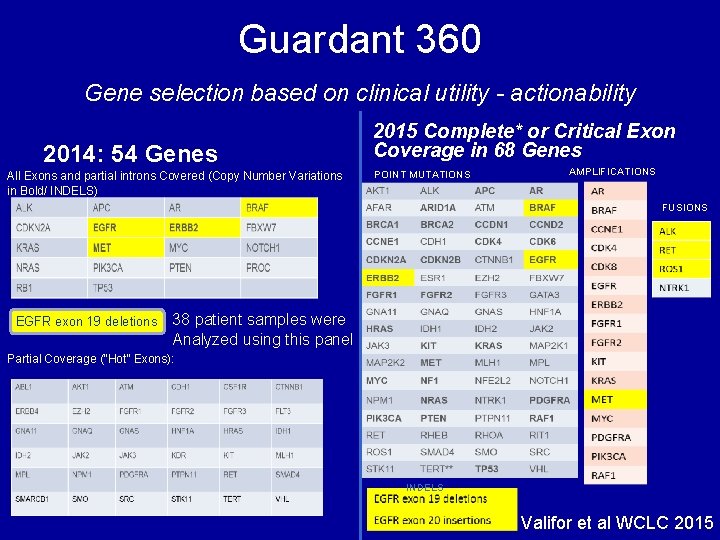
Guardant 360 Gene selection based on clinical utility - actionability 2014: 54 Genes All Exons and partial introns Covered (Copy Number Variations in Bold/ INDELS) 2015 Complete* or Critical Exon Coverage in 68 Genes POINT MUTATIONS AMPLIFICATIONS FUSIONS EGFR exon 19 deletions 38 patient samples were Analyzed using this panel Partial Coverage (“Hot” Exons): INDELS Valifor et al WCLC 2015
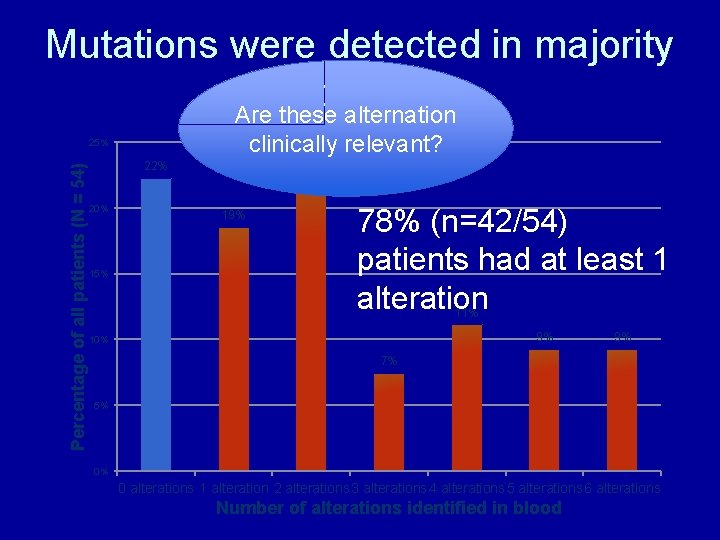
Mutations were detected in majority of patients Are these alternation clinically relevant? Percentage of all patients (N = 54) 25% 22% 20% 15% 22% 19% 78% (n=42/54) patients had at least 1 alteration 11% 9% 10% 9% 7% 5% 0% 0 alterations 1 alteration 2 alterations 3 alterations 4 alterations 5 alterations 6 alterations Number of alterations identified in blood
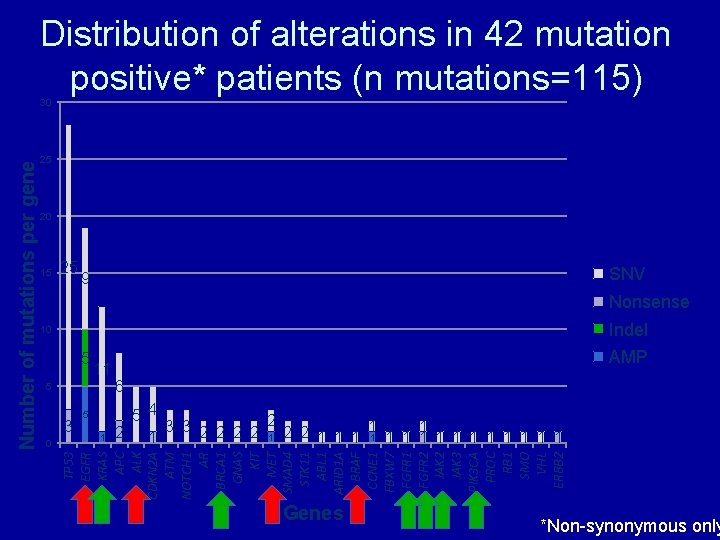
Distribution of alterations in 42 mutation positive* patients (n mutations=115) 25 20 15 25 SNV 9 Nonsense Indel 10 5 11 5 3 0 5 1 AMP 6 5 4 3 3 2 2 2 2 1 1 1 1 1 TP 53 EGFR KRAS APC ALK CDKN 2 A ATM NOTCH 1 AR BRCA 1 GNAS KIT MET SMAD 4 STK 11 ABL 1 ARID 1 A BRAF CCNE 1 FBXW 7 FGFR 1 FGFR 2 JAK 3 PIK 3 CA PROC RB 1 SMO VHL ERBB 2 Number of mutations per gene 30 Genes *Non-synonymous only
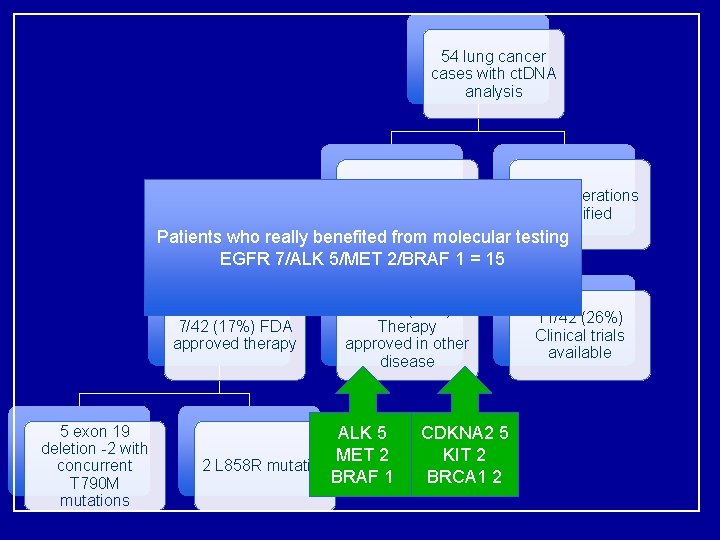
54 lung cancer cases with ct. DNA analysis 42 with 1 or more alteration 12 no alterations identified Patients who really benefited from molecular testing EGFR 7/ALK 5/MET 2/BRAF 1 = 15 7/42 (17%) FDA approved therapy 5 exon 19 deletion -2 with concurrent T 790 M mutations 17/42 (40%) Therapy approved in other disease ALK 5 MET 2 2 L 858 R mutation BRAF 1 CDKNA 2 5 KIT 2 BRCA 1 2 11/42 (26%) Clinical trials available

NGS is to eat the whole genomic pie in one go

Alternative approach is to rule out EGFR mutation first
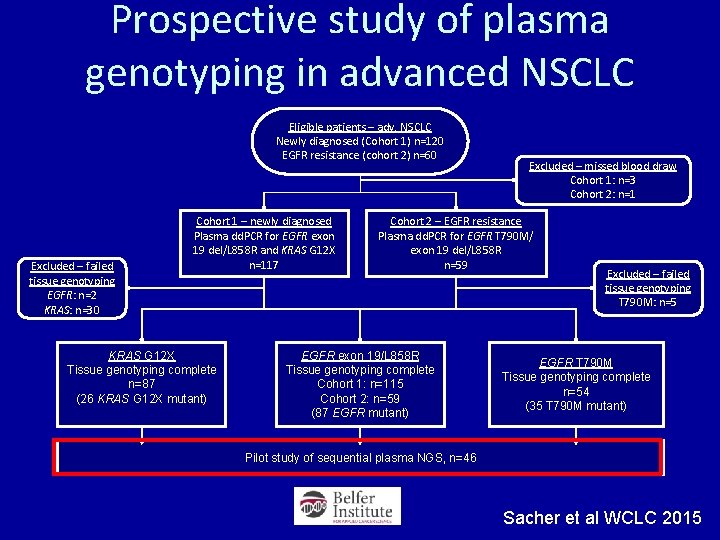
Prospective study of plasma genotyping in advanced NSCLC Eligible patients – adv. NSCLC Newly diagnosed (Cohort 1) n=120 EGFR resistance (cohort 2) n=60 Excluded – failed tissue genotyping EGFR: n=2 KRAS: n=30 Cohort 1 – newly diagnosed Plasma dd. PCR for EGFR exon 19 del/L 858 R and KRAS G 12 X n=117 KRAS G 12 X Tissue genotyping complete n=87 (26 KRAS G 12 X mutant) Excluded – missed blood draw Cohort 1: n=3 Cohort 2: n=1 Cohort 2 – EGFR resistance Plasma dd. PCR for EGFR T 790 M/ exon 19 del/L 858 R n=59 EGFR exon 19/L 858 R Tissue genotyping complete Cohort 1: n=115 Cohort 2: n=59 (87 EGFR mutant) Excluded – failed tissue genotyping T 790 M: n=5 EGFR T 790 M Tissue genotyping complete n=54 (35 T 790 M mutant) Pilot study of sequential plasma NGS, n=46 Sacher et al WCLC 2015
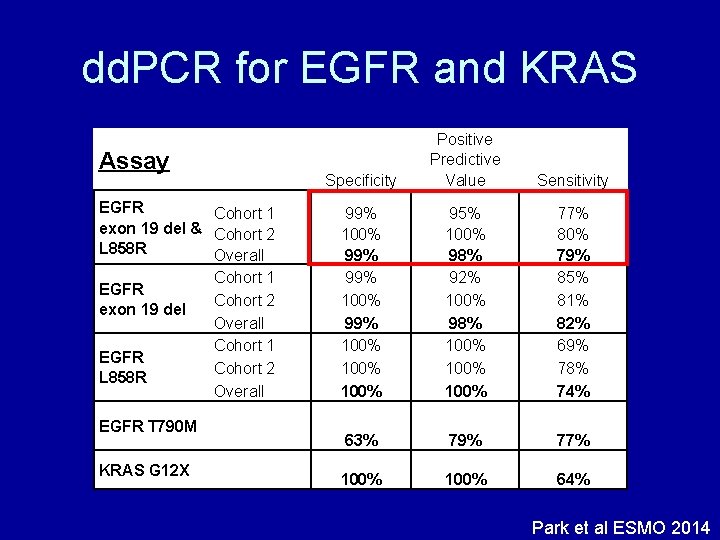
dd. PCR for EGFR and KRAS Assay EGFR Cohort 1 exon 19 del & Cohort 2 L 858 R Overall Cohort 1 EGFR Cohort 2 exon 19 del Overall Cohort 1 EGFR Cohort 2 L 858 R Overall EGFR T 790 M KRAS G 12 X Specificity Positive Predictive Value Sensitivity 99% 100% 100% 95% 100% 98% 92% 100% 98% 100% 77% 80% 79% 85% 81% 82% 69% 78% 74% 63% 79% 77% 100% 64% dd. PCR from plasma cf. DNA in ASPIRATIOM study Park et al ESMO 2014

Sequential plasma dd. PCR and NGS Complex genomic alterations EGFR/KRAS mutant Study ID Tissue Genotype Plasma dd. PCR Plasma NGS 1 KRAS G 12 C Detected 4 EGFR exon 19 del Detected 18 ALK Negative Detected 11 EGFR exon 19 del Detected 22 EGFR G 719 A Negative Detected 17 EGFR exon 19 del Detected 36 EGFR G 719 A Negative Detected 39 EGFR exon 19 del Detected 81 PIK 3 CA E 545 K Negative Detected 44 EGFR exon 19 del Detected 107 RET, p 53 Negative Detected 53 EGFR exon 19 del Detected 127 ROS 1 Negative Detected 70 KRAS G 12 C Detected 130 EGFR exon 20 ins, p 53 Negative Detected 74 EGFR exon 19 del Detected 133 KRAS Q 61 H Negative Detected 81 KRAS G 12 C Detected 137 RET Negative Detected 91 EGFR exon 19 del Detected 95 EGFR exon 19 del Detected 145 EGFR exon 20 ins Negative Detected 105 EGFR exon 19 del Detected 202 ALK Negative Detected 116 EGFR exon 19 Detected 250 EGFR exon 19 del/ins Negative Detected 120 EGFR exon 19/T 790 M Detected 258 None Negative None 179 KRAS G 12 C Detected 89 KRAS G 13 D Negative 195 EGFR exon 19 Detected 108 BRAF V 600 E Negative 200 KRAS G 12 C Detected 115 KRAS Q 61 L Negative 232 EGFR L 858 R/T 790 M Detected 169 RET Negative 48 EGFR L 858 R Negative Detected 209 ALK Negative 244 EGFR exon 19 Negative Detected 8 KRAS G 12 C Detected Negative Plasma NGS detected a diversity of complex genomic 28 EGFR L 858 R Detected Negative Plasma NGS detected alterations in patients with negative plasma dd. PCR (n=19) 13/19 (74%) 61 KRAS G 12 C Detected Negative complex with 74% sensitivity. This assay was ablegenomic to detect alternation and 45 8/13 EGFR exon 19 del Negative 94 KRAS G 12 V Negative patients had FDA approved treatment EGFR/KRAS mutations confirmed by orthogonal plasma 109 EGFR exon 19 del Negative Study ID 15 Tissue Genotype ROS 1 Plasma dd. PCR Plasma NGS Negative Detected dd. PCR with 78% sensitivity (n=27).

This approach is to take a fast big bite of the genomic pie with dd. PCR
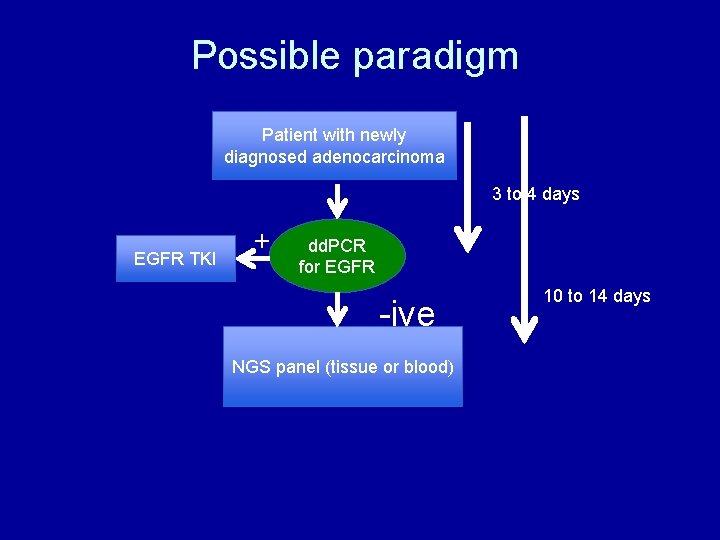
Possible paradigm Patient with newly diagnosed adenocarcinoma 3 to 4 days EGFR TKI + dd. PCR for EGFR -ive NGS panel (tissue or blood) 10 to 14 days

Summary • Finding EGFR mutation from cf. DNA – Finding EGFR mutation from cf. DNA is feasible – Sensitivity 70 to 80%, highly specific – dd. PCR or BEAM • Monitoring progress – Potentially could be used to monitor the EGFR mutation status, but clinical relevance is lacking • Detecting T 790 M – Digital PCR is relatively sensitive – BEAMing for T 790 M is associated with clinical response • NGS – Multiple technology platform is being developed – Sensible to perform NGS only in patients without EGFR mutation in plasma

A liquid can be very solid Liquid Crystal
- Slides: 53