BLAT The BLASTLike Alignment Tool Kent W J

BLAT – The BLAST-Like Alignment Tool Kent, W. J. Genome Res. 2002 12: 656 -664 Presenter: 巨彥霖 田知本

BLAT overview • Use an index to find regions in genome homologous to query. • Do a detailed alignment between query and homologous regions. • Use dynamic programming to stitch together detailed alignments regions into detailed alignment of whole.
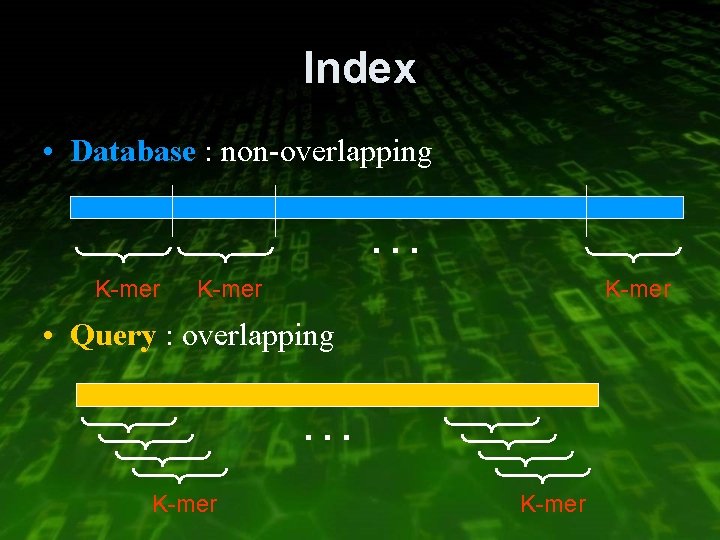
Index • Database : non-overlapping … K-mer • Query : overlapping … K-mer

Example • Database: cacaattatcacgaccgc 3 -mers: cac aat tat cac gac cgc Index: aat 3 gac 12 cac 0, 9 tat 6 cgc 15 • Query: aattctcac 3 -mers: aat att ttc tct ctc tca cac 0 1 2 3 4 5 6

Search Criteria • Single Perfect Matches • Single Near Perfect Matches • Multiple Perfect Matches

Notation • K : K-mer size • M : The match ratio between homologous area • H : Homologous region size • G : Query sequence size • A : The alphabet size

Single Perfect Matches (1) K-mer Homologous Perfect Match region
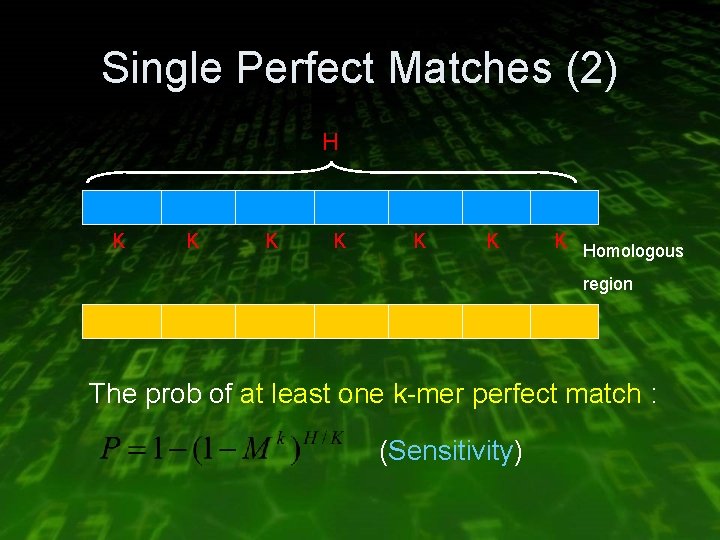
Single Perfect Matches (2) H K K K K Homologous region The prob of at least one k-mer perfect match : (Sensitivity)

Single Perfect Matches (3) • The number of k-mer in the database = G / K • The number of k-mer in the query = Q – K + 1 The number of k-mer that are expected to matched by chance : (Specificity)
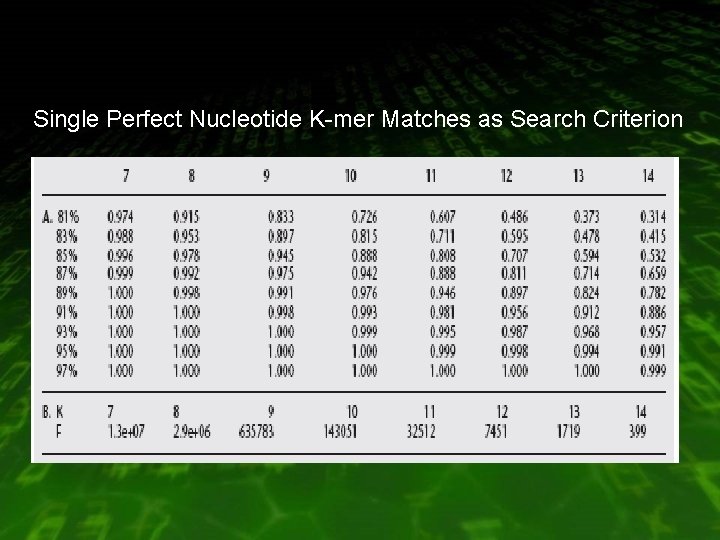
Single Perfect Nucleotide K-mer Matches as Search Criterion

Case (perfect match) • Comparing mouse and human coding sequences at the nucleotide level : H = 100 M = 86% Sensitivity = 0. 99 max K = 7 chance matches = 13078962 (query = 500 , database = 3 billion)

Single Near Perfect Matches (1) Almost Perfect : One letter may mismatch K-mer Homologous Near Perfect Match region

Single Near Perfect Matches (2) • Sensitivity • Specificity

Case (near perfect match) • Comparing mouse and human coding sequences at the nucleotide level : H = 100 M = 86% Sensitivity = 0. 99 max K = 12 chance matches = 275671 (query = 500 , database = 3 billion)
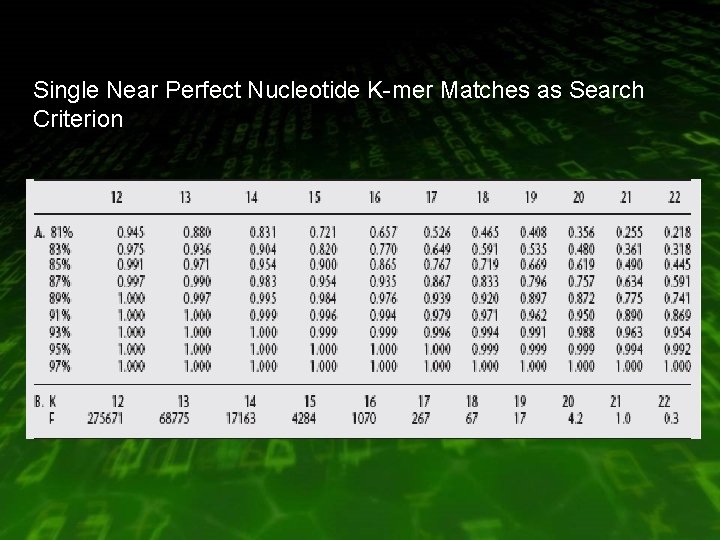
Single Near Perfect Nucleotide K-mer Matches as Search Criterion

Multiple Perfect Matches • Hit is triggered : – there must be N perfect matches – each no further than W letters from each other in the database coordinate – have the same diagonal coordinate

Example Query Coordinate a W b c d Target Coordinate The hits a, b, c, and d are all k letters long. Hits b and d have the same diagonal coordinate within W letters of each other. Therefore, they would match the 2 perfect K-mer search criteria.
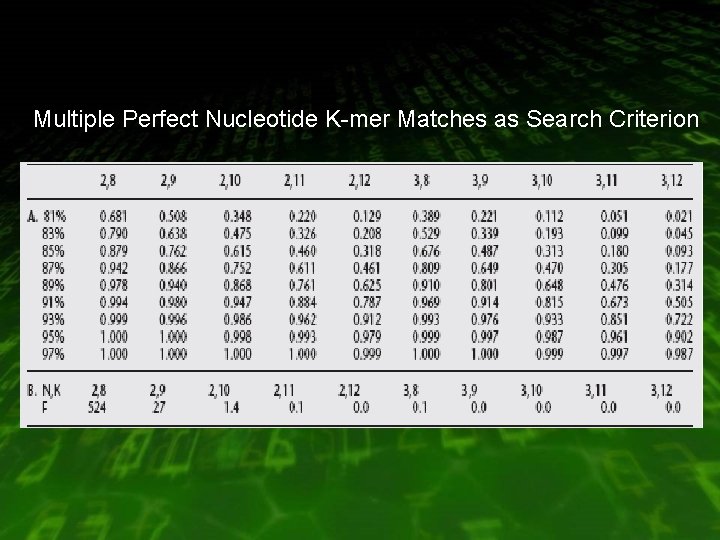
Multiple Perfect Nucleotide K-mer Matches as Search Criterion

Default • Nucleotide – two perfect 11 -mer • Protein – single perfect 5 -mer for standalone version – three perfect 4 -mer for client/server version

BLAST 1) Build the hash table for Sequence A. 2) Scan Sequence B for hits. 3) Extend hits.

BLAST Step 1: Build the hash table for Sequence A. (3 -tuple example) For protein sequences: For DNA sequences: Seq. A = AGATCGAT 12345678 AAA AAC. . AGA. . ATC. . CGA. . GAT. . TCG. . TTT 1 3 5 2 4 6 Seq. A = ELVIS Add xyz to the hash table if Score(xyz, ELV) ≧ T; Add xyz to the hash table if Score(xyz, LVI) ≧ T; Add xyz to the hash table if Score(xyz, VIS) ≧ T;

BLAST Step 2: Scan sequence B for hits.

BLAST Step 2: Scan sequence B for hits. Step 3: Extend hits. BLAST 2. 0 saves the time spent in extension, and hit considers gapped alignments. Terminate if the score of the sxtension fades away. (That is, when we reach a segment pair whose score falls a certain distance below the best score found for shorter extensions. )

Algorithm 1. Search Stage – Use an index to find regions in genome homologous to query 2. Alignment Stage – Do a detailed alignment between query and homologous regions 3. Stitching and Filling In – Use dynamic programming to stitch together detailed alignments regions into detailed alignment of whole
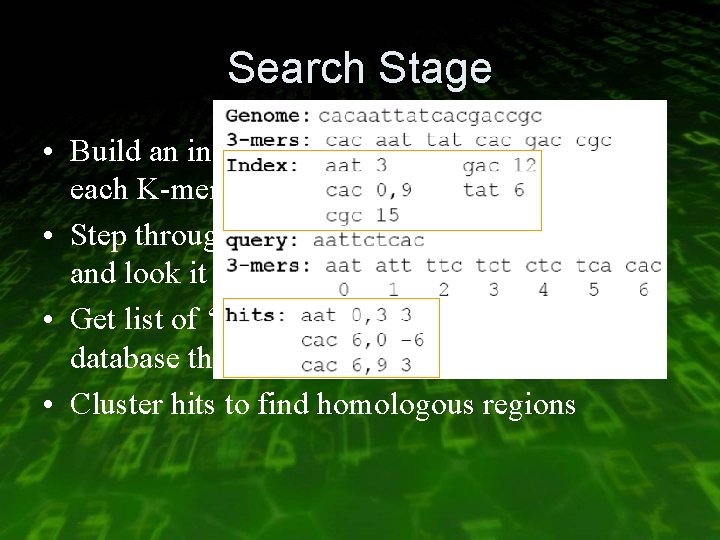
Search Stage • Build an index which contains positions of each K-mer in database. • Step through each overlapping K-mer in query and look it up in index • Get list of ‘hits’ - positions in query and in database that match for K bases • Cluster hits to find homologous regions
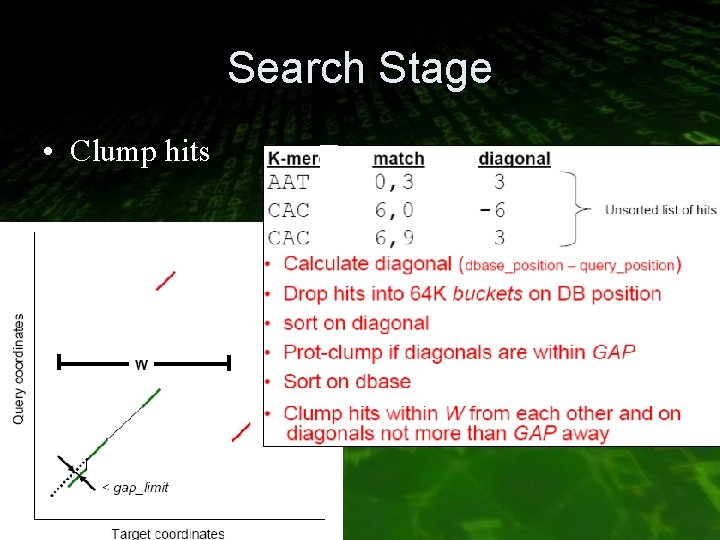
Search Stage • Clump hits
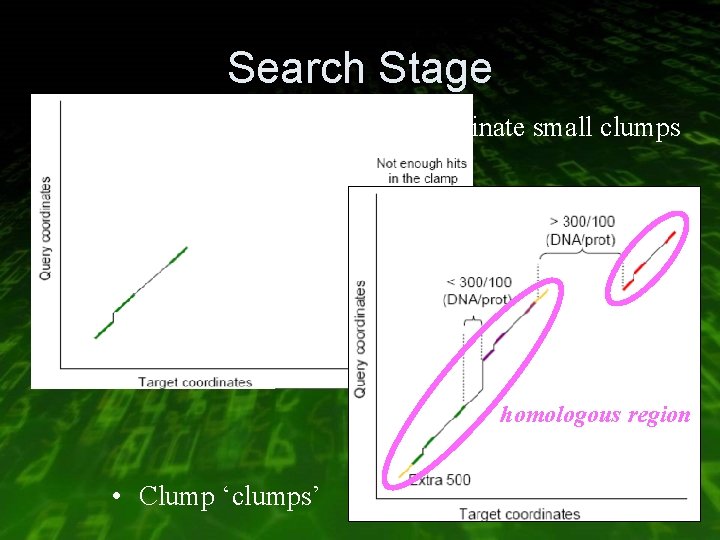
Search Stage • Eliminate small clumps homologous region • Clump ‘clumps’

Alignment Stage (nucleotide) • Start from scratch with regions defined with Kmers • Index on smaller K-mers, but extend each Kmer until it becomes specific • Extend in both direction without mismatches or gaps and merge overlapping or continues alignments • Recurse on gaps with smaller K until gap or hits are eliminated
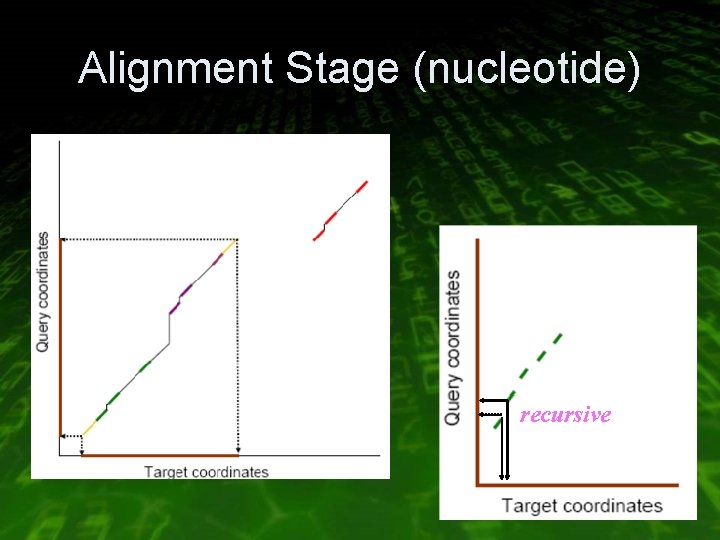
Alignment Stage (nucleotide) recursive

Alignment Stage (protein) • Extend hits into maximal scoring ungapped alignment (HSPs) with +2/-1 scoring scheme • Create a graph of all possible HSP merges • Use dynamic programming to traverse the graph
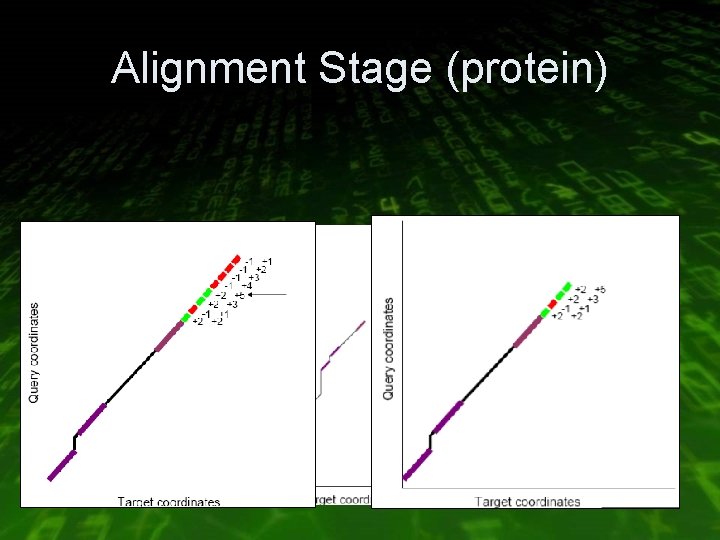
Alignment Stage (protein)
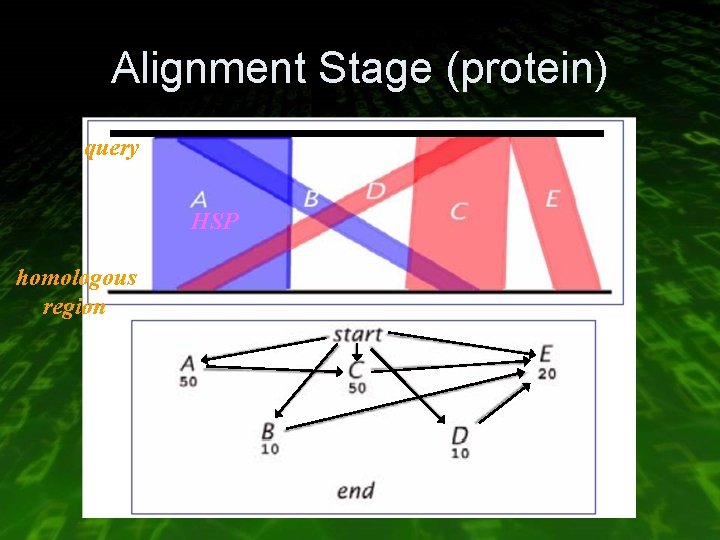
Alignment Stage (protein) query HSP homologous region

Stitching and Filling In • The alignment of gene is often scattered across multiple homologous regions found in the search stage query database
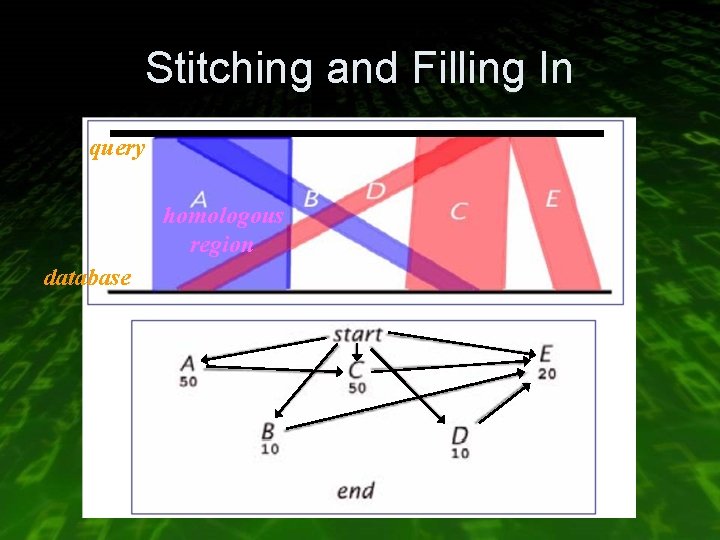
Stitching and Filling In query homologous region database

Evaluation • Comparison with Other Tools: – m. RNA/Genome Alignments – Remapped 713 m. RNAs corresponding to annotated chromosome 22 – BLAT took 26 sec while Sim 4 took 17, 468 sec (almost 5 h) Relative speed Est_genome 1 Sim 4 333 BLAT 223, 000 Base accuracy N/A 99. 66% 99. 99% Gene accuracy 77. 7% 93. 4% 99. 5%
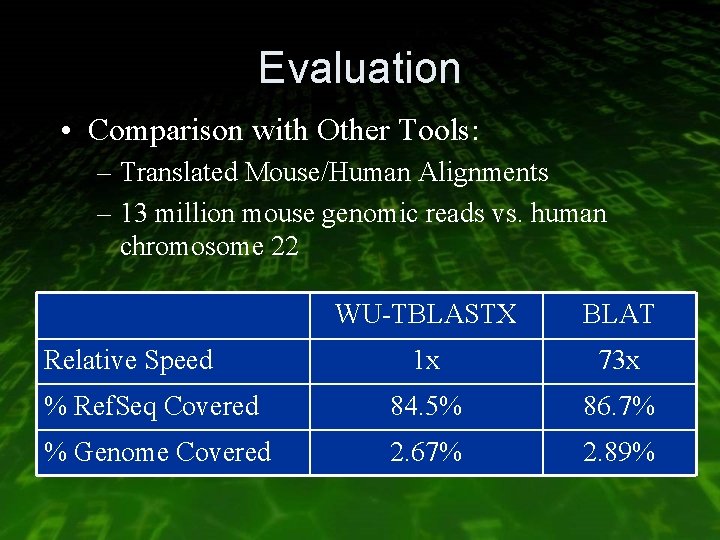
Evaluation • Comparison with Other Tools: – Translated Mouse/Human Alignments – 13 million mouse genomic reads vs. human chromosome 22 WU-TBLASTX BLAT 1 x 73 x % Ref. Seq Covered 84. 5% 86. 7% % Genome Covered 2. 67% 2. 89% Relative Speed

BLAT vs. BLAST • Index – Query vs. Database • Hits – Perfect vs. Near Perfect • Alignment – Separate vs. Together

Magic Time !

Magic 4 No Prediction Great mind! ! 3 !. 5 3 4

Reference • http: //amber. cs. umd. edu/class/838 s 04/nada. ppt • http: //bioportal. weizmann. ac. il/course/ATIB/A TIB 03_lecture 3. print. pdf
- Slides: 40