BLAST et BLAST avanc J S BernardesH Richard

BLAST et BLAST avancé J. S. Bernardes/H. Richard Matériel : Bioinformatics and Functional Genomics Jonathan Pevsner, Wiley-Blackwell ed.

Outline Introduction Definition of Orthology and Paralogy A myoglobin example BLAST search steps Step 1: Specifying Sequence of interest Step 2: Selecting BLAST Program Step 3: Selecting a Database Step 4: Selecting Search Parameters and Formatting Parameters BLAST algorithm uses local alignment search strategy BLAST algorithm parts: list, scan, extend BLAST algorithm: local alignment search statistics and E value Making sense of raw scores with bit scores BLAST algorithm: Relation Between E and p values BLAST search strategies General concepts; principles of BLAST searching How to evaluate the significance of results How to handle too many or few results BLAST searching with multidomain protein: HIV-1 Pol Using BLAST for gene discovery: Find-a-Gene

Learning objectives • Definition of homology, similarity, conservation • Difference between orthologues and paralogues • perform BLAST searches at the NCBI website; • understand how to vary optional BLAST search parameters; • explain the three phases of a BLAST search (compile, scan/extend, trace‐back); • define the mathematical relationship between

Definitions: identity, similarity, conservation Homology Similarity attributed to descent from a common ancestor. Identity The extent to which two (nucleotide or amino acid) sequences are invariant. Similarity The extent to which nucleotide or protein sequences are related. It is based upon identity plus conservation. Conservation B&FG 3 e Page 70 Changes at a specific position of an amino acid or (less commonly, DNA) sequence that preserve the physico-chemical properties of the original residue.
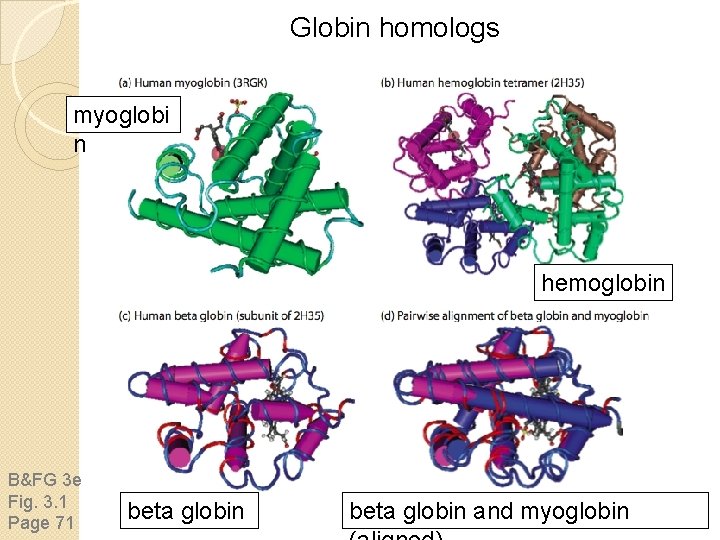
Globin homologs myoglobi n hemoglobin B&FG 3 e Fig. 3. 1 Page 71 beta globin and myoglobin

Definitions: two types of homology Orthologs Homologous sequences in different species that arose from a common ancestral gene during speciation; may or may not be responsible for a similar function. Paralogs Homologous sequences within a single species that arose by gene duplication. B&FG 3 e Fig. 2 -3 Page 22
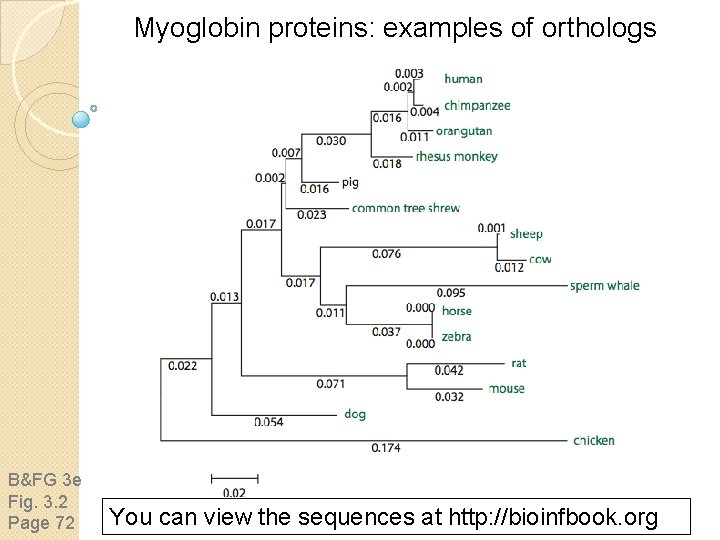
Myoglobin proteins: examples of orthologs B&FG 3 e Fig. 3. 2 Page 72 You can view the sequences at http: //bioinfbook. org
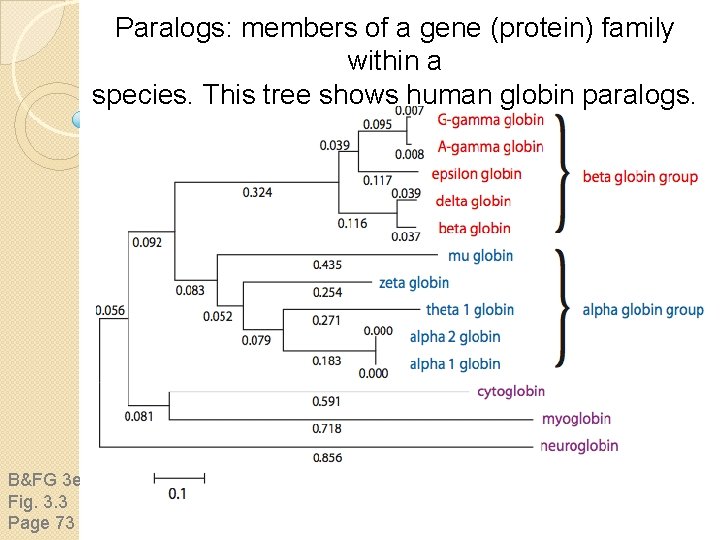
Paralogs: members of a gene (protein) family within a species. This tree shows human globin paralogs. B&FG 3 e Fig. 3. 3 Page 73
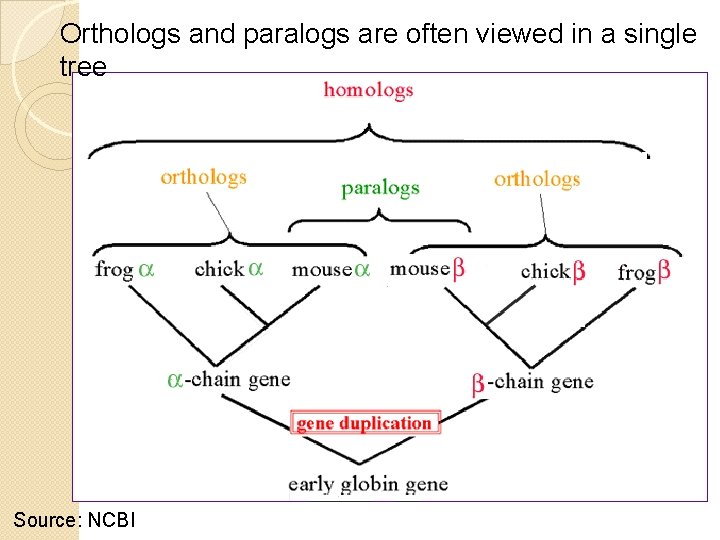
Orthologs and paralogs are often viewed in a single tree Source: NCBI
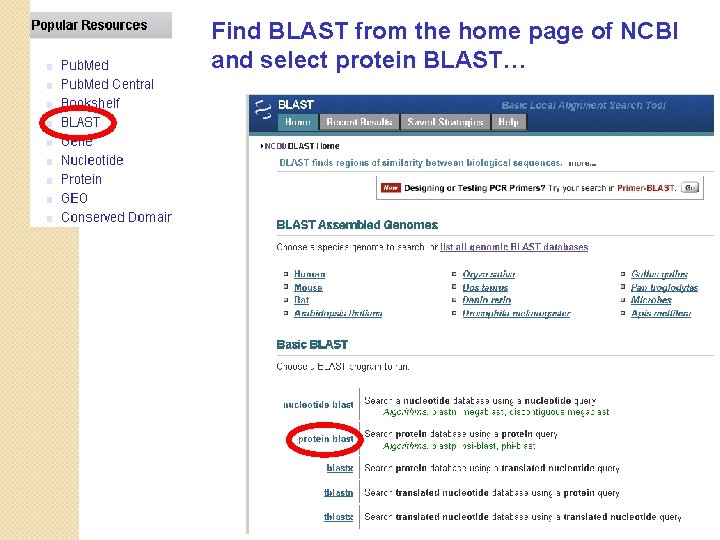
Find BLAST from the home page of NCBI and select protein BLAST…
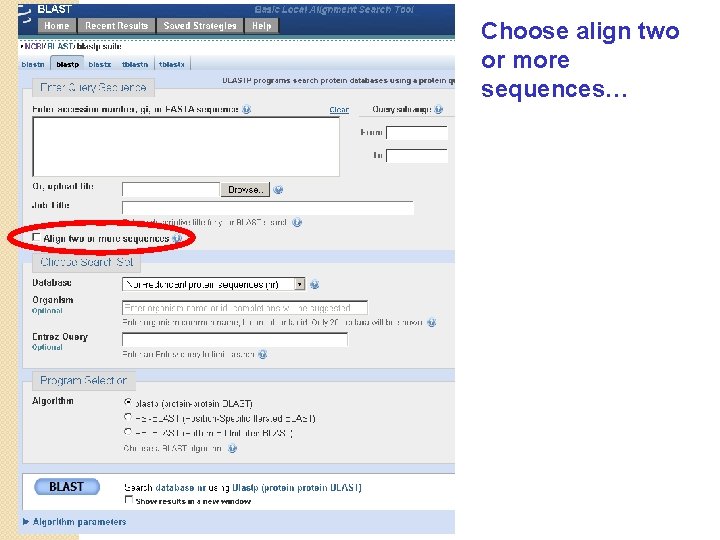
Choose align two or more sequences…
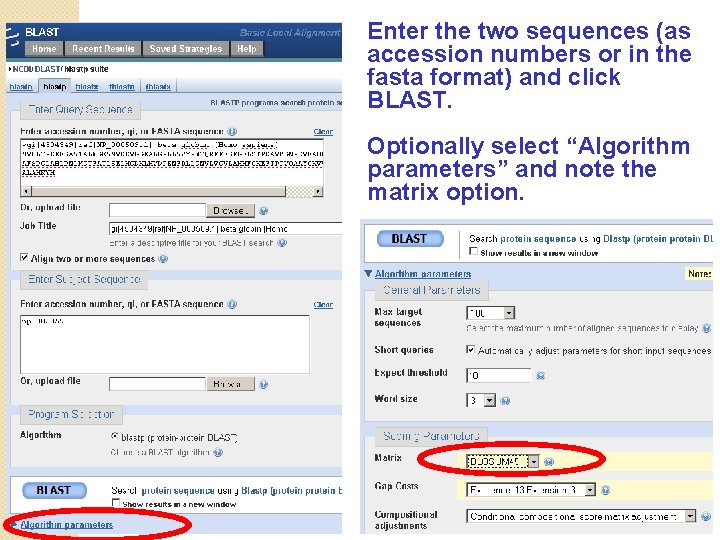
Enter the two sequences (as accession numbers or in the fasta format) and click BLAST. Optionally select “Algorithm parameters” and note the matrix option.
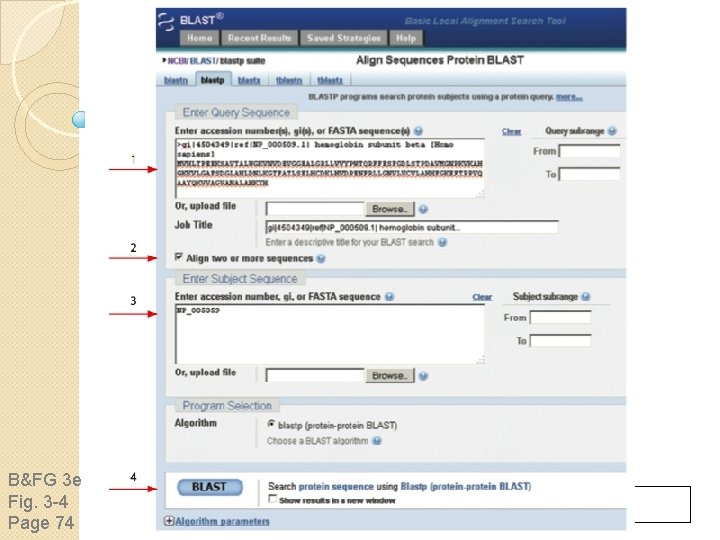
sequence B&FG 3 e Fig. 3 -4 Page 74 Year

Pairwise alignment of human beta globin (the “query”) and myoglobin (the “subject”) B&FG 3 e Fig. 3 -5 Page 75 We’ll examine the highlighted green region of the alignment in more detail.

How raw scores are calculated: an example B&FG 3 e Fig. 3 -5 Page 75 For a set of aligned residues we assign scores based on matches, mismatches, gap open penalties, and gap extension penalties. These scores add up to the total raw score.

Why use BLAST? BLAST searching is fundamental to understanding the relatedness of any favorite query sequence to other known proteins or DNA sequences. Applications include • identifying orthologs and paralogs • discovering new genes or proteins • discovering variants of genes or proteins • investigating expressed sequence tags (ESTs) • exploring protein structure and function
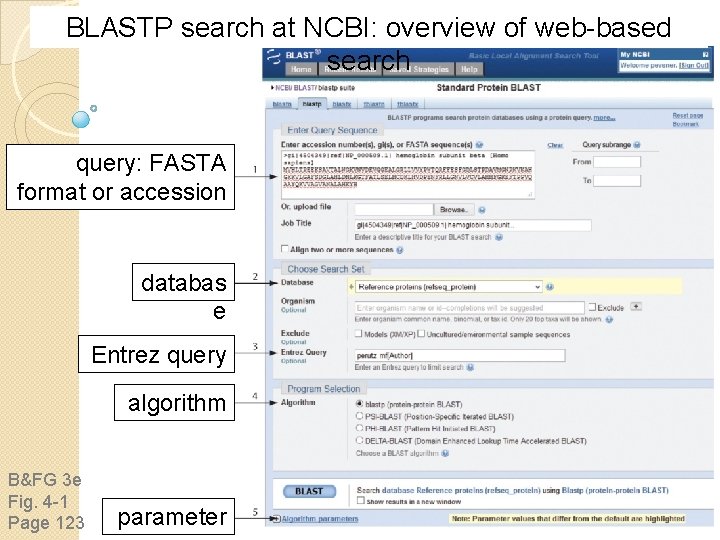
BLASTP search at NCBI: overview of web-based search query: FASTA format or accession databas e Entrez query algorithm B&FG 3 e Fig. 4 -1 Page 123 parameter

Outline Introduction Definition of Orthology and Paralogy A myoglobin example BLAST search steps Step 1: Specifying Sequence of interest Step 2: Selecting BLAST Program Step 3: Selecting a Database Step 4: Selecting Search Parameters and Formatting Parameters BLAST algorithm uses local alignment search strategy BLAST algorithm parts: list, scan, extend BLAST algorithm: local alignment search statistics and E value Making sense of raw scores with bit scores BLAST algorithm: Relation Between E and p values BLAST search strategies General concepts; principles of BLAST searching How to evaluate the significance of results How to handle too many or few results BLAST searching with multidomain protein: HIV-1 Pol Using BLAST for gene discovery: Find-a-Gene

Step 1: Choose your sequence Sequence can be input in FASTA format or as accession number
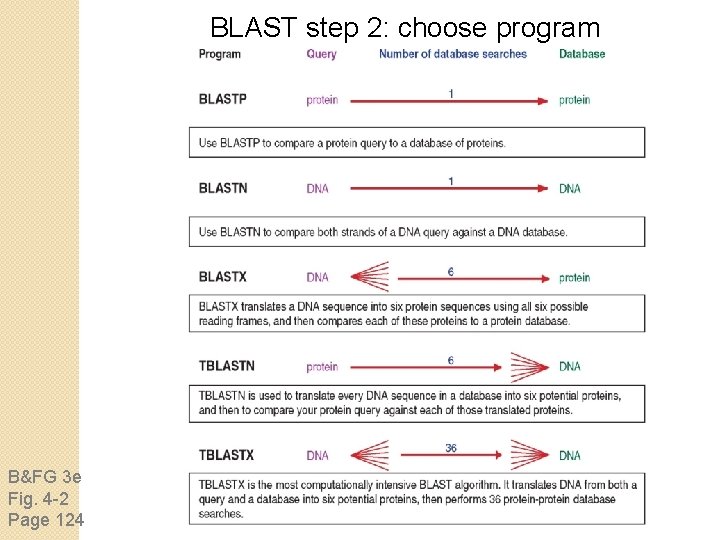
BLAST step 2: choose program B&FG 3 e Fig. 4 -2 Page 124
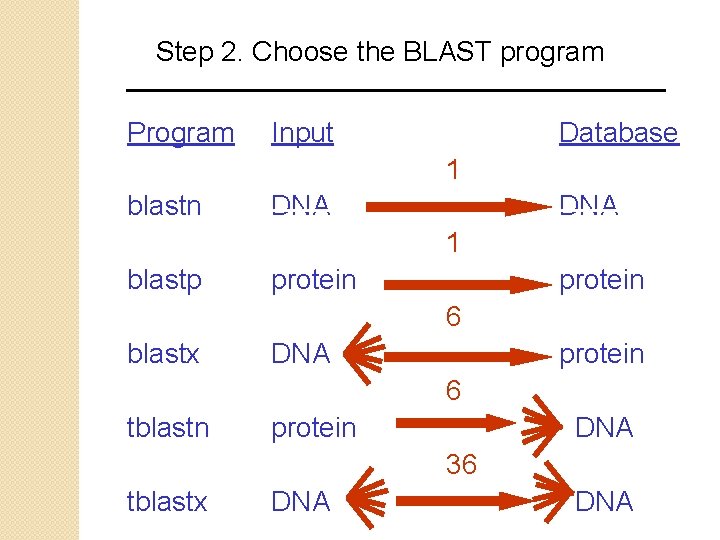
Step 2. Choose the BLAST program Program Input Database 1 blastn DNA 1 blastp protein 6 blastx DNA protein 6 tblastn protein DNA 36 tblastx DNA
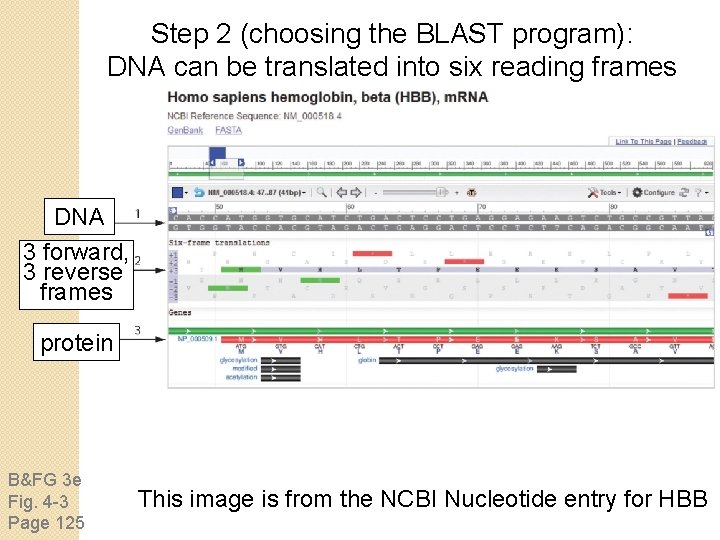
Step 2 (choosing the BLAST program): DNA can be translated into six reading frames DNA 3 forward, 3 reverse frames protein B&FG 3 e Fig. 4 -3 Page 125 This image is from the NCBI Nucleotide entry for HBB

Step 3: choose a database to search (protein database B&FG 3 e Table 4 -1 Page 126

Step 3: choose a database to search (nucleotide) B&FG 3 e Table 4 -2 Page 127

Step 4: optional parameters You can. . . • choose the organism to search • turn filtering on/off • change the substitution matrix • change the expect (e) value • change the word size • change the output format Example: BLASTP human insulin (NP_000198) against a C. elegans Ref. Seq database. Varying some parameters (filtering, compositional adjustments) can greatly affect the alignment itself.
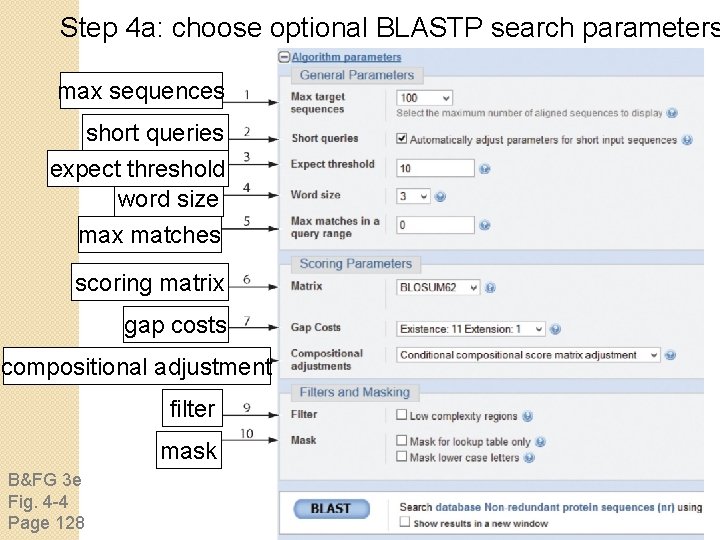
Step 4 a: choose optional BLASTP search parameters max sequences short queries expect threshold word size max matches scoring matrix gap costs compositional adjustment filter mask B&FG 3 e Fig. 4 -4 Page 128

Step 4 a: compositional adjustment influences score, expect value search results expect = 0. 05 Default: conditional compositional score matrix adjustment expect = 0. 09 no adjustment expect = 1 e-04 B&FG 3 e Fig. 4 -5 Page 129 compositionbased statistics

Step 4 b: formatting options The top of the BLAST output summarizes the query, database, and BLAST algorithm. Click to access a summary of the search parameters or a taxonomic report. B&FG 3 e Fig. 4 -6 Page 132
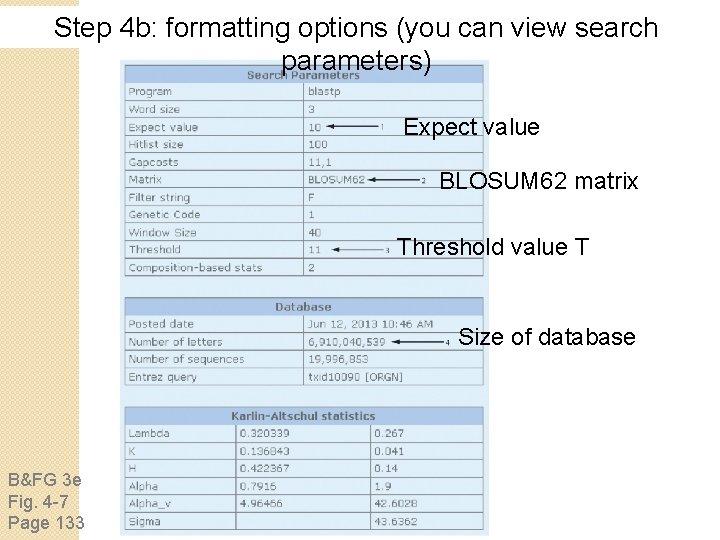
Step 4 b: formatting options (you can view search parameters) Expect value BLOSUM 62 matrix Threshold value T Size of database B&FG 3 e Fig. 4 -7 Page 133
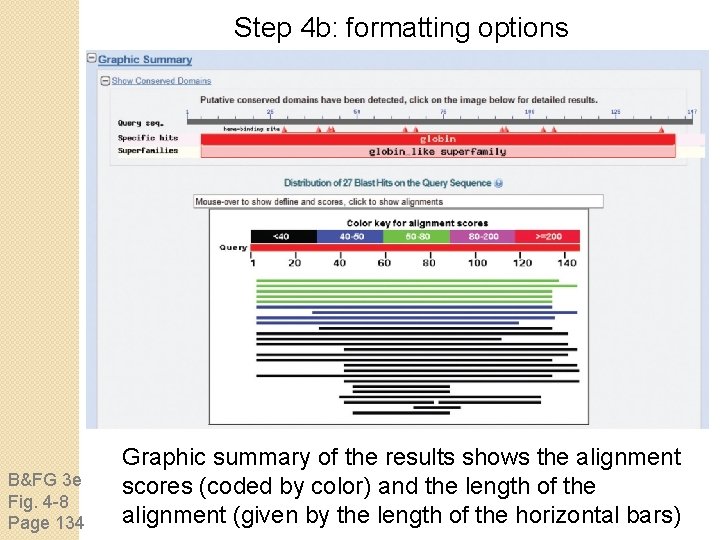
Step 4 b: formatting options B&FG 3 e Fig. 4 -8 Page 134 Graphic summary of the results shows the alignment scores (coded by color) and the length of the alignment (given by the length of the horizontal bars)

BLASTP output includes list of matches; links to the NCBI protein entry; bit score and E value; and download options B&FG 3 e Fig. 4 -9 Page 134
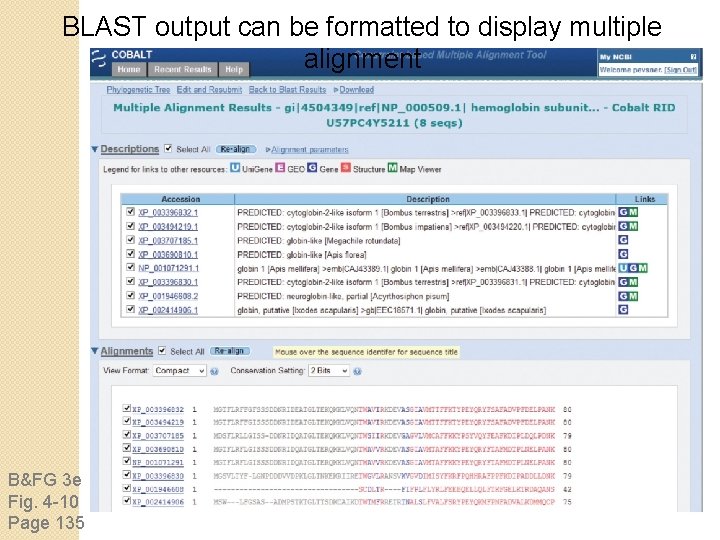
BLAST output can be formatted to display multiple alignment B&FG 3 e Fig. 4 -10 Page 135
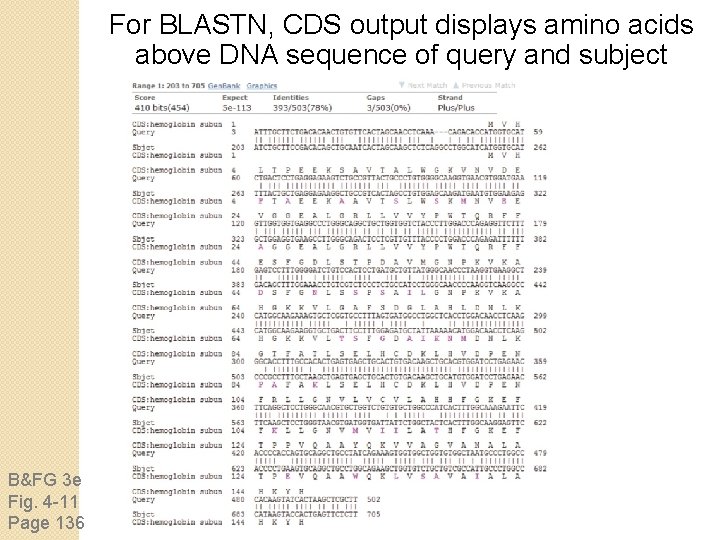
For BLASTN, CDS output displays amino acids above DNA sequence of query and subject B&FG 3 e Fig. 4 -11 Page 136

Outline Introduction Definition of Orthology and Paralogy A myoglobin example BLAST search steps Step 1: Specifying Sequence of interest Step 2: Selecting BLAST Program Step 3: Selecting a Database Step 4: Selecting Search Parameters and Formatting Parameters BLAST algorithm uses local alignment search strategy BLAST algorithm parts: list, scan, extend BLAST algorithm: local alignment search statistics and E value Making sense of raw scores with bit scores BLAST algorithm: Relation Between E and p values BLAST search strategies General concepts; principles of BLAST searching How to evaluate the significance of results How to handle too many or few results BLAST searching with multidomain protein: HIV-1 Pol Using BLAST for gene discovery: Find-a-Gene

How a BLAST search works “The central idea of the BLAST algorithm is to confine attention to segment pairs that contain a word pair of length w with a score of at least T. ” Altschul et al. (1990)

How the original BLAST algorithm works: three phases Phase 1: compile a list of word pairs (w=3) above threshold T Example: for a human RBP query …FSGTWYA… (query word is in green) A list of words (w=3) is: FSG SGT GTW TWY WYA YSG TGT ATW SWY WFA FTG SVT GSW TWF WYS. . .
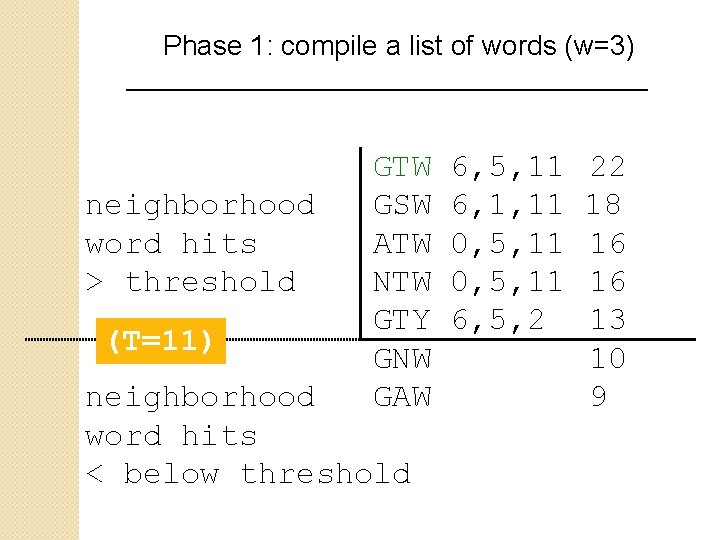
Phase 1: compile a list of words (w=3) neighborhood word hits > threshold (T=11) GTW GSW ATW NTW GTY GNW GAW neighborhood word hits < below threshold 6, 5, 11 6, 1, 11 0, 5, 11 6, 5, 2 22 18 16 16 13 10 9 Fig. 4. 11 page 116

B&FG 3 e Fig. 4 -12 Page 139

Phase 2: scan the database for matches and exten B&FG 3 e Fig. 4 -12 Page 139

Phase 3: Traceback to generate gapped alignme B&FG 3 e Fig. 4 -12 Page 139

How a BLAST search works: threshold You can locally install BLAST and modify the threshold parameter. The default value for BLASTP is 11. To change it, enter “-f 16” or “-f 5” in the advanced options of BLAST+.
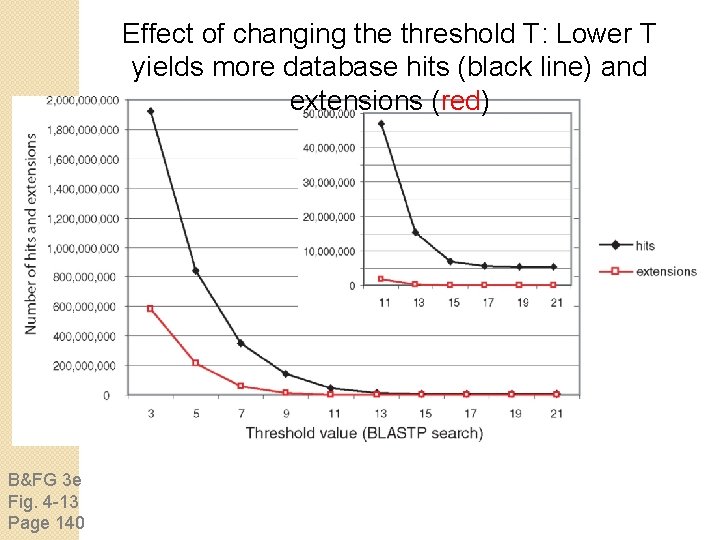
Effect of changing the threshold T: Lower T yields more database hits (black line) and extensions (red) B&FG 3 e Fig. 4 -13 Page 140

For BLASTN, the word size is typically 7, 11, or 15 (EXACT match). Changing word size is like changing threshold of proteins. w=15 gives fewer matches and is faster than w=11 or w=7. For mega. BLAST (see below), the word size is 28 and can be adjusted to 64. What will this do? Mega. BLAST is VERY fast for finding closely related DNA sequences!

How to interpret a BLAST search: expect value It is important to assess the statistical significance of search results. For global alignments, the statistics are poorly understoo For local alignments (including BLAST search results), the statistics are well understood. The scores follow an extreme value distribution (EVD) rather than a normal distribution.
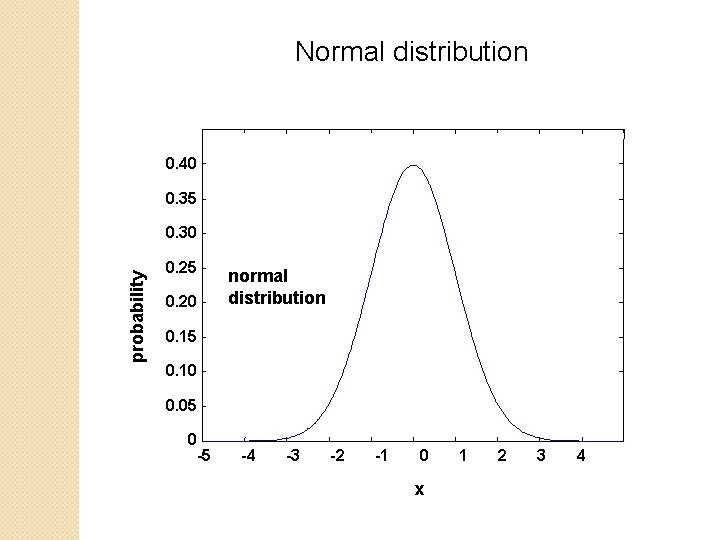
Normal distribution 0. 40 0. 35 probability 0. 30 0. 25 0. 20 normal distribution 0. 15 0. 10 0. 05 0 -5 -4 -3 -2 -1 0 x 1 2 3 4
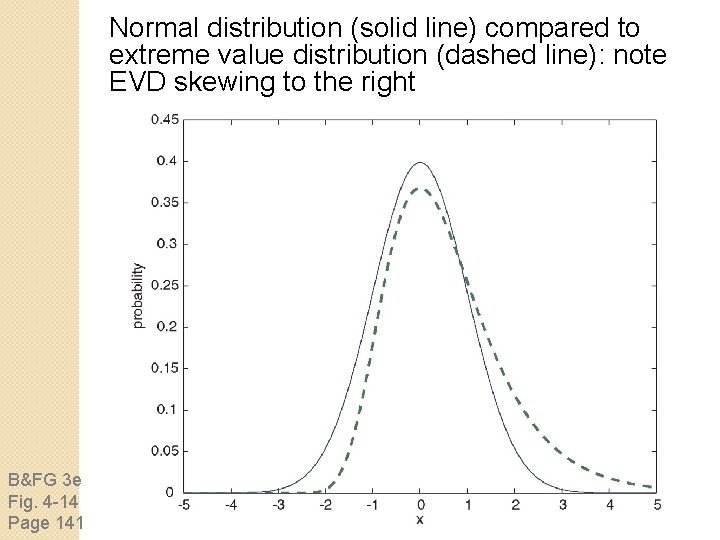
Normal distribution (solid line) compared to extreme value distribution (dashed line): note EVD skewing to the right B&FG 3 e Fig. 4 -14 Page 141

How to interpret a BLAST search: expect value The expect value E is the number of alignments with scores greater than or equal to score S that are expected to occur by chance in a database search. An E value is related to a probability value p. The key equation describing an E value is: E = Kmn e-l. S

E = Kmn e-l. S This equation is derived from a description of the extreme value distribution S = the score E = the expect value = the number of highscoring segment pairs (HSPs) expected to occur with a score of at least S m, n = the length of two sequences l, K = Karlin Altschul statistics

Some properties of the equation E = Kmn e-l. S • The value of E decreases exponentially with increasing (higher S values correspond to better alignments). Very high scores correspond to very low E values. • The E value for aligning a pair of random sequences mus be negative! Otherwise, long random alignments would acquire great scores • Parameter K describes the search space (database). • For E=1, one match with a similar score is expected to occur by chance. For a very much larger or smaller database, you would expect E to vary accordingly

From raw scores to bit scores • There are two kinds of scores: raw scores (calculated from a substitution matrix) and bit scores (normalized scores) • Bit scores are comparable between different searches because they are normalized to account for the use of different scoring matrices and different database sizes S’ = bit score = (l. S - ln. K) / ln 2 The E value corresponding to a given bit score is: E = mn 2 -S’ B&FG 3 e Page 143 Bit scores allow you to compare results between

How to interpret BLAST: E values and p values The expect value E is the number of alignments with scores greater than or equal to score S that are expected to occur by chance in a database search. A p value is a different way of representing the significance of an alignment. p = 1 - e-E B&FG 3 e Page 143

How to interpret BLAST: E values and p values E values of about 1 to 10 are far easier to interpret than corresponding p values. Very small E values are very similar to p values. __E 10 5 2 1 0. 05 0. 001 0. 0001 ____p 0. 99995460 0. 99326205 0. 86466472 0. 63212056 0. 09516258 (about 0. 1) 0. 04877058 (about 0. 05) 0. 00099950 (about 0. 001) 0. 00010000 E values are comparable to p values, and are designed to be more convenient to interpret.

Outline Introduction Definition of Orthology and Paralogy A myoglobin example BLAST search steps Step 1: Specifying Sequence of interest Step 2: Selecting BLAST Program Step 3: Selecting a Database Step 4: Selecting Search Parameters and Formatting Parameters BLAST algorithm uses local alignment search strategy BLAST algorithm parts: list, scan, extend BLAST algorithm: local alignment search statistics and E value Making sense of raw scores with bit scores BLAST algorithm: Relation Between E and p values BLAST search strategies General concepts; principles of BLAST searching How to evaluate the significance of results How to handle too many or few results BLAST searching with multidomain protein: HIV-1 Pol Using BLAST for gene discovery: Find-a-Gene
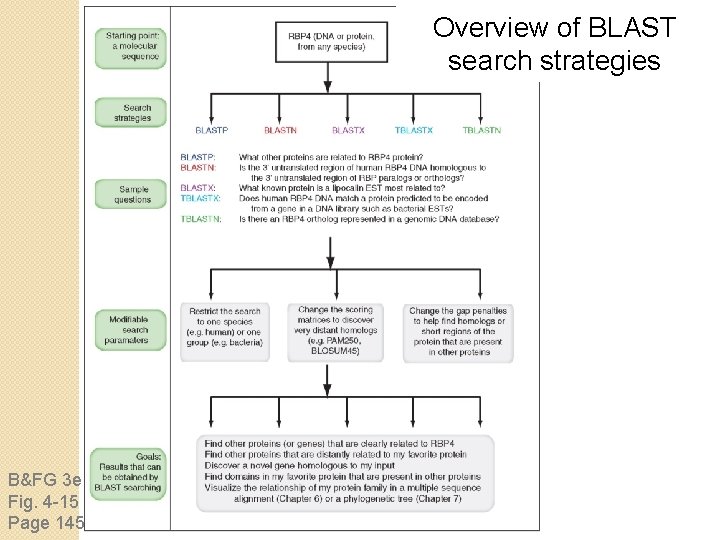
Overview of BLAST search strategies B&FG 3 e Fig. 4 -15 Page 145

BLASTP search: human RBP 4 query, human Ref. Seq database Results include matches (such as CG 8) with high E values and limited identity to the query B&FG 3 e Fig. 4 -16 Page 147

“Recipricol” BLASTP search with CG 8 as query includes RBP 4 and other lipocalins B&FG 3 e Fig. 4 -17 Page 149 This confirms that the finding of CG 8 using RBP 4 as a query was a true positive

Pairwise alignment of CG 8 with non-homologous proteins B&FG 3 e Fig. 4 -17 Page 149 • • Query and subject are very different lengths E values are not significant Matches lack GXW motif Subjects are not annotated as lipocalins
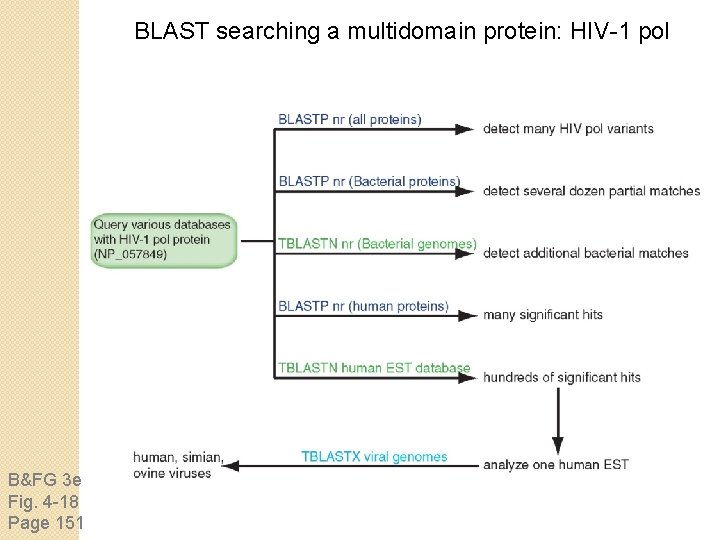
BLAST searching a multidomain protein: HIV-1 pol B&FG 3 e Fig. 4 -18 Page 151
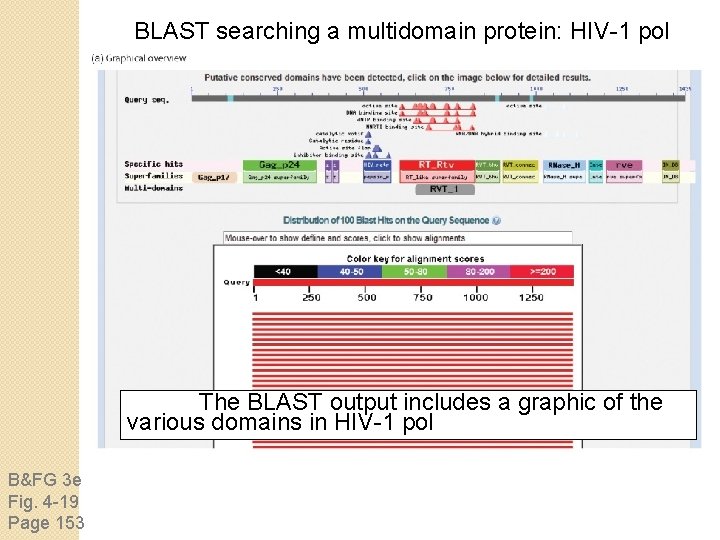
BLAST searching a multidomain protein: HIV-1 pol The BLAST output includes a graphic of the various domains in HIV-1 pol B&FG 3 e Fig. 4 -19 Page 153
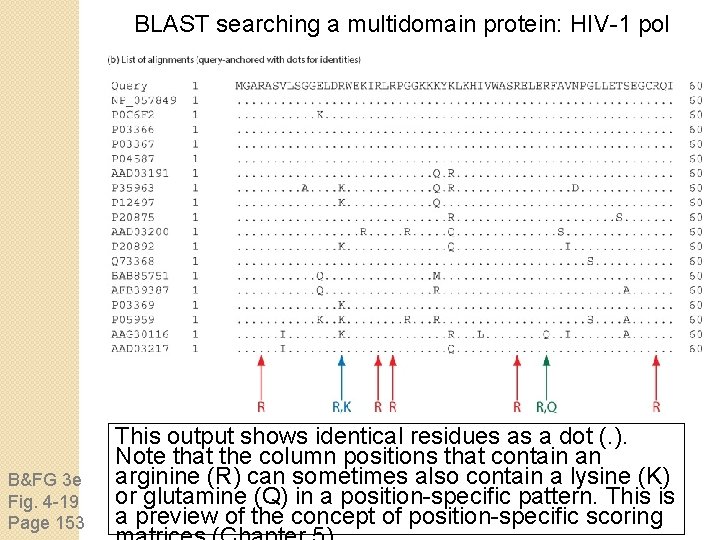
BLAST searching a multidomain protein: HIV-1 pol B&FG 3 e Fig. 4 -19 Page 153 This output shows identical residues as a dot (. ). Note that the column positions that contain an arginine (R) can sometimes also contain a lysine (K) or glutamine (Q) in a position-specific pattern. This is a preview of the concept of position-specific scoring

Taxonomy report for a BLAST searching HIV-1 pol Most of the matches are to viruses, but there also matches to rabbit, fungal, pig, and insect sequences. B&FG 3 e Fig. 4 -20 Page 154
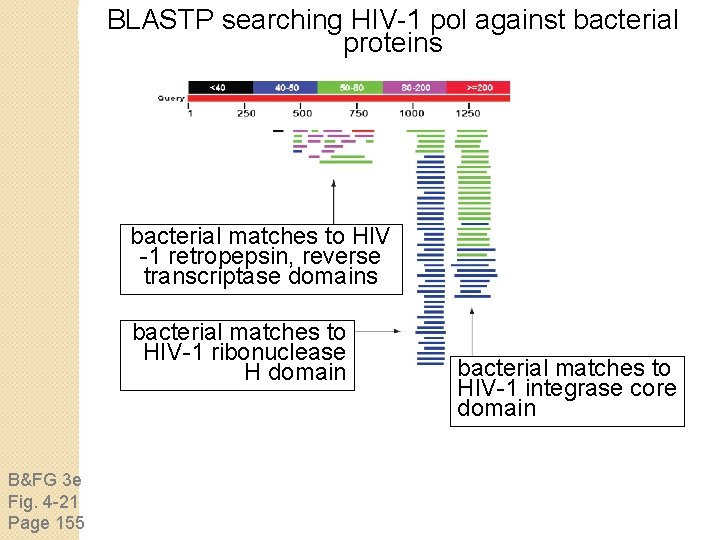
BLASTP searching HIV-1 pol against bacterial proteins bacterial matches to HIV -1 retropepsin, reverse transcriptase domains bacterial matches to HIV-1 ribonuclease H domain B&FG 3 e Fig. 4 -21 Page 155 bacterial matches to HIV-1 integrase core domain
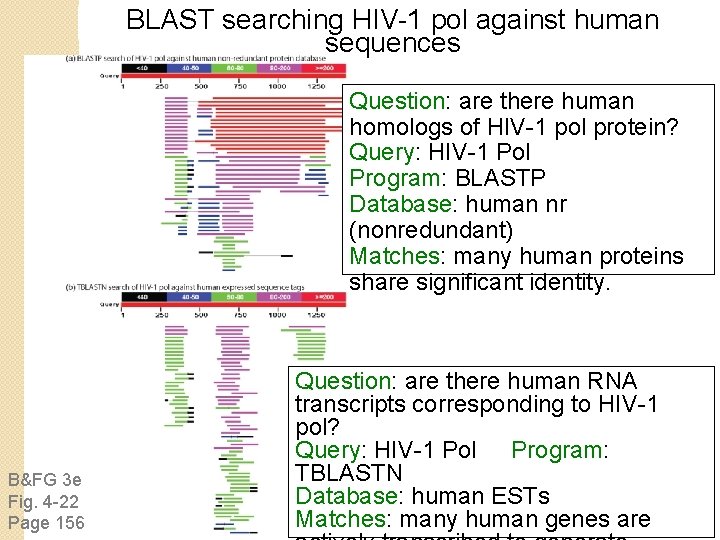
BLAST searching HIV-1 pol against human sequences Question: are there human homologs of HIV-1 pol protein? Query: HIV-1 Pol Program: BLASTP Database: human nr (nonredundant) Matches: many human proteins share significant identity. B&FG 3 e Fig. 4 -22 Page 156 Question: are there human RNA transcripts corresponding to HIV-1 pol? Query: HIV-1 Pol Program: TBLASTN Database: human ESTs Matches: many human genes are

Outline Introduction BLAST search steps Step 1: Specifying sequence of interest Step 2: Selecting BLAST program Step 3: Selecting a database Step 4: Selecting search parameters and formatting parameters Stand-alone BLAST algorithm uses local alignment search strategy BLAST algorithm parts: list, scan, extend BLAST algorithm: local alignment search statistics and E value Making sense of raw scores with bit scores BLAST algorithm: relation between E and p values BLAST search strategies General concepts; principles of BLAST searching How to evaluate the significance of results How to handle too many or too few results BLAST searching with multidomain protein: HIV-1 Pol
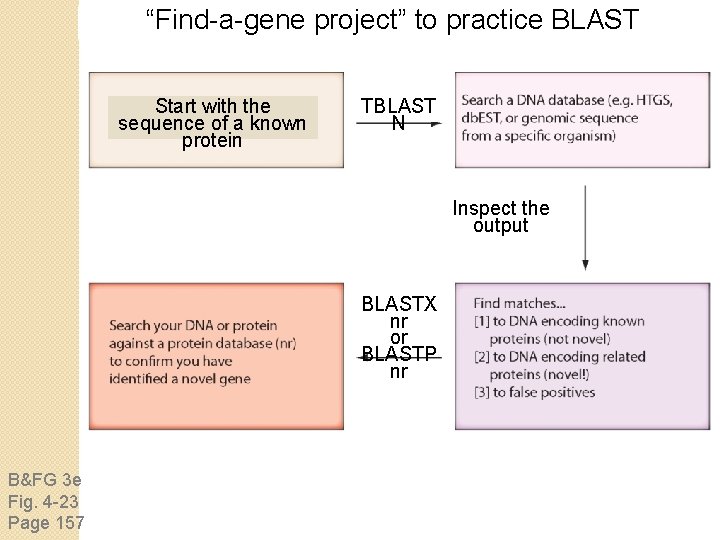
“Find-a-gene project” to practice BLAST Start with the sequence of a known protein TBLAST N Inspect the output BLASTX nr or BLASTP nr B&FG 3 e Fig. 4 -23 Page 157

“Find-a-gene project” example: novel globin Query: NP_000509 Program: TBLASTN Database: EST (nematodes) Match: novel globin B&FG 3 e Fig. 4 -24 Page 158 Confirmation Query: nematode EST Program: BLASTX Best match: a globin, but not a previously annotated globin

“Find-a-gene project” • The find-a-gene project is meant to be a very focused, specific project to help you understand how to use various BLAST tools (e. g. TBLASTN, BLASTX, BLASTP) and various databases. • You can start with (almost) any protein, from the organism of your choice, and discover a “novel” gene in another organism that is homologous but has never been annotated before as related to your query. Therefore you are discovering a new gene. • You can take your new gene/protein, name it, then search it against databases to confirm it has not been described before. • You can further perform multiple sequence alignment (Chapter 6), phylogeny (Chapter 7), and predict its protein structure (Chapter 13) and its function (Chapter 14).

Outline Introduction BLAST search steps Step 1: Specifying sequence of interest Step 2: Selecting BLAST program Step 3: Selecting a database Step 4: Selecting search parameters and formatting parameters Stand-alone BLAST algorithm uses local alignment search strategy BLAST algorithm parts: list, scan, extend BLAST algorithm: local alignment search statistics and E value Making sense of raw scores with bit scores BLAST algorithm: relation between E and p values BLAST search strategies General concepts; principles of BLAST searching How to evaluate the significance of results How to handle too many or too few results BLAST searching with multidomain protein: HIV-1 Pol
![Three problems standard BLAST cannot solve [1] Use human beta globin as a query Three problems standard BLAST cannot solve [1] Use human beta globin as a query](http://slidetodoc.com/presentation_image_h2/4bd99b349f9089d951029c83b9f90bcb/image-69.jpg)
Three problems standard BLAST cannot solve [1] Use human beta globin as a query against human Ref. Seq proteins, and BLASTP does not “find” human myoglobin. This is because the two proteins are too distantly related. PSI-BLAST at NCBI as well as hidden Markov models easily solve this problem. [2] How can we search using 10, 000 base pairs as a query, or even millions of base pairs? Many BLASTlike tools for genomic DNA are available such as Pattern. Hunter, Megablast, BLAT, and LASTZ. [3] How can we align tens of millions of short reads to a reference genome?

Position specific iterated BLAST: PSI-BLAST The purpose of PSI-BLAST is to look deeper into the database for matches to your query protein sequence by employing a scoring matrix that is customized to your query.
![PSI-BLAST is performed in five steps [1] Select a query and search it against PSI-BLAST is performed in five steps [1] Select a query and search it against](http://slidetodoc.com/presentation_image_h2/4bd99b349f9089d951029c83b9f90bcb/image-71.jpg)
PSI-BLAST is performed in five steps [1] Select a query and search it against a protein databas B&FG 3 e Page 172
![PSI-BLAST is performed in five steps [1] Select a query and search it against PSI-BLAST is performed in five steps [1] Select a query and search it against](http://slidetodoc.com/presentation_image_h2/4bd99b349f9089d951029c83b9f90bcb/image-72.jpg)
PSI-BLAST is performed in five steps [1] Select a query and search it against a protein databas [2] PSI-BLAST constructs a multiple sequence alignment then creates a “profile” or specialized position-specific scoring matrix (PSSM) B&FG 3 e Page 172

Inspect the BLASTP output to identify empirical “rules” regarding amino acids tolerated at each position R, I, K C D, E, T K, R, T N, L, Y, G

1 M 2 K 3 W 4 V 5 W 6 A 7 L 8 L 9 L 10 L 11 A 12 A 13 W 14 A 15 A 16 A. . . 37 S 38 G 39 T 40 W 41 Y 42 A A -1 -1 -3 0 -3 5 -2 -1 -1 -2 5 5 -2 3 2 4 R N D C Q E G H I L K M F -2 -2 -3 -2 -1 -2 -3 -2 1 2 -2 6 0 1 -4 2 4 -2 0 -3 -3 3 -2 -4 -3 -4 -5 -3 -2 -3 -2 1 20 amino acids -3 -3 -4 -1 -3 -3 -4 -4 3 1 -1 -3 -4 -5 -3 -2 -3 -2 1 -2 -2 -2 -1 -1 -1 0 -2 -2 -2 -1 -1 -3 -2 -4 -4 -1 -2 -3 -4 -3 2 0 -3 -3 -4 -1 -3 -3 -4 -3 2 2 -3 1 3 -3 -4 -4 -1 -2 -3 -4 -3 2 0 -2 -2 -2 -1 -1 -1 0 -2 -2 -2 -1 -1 -3 all the amino acids -3 -4 -4 -2 -2 -3 -4 -3 1 4 -3 2 1 from 1 to 4 the -2 -1 position -2 -1 -1 -2 -2 -1 -2 -3 -1 0 -1 -2 2 0 2 -1 -3 -3 0 -2 -3 end of your PSI-2 -1 -1 -1 3 -2 -2 -2 -1 -1 -3 P -3 -1 -4 -3 -4 -1 -3 -3 -1 -1 -1 W -2 -3 12 -3 -2 -2 -3 -3 7 -3 -3 -3 Y -1 -2 2 -1 2 -2 -1 0 -1 -1 -2 -2 0 -3 -2 -2 V 1 -3 -3 4 -3 0 1 3 2 1 0 0 0 -1 -2 -1 2 0 0 -3 -2 4 -1 -3 -2 -2 -1 4 1 -3 -2 0 -2 -3 -1 1 5 -3 -4 -3 -3 12 -3 -2 -2 2 -1 1 0 -3 -2 2 7 -2 -2 -4 0 -3 -1 0 BLAST query protein 0 -1 0 -4 -2 -2 -1 -5 -3 -2 -1 -3 -3 -1 0 -2 -1 -2 -2 -1 0 0 -1 -2 -3 -2 6 -2 -4 -4 -1 -2 -2 -1 -1 -3 -3 -2 -2 -3 2 -2 -1 -1 0 -2 -2 -2 0 -2 -1 -3 -2 -1 -2 -3 -1 -2 -1 -1 -3 -4 -2 1 3 -3 S -2 0 -3 -2 -3 1 -3 -2 -3 -3 1 1 -3 1 T -1 -1 -3 0 -1 -1 0 0 -2 -1 0 0

1 M 2 K 3 W 4 V 5 W 6 A 7 L 8 L 9 L 10 L 11 A 12 A 13 W 14 A 15 A 16 A. . . 37 S 38 G 39 T 40 W 41 Y 42 A A -1 -1 -3 0 -3 5 -2 -1 -1 -2 5 5 -2 3 2 4 R -2 1 -3 -3 -3 -2 -2 -2 -3 -2 -1 -2 N -2 0 -4 -3 -4 -2 -4 -3 -4 -4 -2 -2 -4 -1 0 -1 D -3 1 -5 -4 -5 -2 -4 -4 -2 -2 -4 -2 -1 -2 C -2 -4 -3 -1 -1 -2 -1 Q -1 2 -2 -3 -2 -1 -2 -3 -2 -2 -1 -1 -2 -1 E -2 4 -3 -3 -3 -1 -1 -3 -2 0 -1 G -3 -2 -3 -4 -3 0 -4 -4 0 0 -4 4 2 3 H -2 0 -3 -4 -3 -2 -3 -3 -2 -2 -3 -2 -1 -2 I 1 -3 -3 -2 2 2 -2 -2 1 -2 -3 -2 L 2 -3 -2 1 -2 -2 4 4 -2 -2 4 -2 -3 -2 K -2 3 -3 -3 -3 -1 -1 -3 -1 0 -1 M 6 -2 -2 1 -2 -1 2 2 -1 -1 2 -2 -2 -1 F 0 -4 1 -1 1 -3 0 0 -3 -3 1 -3 -3 -3 P -3 -1 -4 -3 -4 -1 -3 -3 -1 -1 -1 S -2 0 -3 -2 -3 1 -3 -2 -3 -3 1 1 -3 1 T -1 -1 -3 0 -1 -1 0 0 -2 -1 0 0 W -2 -3 12 -3 -2 -2 -3 -3 7 -3 -3 -3 Y -1 -2 2 -1 2 -2 -1 0 -1 -1 -2 -2 0 -3 -2 -2 V 1 -3 -3 4 -3 0 1 3 2 1 0 0 0 -1 -2 -1 2 0 0 -3 -2 4 -1 -3 -2 -2 0 -1 0 -4 -2 -2 -1 -5 -3 -2 -1 -3 -3 -1 0 -2 -1 -2 -2 -1 0 0 -1 -2 -3 -2 6 -2 -4 -4 -1 -2 -2 -1 -1 -3 -3 -2 -2 -3 2 -2 -1 -1 0 -2 -2 -2 0 -2 -1 -3 -2 -1 -2 -3 -1 -2 -1 -1 -3 -4 -2 1 3 -3 -1 4 1 -3 -2 0 -2 -3 -1 1 5 -3 -4 -3 -3 12 -3 -2 -2 2 -1 1 0 -3 -2 2 7 -2 -2 -4 0 -3 -1 0
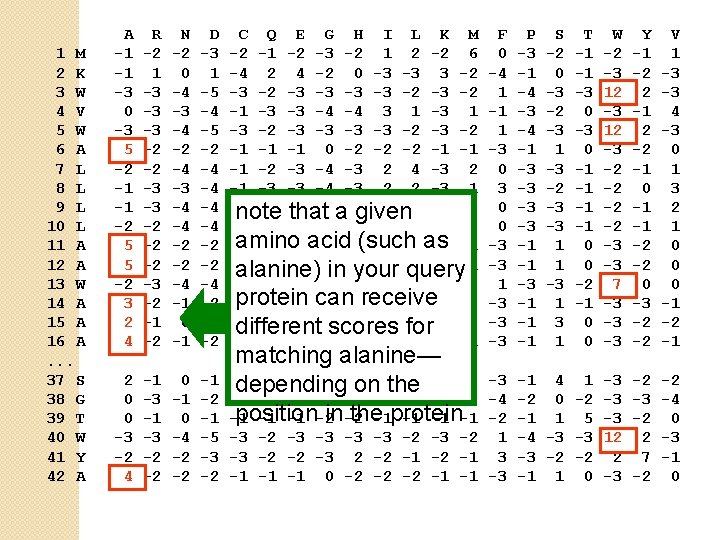
1 M 2 K 3 W 4 V 5 W 6 A 7 L 8 L 9 L 10 L 11 A 12 A 13 W 14 A 15 A 16 A. . . 37 S 38 G 39 T 40 W 41 Y 42 A A -1 -1 -3 0 -3 5 -2 -1 -1 -2 5 5 -2 3 2 4 R -2 1 -3 -3 -3 -2 -2 -2 -3 -2 -1 -2 N -2 0 -4 -3 -4 -2 -4 -3 -4 -4 -2 -2 -4 -1 0 -1 D -3 1 -5 -4 -5 -2 -4 -4 -2 -2 -4 -2 -1 -2 C Q E G H I L K M -2 -1 -2 -3 -2 1 2 -2 6 -4 2 4 -2 0 -3 -3 3 -2 -3 -3 -2 -1 -3 -3 -4 -4 3 1 -3 -2 -3 -2 -1 -1 -1 0 -2 -2 -2 -1 -1 -1 -2 -3 -4 -3 2 -1 -3 -3 -4 -3 2 2 -3 1 -1 -2 -3 -4 a-3 2 4 -3 2 note that given -1 -2 -3 -4 -3 2 amino -1 -1 -1 acid 0 -2(such -2 -2 as -1 -1 -1 0 in-2 -2 -2 -1 -1 alanine) your query -2 -2 -3 -4 -3 1 4 -3 2 protein -1 -1 -2 can 4 -2 receive -2 -2 -1 -2 -2 2 0 2 scores -1 -3 -3 different for 0 -2 -1 -1 -1 3 -2 -2 -2 -1 -1 2 0 0 -3 -2 4 -1 -3 -2 -2 0 -1 0 -4 -2 -2 -1 -3 -2 -4 -1 -2 -5 -3 -2 -3 -2 1 -3 -3 -2 -2 -3 2 -2 -1 3 -2 -1 -1 -1 0 -2 -2 -2 -1 -1 -3 matching alanine— -1 0 0 0 -1 -3 0 -2 depending on-2 the -3 -2 -2 6 -2 -4 -4 -2 -3 position in -2 the-1 protein -1 -1 -1 -2 -1 -1 -1 F 0 -4 1 -1 1 -3 0 0 -3 -3 1 -3 -3 -3 P -3 -1 -4 -3 -4 -1 -3 -3 -1 -1 -1 S -2 0 -3 -2 -3 1 -3 -2 -3 -3 1 1 -3 1 T -1 -1 -3 0 -1 -1 0 0 -2 -1 0 0 W -2 -3 12 -3 -2 -2 -3 -3 7 -3 -3 -3 Y -1 -2 2 -1 2 -2 -1 0 -1 -1 -2 -2 0 -3 -2 -2 V 1 -3 -3 4 -3 0 1 3 2 1 0 0 0 -1 -2 -1 -1 4 1 -3 -2 0 -2 -3 -1 1 5 -3 -4 -3 -3 12 -3 -2 -2 2 -1 1 0 -3 -2 2 7 -2 -2 -4 0 -3 -1 0
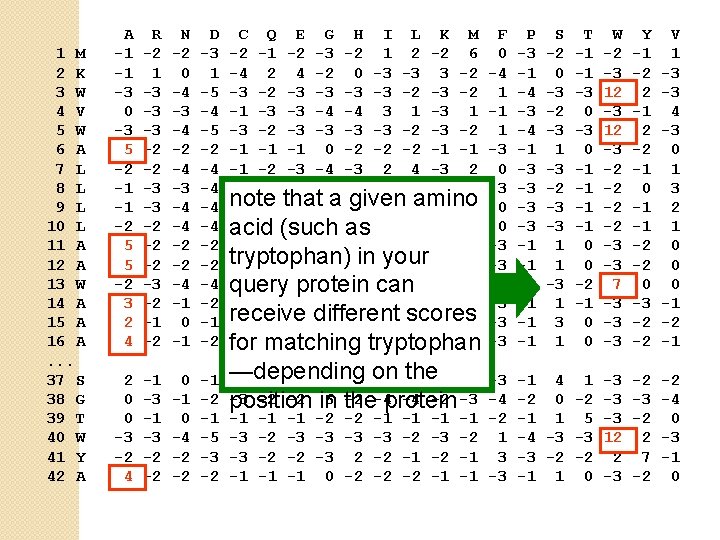
1 M 2 K 3 W 4 V 5 W 6 A 7 L 8 L 9 L 10 L 11 A 12 A 13 W 14 A 15 A 16 A. . . 37 S 38 G 39 T 40 W 41 Y 42 A A -1 -1 -3 0 -3 5 -2 -1 -1 -2 5 5 -2 3 2 4 R -2 1 -3 -3 -3 -2 -2 -2 -3 -2 -1 -2 N -2 0 -4 -3 -4 -2 -4 -3 -4 -4 -2 -2 -4 -1 0 -1 D -3 1 -5 -4 -5 -2 -4 -4 -2 -2 -4 -2 -1 -2 C Q E G H I L K M -2 -1 -2 -3 -2 1 2 -2 6 -4 2 4 -2 0 -3 -3 3 -2 -3 -3 -2 -1 -3 -3 -4 -4 3 1 -3 -2 -3 -2 -1 -1 -1 0 -2 -2 -2 -1 -1 -1 -2 -3 -4 -3 2 -1 -3 -3 -4 -3 2 2 -3 1 note given -1 -2 that -3 -4 a -3 2 4 amino -3 2 -1 -2 (such -3 -4 -3 acid as 2 4 -3 2 -1 -1 -1 0 -2 -2 -2 -1 -1 tryptophan) your -1 -1 -1 0 -2 in-2 -2 -1 -1 -2 -2 -3 -4 -3 can 1 4 -3 2 query protein -1 -1 -2 4 -2 -2 -2 -1 -2 receive scores -2 2 0 different 2 -1 -3 -3 0 -2 -1 -1 3 -2 tryptophan -2 -2 -1 -1 for -1 matching F 0 -4 1 -1 1 -3 0 0 -3 -3 1 -3 -3 -3 P -3 -1 -4 -3 -4 -1 -3 -3 -1 -1 -1 S -2 0 -3 -2 -3 1 -3 -2 -3 -3 1 1 -3 1 T -1 -1 -3 0 -1 -1 0 0 -2 -1 0 0 W -2 -3 12 -3 -2 -2 -3 -3 7 -3 -3 -3 Y -1 -2 2 -1 2 -2 -1 0 -1 -1 -2 -2 0 -3 -2 -2 V 1 -3 -3 4 -3 0 1 3 2 1 0 0 0 -1 -2 -1 2 0 0 -3 -2 4 -1 -3 -2 -2 0 -1 0 -4 -2 -2 -1 -5 -3 -2 —depending the 0 -2 -1 0 0 0 -1 on -2 -3 -3 -2 -2 in 6 the -2 -4 -4 -2 -3 position protein -1 -1 -1 -2 -2 -1 -1 -3 -2 -3 -2 -2 -3 2 -2 -1 -1 0 -2 -2 -2 -1 -1 -3 -4 -2 1 3 -3 -1 4 1 -3 -2 0 -2 -3 -1 1 5 -3 -4 -3 -3 12 -3 -2 -2 2 -1 1 0 -3 -2 2 7 -2 -2 -4 0 -3 -1 0
![PSI-BLAST is performed in five steps [1] Select a query and search it against PSI-BLAST is performed in five steps [1] Select a query and search it against](http://slidetodoc.com/presentation_image_h2/4bd99b349f9089d951029c83b9f90bcb/image-78.jpg)
PSI-BLAST is performed in five steps [1] Select a query and search it against a protein databas [2] PSI-BLAST constructs a multiple sequence alignment then creates a “profile” or specialized position-specific scoring matrix (PSSM) [3] The PSSM is used as a query against the database B&FG 3 e Page 172
![PSI-BLAST is performed in five steps [1] Select a query and search it against PSI-BLAST is performed in five steps [1] Select a query and search it against](http://slidetodoc.com/presentation_image_h2/4bd99b349f9089d951029c83b9f90bcb/image-79.jpg)
PSI-BLAST is performed in five steps [1] Select a query and search it against a protein databas [2] PSI-BLAST constructs a multiple sequence alignment then creates a “profile” or specialized position-specific scoring matrix (PSSM) [3] The PSSM is used as a query against the database [4] PSI-BLAST estimates statistical significance (E value B&FG 3 e Page 172

![PSI-BLAST is performed in five steps [1] Select a query and search it against PSI-BLAST is performed in five steps [1] Select a query and search it against](http://slidetodoc.com/presentation_image_h2/4bd99b349f9089d951029c83b9f90bcb/image-81.jpg)
PSI-BLAST is performed in five steps [1] Select a query and search it against a protein databas [2] PSI-BLAST constructs a multiple sequence alignment then creates a “profile” or specialized position-specific scoring matrix (PSSM) [3] The PSSM is used as a query against the database [4] PSI-BLAST estimates statistical significance (E value [5] Repeat steps [3] and [4] iteratively, typically 5 times. At each new search, a new profile is used as the query. B&FG 3 e Page 172

Position-specific scoring matrix (PSSM) B&FG 3 e Fig. 5. 3 Page 173

PSI-BLAST: dramatic increase in number of hits Given this query, a standard BLASTP search would produce B&FG 3 e about 9 hits with low expect values. This PSI-BLAST search Table 5. 2 produces >200 hits after 3 or 4 iterations. Page 174

Note that PSI-BLAST E values can improve dramatically! After 1 st iteration: Expect = 4 e-04 Alignment length = 87 amino acids After 2 nd iteration: Expect = 1 e-36 Alignment length = 110 amino acids After 3 rd iteration: Expect = 2 e-33 Alignment length = 146 amino acids B&FG 3 e Fig. 5. 4 Page 175
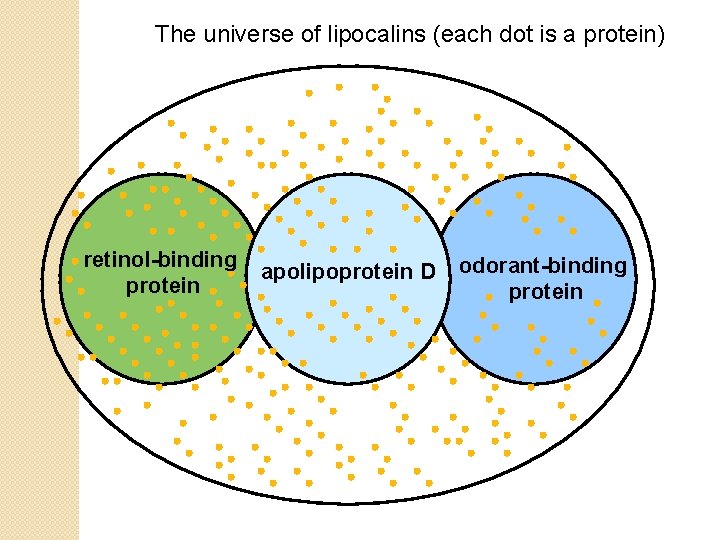
The universe of lipocalins (each dot is a protein) retinol-binding protein apolipoprotein D odorant-binding protein
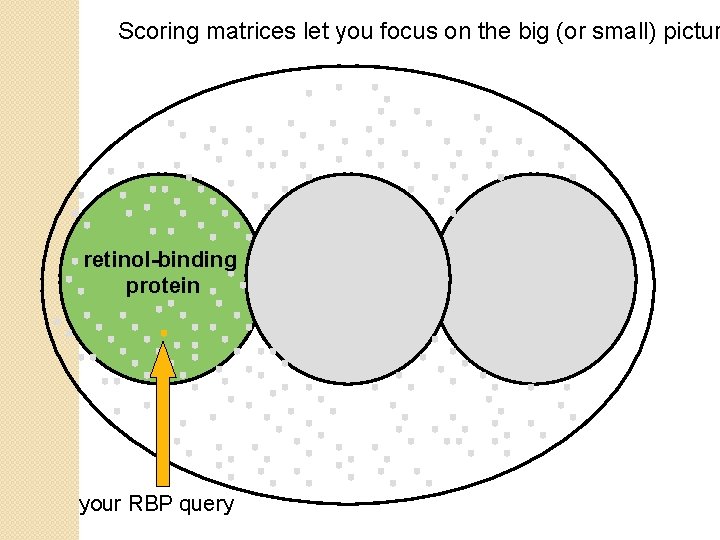
Scoring matrices let you focus on the big (or small) pictur retinol-binding protein your RBP query
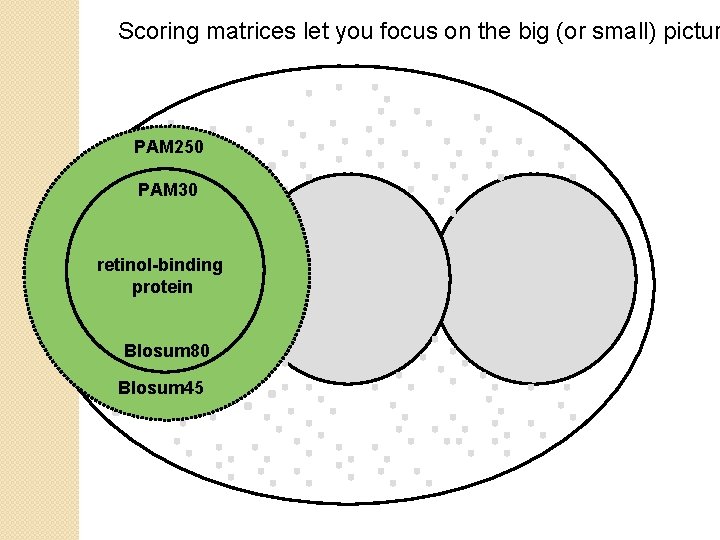
Scoring matrices let you focus on the big (or small) pictur PAM 250 PAM 30 retinol-binding protein Blosum 80 Blosum 45
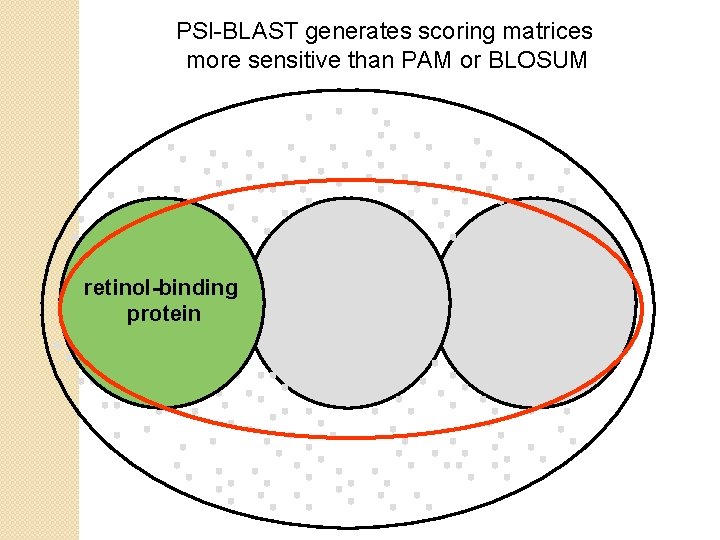
PSI-BLAST generates scoring matrices more sensitive than PAM or BLOSUM retinol-binding protein
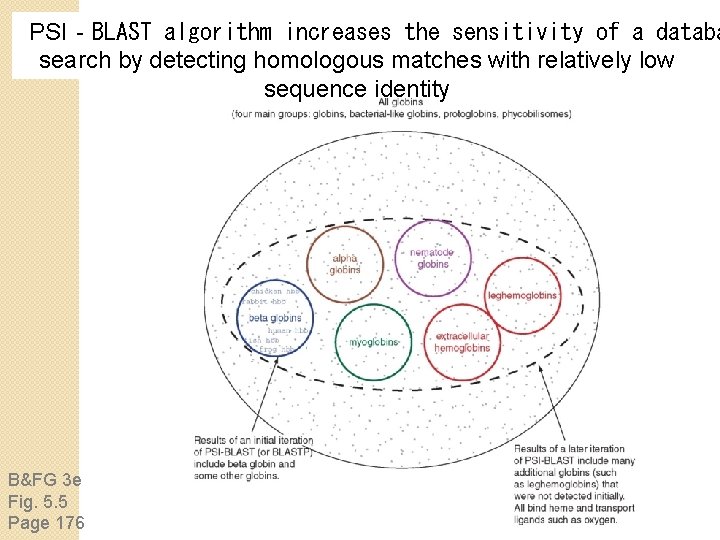
PSI‐BLAST algorithm increases the sensitivity of a databa search by detecting homologous matches with relatively low sequence identity B&FG 3 e Fig. 5. 5 Page 176

PSI-BLAST: the problem of corruption In PSI-BLAST once a match is incorporated into a PSSM it will never be removed, even if it is wrong (i. e. even if it is a false positive that is not truly homologous to the query). Not only will it stay, it may lead to the inclusion of many other related false positive hits. There are three main approaches to removing false positives: B&FG 3 e Page 177 (1) Filter biased amino acid regions. (This is an option in BLAST. ) (2) Lower the expect value threshold to make the search more stringent. (3) Visually inspect the output from each PSI-BLAST iteration and remove suspicious matches (by unchecking the corresponding boxes).
- Slides: 90