BisulfiteSequencing Theory and Quality Control Felix Krueger felix

Bisulfite-Sequencing Theory and Quality Control Felix Krueger felix. krueger@babraham. ac. uk November 2017

• • 9: 30 -10: 30 -11: 15 11: 30 -12: 30 13: 00 -14: 00 -14: 30 -15: 15 -17: 00 practical 2 Bisulfite-Seq theory and QC Mapping and QC practical Visualising and Exploring talk Lunch Methylation tools in Seq. Monk Visualising and Exploring practical Differential methylation talk &
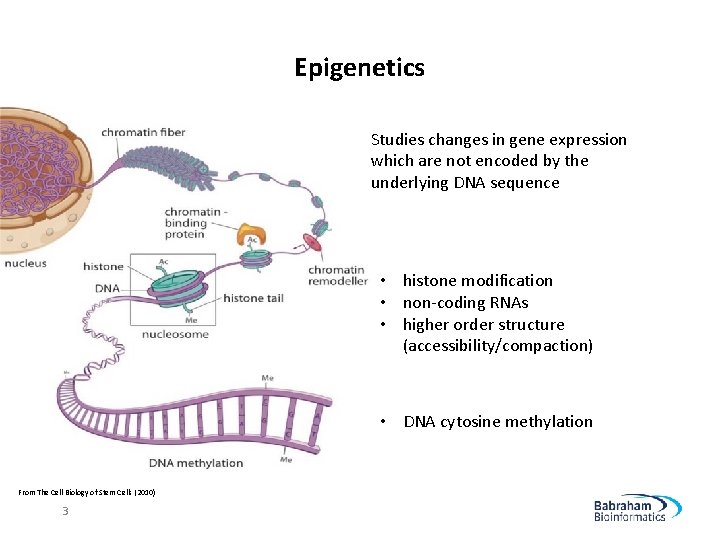
Epigenetics Studies changes in gene expression which are not encoded by the underlying DNA sequence • histone modification • non-coding RNAs • higher order structure (accessibility/compaction) • DNA cytosine methylation From The Cell Biology of Stem Cells (2010) 3
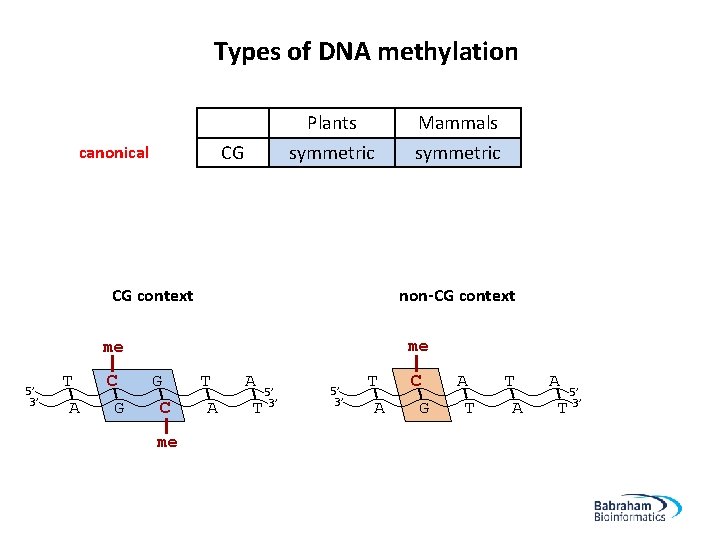
Types of DNA methylation canonical non-canonical (mammals) CG CHH Plants symmetric asymmetric Mammals symmetric asymmetric H = T/A/C CG context non-CG context me me 5’ 3’ T A C G G C me T A A 5’ T 3’ 5’ 3’ T A C G A T T A A 5’ T 3’
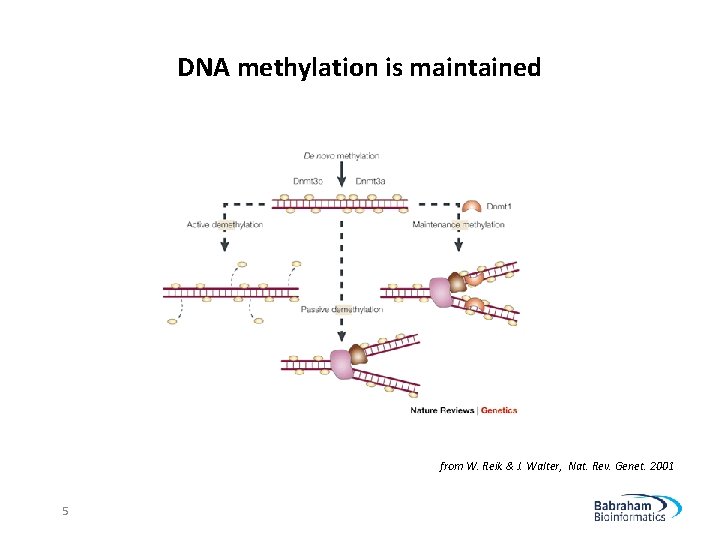
DNA methylation is maintained from W. Reik & J. Walter, Nat. Rev. Genet. 2001 5
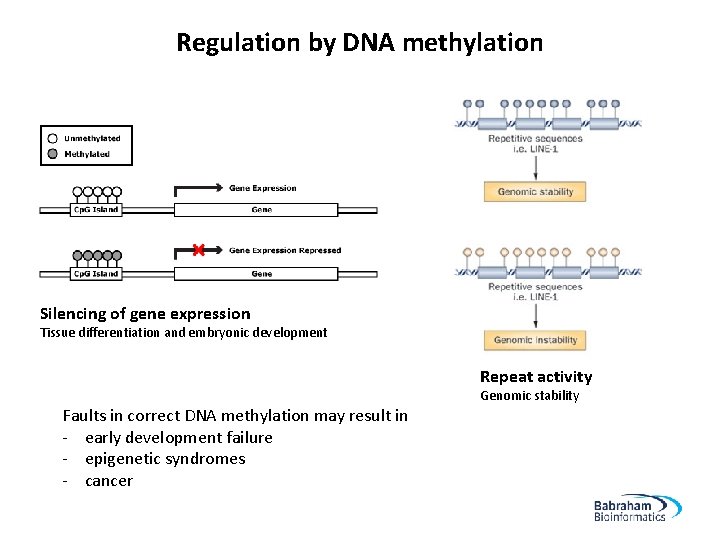
Regulation by DNA methylation Silencing of gene expression Tissue differentiation and embryonic development Repeat activity Genomic stability Faults in correct DNA methylation may result in - early development failure - epigenetic syndromes - cancer
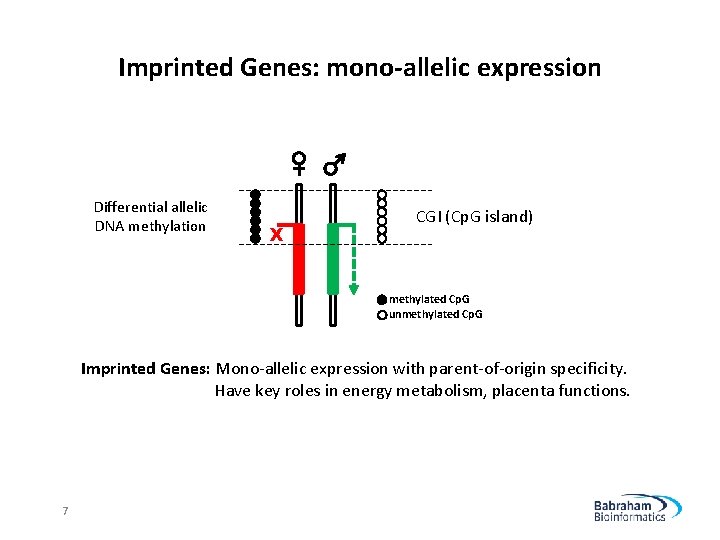
Imprinted Genes: mono-allelic expression Differential allelic DNA methylation X CGI (Cp. G island) methylated Cp. G unmethylated Cp. G Imprinted Genes: Mono-allelic expression with parent-of-origin specificity. Have key roles in energy metabolism, placenta functions. 7
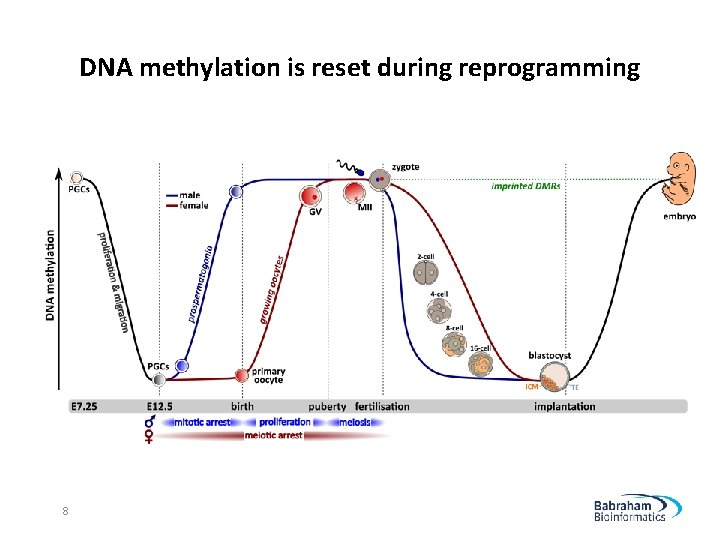
DNA methylation is reset during reprogramming 8
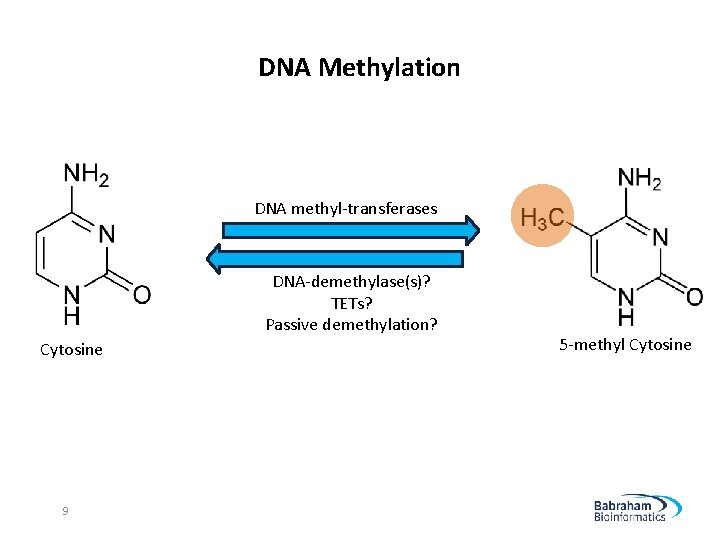
DNA Methylation DNA methyl-transferases DNA-demethylase(s)? TETs? Passive demethylation? Cytosine 9 5 -methyl Cytosine
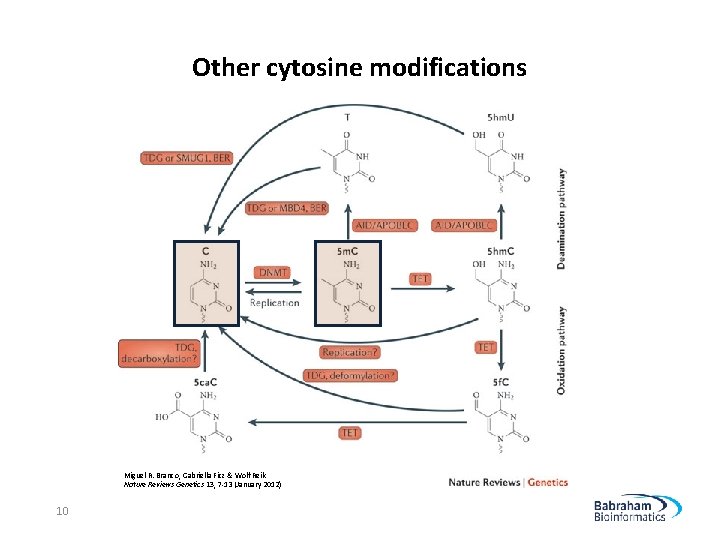
Other cytosine modifications Miguel R. Branco, Gabriella Ficz & Wolf Reik Nature Reviews Genetics 13, 7 -13 (January 2012) 10
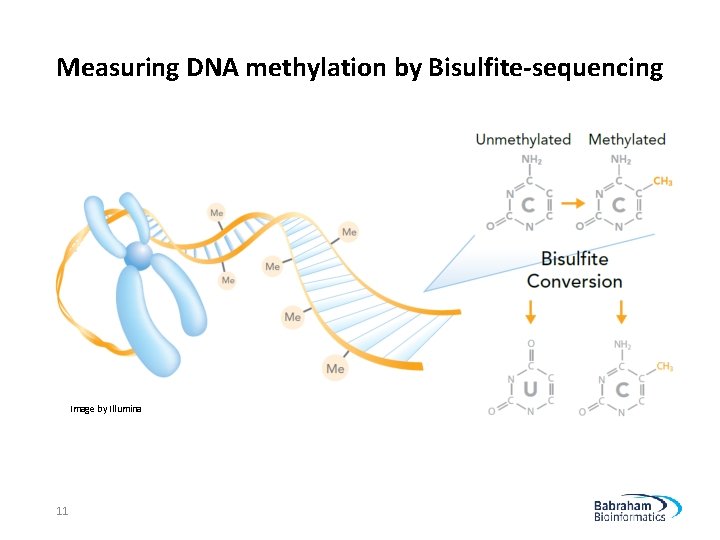
Measuring DNA methylation by Bisulfite-sequencing Image by Illumina 11
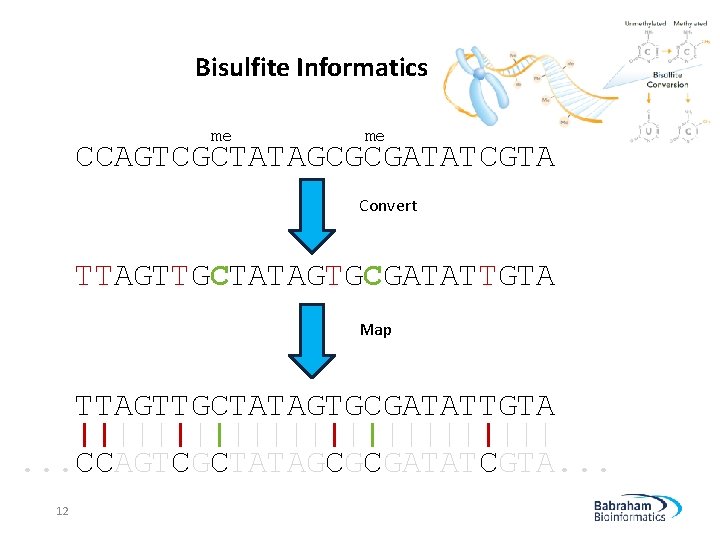
Bisulfite Informatics me me CCAGTCGCTATAGCGCGATATCGTA Convert TTAGTTGCTATAGTGCGATATTGTA Map TTAGTTGCTATAGTGCGATATTGTA |||||||||||||. . . CCAGTCGCTATAGCGCGATATCGTA. . . 12
BS-Seq Analysis Workflow Explore and understand your data Sequencing 13 Processing pipeline Methylation Analysis
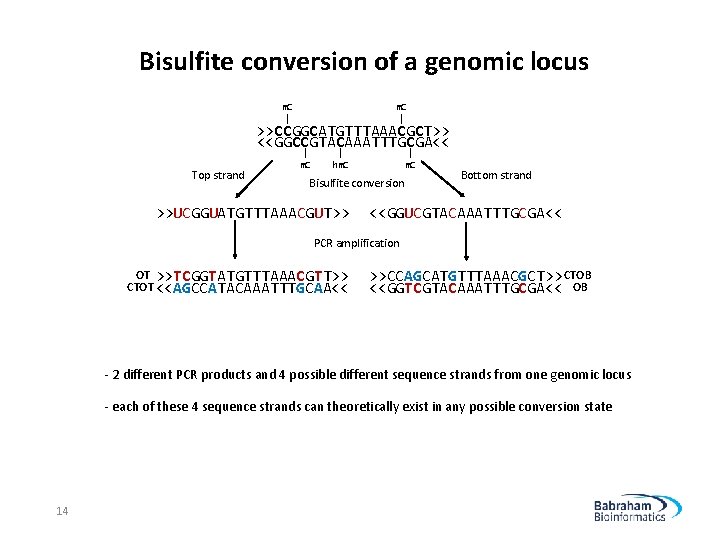
Bisulfite conversion of a genomic locus m. C | >>CCGGCATGTTTAAACGCT>> <<GGCCGTACAAATTTGCGA<< Top strand | m. C | hm. C | m. C Bisulfite conversion >>UCGGUATGTTTAAACGUT>> Bottom strand <<GGUCGTACAAATTTGCGA<< PCR amplification OT >>TCGGTATGTTTAAACGTT>> CTOT <<AGCCATACAAATTTGCAA<< >>CCAGCATGTTTAAACGCT>> CTOB <<GGTCGTACAAATTTGCGA<< OB - 2 different PCR products and 4 possible different sequence strands from one genomic locus - each of these 4 sequence strands can theoretically exist in any possible conversion state 14
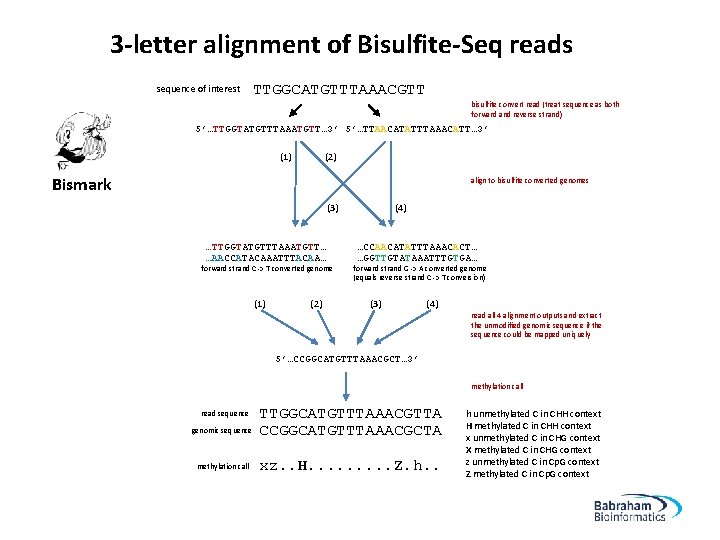
3 -letter alignment of Bisulfite-Seq reads sequence of interest TTGGCATGTTTAAACGTT bisulfite convert read (treat sequence as both forward and reverse strand) 5’…TTGGTATGTTTAAATGTT… 3’ 5’…TTAACATATTTAAACATT… 3’ (1) (2) Bismark align to bisulfite converted genomes (3) …TTGGTATGTTTAAATGTT… …AACCATACAAATTTACAA… forward strand C -> T converted genome (1) (2) (4) …CCAACATATTTAAACACT… …GGTTGTATAAATTTGTGA… forward strand G -> A converted genome (equals reverse strand C -> T conversion) (3) (4) read all 4 alignment outputs and extract the unmodified genomic sequence if the sequence could be mapped uniquely 5’…CCGGCATGTTTAAACGCT… 3’ methylation call read sequence genomic sequence methylation call TTGGCATGTTTAAACGTTA CCGGCATGTTTAAACGCTA xz. . H. . Z. h. . h unmethylated C in CHH context H methylated C in CHH context x unmethylated C in CHG context X methylated C in CHG context z unmethylated C in Cp. G context Z methylated C in Cp. G context
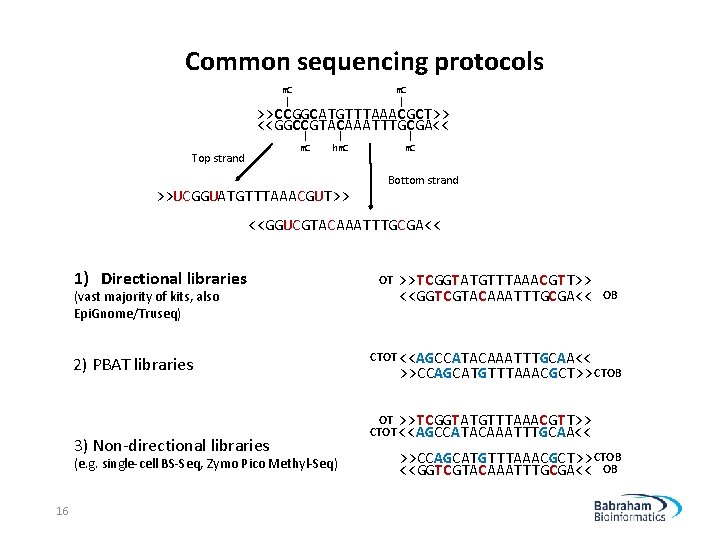
Common sequencing protocols m. C | >>CCGGCATGTTTAAACGCT>> <<GGCCGTACAAATTTGCGA<< | m. C Top strand | hm. C >>UCGGUATGTTTAAACGUT>> | m. C Bottom strand <<GGUCGTACAAATTTGCGA<< 1) Directional libraries (vast majority of kits, also Epi. Gnome/Truseq) 2) PBAT libraries 3) Non-directional libraries (e. g. single-cell BS-Seq, Zymo Pico Methyl-Seq) 16 OT >>TCGGTATGTTTAAACGTT>> <<GGTCGTACAAATTTGCGA<< OB CTOT <<AGCCATACAAATTTGCAA<< >>CCAGCATGTTTAAACGCT>> CTOB OT >>TCGGTATGTTTAAACGTT>> CTOT <<AGCCATACAAATTTGCAA<< >>CCAGCATGTTTAAACGCT>> CTOB <<GGTCGTACAAATTTGCGA<< OB
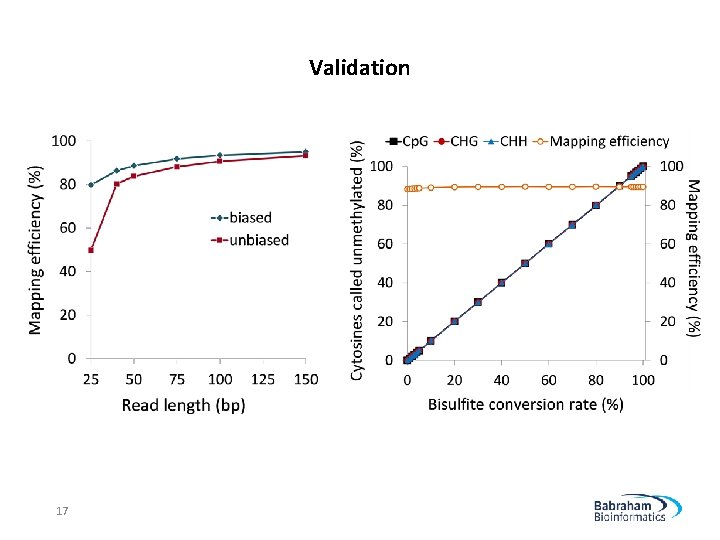
Validation 17
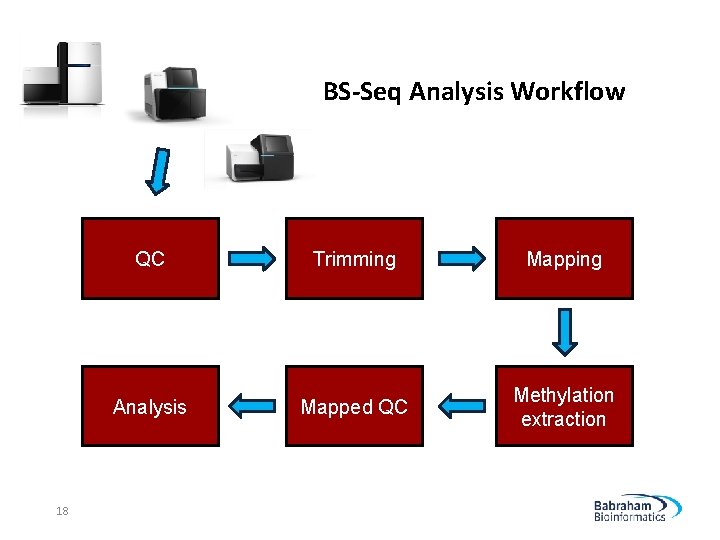
BS-Seq Analysis Workflow 18 QC Trimming Mapping Analysis Mapped QC Methylation extraction

Raw Sequence Data . . . up to 1, 000, 000 lines per lane 19

Part I: Initial QC What does QC tell you about your library? # of sequences Basecall qualities Base composition Potential contaminants Expected duplication rate • • • 20
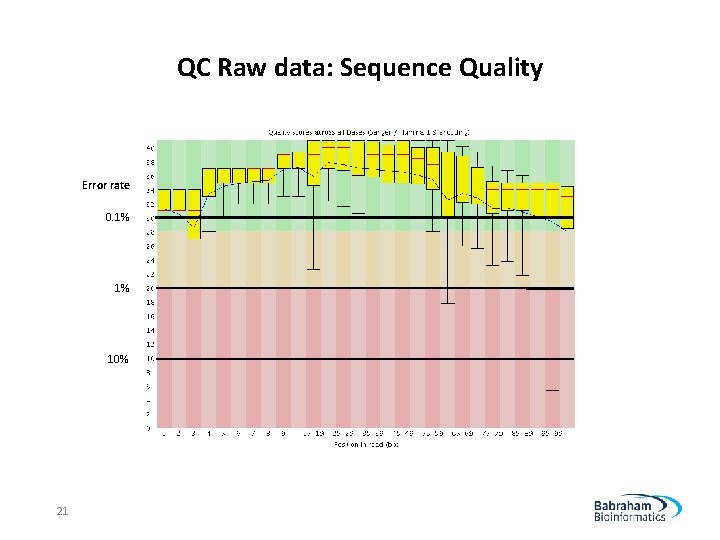
QC Raw data: Sequence Quality Error rate 0. 1% 10% 21
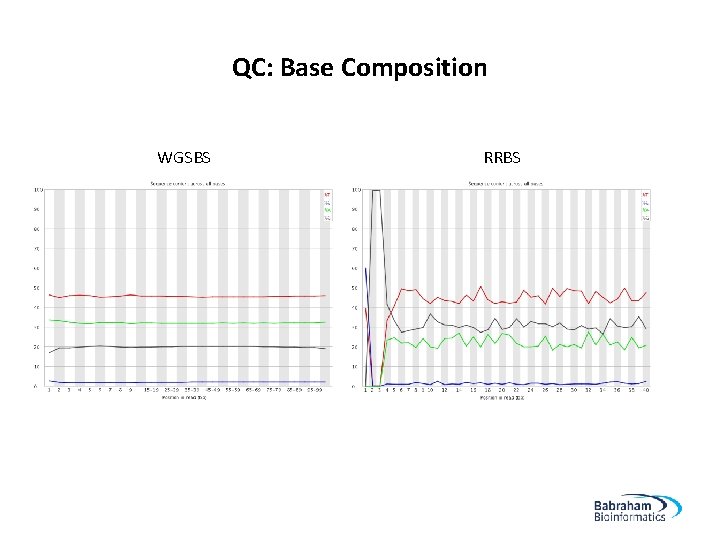
QC: Base Composition WGSBS RRBS 22
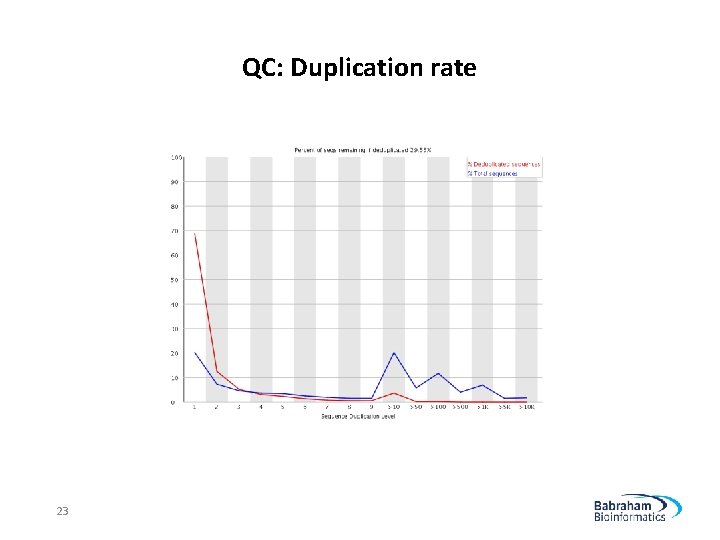
QC: Duplication rate 23
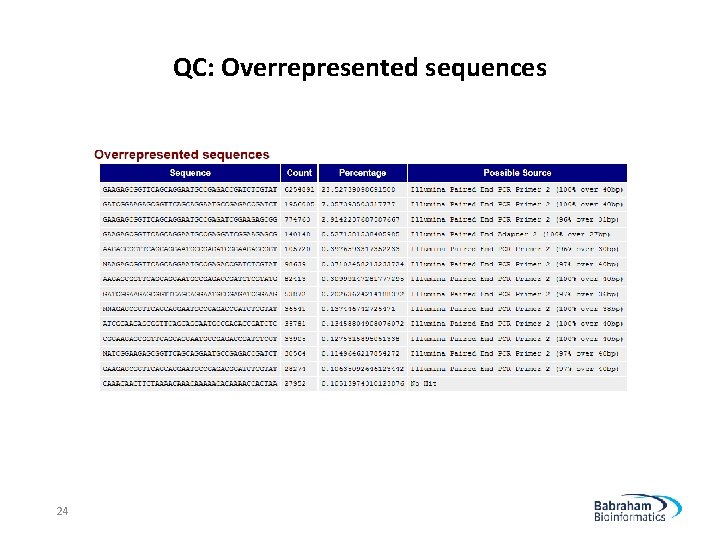
QC: Overrepresented sequences 24
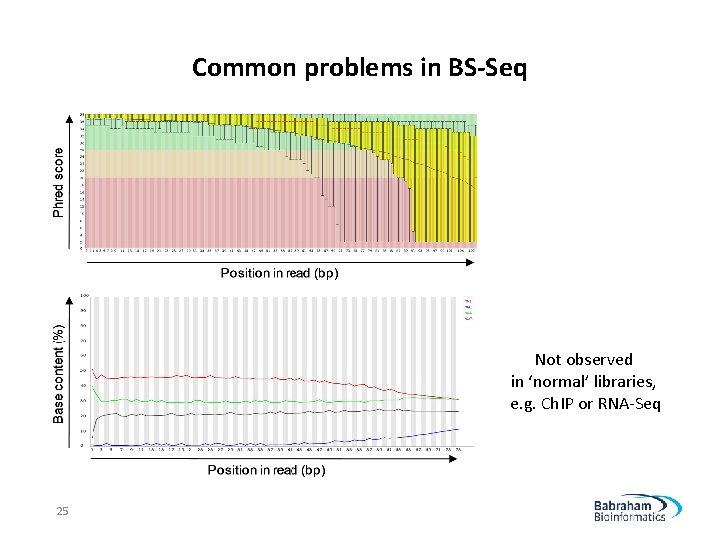
Common problems in BS-Seq Not observed in ‘normal’ libraries, e. g. Ch. IP or RNA-Seq 25
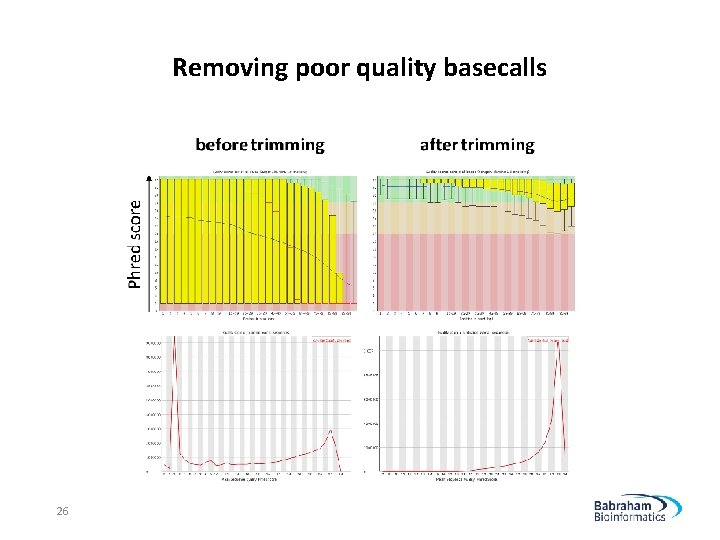
Removing poor quality basecalls 26
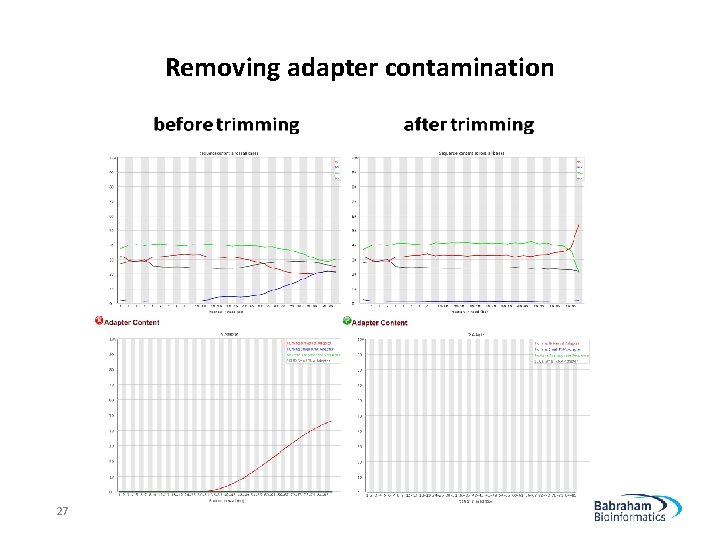
Removing adapter contamination 27

Adapter trimming (Illumina adapter: AGATCGGAAGAGC) B: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAGGAT A: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAGGAT partial match full match B: AGATCTTTTATTCGGTAGGATAGATCGGAAGAGCXXXXXXXX A: AGATCTTTTATTCGGTAGGAT B: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAGATC A: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGG B: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAGAGA A: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAG B: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAGGAG A: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAGG B: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAGGAA A: AGATCTTTTATTCGGTAGGATTAGCGGTAGTTATTTTGGAGGA 28

Summary Adapter/Quality Trimming Important to trim because failure to do so might result in: § Low mapping efficiency § Mis-alignments § Errors in methylation calls since adapters are methylated § Basecall errors tend toward 50% (C: m. C) 29

Part II: Sequence alignment – Bismark primary alignment output (BAM file) chromosome position Read 1 HISEQ 2000 -06: 366: C 3 G 4 NACXX: 3: 1101: 1316: 2067_1: N: 0: 99 16 71322125 255 100 M = 71322232 207 sequence NTTATTTAGTTTTTTAGGGTTTGTGTGTAGGAGTGTGGGAATTATGTTTTTTATGGTTGATATTTAAAAGTGAGTATAAATTATATTTTTTT #1=DDDDDAAFFHIIIA: <FGHCCEFGHD? CFFBBBGEHHGHIII<FEHIIIII==DE? ? EHHFHEEEEEEEC>; >66; @CDEEEDCEEEEEEEDDDCBB quality NM: i: 14 XX: Z: G 8 C 2 C 7 C 21 C 13 C 6 CC 1 C 17 CC 3 C 4 CC 4 XM: Z: . . h. . . . x. . . . . h. . . x. . . hh. h. . . . hh. . XR: Z: CT XG: Z: CT XA: Z: 1 HISEQ 2000 -06: 366: C 3 G 4 NACXX: 3: 1101: 1316: 2067_1: N: 0: 147 16 71322232 255 100 M = 71322125 -207 GGTTATTTAGGGTTATTGTTTTAGAGTTTTATTGTTGTGAACAGATATATGATTAAGGTAATTTTTATAAGGATAATATTTAATTGGAGTTGGTT CCCEEECADCFFFFHHGHGHIIGIHFIJJIJIHFGHGGGEHIJIIJGIGFJJJJJIGJJJJIIIJJIJIJJJJJJIJHHHHHFFFFFCCC NM: i: 21 XX: Z: 2 G 2 CC 1 C 11 C 11 C 2 C 10 C 1 C 4 C 2 C 1 C 3 C 5 C 2 C 12 C 3 C 1 XM: Z: . . . hh. h. h. x. . . h. . . x. . . . X. . . h. h. . . x. . . h. XR: Z: GA XG: Z: CT XB: Z: 1 Read 2 methylation call 30
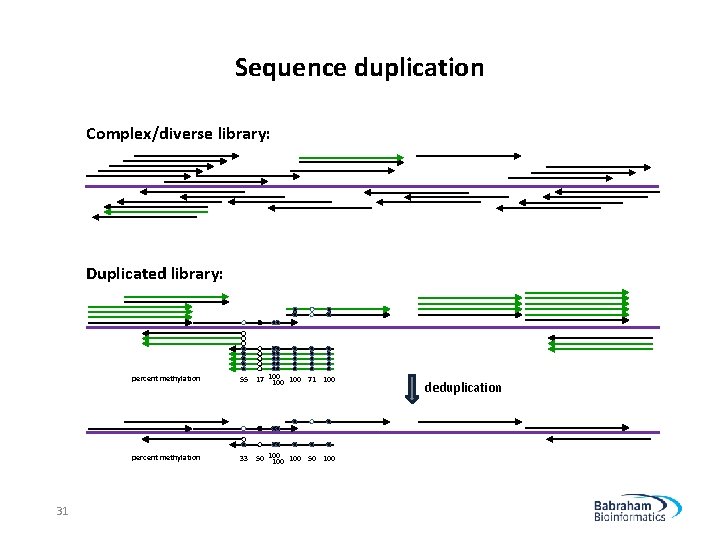
Sequence duplication Complex/diverse library: Duplicated library: 31 percent methylation 55 17 100 100 71 100 percent methylation 33 50 100 100 50 100 deduplication

Deduplication - considerations Advisable for large genomes and moderate coverage - unlikely to sequence several genuine copies of the same fragment amongst >5 bn possible fragments with different start sites - maximum coverage with duplication may still be (read length)-fold (even more with paired-end reads) NOT advisable for RRBS or other target enrichment methods where higher coverage is either desired or expected RRBS CCGG deduplication 32
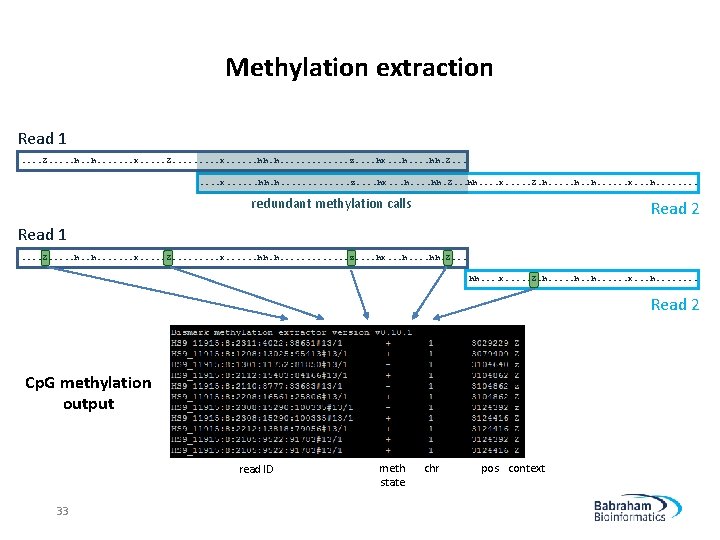
Methylation extraction Read 1. . Z. . . h. . . . x. . . Z. . x. . . hh. h. . . . z. . hx. . . hh. Z. . . hh. . x. . . Z. h. . . . x. . . h. . . . redundant methylation calls Read 2 Read 1. . Z. . . h. . . . x. . . Z. . x. . . hh. h. . . z. . hx. . . hh. Z. . . hh. . x. . . Z. h. . . . x. . . h. . . . Read 2 Cp. G methylation output read ID 33 meth state chr pos context

Methylation extraction I Cp. G methylation output bismark 2 bed. Graph/coverage output chr 34 pos methylation percentage meth unmeth

Methylation extraction II coverage output coverage 2 cytosine optional: merge into Cp. G dinucleotide entities Genome wide Cp. G report chr 35 pos strand meth unmeth di-nuc tri-nuc
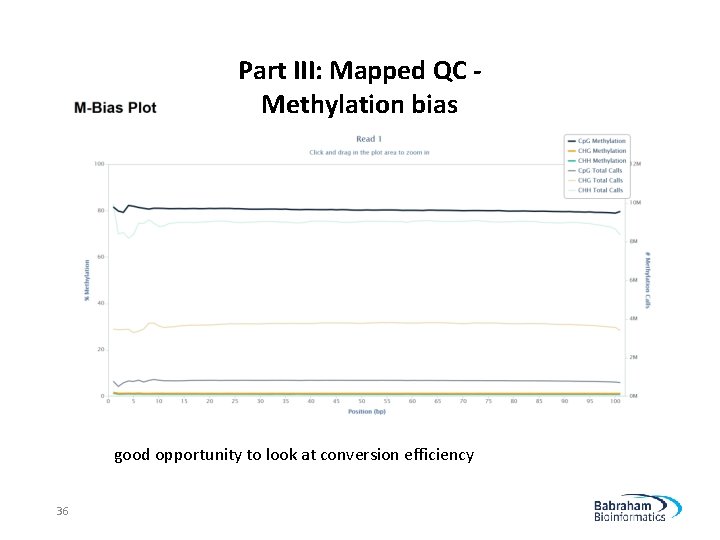
Part III: Mapped QC Methylation bias good opportunity to look at conversion efficiency 36
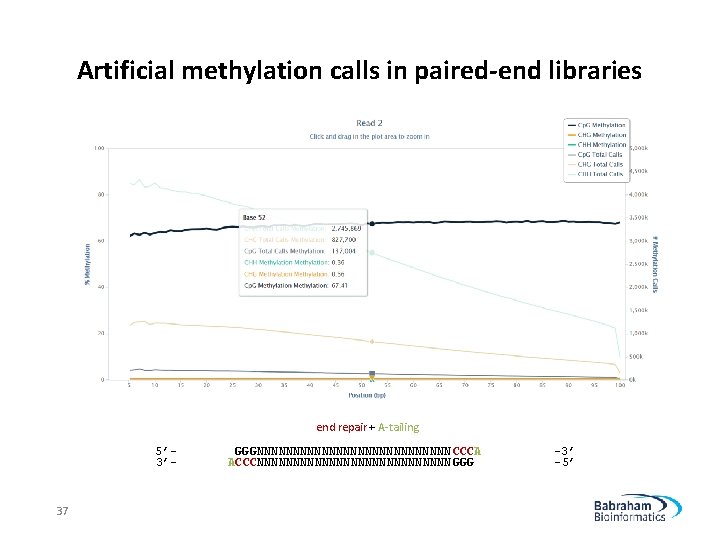
Artificial methylation calls in paired-end libraries end repair + A-tailing 5’ 3’- 37 GGGNNNNNNNNNNNNNN CCCA ACCCNNNNNNNNNNNNNN GGG -3’ -5’
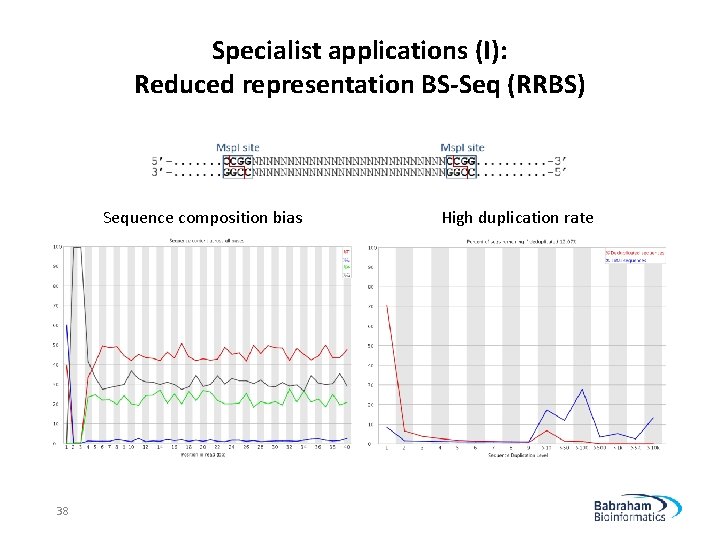
Specialist applications (I): Reduced representation BS-Seq (RRBS) Sequence composition bias 38 High duplication rate
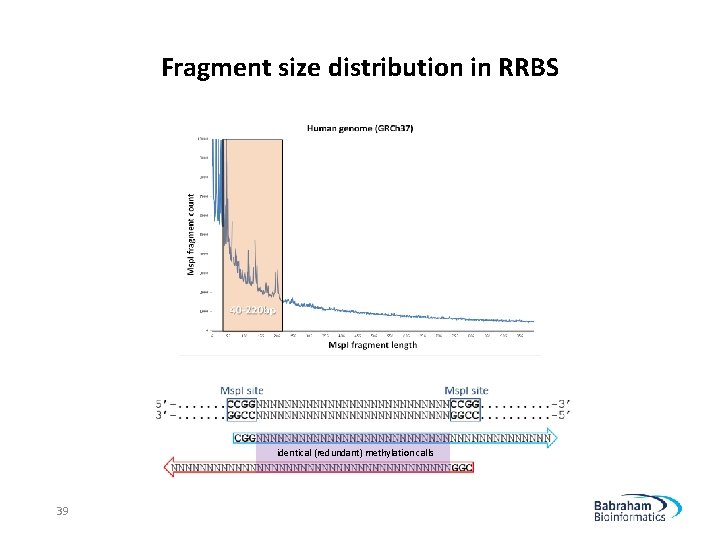
Fragment size distribution in RRBS identical (redundant) methylation calls 39
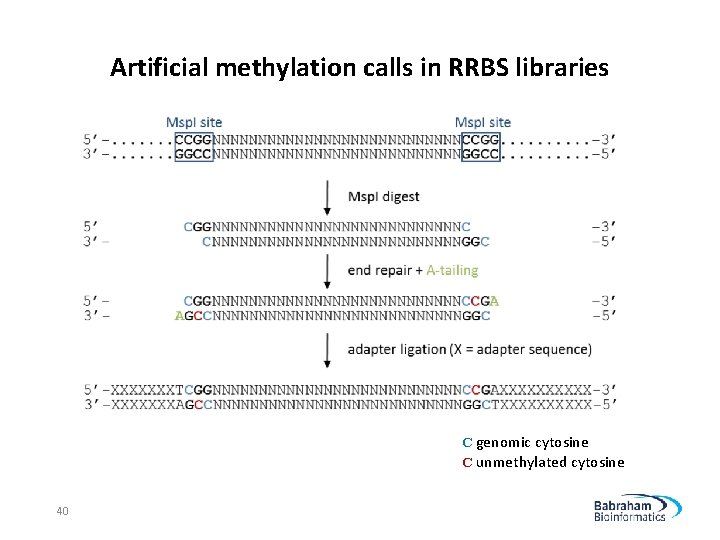
Artificial methylation calls in RRBS libraries C genomic cytosine C unmethylated cytosine 40
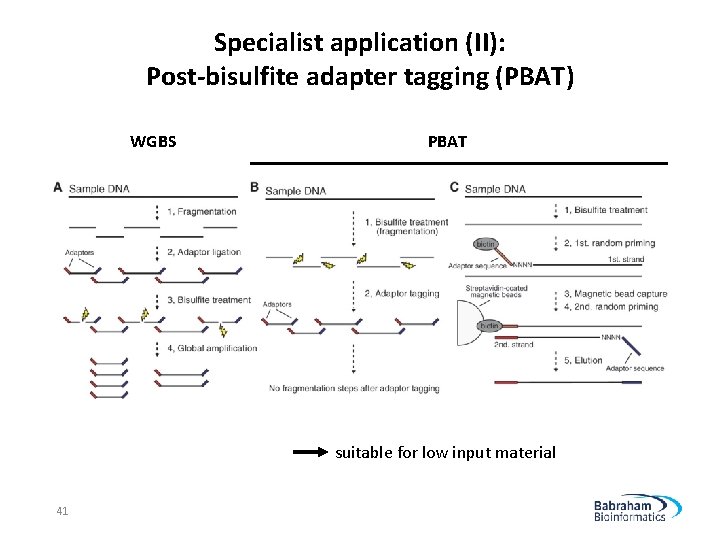
Specialist application (II): Post-bisulfite adapter tagging (PBAT) WGBS PBAT suitable for low input material 41
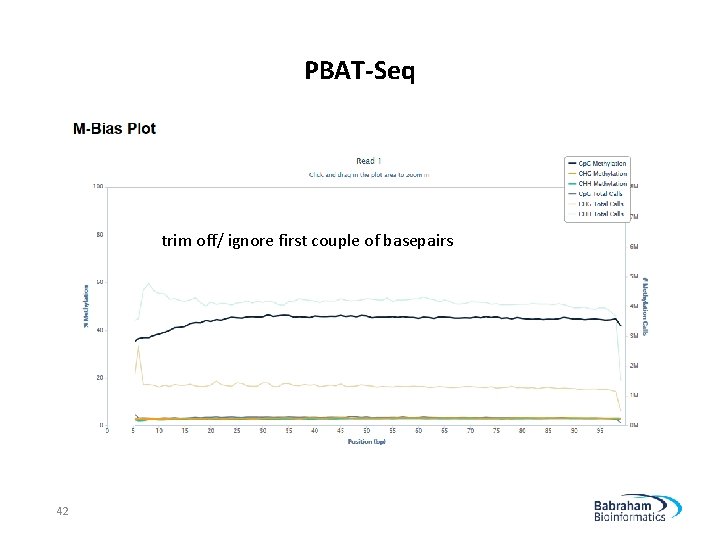
PBAT-Seq trim off/ ignore first couple of basepairs 42

Bismark User Guide https: //rawgit. com/Felix. Krueger/Bismark/master/Docs/Bismark_User_Guide. html 43
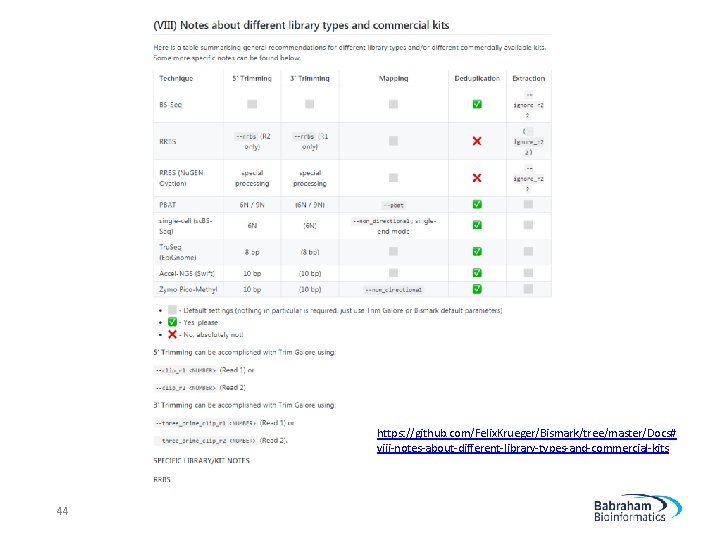
https: //github. com/Felix. Krueger/Bismark/tree/master/Docs# viii-notes-about-different-library-types-and-commercial-kits 44

https: //sequencing. qcfail. com/ 45

Bismark workflow Pre Alignment Fast. QC Trim Galore Alignment Bismark Initial quality control Adapter/quality trimming using Cutadapt; handles RRBS and paired-end reads; Trim Galore and RRBS User guide Output BAM Post Alignment Deduplication Methylation extractor optional Output individual cytosine methylation calls; optionally bed. Graph or genome-wide cytosine report M-bias analysis bismark 2 report Graphical HTML report generation Example: http: //www. bioinformatics. babraham. ac. uk/projects/bismark/PE_report. html protocol: Quality Control, trimming and alignment of Bisulfite-Seq data 46

Useful links • Fast. QC www. bioinformatics. babraham. ac. uk/projects/fastqc/ • Trim Galore www. bioinformatics. babraham. ac. uk/projects/trim_galore/ • Cutadapt https: //code. google. com/p/cutadapt/ • Bismark www. bioinformatics. babraham. ac. uk/projects/bismark/ • Bowtie http: //bowtie-bio. sourceforge. net/ • Bowtie 2 http: //bowtie-bio. sourceforge. net/bowtie 2/ • Seq. Monk www. bioinformatics. babraham. ac. uk/projects/seqmonk/ • Cluster Flow www. bioinformatics. babraham. ac. uk/projects/clusterflow/ protocol: Quality control, trimming and alignment of Bisulfite-Seq data http: //www. epigenesys. eu/en/protocols/bio-informatics/483 -quality-control-trimming-and-alignment-of-bisulfite-seq-data-prot-57 https: //sequencing. qcfail. com/ 47

Sierra: A web-based LIMS system for small sequencing facilities Seq. Monk: Genome browser, quantitation and data analysis Trim Galore! Quality and adapter trimming for (RRBS) libraries Fast. Q Screen: organism and contamination detection Bismark: Bisulfite-sequencing alignments and methylation calls Hi-C mapping ASAP: Allele-specific alignments Fast. QC: quality control for high throughput sequencing 48 www. bioinformatics. babraham. ac. uk
- Slides: 48