BIRN Image Ontology Requirements Maryann Martone Jeff Grethe

BIRN Image Ontology Requirements Maryann Martone & Jeff Grethe (UCSD) Bill Bug (Drexel U. College of Medicine) NCBO/NCOR Image Ontology Mtg - 3/2006

Biomedical Informatics Research Network A shared biomedical IT infrastructure to hasten the derivation of new understanding and treatment of disease through use of distributed knowledge • Collaboration between groups with different expertise and resources (technical, scientific, social and political) • Technical infrastructure to support collaboration (designed to be extensible to other biomedical communities) • Open access and dissemination of data and tools (i. e. Open Source) • Bringing transparent GRID Computing to Biomedical Research

BIRN Image Related Ontology Use Critical to BIRN-wide: • data integration • automated data validity & integrity • data pooling/binning/sorting/modes • data reduction • analysis • knowledge extraction across BIRN sites (29 locations - multiple labs/location) and with the rest of the world human experts & machine-based

BIRN Image Related Ontology Use A work in progress • BIRN IT Infrastructure maturing • Intra-BIRN and external collaborative practice maturing • Broad spectrum of Image-related tools & data • software & APIs for handling files - many image-specific • spatially-mapped neuro-data sets • visualization, spatial normalization & analysis • workflow - models of computation (e. g. , Ptolemy / Kepler)

BIRN Image Related Ontology Use A work in progress • Ontology tools & ontology use rapidly evolving • BONFIRE (tool) • BIRNLex (lexicon) • MIND (ontology) - only what we cannot re-use • BIRN ontology “best practices” - social engineering • BIRN Ontology use • expect to make mistakes • have a huge task - must finish start yesterday • BIRN Project - comprehensive Use Case set • research use of biomedical image related ontologies

BIRN Image Related Ontology Use We need your help • Image Ontology Consortium (burgeoning) • Beat us up • BUT BIRN will need • triage BIRN • chronic Rx Image Ontology Consortium

BIRN Image-Related Ontology Requirements Biological Specimen Imaging real-world bio-entities: a taxonomy of parts • molecules • multi-subunit macromolecular complexes • small : pre-promelanocortin, etc. • large: transcriptional apparatus, etc • sub-cellular components • small: Golgi apparatus, lysosomes, dendritic spines, transmitter vesicles, etc. • large: dendrites, axon hillock, nodes of Ranvier, presynaptic compartment, etc. • cells • mesoscopic cellular complexes • Purkinje cell body layer in the cerebellar lobes, gomeruli of the olfactory bulb, etc. • brain nuclei • macroscopic brain regions • organs & body regions • whole subject

BIRN Image-Related Ontology Requirements Imaging the Biological Specimen material entity indirectly observed via the detector • Resulting image combination of: • physical scale of the object imaged • physical scale of the imaging detector • physical laws governing interaction between: • incident radiation source and applied fields (EMF) • device components effecting incident fields (lenses, filters) • specimen • device components downstream of specimen (lenses, filters, detector(s) )

BIRN Image-Related Ontology Requirements Imaging the Biological Specimen Physical Properties • interaction with incident/ stimulus EMF Contrast Enhancement • Better living through chemistry • Histochemical treatment • differentially alter specimen light-scattering properties to enhance reflected or transmitted light differential • Emission-based probes • highly selective affinity for specimen molecular determinant • radio-isotopes • fluorescent probes • Effect NMR profile • MRI contrast agents

BIRN Image-Related Ontology Requirements • ultrasound/echo • MRI Imaging Modalities = not in BIRN yet • standard structural MRI • diffusion-tensor imaging MRI (DTI-MRI) • functional MRI (f. MRI) • PET • optical Imaging • transmittance/absorbance • brightfield, phase-contrast • reflectance • fluorescence • standard wide-field, confocal (optical/LSCM, digital deconvolution), multi-photon • electron microscopy • TEM • 2 D TEM & 3 D TEM (EM tomography) • SEM • video • vital cellular imaging • behaving subjects • provenance data varies widely by technique • typical lab likely producing only 1 or 2 of these

BIRN Image-Related Ontology Requirements Cross Image Comparison Provenance (KEY) • image acquisition use Image Ontology • device characteristics & image protocol parameters • image manipulation use Computation/SW Ontology • algorithm(s) / workflow • models of computation background vs. signal • only a subset of specimen determinants are under study intra-modality comparison • highly controlled assay / protocol • calibration via controlled specimen mimics • standards: Air Force target, uranium glass, fluorescent beads (size & intensity) • phantoms inter-modality comparison • ? - need ontologies beyond Image & Computation

Ways of Representing Knowing Neuroanatomy Neuroscience Lexicon • Neuro. Names, BIRNLex, Rad. Lex, NIF, etc. • “control” vocabulary use? • manage lexical exceptions • homographic homonyms, synonyms, eponyms, etc. • NLP KR extraction Ontologies • FMA, Rad. IO, etc. • support mereotopological reasoning Voxel-based data analysis & abstraction • feature (fiducial)-based & voxel-based spatial normalization • BIRN-wide morphological & stereological analysis • across sites, specimens, scales & imaging modalities • analysis of spatially-mapped neuro-data sets

Ways of Representing Neuroanatomy Must integrate these 3 informatic threads - and eventualy • literature informatics (The Bibliome) • biological models • parameterized view of 3 D data sets can link ontology view to voxel view • BIRN sponsored meeting? • contact Maryann Martone or Jeff Grethe Integrate Images Across • distinct spatial scales • distinct imaging modalities • distinct subject image sets (species/strain/sub-strain/…/subject)

BIRN Image-Related Ontology Requirements BIRN Use Cases for Image-Related Ontology
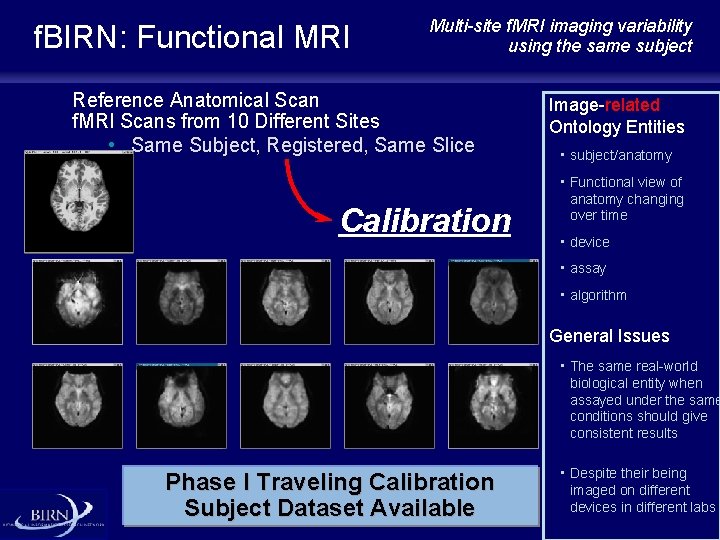
f. BIRN: Functional MRI Multi-site f. MRI imaging variability using the same subject Reference Anatomical Scan f. MRI Scans from 10 Different Sites • Same Subject, Registered, Same Slice Calibration Image-related Ontology Entities • subject/anatomy • Functional view of anatomy changing over time • device • assay • algorithm General Issues • The same real-world biological entity when assayed under the same conditions should give consistent results Phase I Traveling Calibration Subject Dataset Available • Despite their being imaged on different devices in different labs
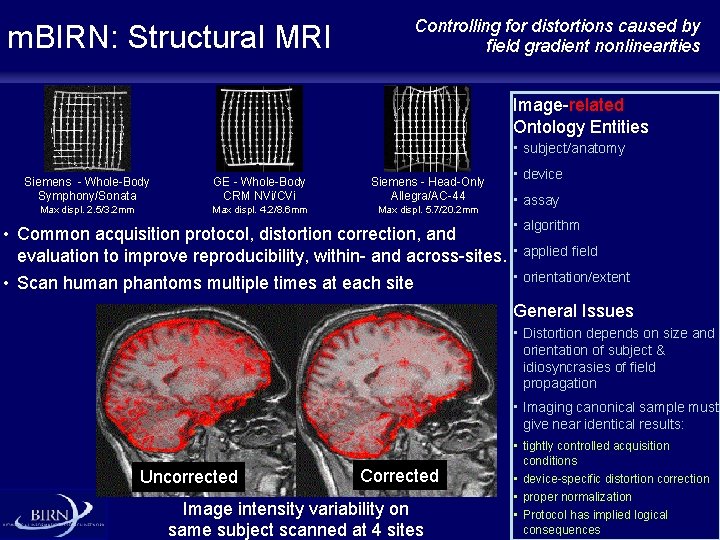
m. BIRN: Structural MRI Controlling for distortions caused by field gradient nonlinearities Image-related Ontology Entities • subject/anatomy Siemens - Whole-Body Symphony/Sonata GE - Whole-Body CRM NVi/CVi Siemens - Head-Only Allegra/AC-44 Max displ. 2. 5/3. 2 mm Max displ. 4. 2/8. 6 mm Max displ. 5. 7/20. 2 mm • Common acquisition protocol, distortion correction, and evaluation to improve reproducibility, within- and across-sites. • Scan human phantoms multiple times at each site • device • assay • algorithm • applied field • orientation/extent General Issues • Distortion depends on size and orientation of subject & idiosyncrasies of field propagation • Imaging canonical sample must give near identical results: Uncorrected Corrected Image intensity variability on same subject scanned at 4 sites • tightly controlled acquisition conditions • device-specific distortion correction • proper normalization • Protocol has implied logical consequences
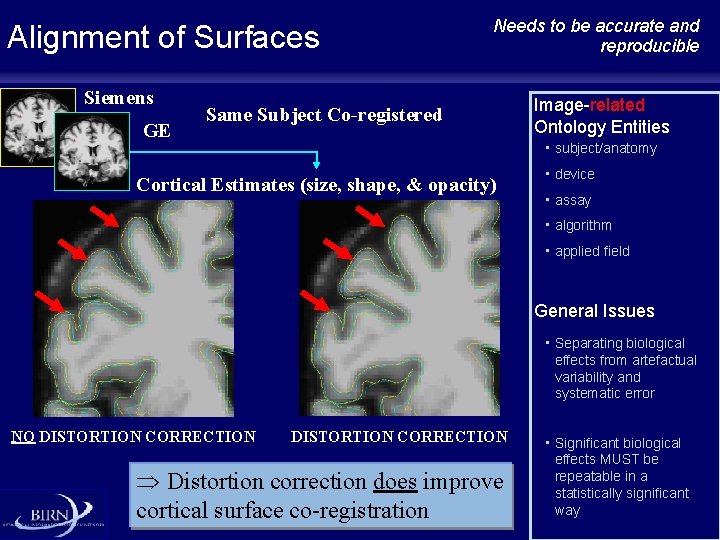
Alignment of Surfaces Siemens GE Needs to be accurate and reproducible Same Subject Co-registered Image-related Ontology Entities • subject/anatomy Cortical Estimates (size, shape, & opacity) • device • assay • algorithm • applied field General Issues • Separating biological effects from artefactual variability and systematic error NO DISTORTION CORRECTION Þ Distortion correction does improve cortical surface co-registration • Significant biological effects MUST be repeatable in a statistically significant way

Developing a White Matter Atlas Parcellation • Freesurfer (MGH) Tractography • Do. DTI (H. J. Park) Visualization • Slicer (BWH) Using Diffusion Tensor Imaging MRI (DTI-MRI) Image-related Ontology Entities • subject/anatomy • boundaries • device • DTI-specific • assay • algorithm General Issues • Defining tract boundaries & spatial relations very difficult • Need to relate to functional brain maps • DTI - specific hypothesis about biological reality postulated based on assumptions about relative H 2 O mobility
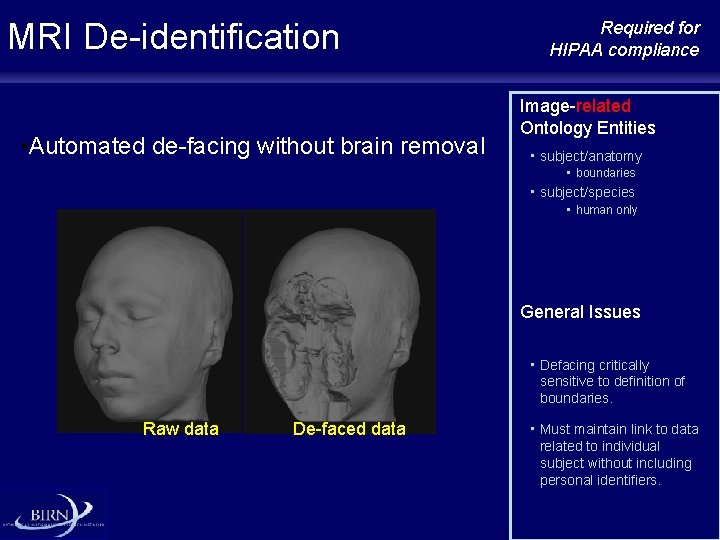
MRI De-identification • Automated de-facing without brain removal Required for HIPAA compliance Image-related Ontology Entities • subject/anatomy • boundaries • subject/species • human only General Issues • Defacing critically sensitive to definition of boundaries. Raw data De-faced data • Must maintain link to data related to individual subject without including personal identifiers.

Automated Upload Pipeline Automatic network-based, processing and validation Pipeline for bulk network file management • robust & automated • upload: • MRI Storage Request Broker (SRB) network file storage • metadata • integration & interoperability: SRB & database warehouse Image-related Ontology Entities • subject/anatomy • boundaries • QA • based on real-world criteria • diverse inputs • sharable network repository • BIRN-wide common data access stack (BIRN Rocks) • distributed GRID architecture (BIRN Racks) • Automated data handling Generate and/or validate: • De-identification (HIPAA) • Quality Assurance (QA) • Image format • BIRN ID General Issues • QA criteria: SRB BIRN warehouse • reliably represent biological entities across sites, devices and subjects • can have very complex assumptions with significant ontological component
Mouse BIRN Animal models of human neurological disorders Study animal models to test hypotheses about human neurological disorders • across dimensional scales • across imaging modalities Experimental Allergic Encephalomyelitis (EAE) mouse model Multiple Sclerosis (MS) Dopamine Transporter (DAT) knockout mouse schizophrenia, attention-deficit hyperactivity disorder (ADHD), Tourette’s disorder, and substance abuse Alpha-synuclein mouse model symptoms/pathology of Parkinson’s Disease Cancer animal models consortium with astrocytoma mouse model NCI supported with Terry Van Dyke @ Duke Cal Tech - Duke - UCLA - UCSD - U. Tenn. /Drexel U.
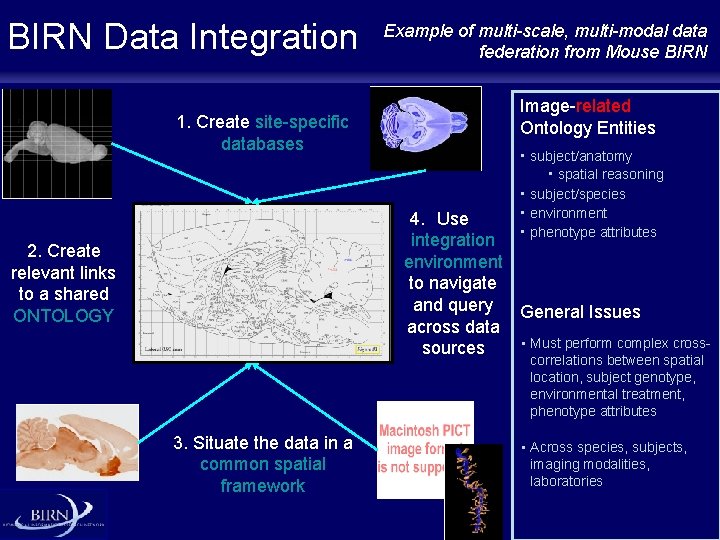
BIRN Data Integration 1. Create site-specific databases Example of multi-scale, multi-modal data federation from Mouse BIRN Image-related Ontology Entities • subject/anatomy • spatial reasoning • subject/species • environment • phenotype attributes 4. Use integration environment to navigate and query General Issues across data • Must perform complex crosssources 2. Create relevant links to a shared ONTOLOGY correlations between spatial location, subject genotype, environmental treatment, phenotype attributes 3. Situate the data in a common spatial framework • Across species, subjects, imaging modalities, laboratories

Semi-automatic spatial normalization across a large, multifarious neuroimage repository Spatial Registration Workflow Register: • image slice series and volumetric atlases Correlate: • cellular and subcellular changes with non-invasive imaging Image-related Ontology Entities • subject/anatomy • spatial reasoning • device • assay • algorithm / workflow General Issues Brain volume spatial • Algorithms must be specified via models of computation registration (e. g. , Ptolemy) to support full processing stream automation • pipelines: • LONI • Neuro. Terrain • GRID: • GRIDSphere • Kepler • Gray-scale voxel-based analysis (feature-based & numerical) must seamlessly combine with mereotopological & histo-cellular models of neuroanatomy
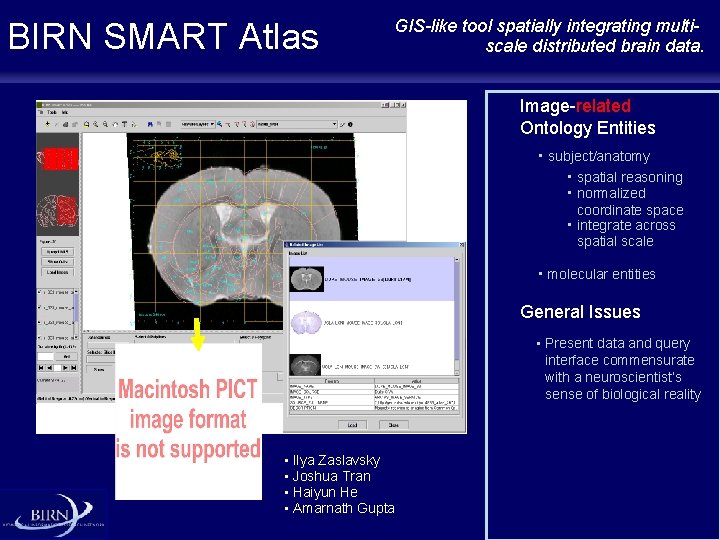
BIRN SMART Atlas GIS-like tool spatially integrating multiscale distributed brain data. Image-related Ontology Entities • subject/anatomy • spatial reasoning • normalized coordinate space • integrate across spatial scale • molecular entities General Issues • Present data and query interface commensurate with a neuroscientist’s sense of biological reality • Ilya Zaslavsky • Joshua Tran • Haiyun He • Amarnath Gupta

Mediated database interoperability and distributed query architecture BIRN SMART Atlas Image-related Ontology Entities • subject/anatomy • distributed spatial reasoning • normalized coordinate space • integrate across spatial scale • gene/protein • spatially-mapped m. RNA & protein distribution General Issues Mediator wrapper wrapper SRB UCSD Cal Tech Duke UCLA UT/Drexel • Data mediation infrastructure must be ontology-centric to support sematically-based queries • Source database data models must be mapped ontologically in order to: • determine field semantic equivalency • search content in appropriate, shared semantic context

BIRN Image-Related Ontology Use In closing HELP!
- Slides: 26