Biotech Continued 3 5 Biotech 7 1 p

Biotech Continued…. 3. 5 Biotech 7. 1 p 351 -4: DNA Profiling & DNA Sequencing 2. 7 p 115 -6: PCR – the polymerase chain reaction 7. 3 p 368 Bioinformatics

n How do forensic scientists determine who’s blood has been left at a crime scene? n How are paternity tests conducted? n ANSWER: Using restriction enzymes, a technique called Gel Electrophoresis, and a concept called RFLP (restriction fragment length polymorphism)

n Individuals sequences. have differences in their DNA n Differences in coding regions (exons) may result in a mutation (i. e. . Sickle cell anemia) n Differences in noncoding regions (introns) may exist in the form of variations in the number of repeating units

VNTR n VNTR: variable number tandem repeats n These are short nucleotide sequences that repeat themselves within the genome n Ex: 5’ CAGATAGATAGATAAGC 3’ n In this fragment of DNA, GATA is a repeating sequence that appears 5 times. n These sequence appears in intron regions and the number of repeats is variable in different people

VNTRs n You have similar VNTRs to your relatives. n Half your chromosomes (i. e. half your DNA) comes from each of your parents half of your VNTRs are the same as your mom and half are the same as your dad. n (Well, actually more than half since your mom and dad are bound to have some that are the same even though they’re not related)

RFLP analysis n If a sample of DNA is digested by a restriction enzyme, the result would be the formation of a number of DNA fragments of various lengths. n Since we all have different DNA* exposure to the same restriction enzyme would produce different numbers and lengths of fragments.

RFLP Analysis n I. E. If Ana’s chromosome 3 has recognitions sites for the restriction enzyme, then it will be cut into 4 pieces. n If Sarah has 5 recognition sites, then her chromosome will be cut into 6 pieces. n This is known as RESTRICTION FRAGMENT LENGTH POLYMORPHISM
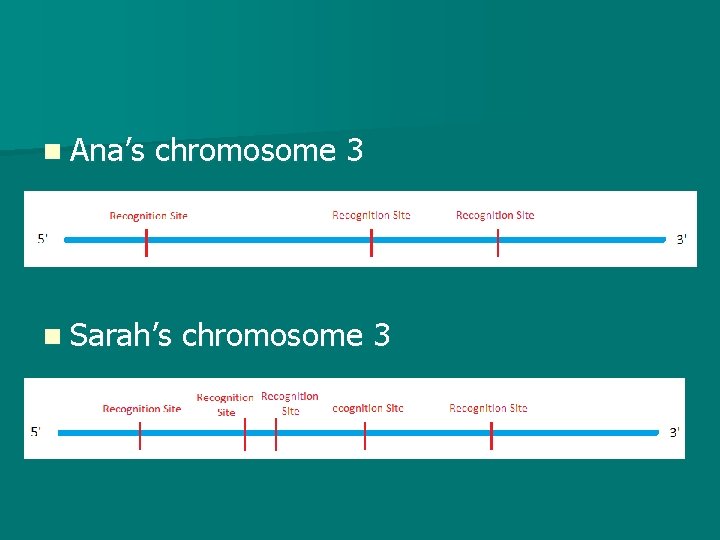
n Ana’s chromosome 3 n Sarah’s chromosome 3
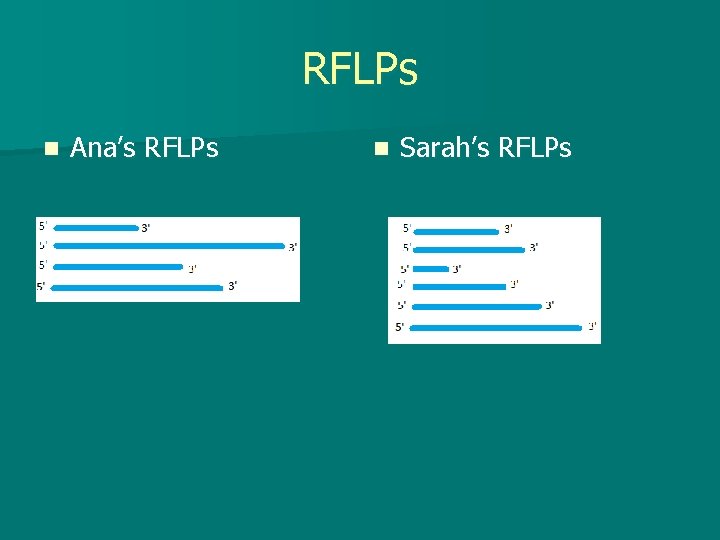
RFLPs n Ana’s RFLPs n Sarah’s RFLPs

n RFLP can be used to identify individuals. n *NOTE: Most of the DNA among humans is the same. For most of your genes, we have the exact same sequence of nucleotides. n Aside from a few variations in alleles, the differences in one human’s DNA to another’s is in the introns – in particular the number of VNTR

n Scenario: A crime is committed and there are 3 suspects. n If the criminal left a sample of blood at the crime scene, RFLPs from this sample can be compared to RFLPs from blood of all suspects to determine who the criminal is.

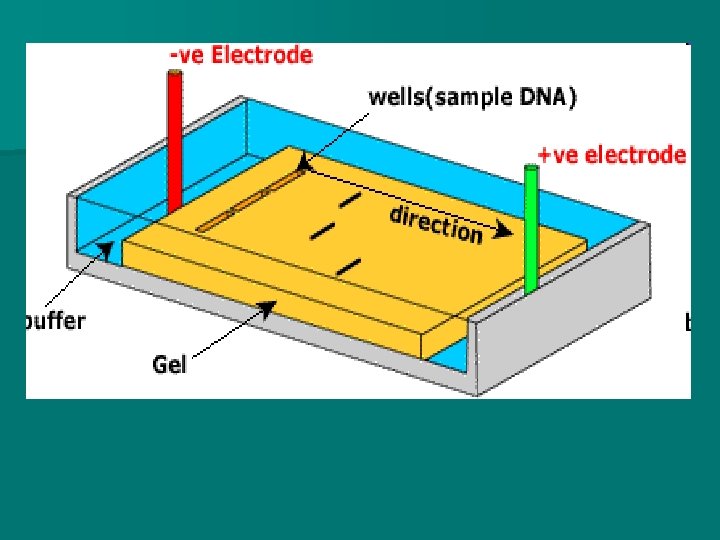
Gel Electrophoresis n The DNA fragments (produced from exposure to a restriction enzyme) must be separated and purified from each other for analysis using a process called gel electrophoresis n In this process, the DNA fragments are placed in wells on a “jello-like gel- called agarose (or agar) The agarose is fibrous and acts like a net The gel is immersed into a conducting fluid n n

Gel Electrophoresis n An gel electrical current is passed through the n Since DNA is negatively charged, the fragments of DNA are attracted to the positive end.
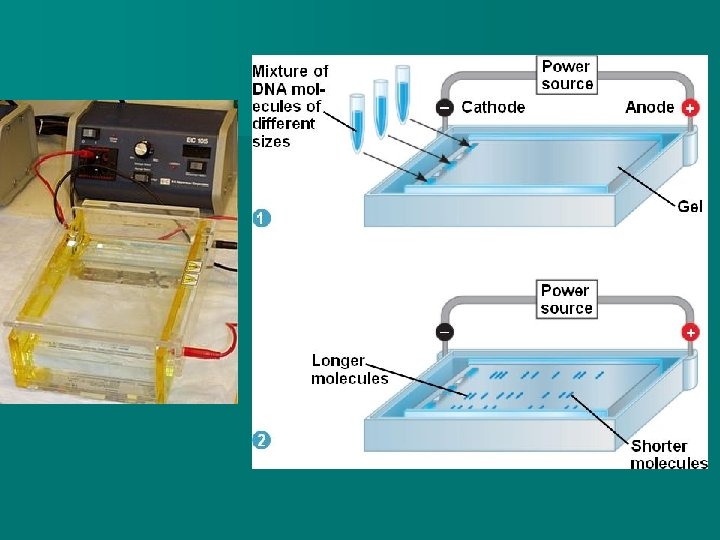

n As the DNA fragments move through the agar, the small fragments move quickly, but larger fragments move slowly because they get caught up in the fibres of the gel. n. A blue dye is added initially to the DNA fragments. When the dye reaches the positive end, the power is turned off. n In this way, gel electrophoresis is used to separate DNA (or proteins) by size

n. A fluorescent dye is then used to stain the DNA fragments. n Ideally this creates a band pattern that is unique to each individual.
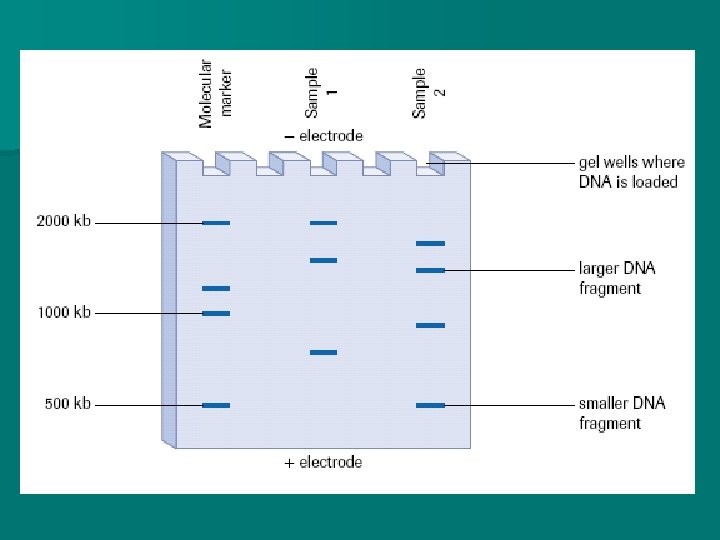
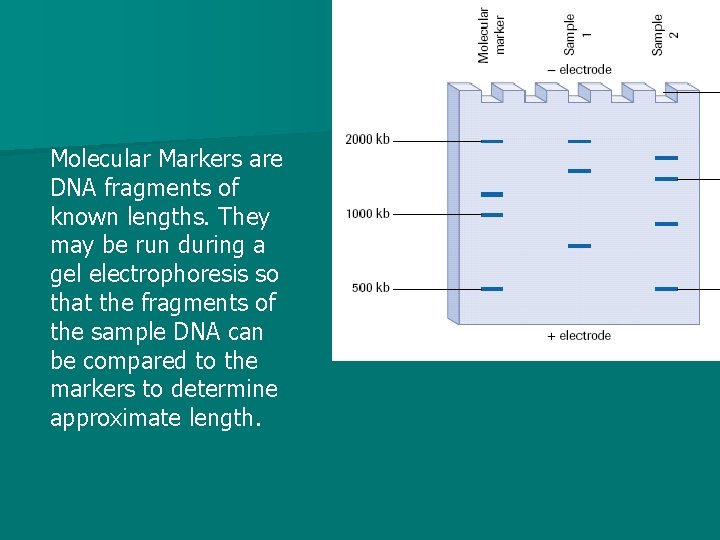
Molecular Markers are DNA fragments of known lengths. They may be run during a gel electrophoresis so that the fragments of the sample DNA can be compared to the markers to determine approximate length.

n In reality, the amount of DNA is usually so large and the bands too numerous that instead of seeing individual bands, a large smear is seen on the gel. n In a process called SOUTHERN BLOTTING, the DNA from the gel can be transferred to a nylon membrane. n The nylon membrane is placed against an X-ray film to read an autoradiogram which will show the band pattern
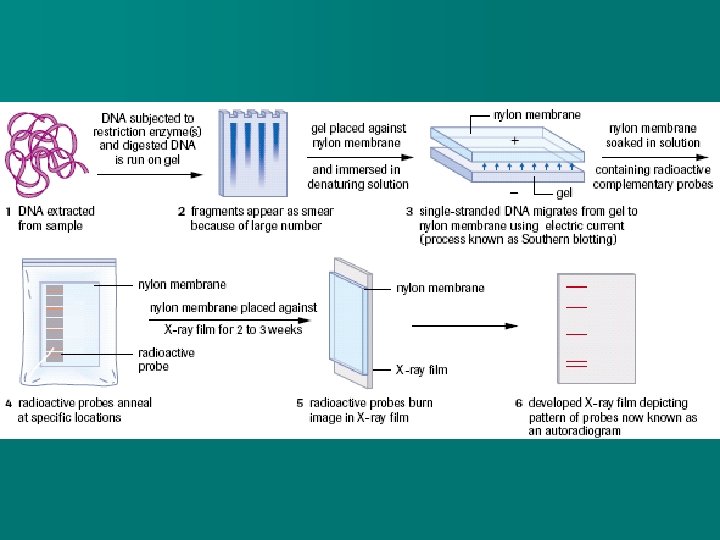

DNA Profiling n DNA profiling involves comparing DNA samples in paternity investigations and forensic science. n The various sample DNA is exposed to the same restriction enzyme (or same group of restriction enzymes) n The band pattern produced is unique to each human (other than identical twins)
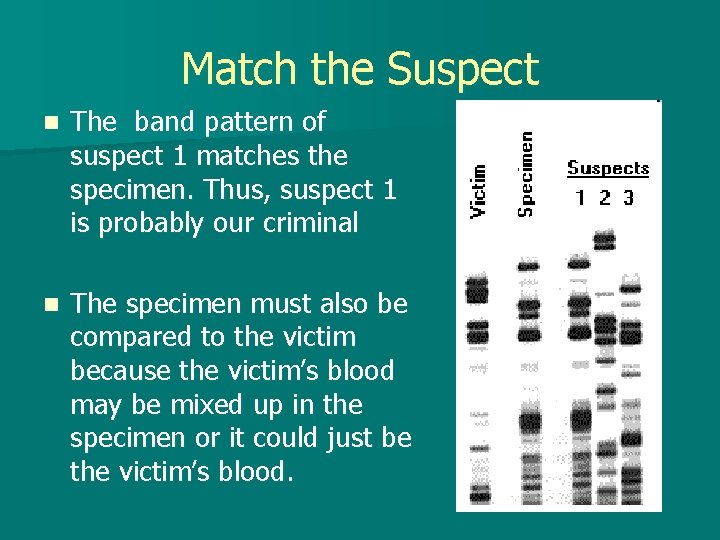
Match the Suspect n The band pattern of suspect 1 matches the specimen. Thus, suspect 1 is probably our criminal n The specimen must also be compared to the victim because the victim’s blood may be mixed up in the specimen or it could just be the victim’s blood.

Who’s your Daddy? ? ? n IN paternity test, the child’s DNA is compared to its mother’s and the possible fathers. Since the child has DNA from both its mom and dad, the bands that match its mother can be ignored. n Look at the bands that do not match the mother’s n Who does it match? n The reason the child doesn’t have the exact same DNA of its parents is because it only receives half of each parent’s chromosomes/DNA

DNA Evidence n Where can DNA evidence be obtained? n Blood stains at the crime scene or on the victim or victim’s clothing n (Remember, it’s really the WBC not the RBC used) n Hair (though requires the follicle) n Semen n Saliva n Others

PCR: Polymerase Chain Reaction n PCR DNA is a technique used to clone (amplify) n Before the technique was developed, if scientists wanted to make multiple copies of a gene fragment, the gene would have to be inserted into a plasmid and the bacterial cell would make more copies when it replicates its plasmids. n Then scientists would have to remove the plasmids and cut out the bacterial genes.

n With PCR, many copies of DNA can be made quickly. n This is particularly useful when only a small sample of original DNA is available. n (Ex: If only a small sample of DNA is obtained from a crime scene, PCR may be used to allow for multiple forensic tests. )

n Before running a DNA through the Gel Electrophoresis for analysis, the sample will undergo PCR to make sure there is enough DNA for adequate testing. n The process of PCR is closely related to DNA replication that occurs within the nucleus of a cell.

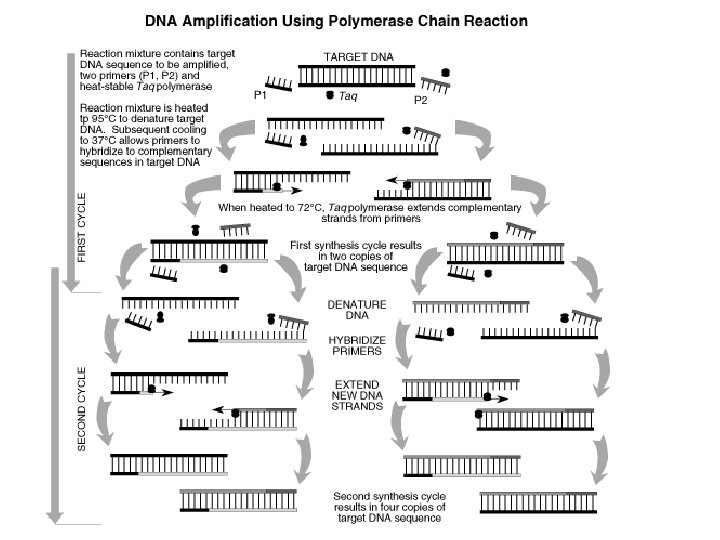
DNA REPLICATION n The 2 DNA strands are separated using enzymes helicase and gyrase PCR The 2 strands of DNA are separated using heat n At a temp of 95°C, the hydrogen bonds between the complimentary strand will break, separating the strands n

DNA REPLICATION n An RNA primer must be added first before DNA nucleotides are added to build a complementary strand to the template strand PCR n A DNA primer is used instead because they are easy to produce in labs. n (The temp is lowered to 54°C to allow for the DNA primers to attach) n One of the primers is known as a forward primer and the other is a reverse primer cause they start synthesis of DNA in opposite directions

DNA REPLICATION n PCR Once the RNA primers n Temp raised to 72°C have been laid down, DNA polymerase III n Taq polymerase – a will build a type of DNA complementary strand polymerase – builds by adding DNA complementary nucleotides strands using DNA nucleotides that have been added to the solution

Taq Polymerase n The Taq polymerase does not denature because it is extracted from a bacteria (Thermus aquaticus) that lives in hot springs (Used to hot temps!) n At 72°C it is at its optimal temperature and can add 1000 nucleotides per second
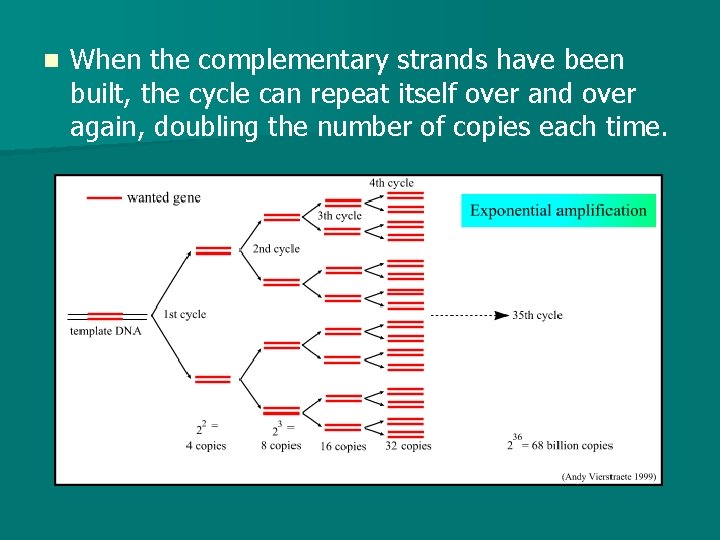
n When the complementary strands have been built, the cycle can repeat itself over and over again, doubling the number of copies each time.

PCR n Thermal Cycler is the name of the machine that amplifies the DNA. n A scientist can insert the DNA sample to amplified and all the require materials (including DNA primers, nucleotides, and Taq polymerase) and the machine will continuously go through the cycle of raising, lowering, and raising the temperature again and again to allow the DNA to be copied.
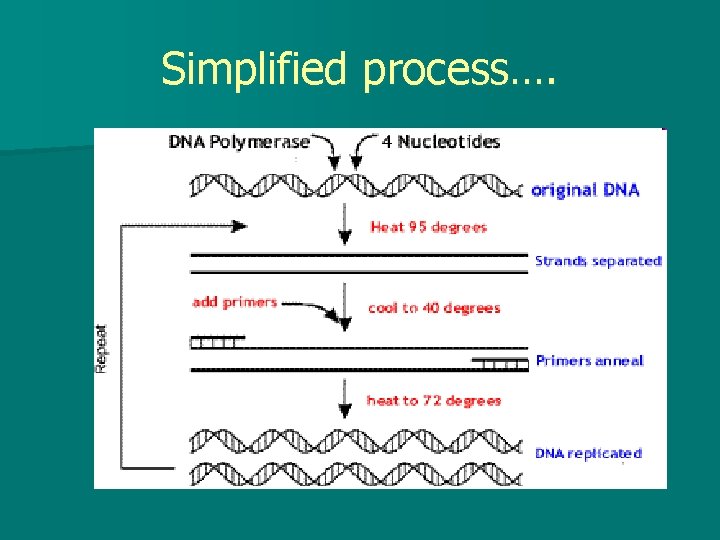
Simplified process….

Animations n http: //highered. mcgraw- hill. com/sites/0072437316/student_view 0/ chapter 16/animations. html#

DNA Sequencing n The process of determining the exact sequence of nucleotides in a DNA molecule. n The DNA sample is amplified with PCR and then cleaved with a restriction enzyme n The many copies of the fragments of DNA are divided into four tubes and placed in the sequencing machine along with all the raw materials of replication (enzymes, primers, nucleotides)

DNA sequencing - Dideoxyribonucleotides In addition, a small quantity dideoxyribonucleotides (dd. NTP) are added to the mixtures. n These nucleotides do not have an OH – on the 3 rd carbon (of deoxyribose) and thus they inhibit DNA polymerase from adding more nucleotides. n n They are also labelled with different fluorescent markers that will make them fluoresce a particular colour

DNA sequencing Dideoxyribonucleotides n dd. C – dideoxyribonucleotide cytosine n dd. A – dideoxyribonucleotide adenine n dd. G – dideoxyribonucleotide guanine n dd. T – dideoxyribonucleotide thymine n Each type of dideoxyribonucleotide is added to a different tube
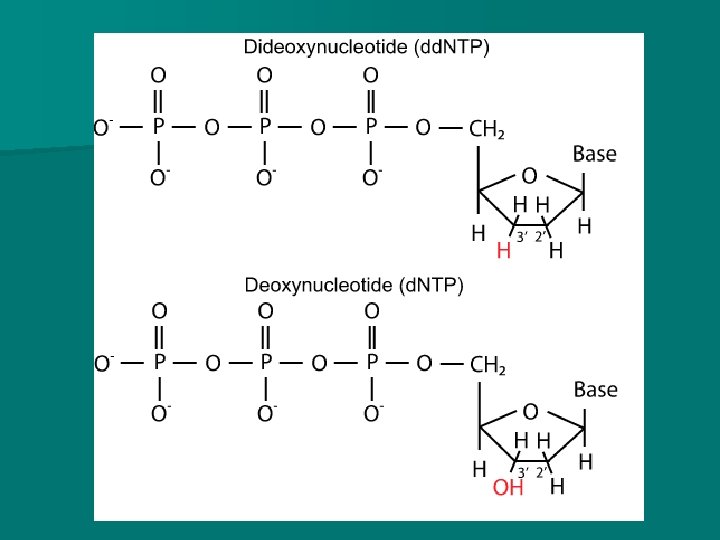

n. A second round of copying (the DNA) will take place, incorporating the dideoxyribonucleotides. n After this round of copying, a different fluorescent dye will be added to each batch of DNA.
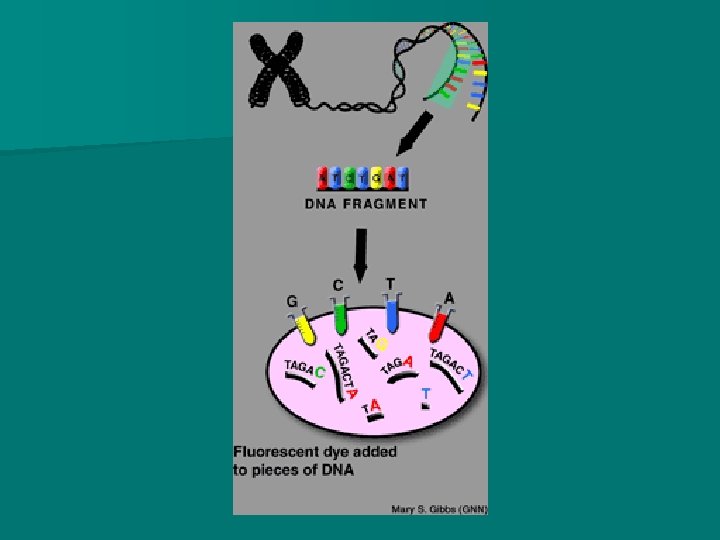

n Suppose you apply that procedure to this sequence of DNA: n TAGACT n At the end of the second round of copying, each batch will contain the following pieces of DNA: n 1: blue-T, blue-TAGACT 2: red-TA, red-TAGA 3: yellow-TAG 4: green-TAGAC
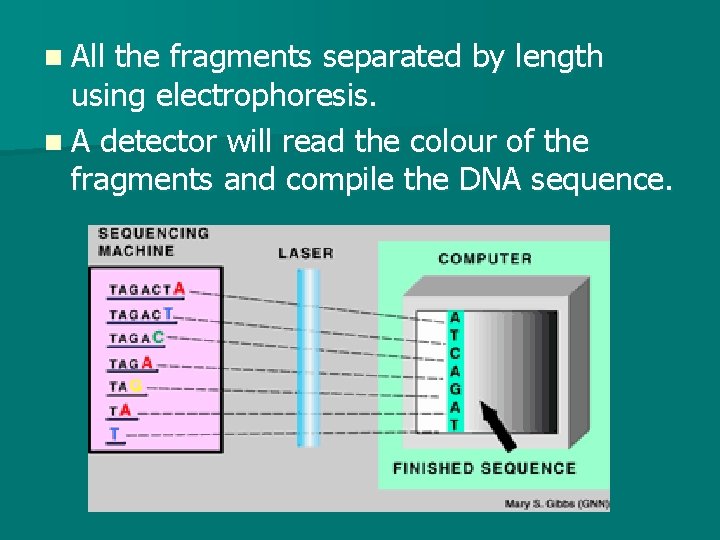
n All the fragments separated by length using electrophoresis. n A detector will read the colour of the fragments and compile the DNA sequence.

Bioinformatics n Bioinformatics: uses computers to analyze biological data n Development in scientific research follow improvements in computing. n The use of computers has enabled scientists to make advances in bioinformatics applications such as locating genes within genomes and identifying conserved sequences

Bioinformatics n Information is often amassed in databases such as Gen. Bank, DDBJ (DNA databank of Japan, EMBL (European Molecular Biology Laboratory) and can be made accessible to other scientists globally.

Bioinformatics n If you are a scientist researching a particular genetic disorder, you could access genetic sequences of multiple people with this disorder. n Then through computer programs, find the amino acid sequence or nucleotide sequence that is similar in all these individuals to determine was is causing the disorder.
- Slides: 48