Biopython What is Biopython tools for computational molecular

Biopython

What is Biopython? • tools for computational molecular biology • to program in python and want to make it as easy as possible to use python for bioinformatics by creating high-quality, reusable modules and scripts 2

What can Biopython do? • • Manipulate DNA and protein sequences Run BLAST Access public databases Manipulate protein structures Population genetics Supervised learning methods Networks of various kinds

Obtaining Biopython • http: //www. biopython. org 4

Making sure it worked >>> new_seq. complement() >>> new_seq. reverse_complement() 5

Working with sequences • A biopython Seq object has two important attributes: – data : as the name implies, this is the actual sequence data string of the sequence – alphabet : an object describing what the individual characters making up the string "mean" and how they should be interpreted • Two advantages 1. this gives an idea of the type of information the data object contains 2. this provides a means of contraining the information you have in the data object, as a means of type checking 6
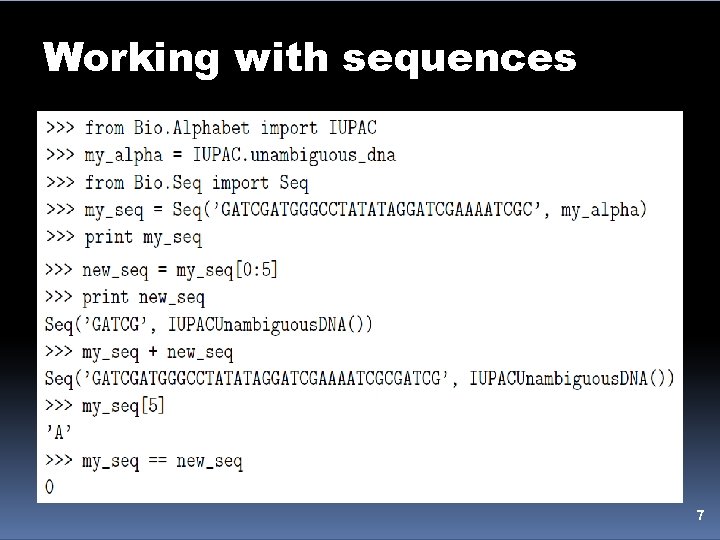
Working with sequences 7

Working with sequences >>> >>> >>> >>> protein_seq = Seq('EVRNAK', IUPAC. protein) dna_seq = Seq('ACGT', IUPAC. unambiguous_dna) protein_seq + dna_seq my_seq. tostring() my_seq[5] = 'G mutable_seq = my_seq. tomutable() print mutable_seq[5] = 'T' print mutable_seq. remove('T') print mutable_seq. reverse() print mutable_seq 8
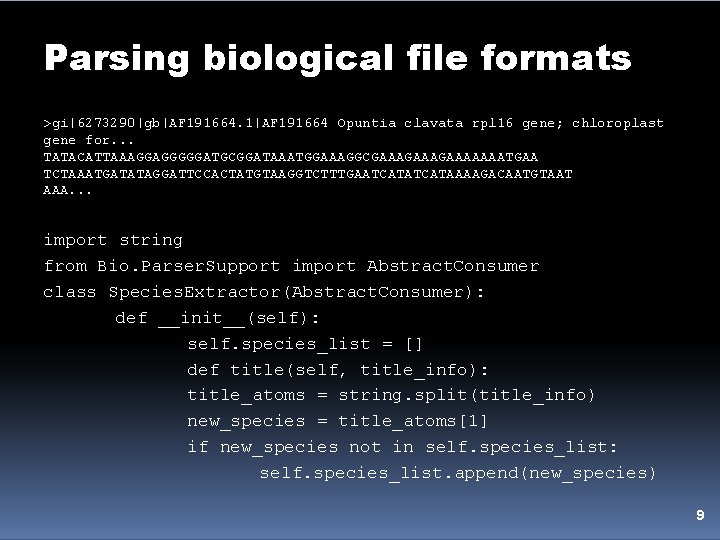
Parsing biological file formats >gi|6273290|gb|AF 191664. 1|AF 191664 Opuntia clavata rpl 16 gene; chloroplast gene for. . . TATACATTAAAGGAGGGGGATGCGGATAAATGGAAAGGCGAAAGAAAAAAATGAA TCTAAATGATATAGGATTCCACTATGTAAGGTCTTTGAATCATAAAAGACAATGTAAT AAA. . . import string from Bio. Parser. Support import Abstract. Consumer class Species. Extractor(Abstract. Consumer): def __init__(self): self. species_list = [] def title(self, title_info): title_atoms = string. split(title_info) new_species = title_atoms[1] if new_species not in self. species_list: self. species_list. append(new_species) 9
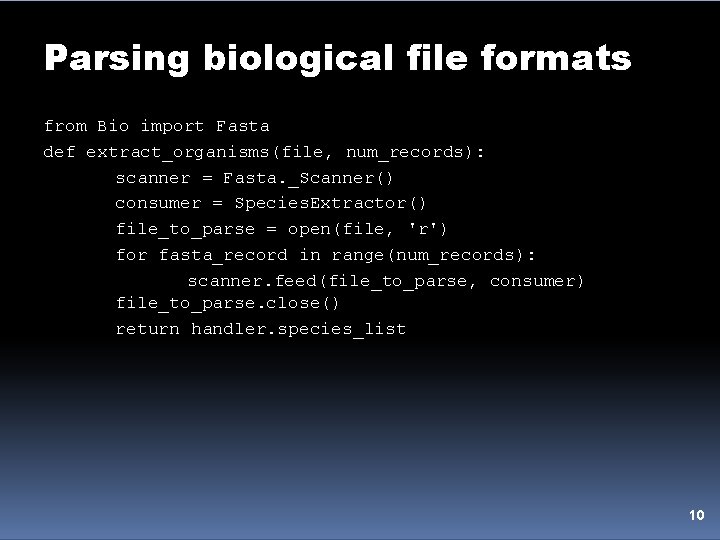
Parsing biological file formats from Bio import Fasta def extract_organisms(file, num_records): scanner = Fasta. _Scanner() consumer = Species. Extractor() file_to_parse = open(file, 'r') for fasta_record in range(num_records): scanner. feed(file_to_parse, consumer) file_to_parse. close() return handler. species_list 10
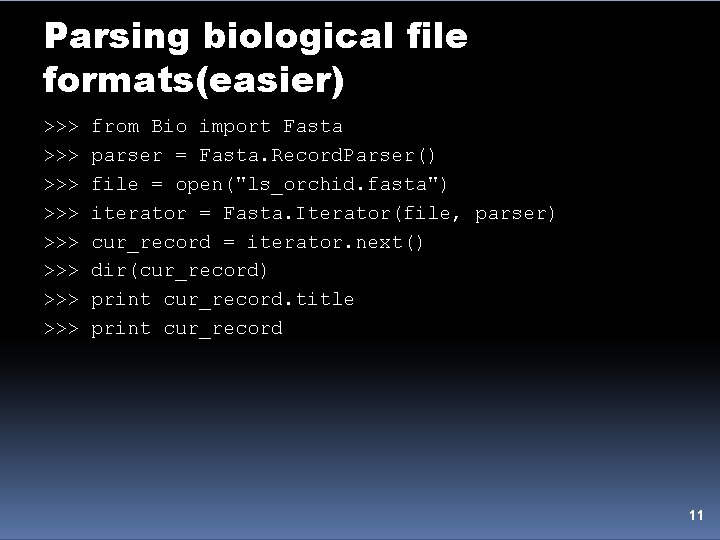
Parsing biological file formats(easier) >>> >>> from Bio import Fasta parser = Fasta. Record. Parser() file = open("ls_orchid. fasta") iterator = Fasta. Iterator(file, parser) cur_record = iterator. next() dir(cur_record) print cur_record. title print cur_record 11
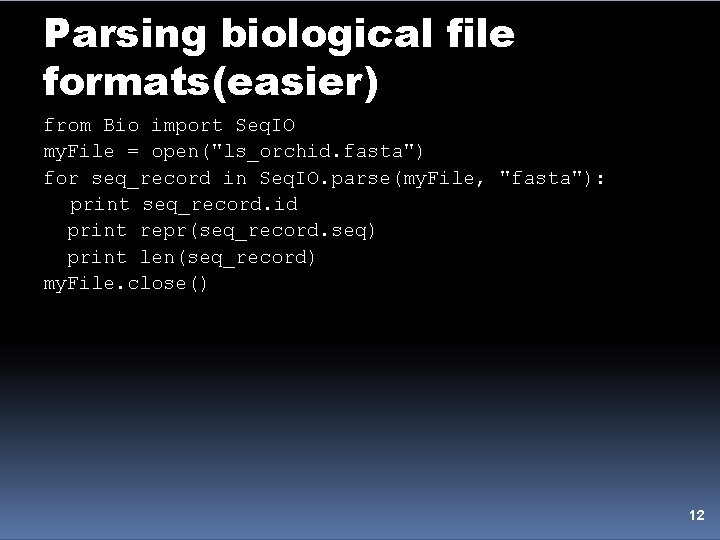
Parsing biological file formats(easier) from Bio import Seq. IO my. File = open("ls_orchid. fasta") for seq_record in Seq. IO. parse(my. File, "fasta"): print seq_record. id print repr(seq_record. seq) print len(seq_record) my. File. close() 12

FASTA files as Dictionaries import string def get_accession_num(fasta_record): title_atoms = string. split(fasta_record. title) # all of the accession number information is stuck in the first element # and separated by '|'s accession_atoms = string. split(title_atoms[0], '|') # the accession number is the 4 th element gb_name = accession_atoms[3] # strip the version info before returning return gb_name[: -2] 13

FASTA files as Dictionaries(easier) >>> from Bio import Fasta >>> Fasta. index_file("ls_orchid. fasta", "my_orchid_dict. idx", get_accession_num) >>> from Bio. Alphabet import IUPAC >>> dna_parser = Fasta. Sequence. Parser(IUPAC. ambiguous_dna) >>> orchid_dict = Fasta. Dictionary("my_orchid_dict. idx", dna_parser) 14
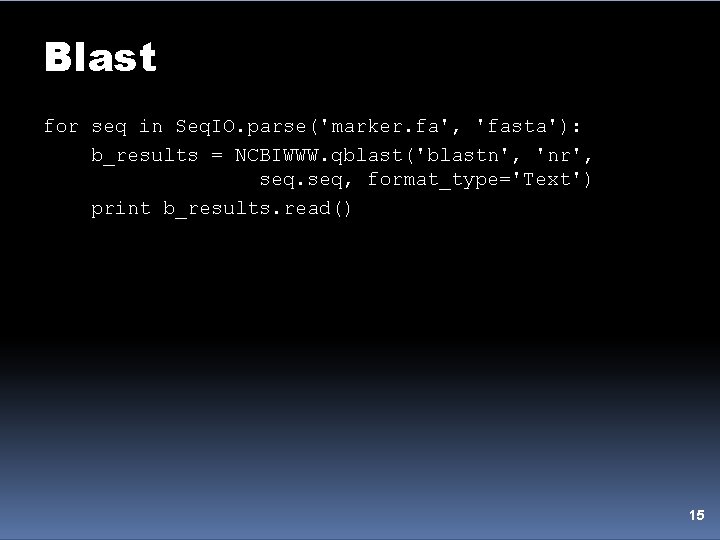
Blast for seq in Seq. IO. parse('marker. fa', 'fasta'): b_results = NCBIWWW. qblast('blastn', 'nr', seq, format_type='Text') print b_results. read() 15

More information http: //www. biopython. org

Problem • Write a program to read a FASTA file and print the number of sequences, number of residues, and minimum, maximum and average lengths of the sequences. > python read-fasta-file. py sample. fa Number of sequences = 7 Number of residues = 285 Minimum length = 21 Maximum length = 94 Average length = 40. 7
- Slides: 17