Biophotonics Winter School 2007 Lecture 1 Biodetection using

Biophotonics Winter School 2007 Lecture 1: Biodetection using Silicon Photonic Bandgap Devices Philippe M. Fauchet University of Rochester Supported in part by the National Science Foundation, the Infotonics Center of Excellence, and the Center for Future Health 1
Where is Rochester? Toronto California >6000 km > 3000 km Italy New York 2

Rochester: the campus and the city 3

Organization • • Long-Term Goal Materials Science of Porous Silicon Sensing Principle using Microcavities Examples of Biosensing In lecture 2: • Ultimate Performance of these Biosensors • Futuristic Application 4

The state of the art…yesterday and today 1860 2002 5

The state of the art…tomorrow 6
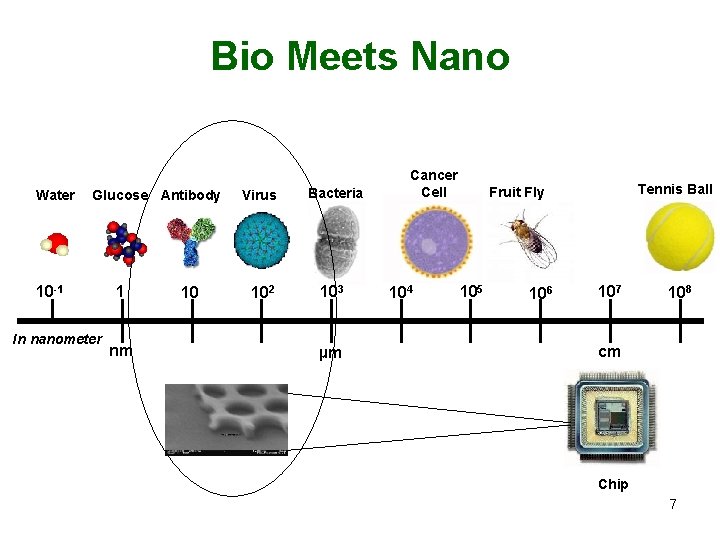
Bio Meets Nano Water Glucose Antibody 10 -1 In nanometer 1 nm 10 Virus Bacteria 102 103 µm Cancer Cell 104 Tennis Ball Fruit Fly 105 106 107 108 cm Chip 7

Objectives n Biosensor platforms capable of detecting the presence of harmful pathogens, including public health hazards and biowarfare agents, are under development. n These biosensors rely on advances in molecular recognition, nanoscience, nanotechnology, and optics. n They are can be used for lab-on-chip applications or form intelligent systems that can be used by untrained personnel. 8

Porous Silicon Materials Science 9
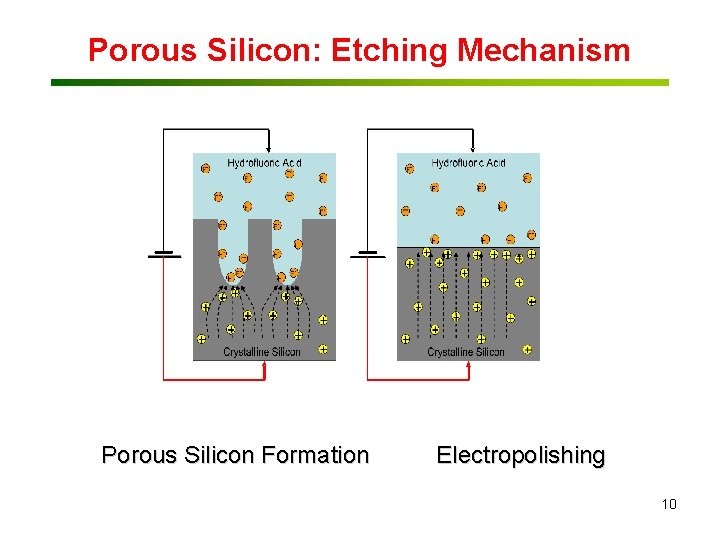
Porous Silicon: Etching Mechanism Porous Silicon Formation Electropolishing 10
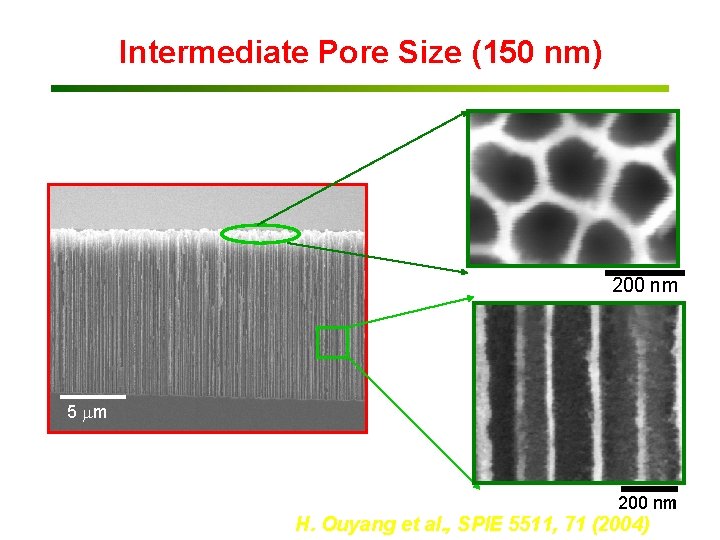
Intermediate Pore Size (150 nm) 200 nm 5 mm ~150 nm diameter ~200 nm pore-to-pore spacing 11 200 nm H. Ouyang et al. , SPIE 5511, 71 (2004)

Material: Porous Silicon Mesopores 20 nm Chemicals, short DNA strands, small molecules Small Macropores 200 nm Macromolecules, proteins Large Macropores 2000 nm Viruses, bacteria H. Ouyang, M. Lee, B. L. Miller, and P. M. Fauchet, in Tuning the Optical Response of Photonic Bandgap Structures II, SPIE Proc. (2005) 12

Pore Size and Morphology Engineering From nanopores to mesopores to macropores and from “spongy” to directional and smooth 100 nm 10 µm 13
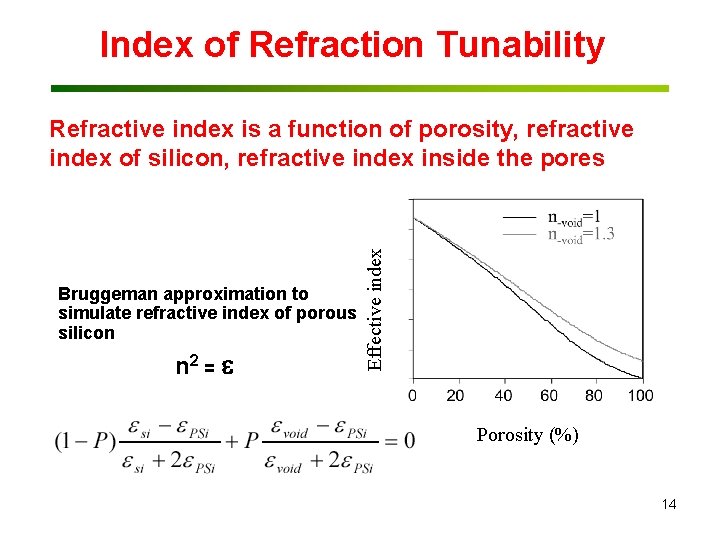
Index of Refraction Tunability Bruggeman approximation to simulate refractive index of porous silicon n 2 = Effective index Refractive index is a function of porosity, refractive index of silicon, refractive index inside the pores Porosity (%) 14

Biosensing Principles 15

Biosensing with PSi Microcavities § The optical properties of a porous silicon microcavity are governed by the refractive index of the porous silicon layer(s) § The refractive index of a porous silicon layer depends on what is inside the pores § The functionalized internal surface of porous silicon can bind the desired biological objects (“targets”) § Binding is detected through a change in refractive index, hence a change in optical properties (luminescence, transmission or reflectivity) 16
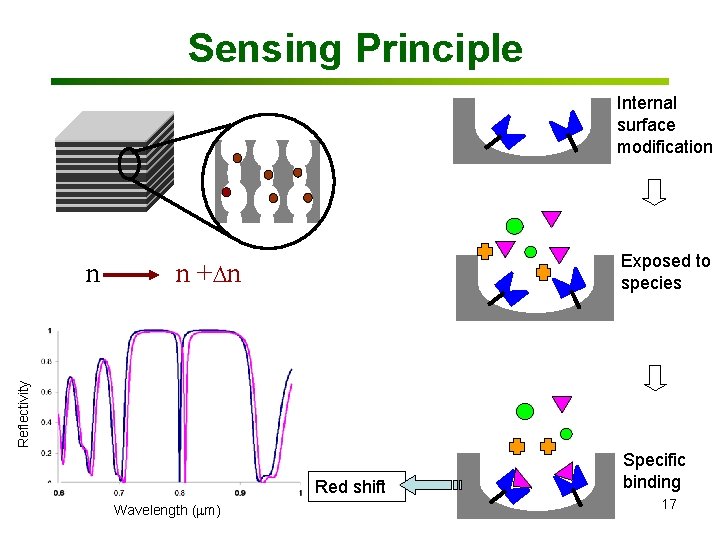
Sensing Principle Internal surface modification n + n Reflectivity n Exposed to species Red shift Wavelength (mm) Specific binding 17
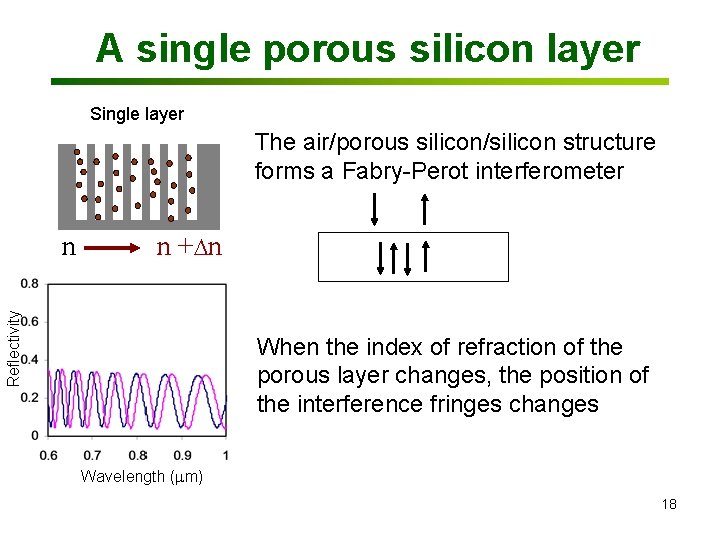
A single porous silicon layer Single layer The air/porous silicon/silicon structure forms a Fabry-Perot interferometer n + n Reflectivity n When the index of refraction of the porous layer changes, the position of the interference fringes changes Wavelength (mm) 18
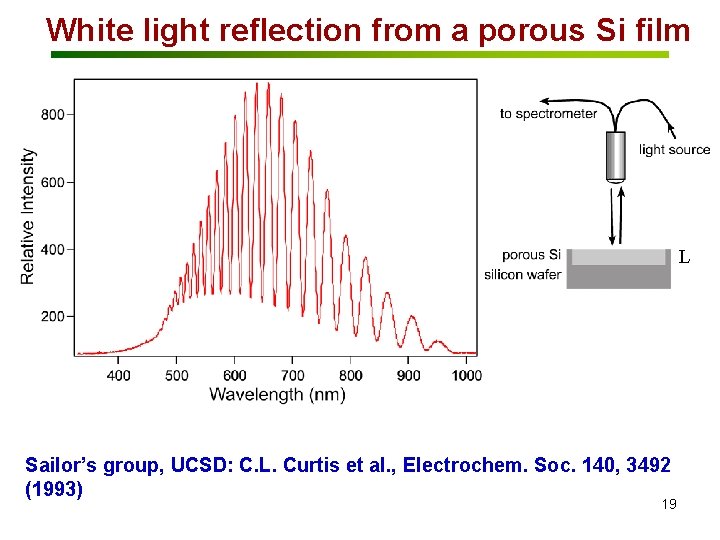
White light reflection from a porous Si film L Sailor’s group, UCSD: C. L. Curtis et al. , Electrochem. Soc. 140, 3492 (1993) 19

Protein A/Human Ig. G Binding to Porous Si A B C Streptavidin D E b-Prot. A F G HI Ig. G Rinse J K LM Ig. G Rinse Sailor’s group, UCSD 20
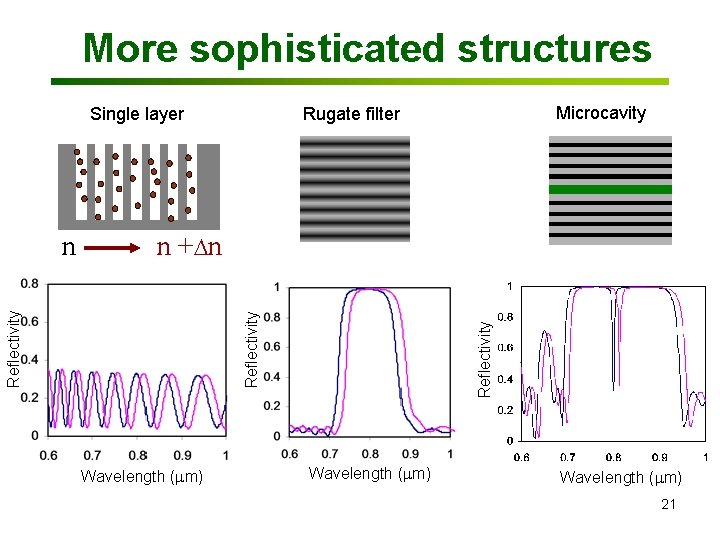
More sophisticated structures Single layer n + n Wavelength (mm) Reflectivity n Microcavity Rugate filter Wavelength (mm) 21
![Multilayer Structures Electrolyte [m. A/cm 2] j [s] t C-Silicon Subtrate H. Ouyang et Multilayer Structures Electrolyte [m. A/cm 2] j [s] t C-Silicon Subtrate H. Ouyang et](http://slidetodoc.com/presentation_image/13e982cc1e2e4b82a6d9c4b95c05e4c8/image-22.jpg)
Multilayer Structures Electrolyte [m. A/cm 2] j [s] t C-Silicon Subtrate H. Ouyang et al, Adv. Funct. Mater. 15, 1851 (2005) 22
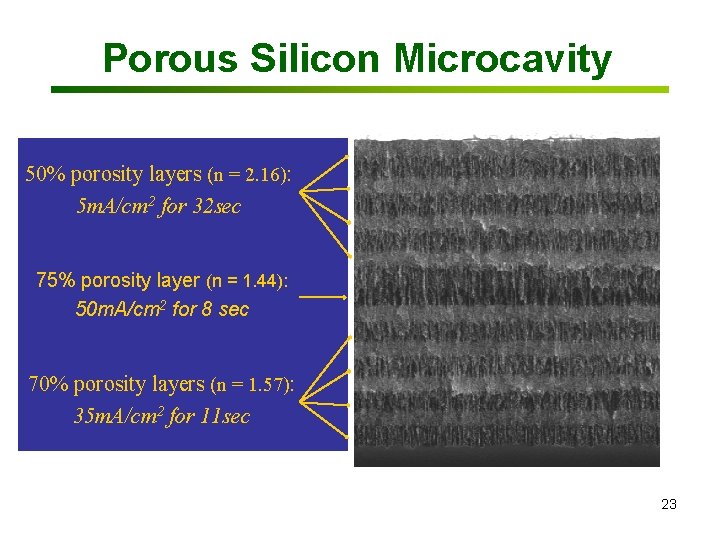
Porous Silicon Microcavity 50% porosity layers (n = 2. 16): 2 for 32 sec 5 m. A/cm Bragg mirror 75% porosity layer (n = 1. 44): 50 m. A/cm 2 for 8 sec Defect layer 70% porosity layers (n = 1. 57): Bragg mirror 35 m. A/cm 2 for 11 sec 2µm 23

Porous Silicon Microcavity CONTROL OVER THREE LENGTH SCALES PSi Multilayer Mirror PSi Central Layer PSi Multilayer Mirror 24
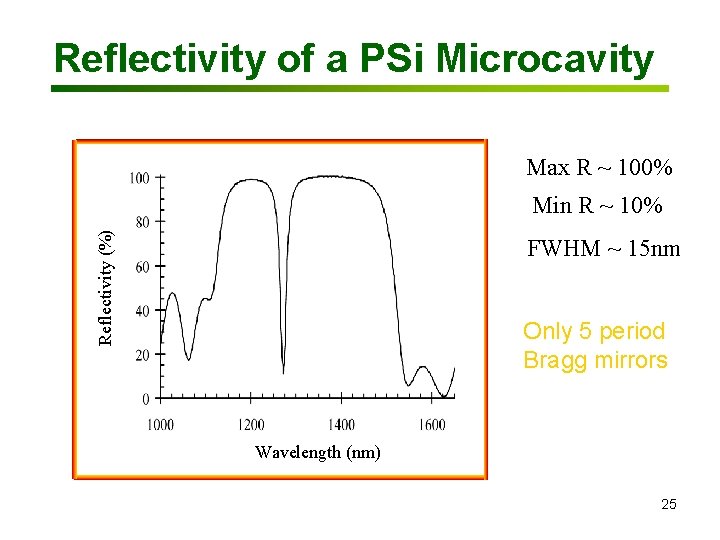
Reflectivity of a PSi Microcavity Max R ~ 100% Reflectivity (%) Min R ~ 10% FWHM ~ 15 nm Only 5 period Bragg mirrors Wavelength (nm) 25
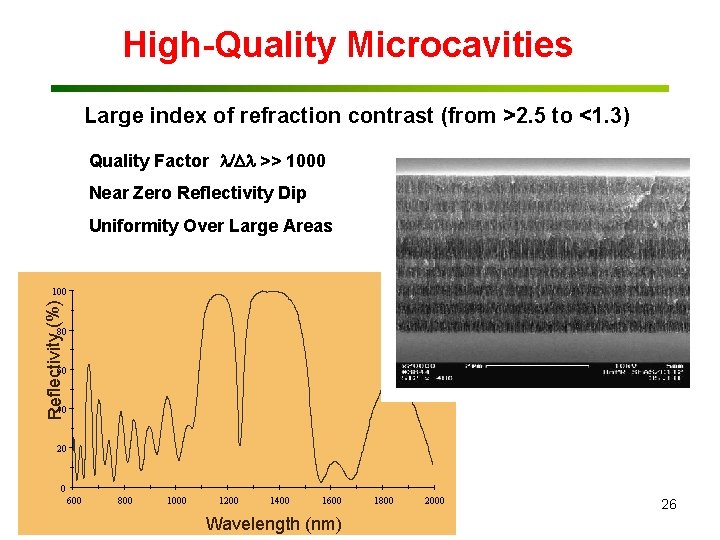
High-Quality Microcavities Large index of refraction contrast (from >2. 5 to <1. 3) Quality Factor >> 1000 Near Zero Reflectivity Dip Uniformity Over Large Areas Reflectivity (%) 100 80 60 40 20 0 600 800 1000 1200 1400 1600 Wavelength (nm) 1800 2000 26
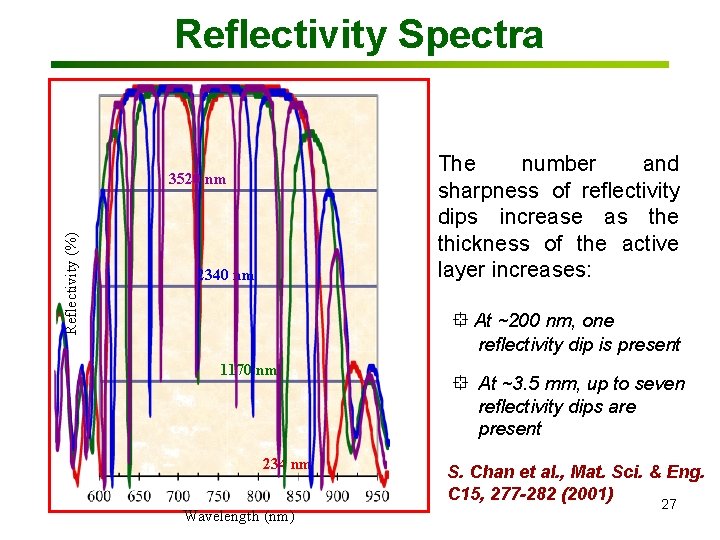
Reflectivity Spectra The number and sharpness of reflectivity dips increase as the thickness of the active layer increases: Reflectivity (%) 3520 nm 2340 nm At ~200 nm, one reflectivity dip is present 1170 nm 234 nm Wavelength (nm) At ~3. 5 mm, up to seven reflectivity dips are present S. Chan et al. , Mat. Sci. & Eng. C 15, 277 -282 (2001) 27
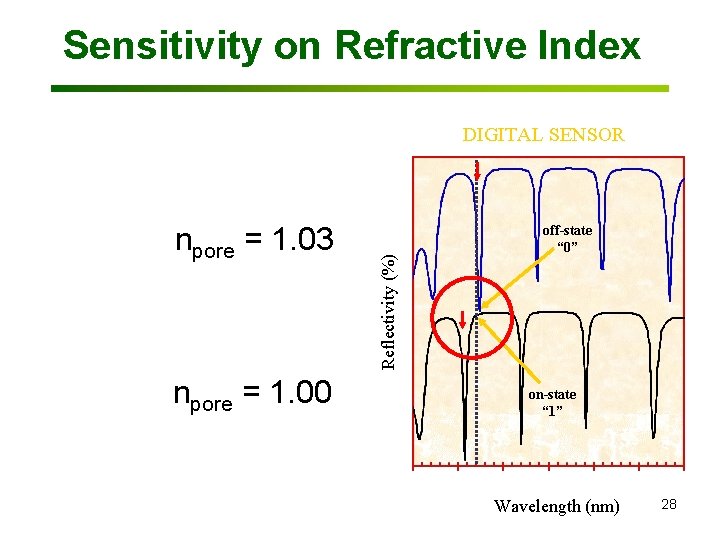
Sensitivity on Refractive Index npore = 1. 03 npore = 1. 00 Reflectivity (%) DIGITAL SENSOR off-state “ 0” on-state “ 1” Wavelength (nm) 28
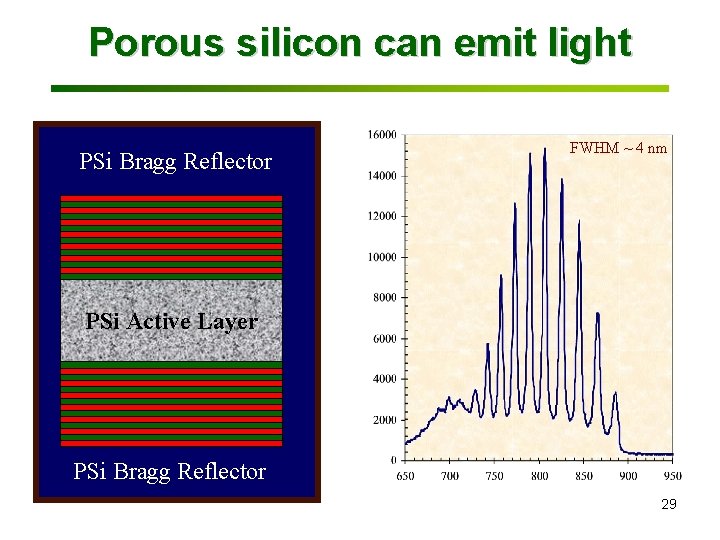
Porous silicon can emit light FWHM ~ 4 nm PSi Active Layer Photoluminescence Intensity (a. u. ) PSi Bragg Reflector Wavelength (nm) 29

Examples of Biosensing § DNA § Proteins § Bacteria 30
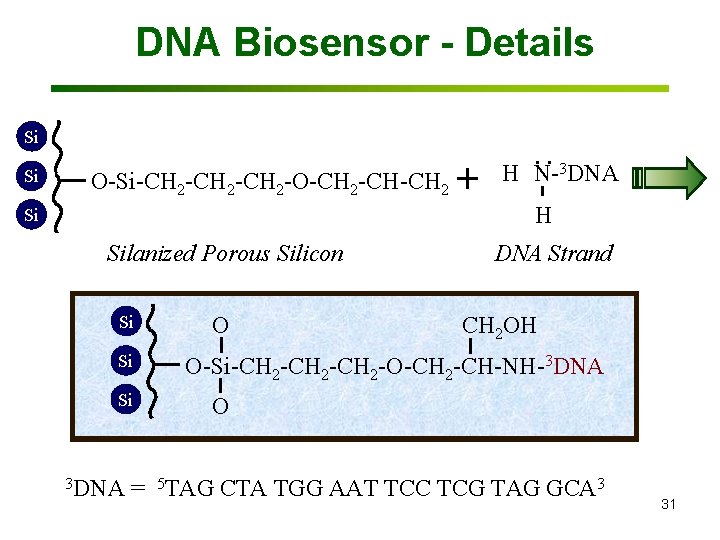
DNA Biosensor - Details Si Si Si O O-Si-CH 2 -CH 2 -O-CH 2 -CH-CH 2 O Silanized Porous Silicon Si Si Si 3 DNA O O + H . . N- DNA 3 H DNA Strand CH 2 OH O-Si-CH 2 -CH 2 -O-CH 2 -CH-NH-3 DNA O = 5 TAG CTA TGG AAT TCC TCG TAG GCA 3 31

Microcavity DNA Biosensor Normalized PL Intensity (a. u. ) PSi / DNA / c. DNA Differential Signal Wavelength (nm) § 50 µM of DNA is exposed to the porous silicon microcavity sensor § 1 µM of c. DNA binds to the DNA sensor for one hour § 7 nm PL red-shift is observed after binding § No PL shifting is observed when two noncomplementary strands of DNA are in contact S. Chan et Phys. al. , Mat. Sci. (a) & Eng. S. et al. , Stat. Sol. C 15, 541 277 -282 182, (2000). (2001) 32
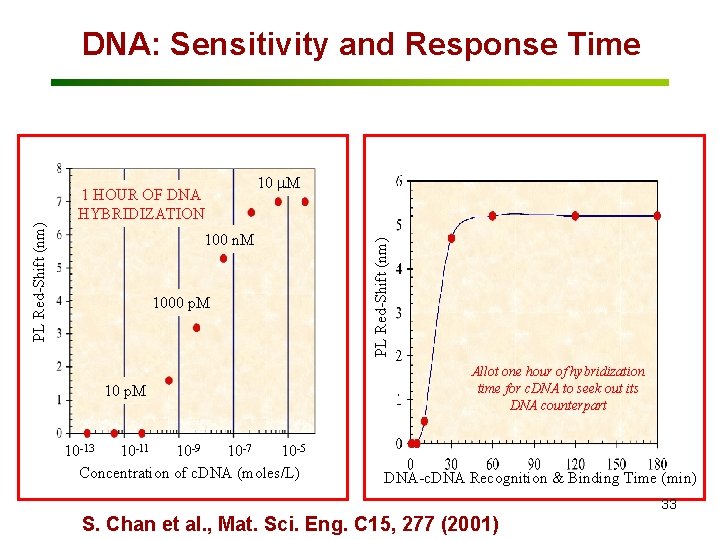
1 HOUR OF DNA HYBRIDIZATION 10 m. M 100 n. M 1000 p. M 10 -13 10 -11 10 -9 10 -7 10 -5 Concentration of c. DNA (moles/L) PL Red-Shift (nm) DNA: Sensitivity and Response Time Allot one hour of hybridization time for c. DNA to seek out its DNA counterpart DNA-c. DNA Recognition & Binding Time (min) 33 S. Chan et al. , Mat. Sci. Eng. C 15, 277 (2001)
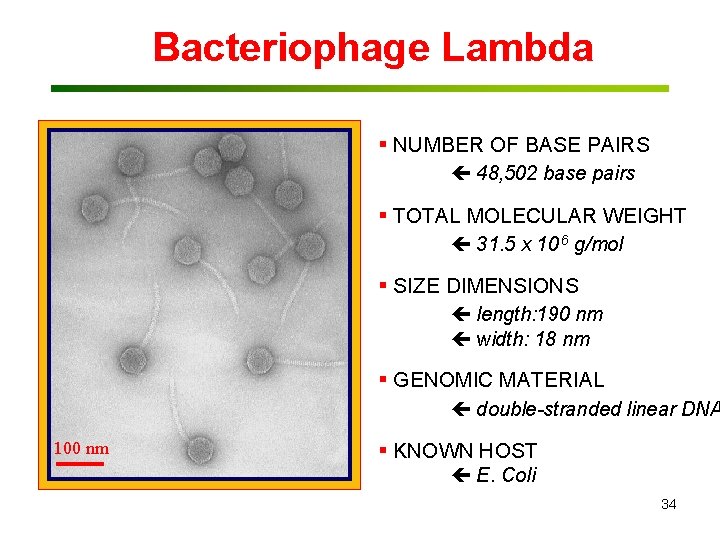
Bacteriophage Lambda § NUMBER OF BASE PAIRS 48, 502 base pairs § TOTAL MOLECULAR WEIGHT 31. 5 x 10 6 g/mol § SIZE DIMENSIONS length: 190 nm width: 18 nm § GENOMIC MATERIAL double-stranded linear DNA 100 nm § KNOWN HOST E. Coli 34
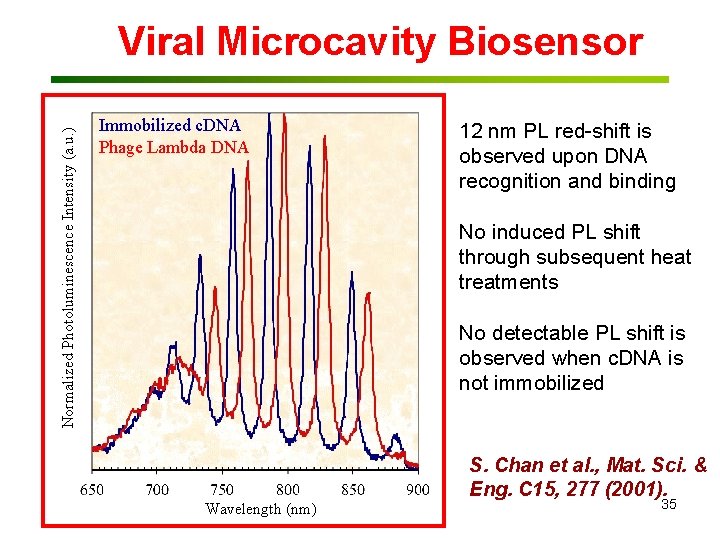
Normalized Photoluminescence Intensity (a. u. ) Viral Microcavity Biosensor Immobilized c. DNA Phage Lambda DNA 12 nm PL red-shift is observed upon DNA recognition and binding No induced PL shift through subsequent heat treatments No detectable PL shift is observed when c. DNA is not immobilized Wavelength (nm) S. Chan et al. , Mat. Sci. & Eng. C 15, 277 (2001). 35
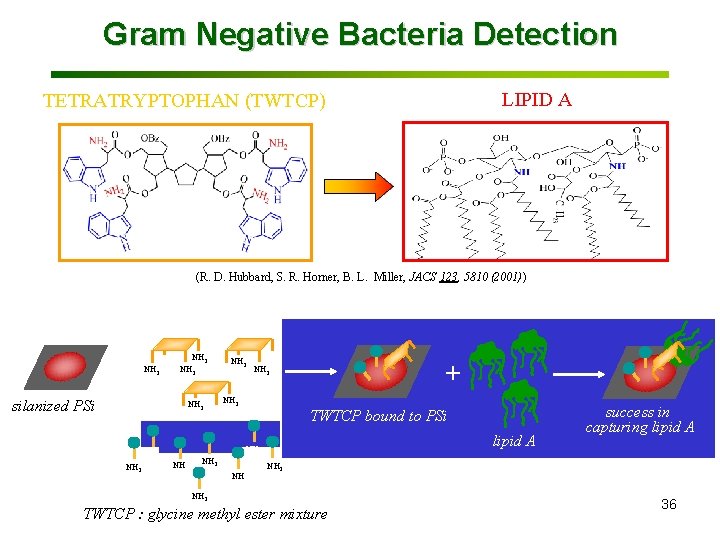
Gram Negative Bacteria Detection LIPID A TETRATRYPTOPHAN (TWTCP) (R. D. Hubbard, S. R. Horner, B. L. Miller, JACS 123, 5810 (2001)) + NH 2 NH 2 NH 2 silanized PSi TWTCP NH 2 ++ TWTCPboundtoto. PSi TWTCP lipid. AA lipid NH 2 failureinto success capturelipid. AA capturing NH 2 TWTCP : glycine methyl ester mixture 36

Gram-Negative Bacteria Detection BACTERIUM CLASS PL RED-SHIFT E. coli Gram-(-) Bacillus subtilis Gram-(+) L. Acidiophilus Gram-(+) Salmonella Gram-(-) Pseudomom. Aeruginosa Gram-(-) E. coli Pseudomonas Aeruginosa 4 nm none detected 3 nm Salmonella Bacillus subtilis Principle: detect Lipid A using TWTCP probe molecules S. Chan et al. , J. Amer. Chem. Soc. 123, 11797 (2001) 37
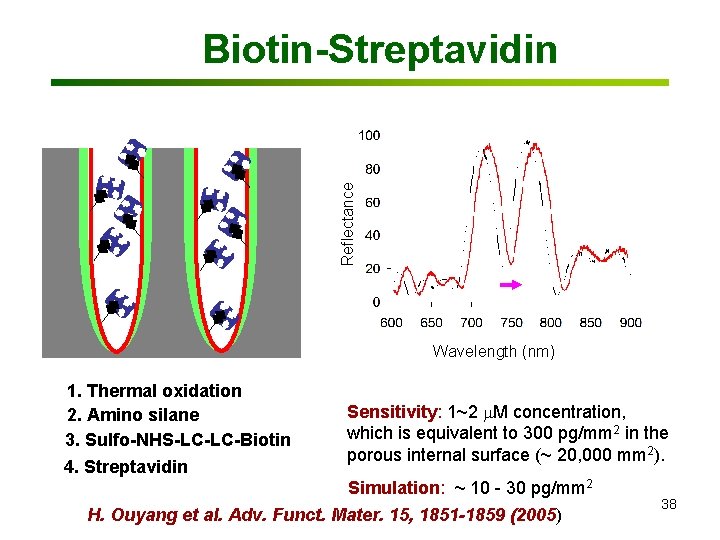
Reflectance Red shift (nm) Biotin-Streptavidin Wavelength (nm)(mg/ml) Biotin concentration 1. Thermal oxidation 2. Amino silane 3. Sulfo-NHS-LC-LC-Biotin 4. Streptavidin Sensitivity: 1~2 m. M concentration, which is equivalent to 300 pg/mm 2 in the porous internal surface (~ 20, 000 mm 2). Simulation: ~ 10 - 30 pg/mm 2 H. Ouyang et al. Adv. Funct. Mater. 15, 1851 -1859 (2005) 38
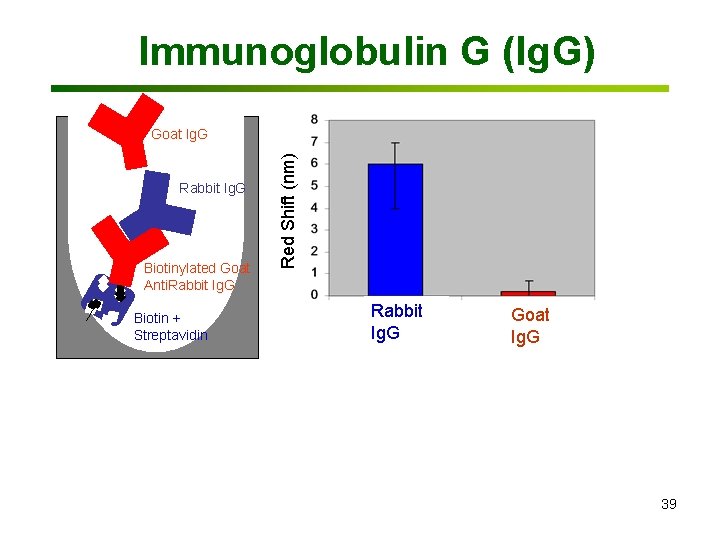
Immunoglobulin G (Ig. G) Rabbit Ig. G Biotinylated Goat Anti. Rabbit Ig. G Biotin + Streptavidin Red Shift (nm) Goat Ig. G Rabbit Ig. G Goat Ig. G 39
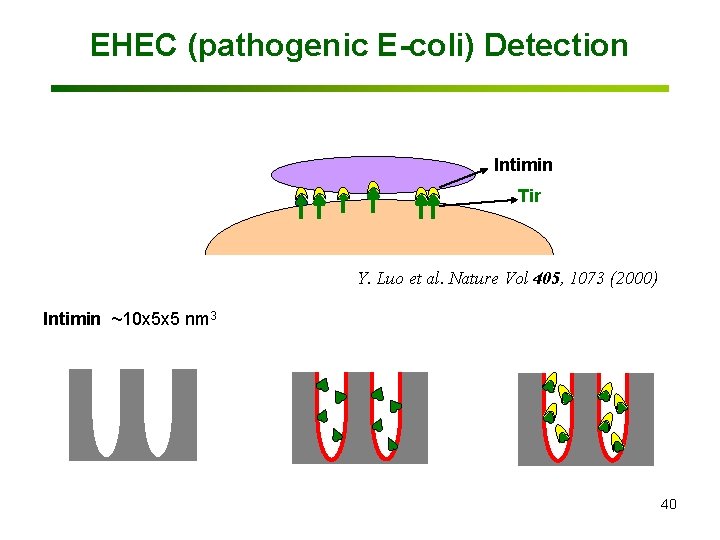
EHEC (pathogenic E-coli) Detection Intimin Tir Y. Luo et al. Nature Vol 405, 1073 (2000) Intimin ~10 x 5 x 5 nm 3 40
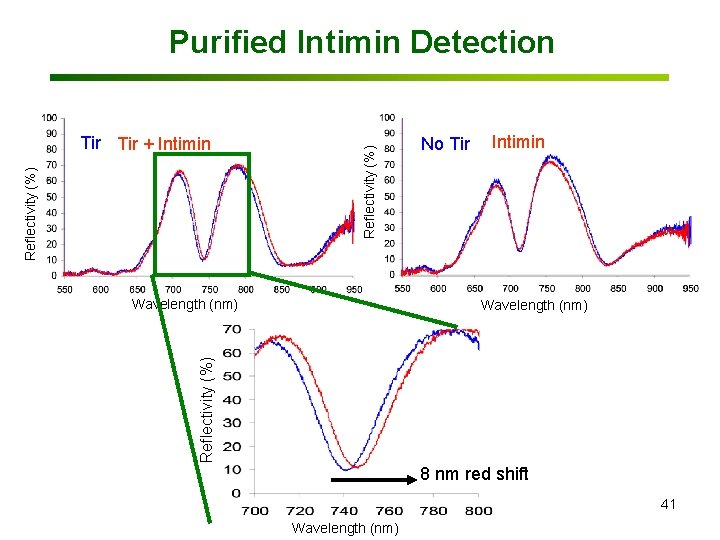
Reflectivity (%) Tir + Intimin Reflectivity (%) Purified Intimin Detection Wavelength (nm) No Tir Intimin Reflectivity (%) Wavelength (nm) 8 nm red shift 41 Wavelength (nm)
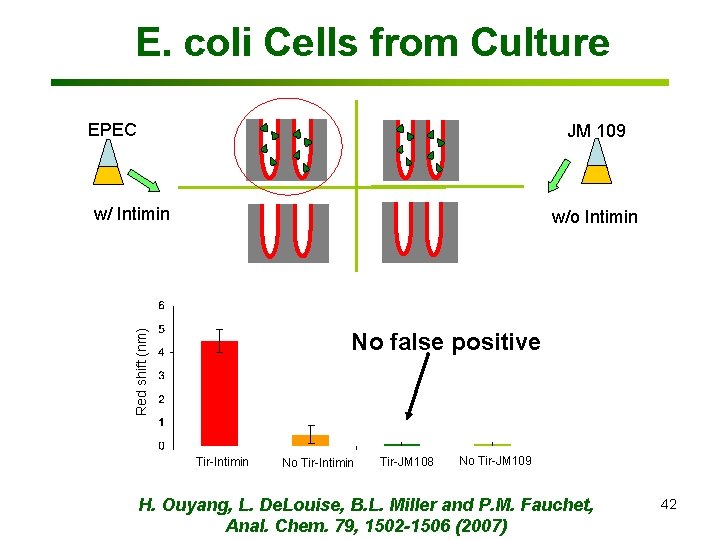
E. coli Cells from Culture EPEC JM 109 w/ Intimin w/o Intimin Red shift (nm) No false positive Tir-Intimin No Tir-Intimin Tir-JM 108 No Tir-JM 109 H. Ouyang, L. De. Louise, B. L. Miller and P. M. Fauchet, Anal. Chem. 79, 1502 -1506 (2007) 42
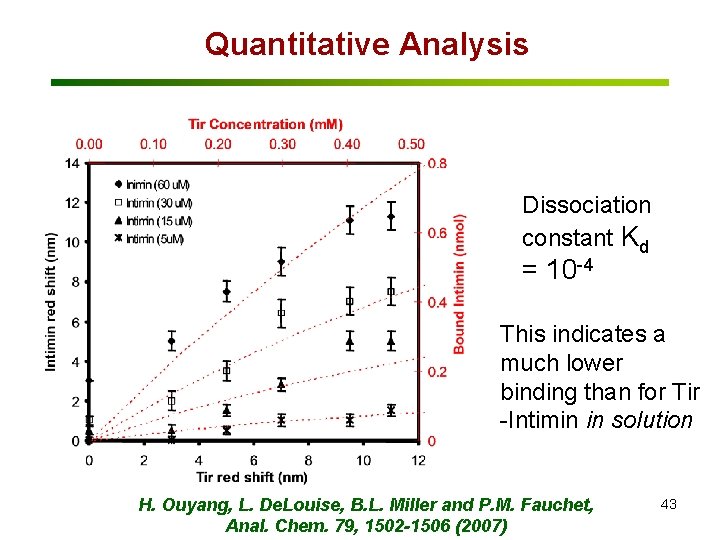
Quantitative Analysis Dissociation constant Kd = 10 -4 This indicates a much lower binding than for Tir -Intimin in solution H. Ouyang, L. De. Louise, B. L. Miller and P. M. Fauchet, Anal. Chem. 79, 1502 -1506 (2007) 43
- Slides: 43