BIONFBENG 203 Functional Genomics Sources of Functional Data

BIONF/BENG 203: Functional Genomics Sources of Functional Data Lectures 1 and 2 Lecture TI 1, 2 Trey Ideker UCSD Departments of Medicine & Bioengineering 1

Instructors l Trey Ideker l Vineet Bafna l Anand Patel (TA) 2

Grading l 40% Problem Sets (best 4 of 5) l 30% Midterm l 30% Final Project 3

Topics Covered By This Course ① ② ③ ④ ⑤ ⑥ ⑦ 4 ⑧ Signal detection in bioinformatics Large-scale data generation platforms Understanding next-gen sequencing data Understanding mass spectrometry data Clustering and Classification Genotype-phenotype association Understanding physical & genetic networks Gene network inference and evolution
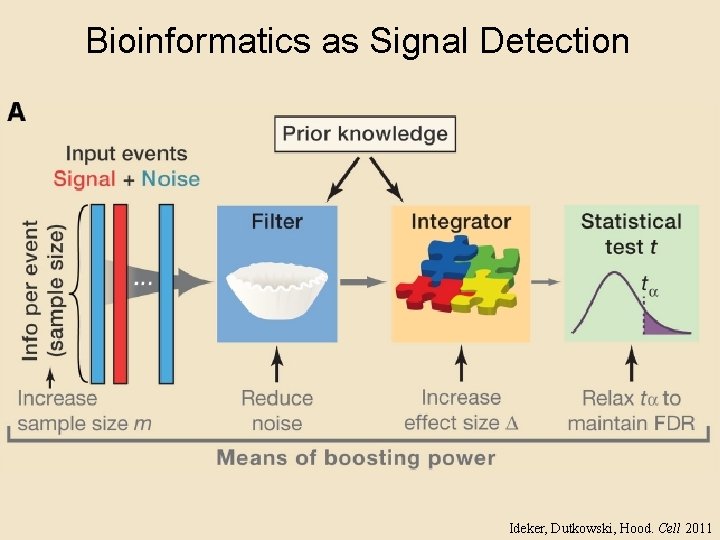
Bioinformatics as Signal Detection Ideker, Dutkowski, Hood. Cell 2011
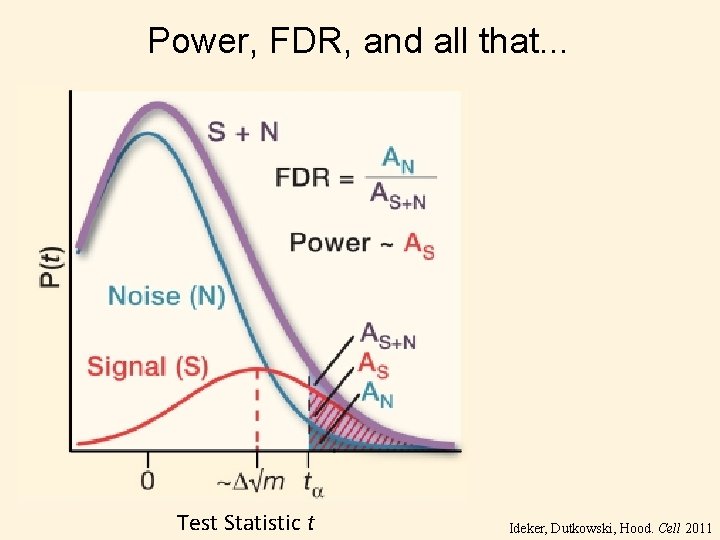
Power, FDR, and all that. . . Test Statistic t Ideker, Dutkowski, Hood. Cell 2011
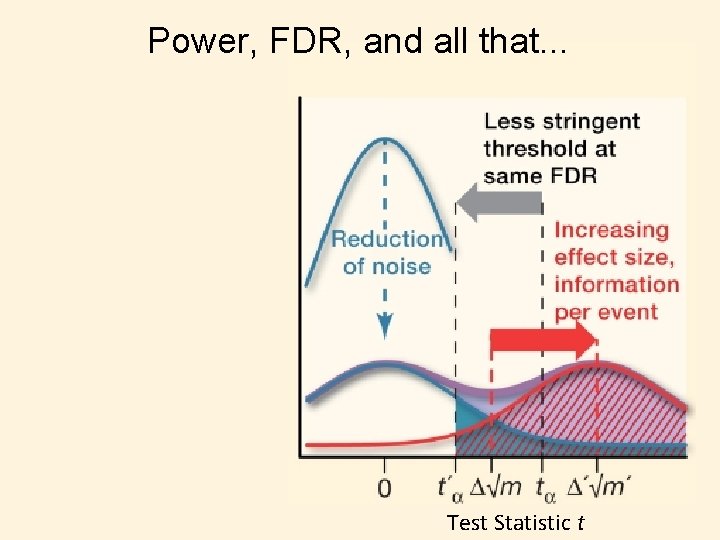
Power, FDR, and all that. . . Test Statistic t
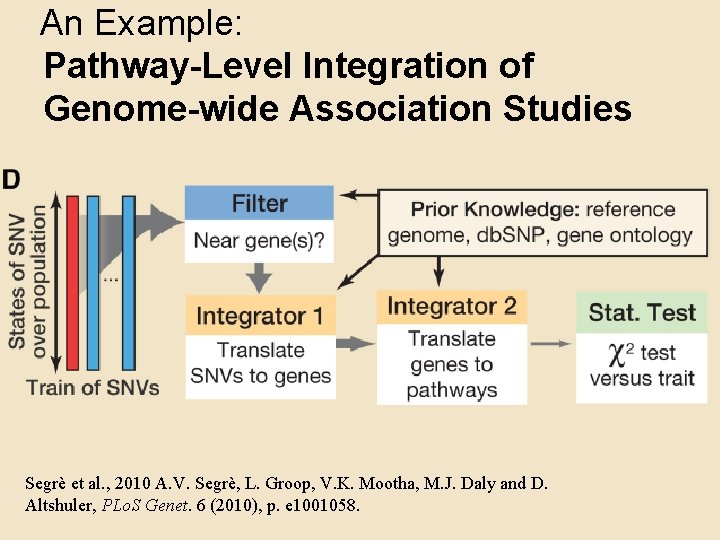
An Example: Pathway-Level Integration of Genome-wide Association Studies Segrè et al. , 2010 A. V. Segrè, L. Groop, V. K. Mootha, M. J. Daly and D. Altshuler, PLo. S Genet. 6 (2010), p. e 1001058.

Classes of biological measurements 1) Molecular States l DNA sequence / genotype: Next-gen sequencing, SNP & CNV arrays l 2) Molecular Networks l Two-hybrid system, co. IP, protein antibody array Gene expression: DNA microarrays, m. RNA sequencing l Protein levels, locations, mods: l l Mass spectrometry, fluorescence microscopy, protein arrays Protein-protein interactions: Protein-DNA interactions: Chromatin IP (chip) sequencing Protein-compound 3) Phenotypic traits Physiological or disease state, binary or quantitative l Growth rate, response to stimulus or stress l Behaviors l
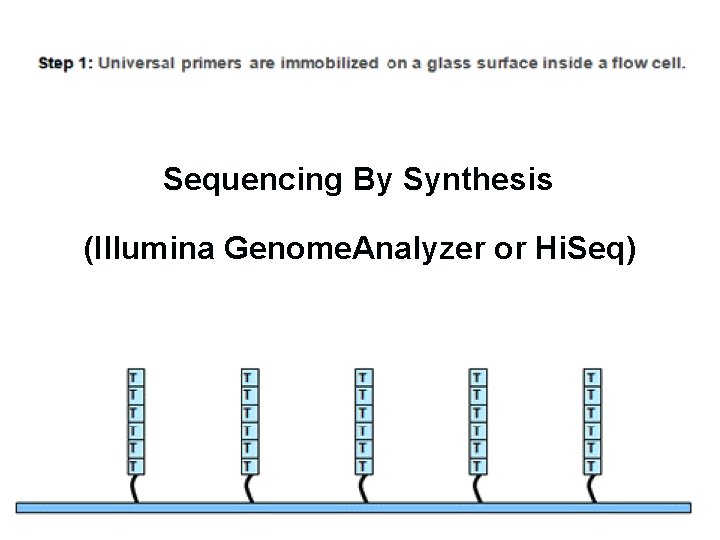
Sequencing By Synthesis (Illumina Genome. Analyzer or Hi. Seq)








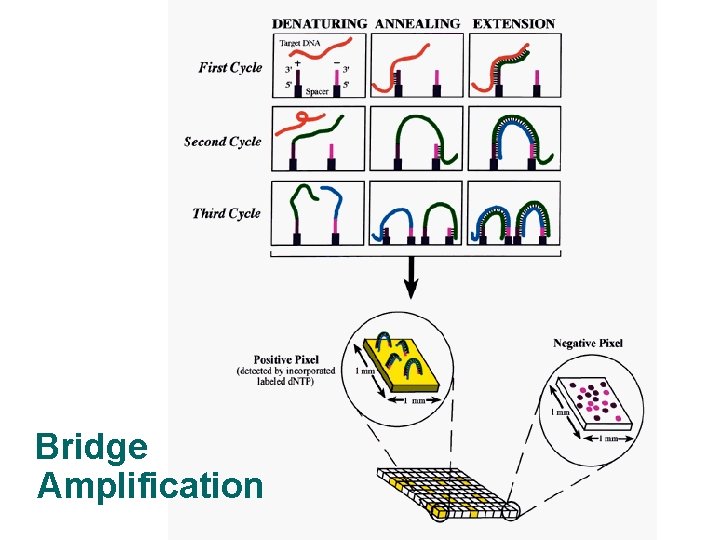
Bridge Amplification

Pyrosequencing Note: No actual houses are burned down in pyrosequencing

Pyrosequencing (Life Sciences / Roche 454) l A luciferase is an enzyme which emits light in the presence of ATP. Several organisms, such as the American firefly and the poisonous Jack-o-lantern mushroom, produce luciferases.

Detecting polymerase activity l l l Recall: Pyrophosphate is also known as PPi, also known as “two phosphate groups stuck together”. During replication, each addition of a d. NTP releases pyrophosphate In the reaction mixture, PPi allows adenosine phosulfate (APS) to be converted to ATP; this ATP allows luciferase to luciferate (emit light). Measures strand extension as it happens

Pyrosequencing cycle l l Add d. ATP. If light is emitted, your sequence starts with A. If not, the d. ATP is degraded (or elutes past immobilized primer). Add d. GTP. If light is emitted, the next base must be a G. Then add T, then C. You now know at least one (maybe more) base of the sequence. Repeat!
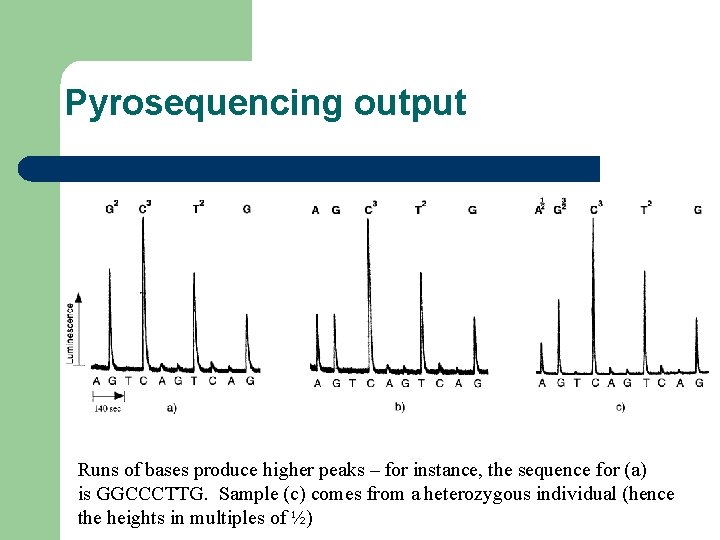
Pyrosequencing output Runs of bases produce higher peaks – for instance, the sequence for (a) is GGCCCTTG. Sample (c) comes from a heterozygous individual (hence the heights in multiples of ½)

The X Prize Foundation In October 2006, the X Prize Foundation established an initiative to promote the development of full genome sequencing technologies, called the Archon X Prize, intending to award $10 million to "the first Team that can build a device and use it to sequence 100 human genomes within 10 days or less, with an accuracy of no more than one error in every 100, 000 bases sequenced, with sequences accurately covering at least 98% of the genome, and at a recurring cost of no more than $10, 000 (US) per genome. ” http: //genomics. xprize. org/

Gene and Protein Expression l l l 26 l The transcriptome is the full complement of RNA molecules produced by a genome The proteome is the full complement of proteins enabled by the transcriptome DNA RNA protein Genome transcriptome proteome 30, 000 genes ? ? ? RNAs ? ? ? proteins? For example, the drosophila gene Dscam can generate 40, 000 distinct transcripts through alternative splicing. What is the minimum number of exons that would be required?

m. RNA Expression: Two dominant approaches RNA sequencing DNA Microarrays 27 Others / older approaches: l EST sequencing l RT-PCR l Differential display l SAGE l Massively parallel signature sequencing (MPSS)

Microarrays Monitors the level of each gene: Is it turned on or off in a particular biological condition? Is this on/off state different between two biological conditions? 28 Microarray is a rectangular grid of spots printed on a glass microscope slide, where each spot contains DNA for a
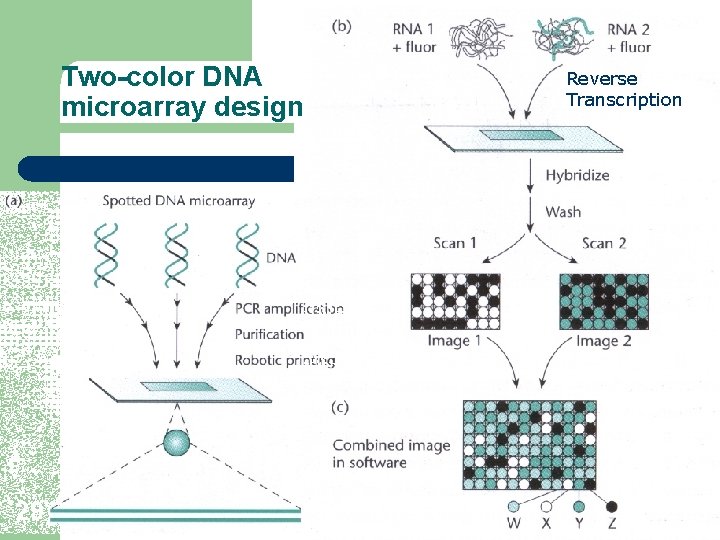
Two-color DNA microarray design 29 Reverse Transcription

Types of microarrays l Spotted (c. DNA) – – l Synthetic (oligo) – – – 30 Robotic transfer of c. DNA clones or PCR products Spotting on nylon membranes or glass slides coated with poly-lysine Direct oligo synthesis on solid microarray substrate Uses photolithography (Affymetrix) or ink-jet printing (Agilent) 100, 000 features per cm 2 l All configurations assume the DNA on the array is in excess of the hybridized sample—thus the kinetics are linear and the spot intensity reflects that amount of hybridized sample. l Labeling can be radioactive, fluorescent (one-color), or two-color

Microarray Spotter 31
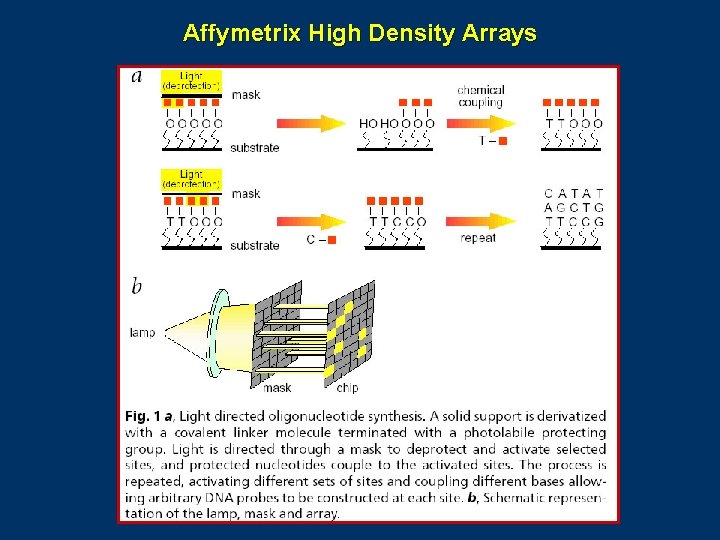
Affymetrix High Density Arrays
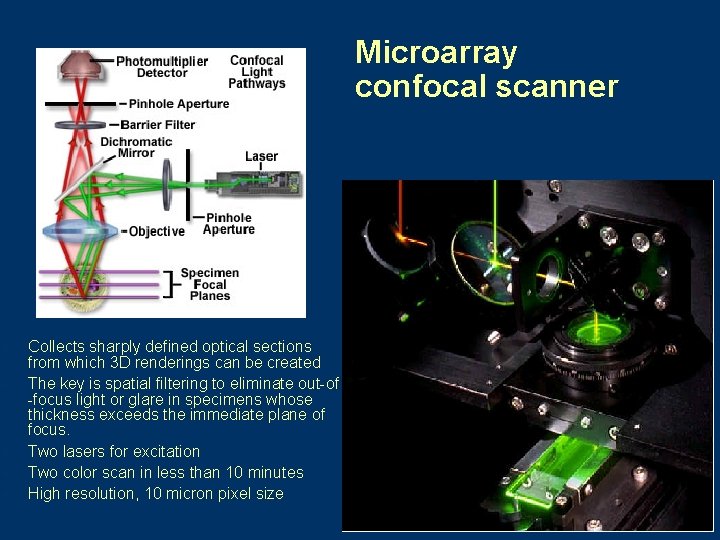
Microarray confocal scanner l l l Collects sharply defined optical sections from which 3 D renderings can be created The key is spatial filtering to eliminate out-of -focus light or glare in specimens whose thickness exceeds the immediate plane of focus. Two lasers for excitation Two color scan in less than 10 minutes High resolution, 10 micron pixel size

Next-Gen Sequencing of m. RNAs c. DNA = complementary or copy DNA EST = Expressed Sequence Tag l l l The microarray could be described as a “closed system” because information about RNAs is limited by the targets available for hybridization. RNAs not represented on the array are not interrogated. Direct sequencing of c. DNAs overcomes this problem by large-scale random sampling of sequences from a wholecell RNA extract Statistical counting of distinct sequences provides a precise estimate of expression level c. DNA library can be normalized to capture rare messages Has been dramatically enabled by large scale sequencing
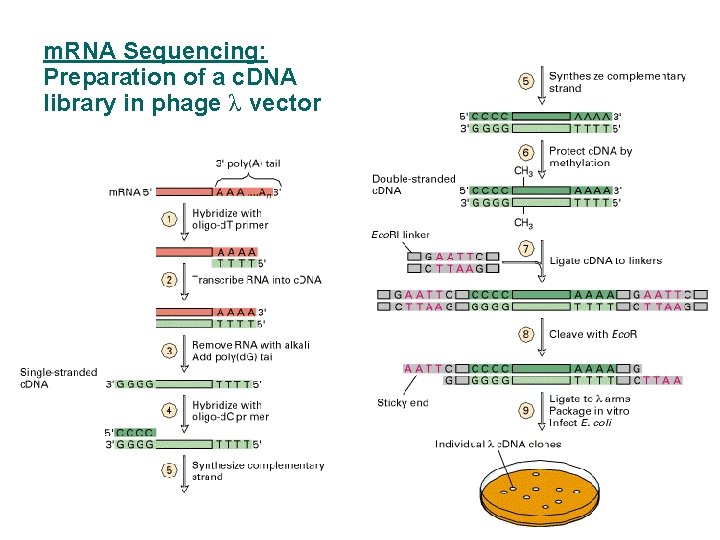
m. RNA Sequencing: Preparation of a c. DNA library in phage λ vector

Proteomics MS / MS 1 D and 2 D SDS PAGE 36

Mass spectrometry Mass spectrometers consist of 3 essential parts – – – 37 Ionization source: Converts peptides into gas-phase ions (MALDI + ESI) Mass analyzer: Separates ions by mass to charge (m/z) ratio (Ion trap, time of flight, quadrupole) Ion detector: Current over time indicates amount of signal at each m/z value
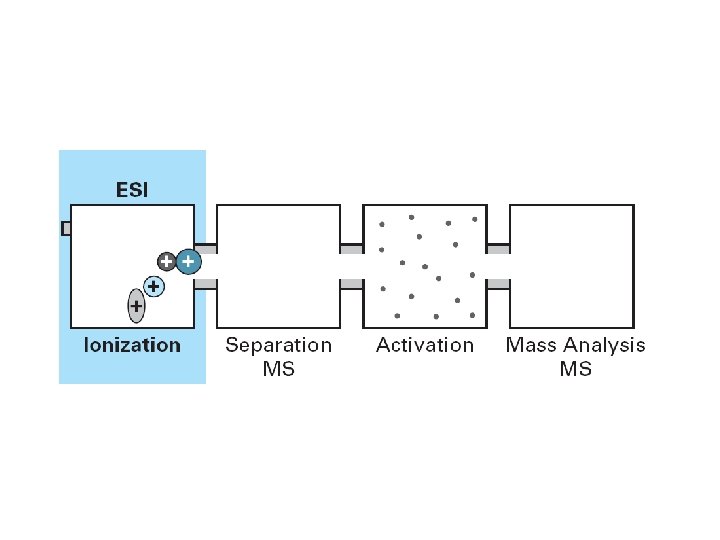
MS/MS Overview
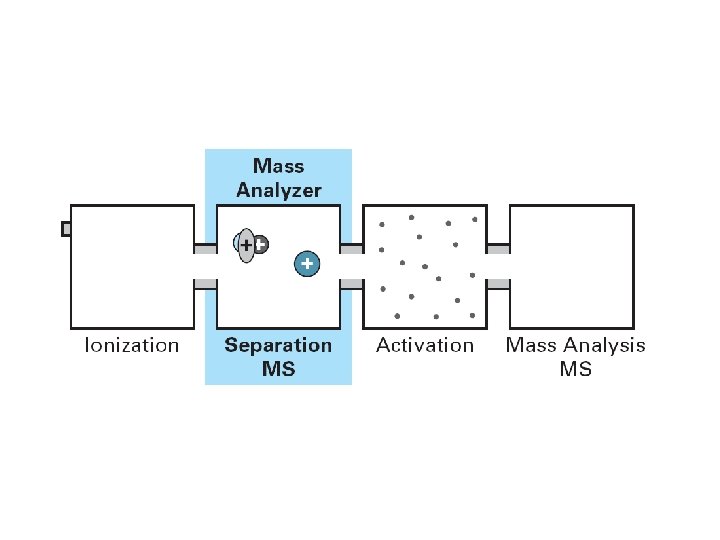
MS/MS Overview


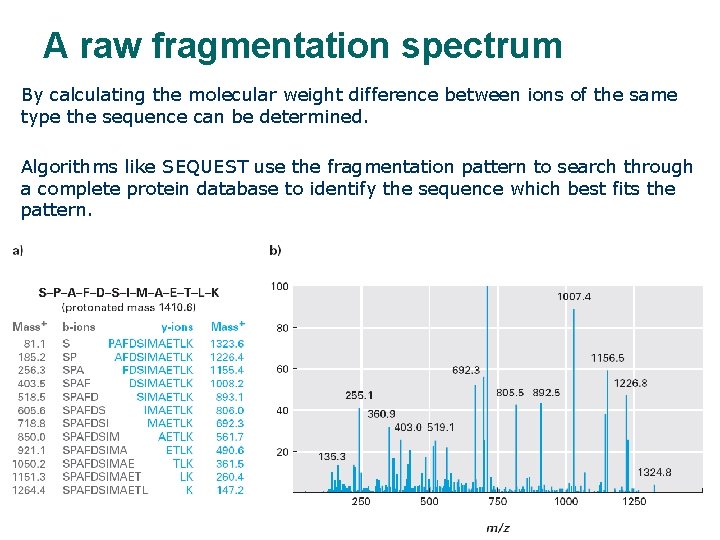
A raw fragmentation spectrum By calculating the molecular weight difference between ions of the same type the sequence can be determined. Algorithms like SEQUEST use the fragmentation pattern to search through a complete protein database to identify the sequence which best fits the pattern.

43 Tandem Mass Spec (MS/MS)
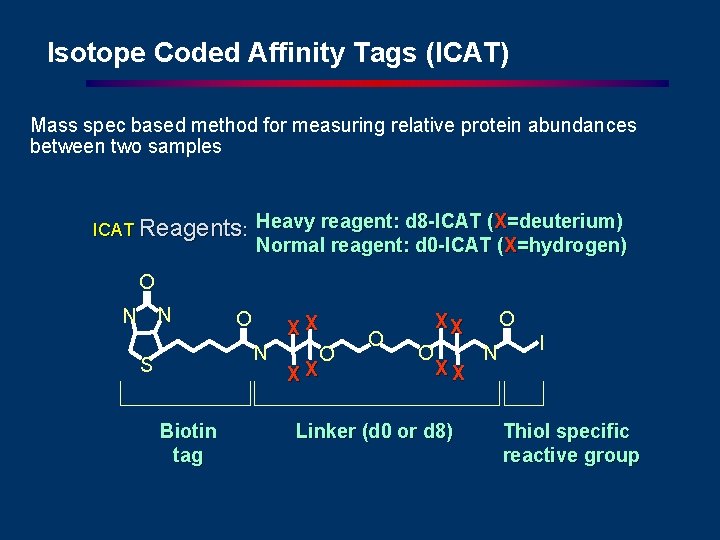
Isotope Coded Affinity Tags (ICAT) Mass spec based method for measuring relative protein abundances between two samples ICAT Reagents: Heavy reagent: d 8 -ICAT (X=deuterium) Normal reagent: d 0 -ICAT (X=hydrogen) O N N O XX N S Biotin tag O XX Linker (d 0 or d 8) N I Thiol specific reactive group
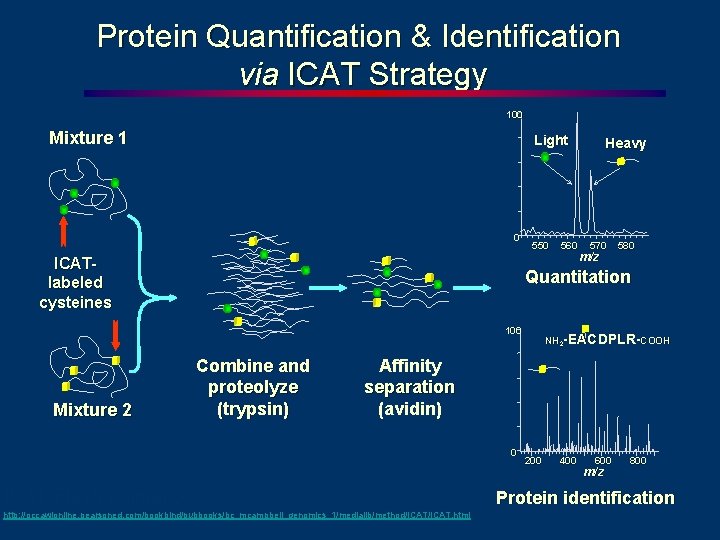
Protein Quantification & Identification via ICAT Strategy 100 Mixture 1 Light 0 ICATlabeled cysteines 550 570 m/z 580 Quantitation 100 Mixture 2 560 Heavy Combine and proteolyze (trypsin) NH 2 -EACDPLR-COOH Affinity separation (avidin) 0 ICAT Flash animation: http: //occawlonline. pearsoned. com/bookbind/pubbooks/bc_mcampbell_genomics_1/medialib/method/ICAT. html 200 400 600 m/z 800 Protein identification
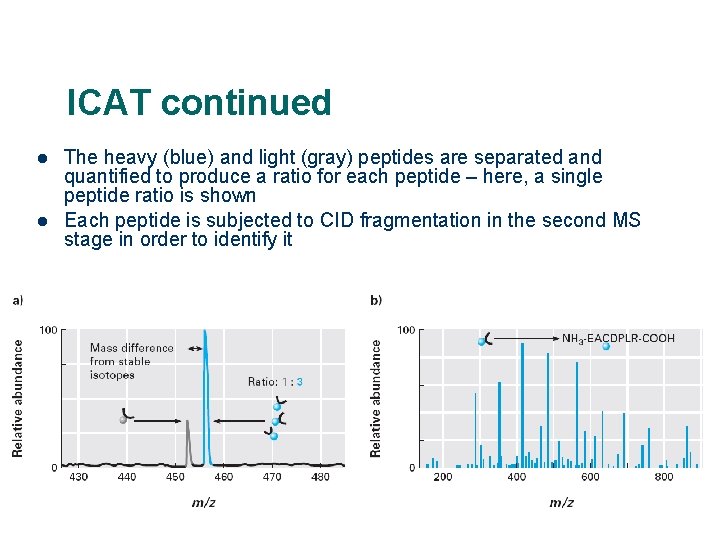
ICAT continued l l The heavy (blue) and light (gray) peptides are separated and quantified to produce a ratio for each peptide – here, a single peptide ratio is shown Each peptide is subjected to CID fragmentation in the second MS stage in order to identify it
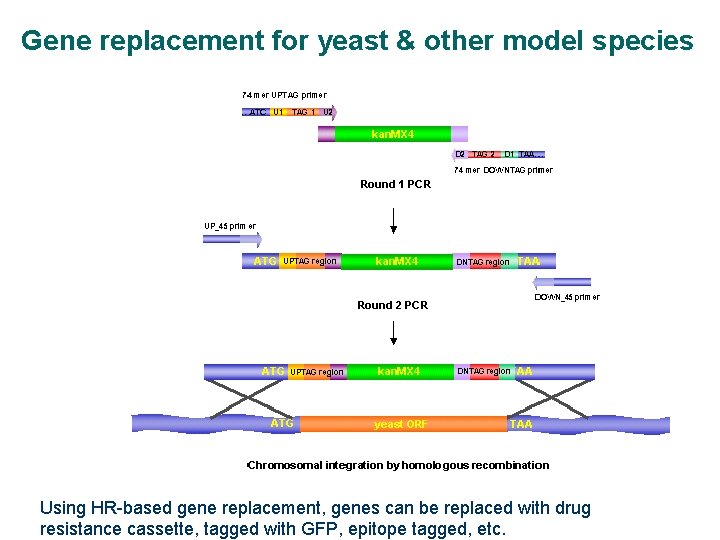
Gene replacement for yeast & other model species Using HR-based gene replacement, genes can be replaced with drug resistance cassette, tagged with GFP, epitope tagged, etc.
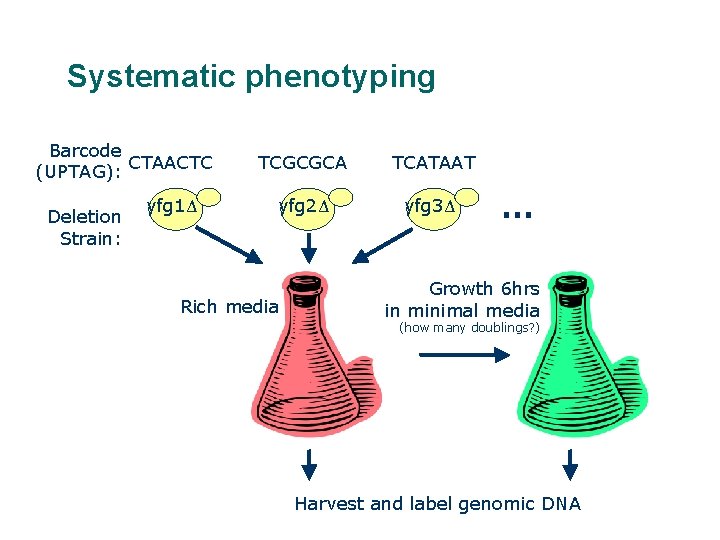
Systematic phenotyping Barcode CTAACTC (UPTAG): Deletion Strain: yfg 1Δ TCGCGCA TCATAAT yfg 2Δ yfg 3Δ Rich media … Growth 6 hrs in minimal media (how many doublings? ) Harvest and label genomic DNA
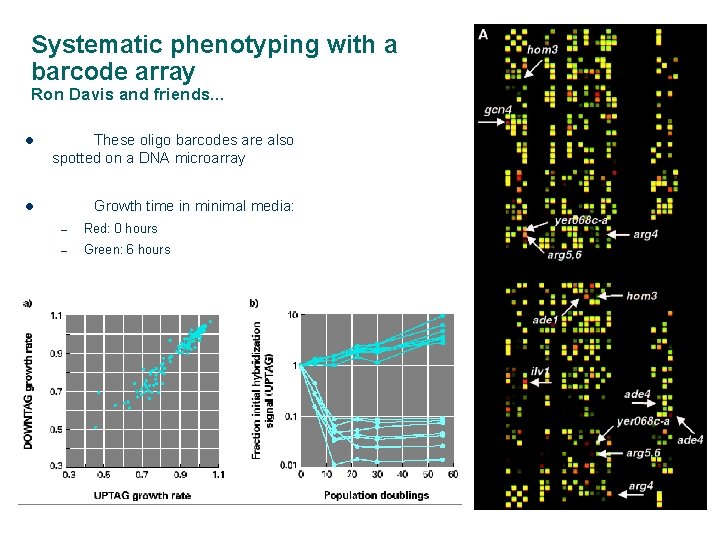
Systematic phenotyping with a barcode array Ron Davis and friends… l These oligo barcodes are also spotted on a DNA microarray l Growth time in minimal media: – Red: 0 hours – Green: 6 hours
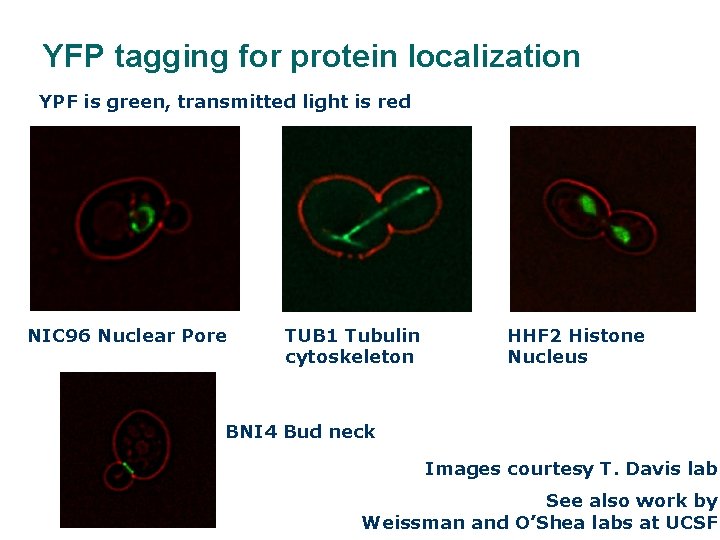
YFP tagging for protein localization YPF is green, transmitted light is red NIC 96 Nuclear Pore TUB 1 Tubulin cytoskeleton HHF 2 Histone Nucleus BNI 4 Bud neck Images courtesy T. Davis lab See also work by Weissman and O’Shea labs at UCSF

Molecular Interactions Among proteins, m. RNA, small molecules, and so on… 51
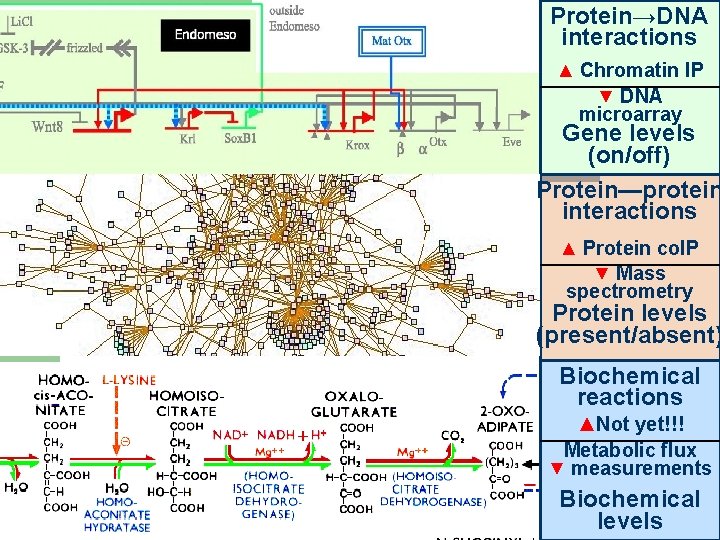
Protein→DNA interactions ▲ Chromatin IP ▼ DNA microarray Gene levels (on/off) Protein—protein interactions ▲ Protein co. IP ▼ Mass spectrometry Protein levels (present/absent) Biochemical reactions ▲Not yet!!! Metabolic flux ▼ measurements 52 Biochemical levels
Measurements of molecular interactions Protein-protein interactions l l l Yeast-two-hybrid Kinase-substrate assays Co-immunoprecipitation w/ mass spec Protein-DNA interactions l Ch. IP-on-chip and Ch. IP-seq Genetic interactions 53 l Systematic Genetic Analysis
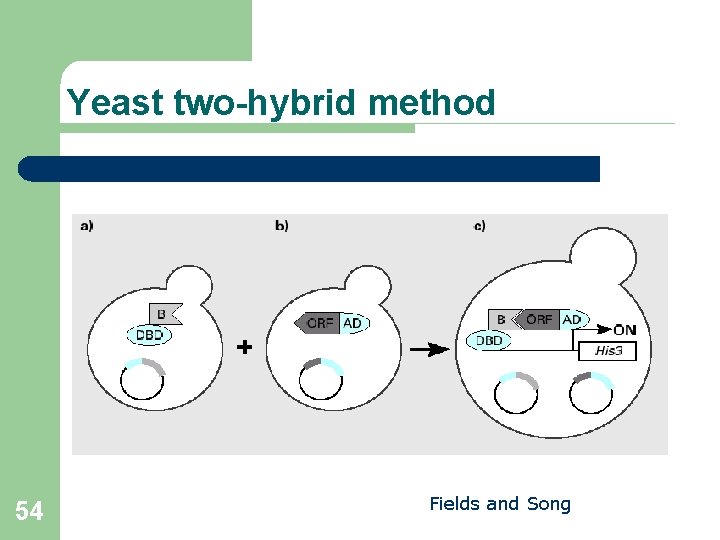
Yeast two-hybrid method 54 Fields and Song
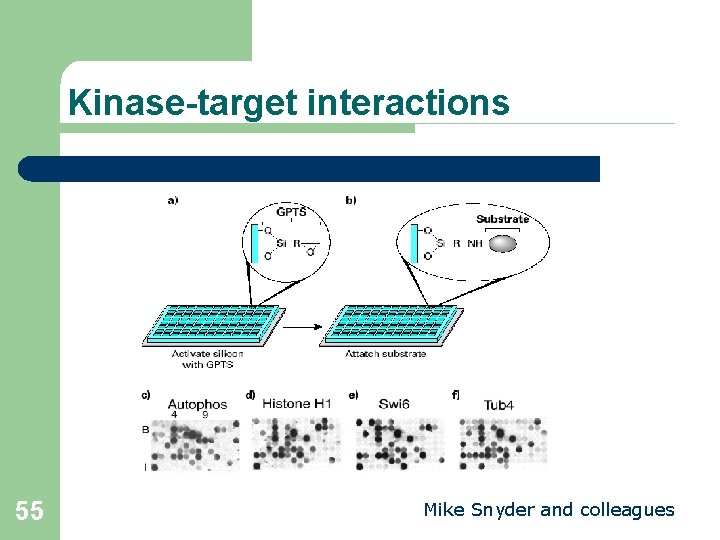
Kinase-target interactions 55 Mike Snyder and colleagues
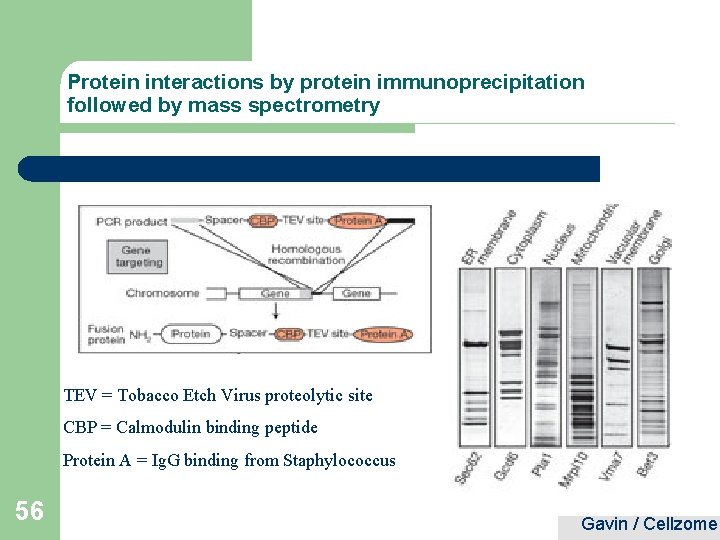
Protein interactions by protein immunoprecipitation followed by mass spectrometry TEV = Tobacco Etch Virus proteolytic site CBP = Calmodulin binding peptide Protein A = Ig. G binding from Staphylococcus 56 Gavin / Cellzome

Ch. IP measurement of protein→DNA interactions From Figure 1 of Simon et al. Cell
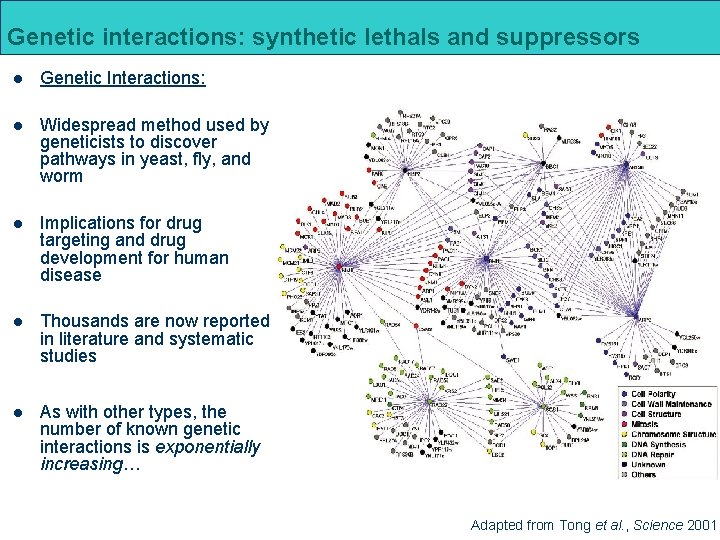
Genetic interactions: synthetic lethals and suppressors l Genetic Interactions: l Widespread method used by geneticists to discover pathways in yeast, fly, and worm l Implications for drug targeting and drug development for human disease l Thousands are now reported in literature and systematic studies l As with other types, the number of known genetic interactions is exponentially increasing… Adapted from Tong et al. , Science 2001
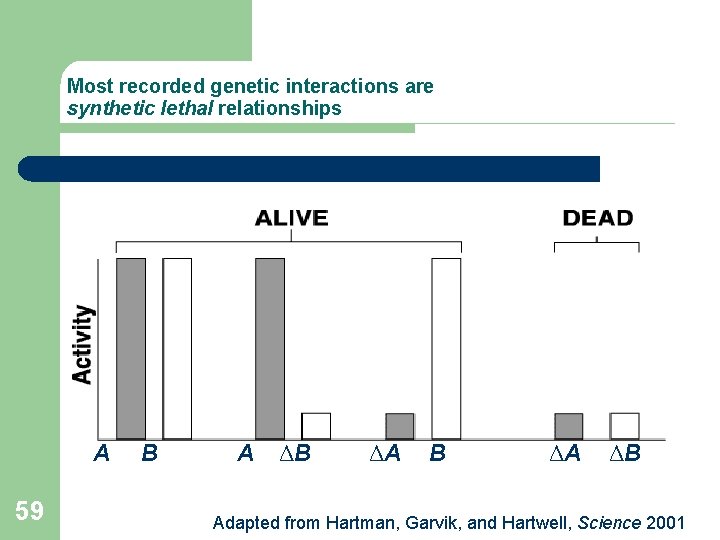
Most recorded genetic interactions are synthetic lethal relationships A 59 B A ΔB ΔA ΔB Adapted from Hartman, Garvik, and Hartwell, Science 2001
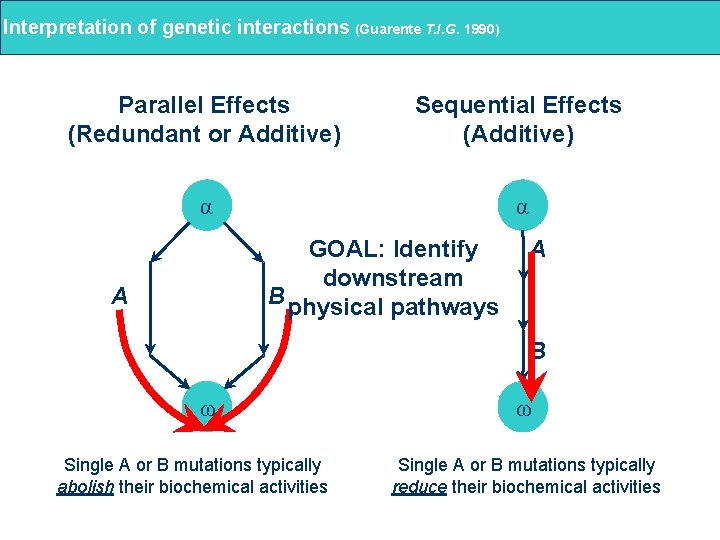
Interpretation of genetic interactions (Guarente T. I. G. 1990) Parallel Effects (Redundant or Additive) Sequential Effects (Additive) α α GOAL: Identify downstream B physical pathways A A B ω Single A or B mutations typically abolish their biochemical activities ω Single A or B mutations typically reduce their biochemical activities
- Slides: 60