Biometrical Genetics Shaun Purcell Twin Workshop March 2004

Biometrical Genetics Shaun Purcell Twin Workshop, March 2004

Single locus model 1. Genetic effects → variance components 2. Genetic effects → familial covariances 3. Variance components → familial covariances
![ADE Model for twin data [0. 25/1] 1 E D e [0. 5/1] 1 ADE Model for twin data [0. 25/1] 1 E D e [0. 5/1] 1](http://slidetodoc.com/presentation_image_h2/001bd1770f294e3590cb451e6fdb7dd6/image-3.jpg)
ADE Model for twin data [0. 25/1] 1 E D e [0. 5/1] 1 1 d PT 1 A a 1 1 1 A D a d PT 2 E e

Some Components of a Genetic Theory • POPULATION MODEL – Allele & genotype frequencies • TRANSMISSION MODEL – Mendelian segregation – Identity by descent & genetic relatedness • PHENOTYPE MODEL – Biometrical model of quantitative traits – Additive & dominance components

MENDELIAN GENETICS
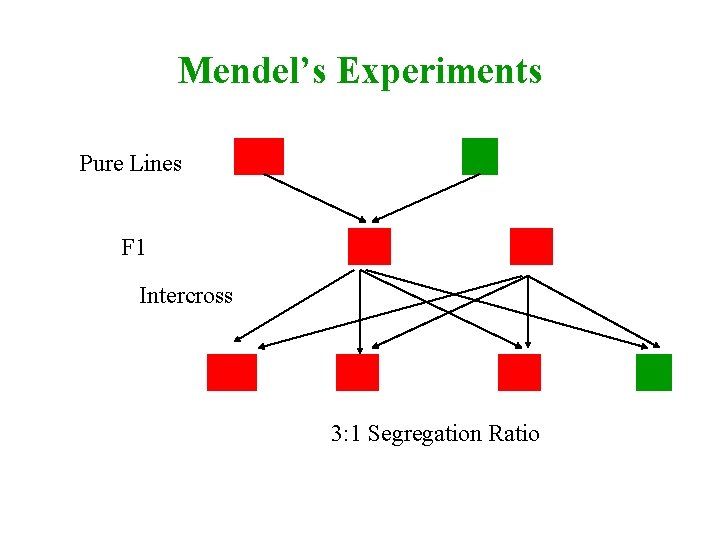
Mendel’s Experiments AA Pure Lines F 1 aa Aa Aa Intercross AA Aa Aa 3: 1 Segregation Ratio aa
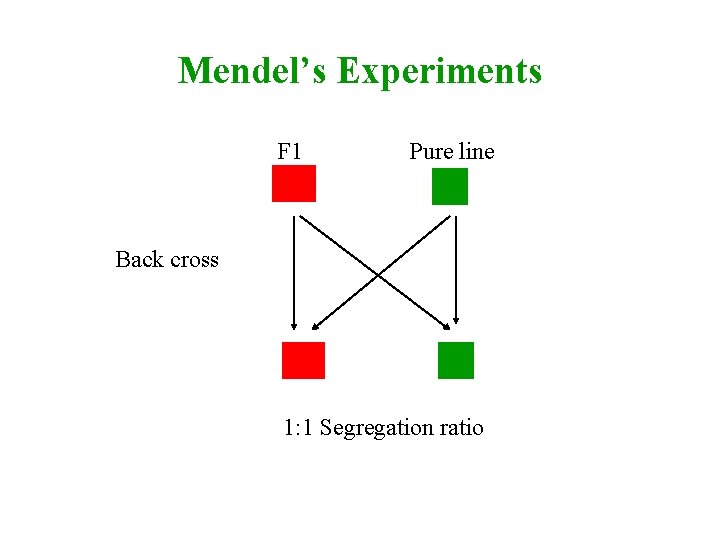
Mendel’s Experiments F 1 Aa Pure line aa Back cross Aa aa 1: 1 Segregation ratio

Mendel’s Experiments AA Pure Lines F 1 aa Aa Aa Intercross AA Aa Aa 3: 1 Segregation Ratio aa
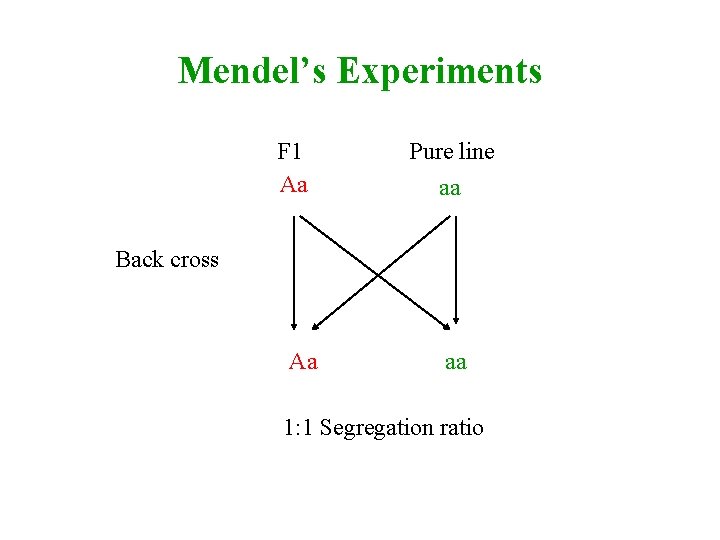
Mendel’s Experiments F 1 Aa Pure line aa Back cross Aa aa 1: 1 Segregation ratio

Mendel’s Law of Segregation Maternal A 3 A 4 Gametes A 1 Paternal A 2 ½ ½ ½ A 1 A 3 ¼ A 1 A 4 ¼ ½ A 2 A 3 ¼ A 2 A 4 Meiosis/Segregation ¼

PHENOTYPE MODEL
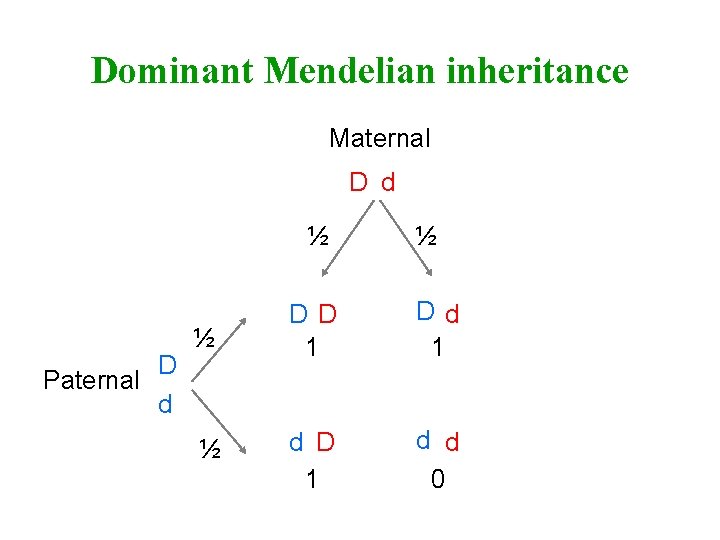
Dominant Mendelian inheritance Maternal D d ½ D Paternal d ½ ½ DD 1 Dd 1 ½ d D 1 d d 0
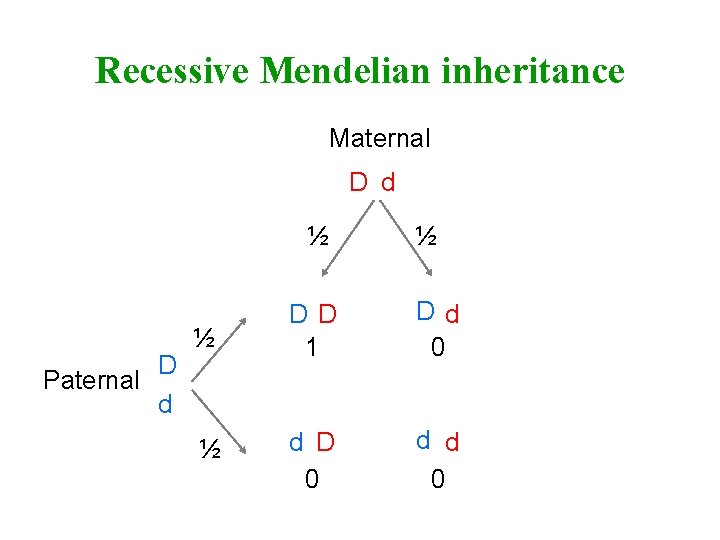
Recessive Mendelian inheritance Maternal D d ½ D Paternal d ½ ½ DD 1 Dd 0 ½ d D 0 d d 0
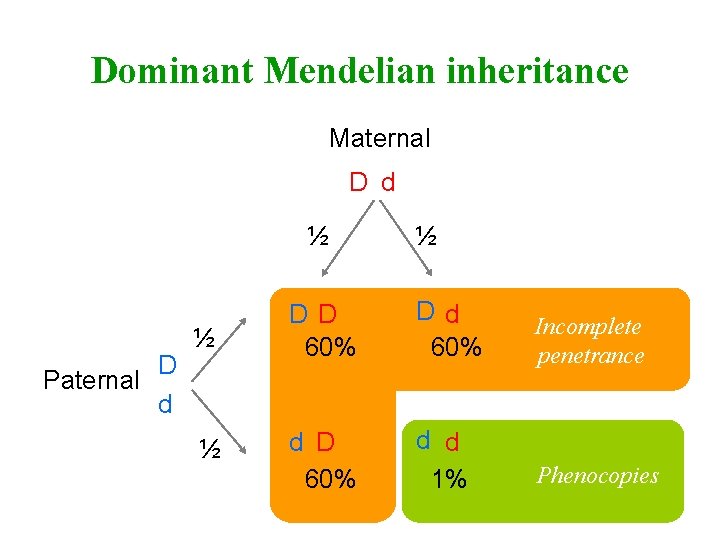
Dominant Mendelian inheritance Maternal D d ½ D Paternal d ½ ½ DD 60% Dd 60% Incomplete penetrance ½ d D 60% d d 1% Phenocopies
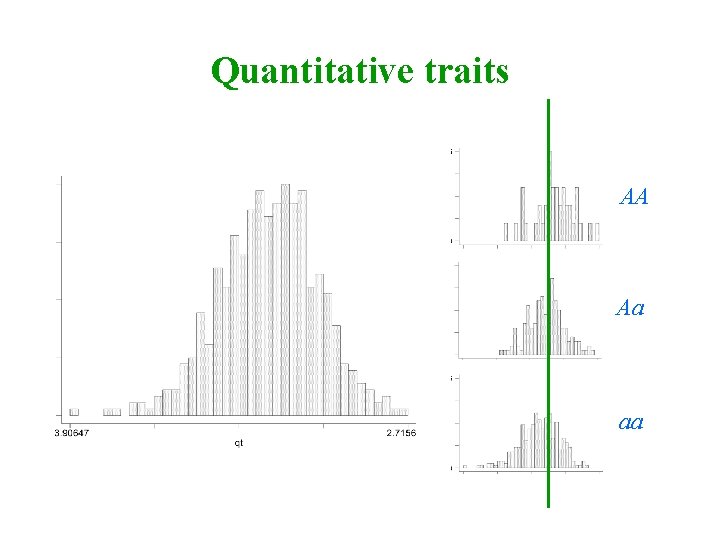
Quantitative traits AA Aa aa
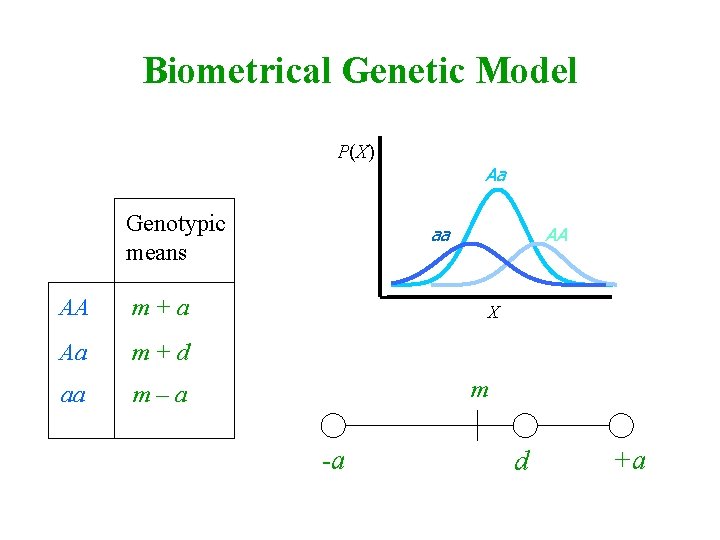
Biometrical Genetic Model P(X) Genotypic means AA m+a Aa m+d aa m–a Aa aa AA X m -a d +a

POPULATION MODEL

Population Frequencies • A single locus, with two alleles – Biallelic / diallelic – Single nucleotide polymorphism, SNP • Alleles A and a – Frequency of A is p – Frequency of a is q = 1 – p • Every individual inherits two copies – A genotype is the combination of the two alleles – e. g. AA, aa (the homozygotes) or Aa (the heterozygote)

Genotype Frequencies (random mating) A a A p 2 pq p a qp q 2 q p q Hardy-Weinberg Equilibrium frequencies P(AA) = p 2 P(Aa) = 2 pq P(aa) = q 2

Before we proceed, some basic statistical tools…
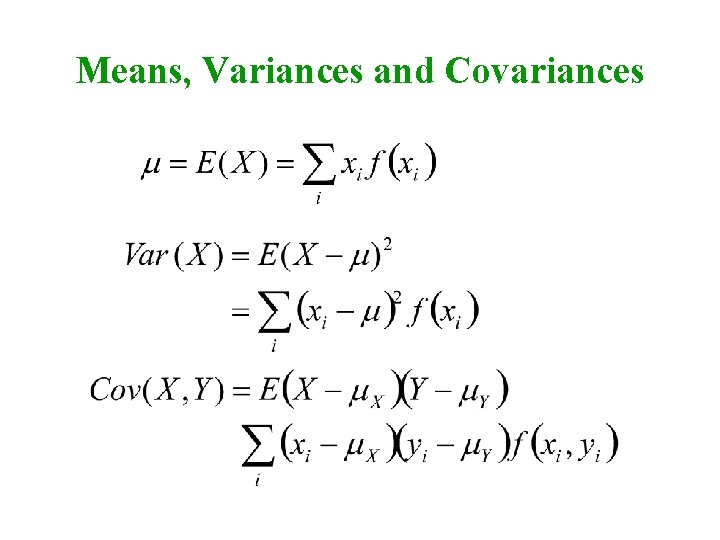
Means, Variances and Covariances

Biometrical Model for Single Locus Genotype AA Aa aa Frequency p 2 2 pq q 2 Effect (x) a d -a Residual var 2 2 2 Mean m = p 2(a) + 2 pq(d) + q 2(-a) = a(p-q) + 2 pqd

Biometrical Model for Single Locus Genotype Frequency (x-m)2 AA p 2 (a-m)2 Aa 2 pq (d-m)2 aa q 2 (-a-m)2 Variance = (a-m)2 p 2 + (d-m)22 pq + (-a-m)2 q 2 = VG (Broad-sense) heritability at this loci = VG / VTOT (Broad-sense) heritability = ΣLVG / VTOT

Additive and dominance effects • Additive effects are the main effects of individual alleles: ‘gene-dosage’ – Parents transmit alleles, not genotypes • Dominance effects represent an interaction between the two alleles – i. e. if the heterozygote is not midway between the two homozygotes
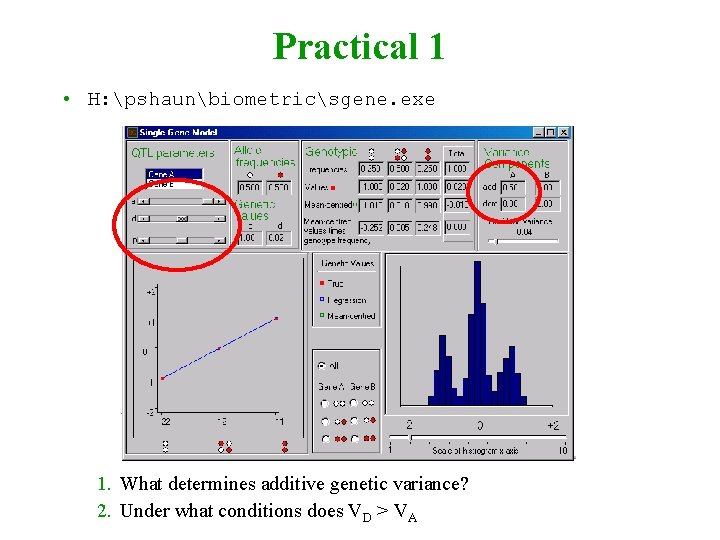
Practical 1 • H: pshaunbiometricsgene. exe 1. What determines additive genetic variance? 2. Under what conditions does VD > VA

Some conclusions 1. Additive genetic variance depends on allele frequency p & additive genetic value a as well as dominance deviation d 2. Additive genetic variance typically greater than dominance variance
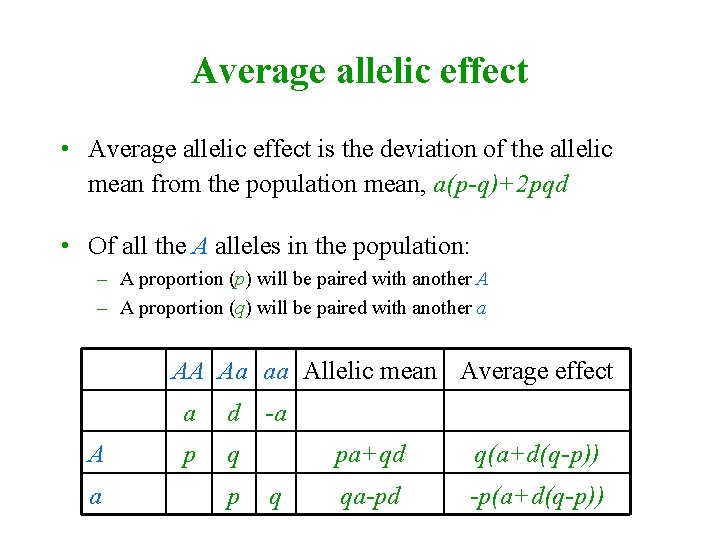
Average allelic effect • Average allelic effect is the deviation of the allelic mean from the population mean, a(p-q)+2 pqd • Of all the A alleles in the population: – A proportion (p) will be paired with another A – A proportion (q) will be paired with another a AA Aa aa Allelic mean Average effect A a a d -a p q pa+qd q(a+d(q-p)) qa-pd -p(a+d(q-p))

Average allelic effect • Denote the average allelic effects as α αA = q(a+d(q-p)) αa = -p(a+d(q-p)) • If only two alleles exist, we can define the average effect of allele substitution α = αA – αa α = (q-(-p))(a+d(q-p)) = (a+d(q-p)) • Therefore, αA = qα and αa = -pα

Additive genetic variance • AA Aa aa The variance of the average allelic effects Freq. p 2 2 pq q 2 Additive effect 2αA αA + αa 2αa = 2 qα = (q-p)α = -2 pα VA = p 2(2 qα)2 + 2 pq((q-p)α)2 + q 2(-2 pα)2 = 2 pqα 2 = 2 pq(a+d(q-p))2

Additive genetic variance • If there is no dominance VA = 2 pqa 2 • If p = q VA = ½a 2
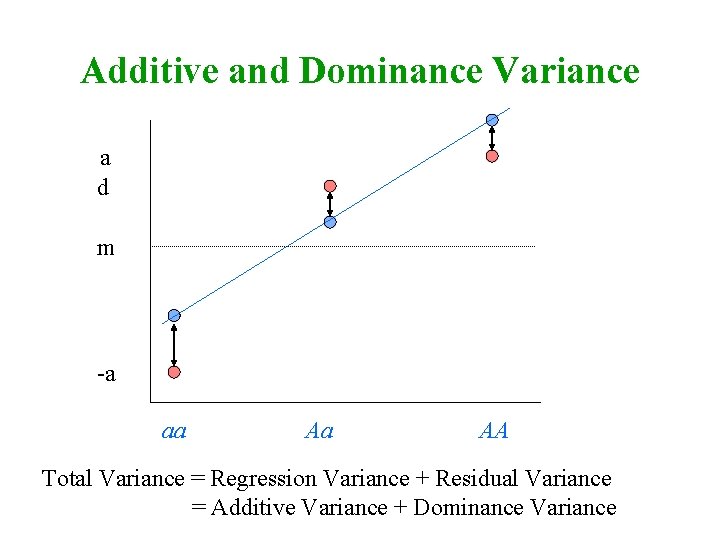
Additive and Dominance Variance a d m -a aa Aa AA Total Variance = Regression Variance + Residual Variance = Additive Variance + Dominance Variance

Biometrical Model for Single Locus Genotype AA Aa aa Frequency p 2 2 pq q 2 (x-m)2 (a-m)2 (d-m)2 (-a-m)2 Variance = (a-m)2 p 2 + (d-m)22 pq + (-a-m)2 q 2 = 2 pq[a+(q-p)d]2 + (2 pqd)2 VG = VA + VD

VA
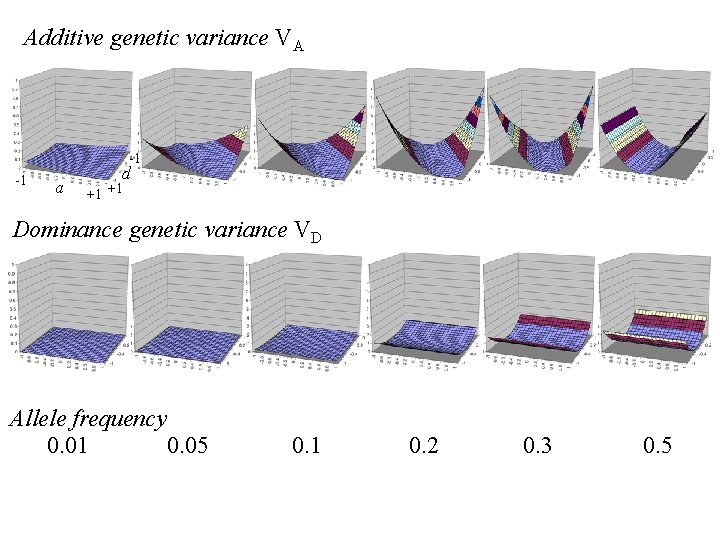
Additive genetic variance VA -1 -1 a d +1 +1 Dominance genetic variance VD Allele frequency 0. 01 0. 05 0. 1 0. 2 0. 3 0. 5
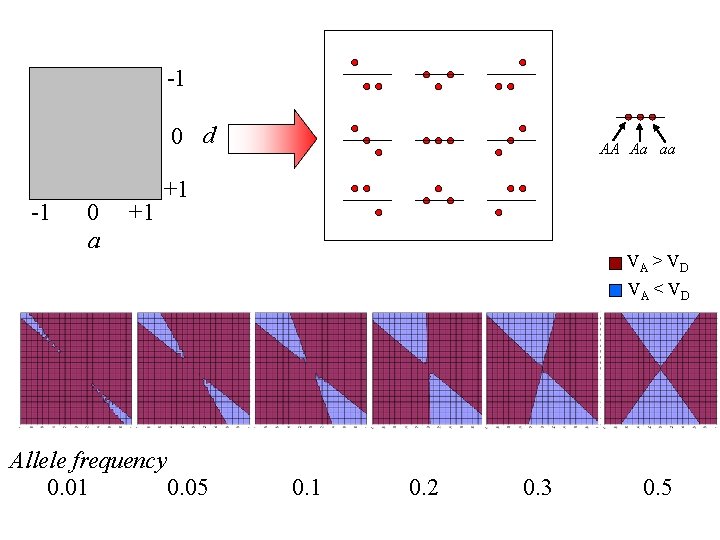
-1 0 d -1 0 a +1 AA Aa aa +1 VA > V D VA < V D Allele frequency 0. 01 0. 05 0. 1 0. 2 0. 3 0. 5

Cross-Products of Deviations for Pairs of Relatives AA Aa aa AA (a-m)2 Aa (a-m)(d-m) aa (a-m)(-a-m)(d-m) (-a-m)2 (d-m)2 The covariance between relatives of a certain class is the weighted average of these cross-products, where each cross-product is weighted by its frequency in that class:

Covariance of MZ Twins AA Aa AA p 2 Aa 0 2 pq aa 0 0 aa q 2 Covariance = (a-m)2 p 2 + (d-m)22 pq + (-a-m)2 q 2 = 2 pq[a+(q-p)d]2 + (2 pqd)2 = VA + VD

Covariance for Parent-offspring (P-O) AA Aa AA ? Aa ? ? aa ? • Exercise 2 : to calculate frequencies of parentoffspring combinations, in terms of allele frequencies p and q.

Exercise 2 • e. g. given an AA father, an AA offspring can come from either AA x AA or AA x Aa parental mating types AA x AA will occur p 2 × p 2 = p 4 and have AA offspring Prob()=1 AA x Aa will occur p 2 × 2 pq = 2 p 3 q and have AA offspring Prob()=0. 5 and have Aa offspring Prob()=0. 5 Therefore, P(AA father & AA offspring) = p 4 + p 3 q = p 3(p+q) = p 3

Covariance for Parent-offspring (P-O) AA Aa AA p 3 Aa ? ? aa ? • AA offspring from AA parents = p 4+p 3 q = p 3(p+q) = p 3
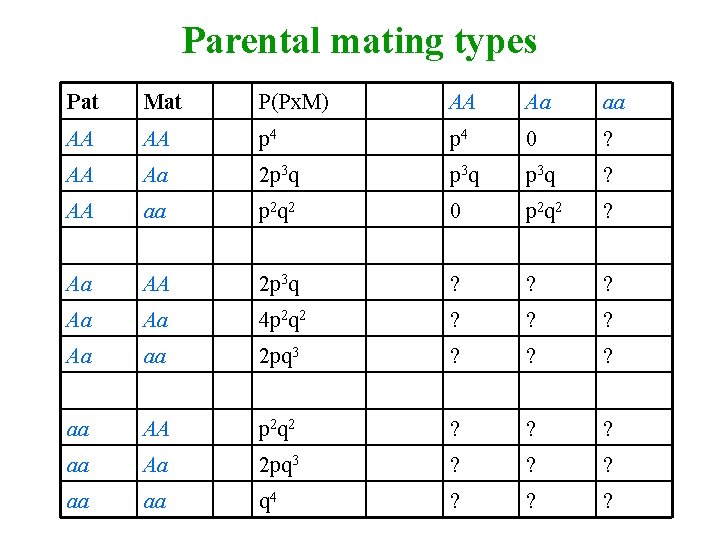
Parental mating types Pat Mat P(Px. M) AA Aa aa AA AA p 4 0 ? AA Aa 2 p 3 q ? AA aa p 2 q 2 0 p 2 q 2 ? Aa AA 2 p 3 q ? ? ? Aa Aa 4 p 2 q 2 ? ? ? Aa aa 2 pq 3 ? ? ? aa AA p 2 q 2 ? ? ? aa Aa 2 pq 3 ? ? ? aa aa q 4 ? ? ?

Covariance for Parent-offspring (P-O) AA Aa AA p 3 Aa p 2 q ? aa ? • AA offspring from AA parents = p 4+p 3 q = p 3(p+q) = p 3 • Aa offspring from AA parents = p 3 q+p 2 q 2 = p 2 q(p+q) = p 2 q
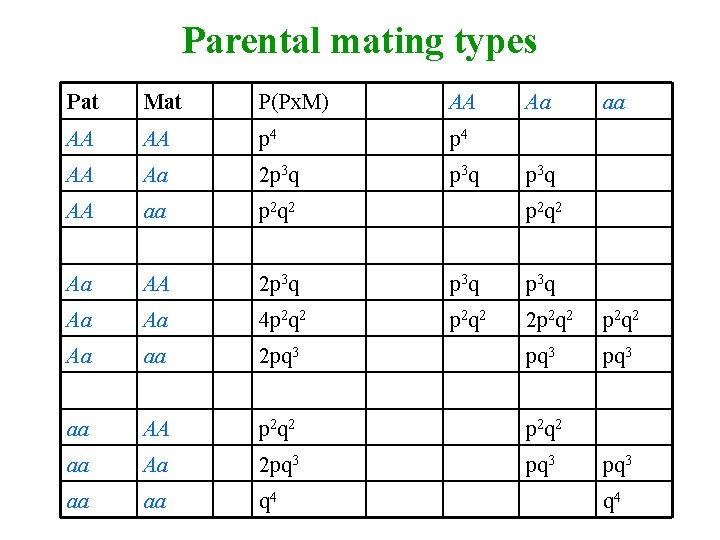
Parental mating types Pat Mat P(Px. M) AA Aa aa AA AA p 4 AA Aa 2 p 3 q AA aa p 2 q 2 Aa AA 2 p 3 q Aa Aa 4 p 2 q 2 2 p 2 q 2 Aa aa 2 pq 3 aa AA p 2 q 2 aa Aa 2 pq 3 aa aa q 4 p 3 q p 2 q 2 pq 3 q 4

Covariance for Parent-offspring (P-O) AA Aa AA p 3 Aa p 2 q pq aa 0 pq 2 aa q 3 Covariance = (a-m)2 p 3 + (d-m)2 pq + (-a-m)2 q 3 + (a-m)(d-m)2 p 2 q + (-a-m)(d-m)2 pq 2 = pq[a+(q-p)d]2 = VA / 2

Covariance for Unrelated Pairs (U) AA Aa AA p 4 Aa 2 p 3 q 4 p 2 q 2 aa p 2 q 2 2 pq 3 aa q 4 Covariance = (a-m)2 p 4 + (d-m)24 p 2 q 2 + (-a-m)2 q 4 + (a-m)(d-m)4 p 3 q + (-a-m)(d-m)4 pq + (a-m)(-a-m)2 p 2 q 2 =0?

IDENTITY BY DESCENT
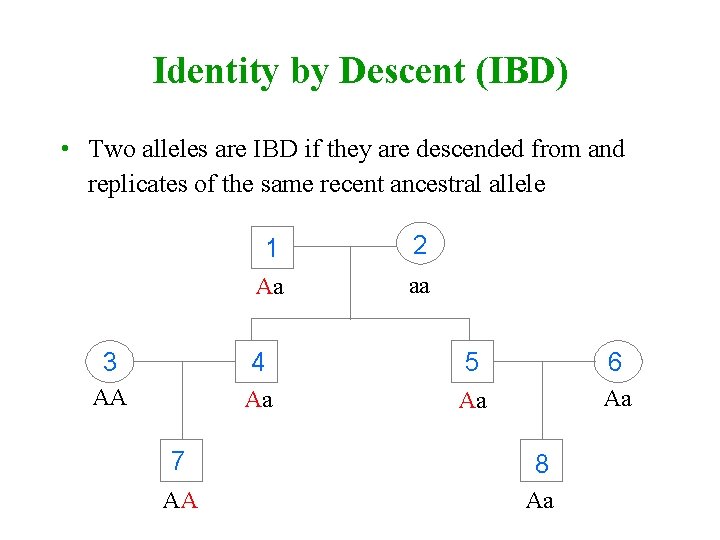
Identity by Descent (IBD) • Two alleles are IBD if they are descended from and replicates of the same recent ancestral allele 1 2 Aa aa 3 4 5 6 AA Aa Aa Aa 7 8 AA Aa
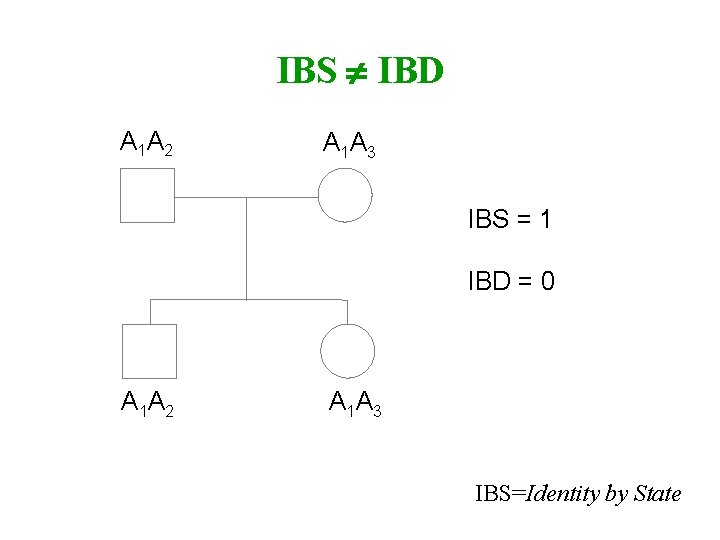
IBS IBD A 1 A 2 A 1 A 3 IBS = 1 IBD = 0 A 1 A 2 A 1 A 3 IBS=Identity by State
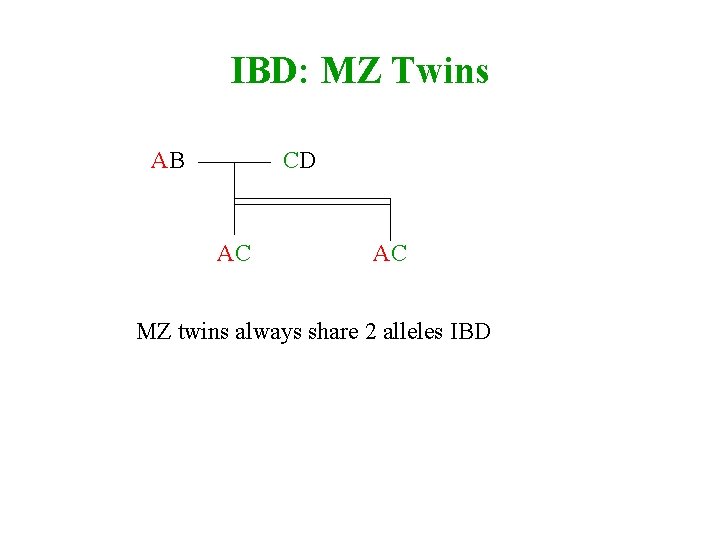
IBD: MZ Twins AB CD AC AC MZ twins always share 2 alleles IBD
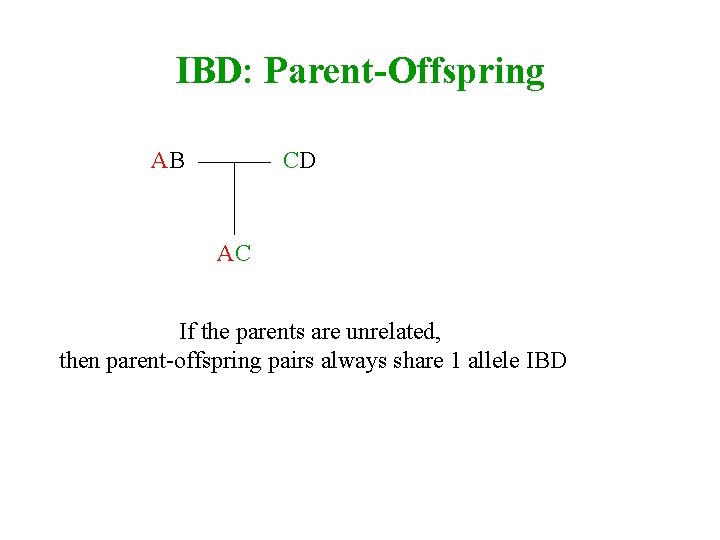
IBD: Parent-Offspring AB CD AC If the parents are unrelated, then parent-offspring pairs always share 1 allele IBD

IBD: Unrelated individuals AB CD If two individuals are unrelated, they always share 0 allele IBD
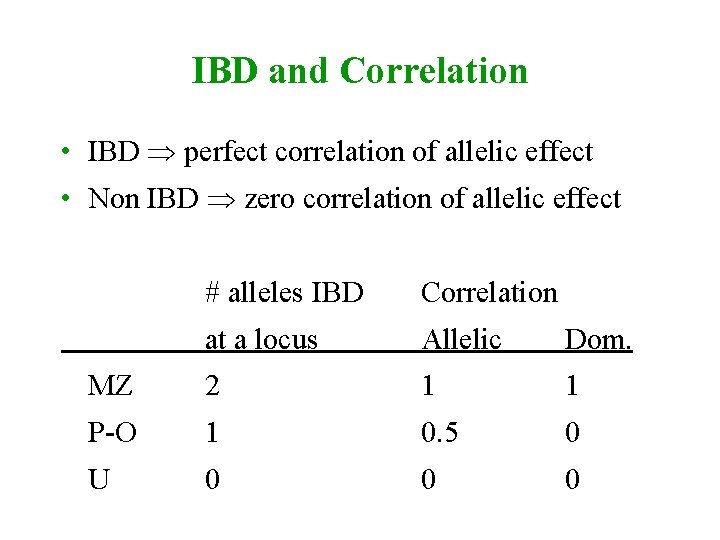
IBD and Correlation • IBD perfect correlation of allelic effect • Non IBD zero correlation of allelic effect # alleles IBD Correlation at a locus Allelic Dom. MZ 2 1 1 P-O 1 0. 5 0 U 0 0 0

Covariance between relatives • Partition of variance Partition of covariance • Overall covariance = sum of covariances of all components • Covariance of component between relatives = correlation of component variance due to component
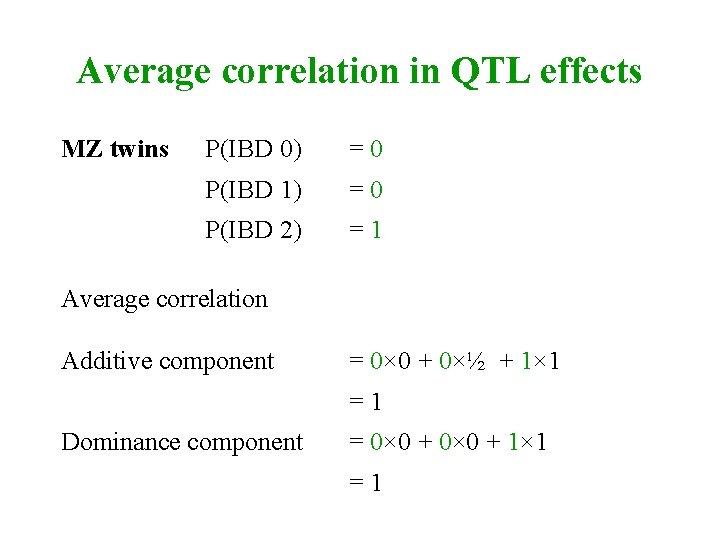
Average correlation in QTL effects MZ twins P(IBD 0) =0 P(IBD 1) =0 P(IBD 2) =1 Average correlation Additive component = 0× 0 + 0×½ + 1× 1 =1 Dominance component = 0× 0 + 1× 1 =1

Average correlation in QTL effects P-O P(IBD 0) =0 P(IBD 1) =1 P(IBD 2) =0 Average correlation Additive component = 0× 0 + 1×½ + 0× 1 =½ Dominance component = 0× 0 + 1× 0 + 0× 1 =0
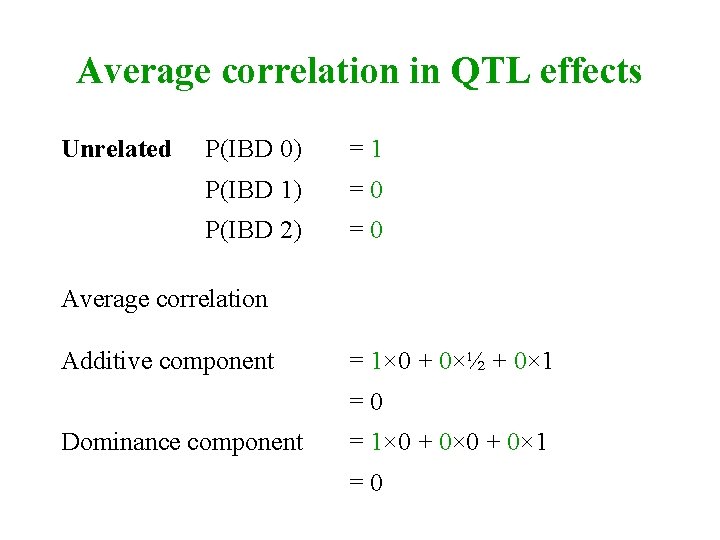
Average correlation in QTL effects Unrelated P(IBD 0) =1 P(IBD 1) =0 P(IBD 2) =0 Average correlation Additive component = 1× 0 + 0×½ + 0× 1 =0 Dominance component = 1× 0 + 0× 1 =0

Mendel’s Law of Segregation Maternal A 3 A 4 ½ A 1 Paternal A 2 ½ ½ A 1 A 3 ¼ A 1 A 4 ¼ ½ A 2 A 3 ¼ A 2 A 4 ¼
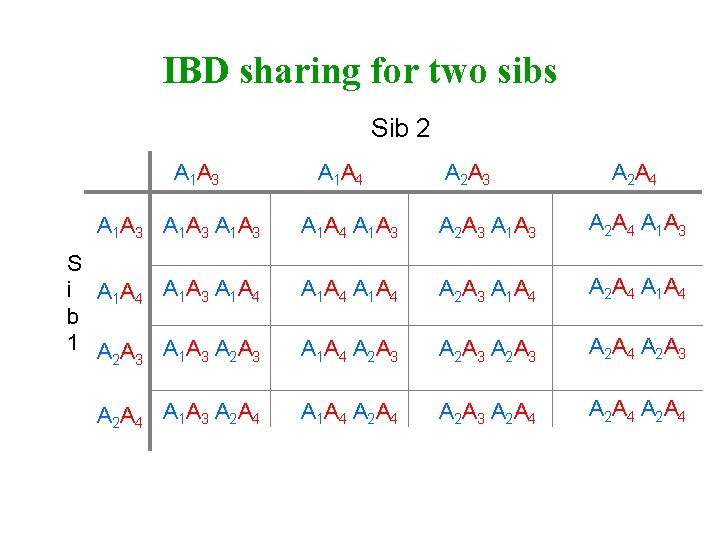
IBD sharing for two sibs Sib 2 A 1 A 3 S i A 1 A 4 A 1 A 3 A 1 A 4 b 1 AA AA AA 1 3 2 3 A 2 A 4 A 1 A 4 A 2 A 3 A 2 A 4 A 1 A 3 A 2 A 3 A 1 A 3 A 2 A 4 A 1 A 3 A 1 A 4 A 2 A 4 A 1 A 4 A 2 A 3 A 2 A 4 A 2 A 3 A 1 A 4 A 2 A 3 A 2 A 4
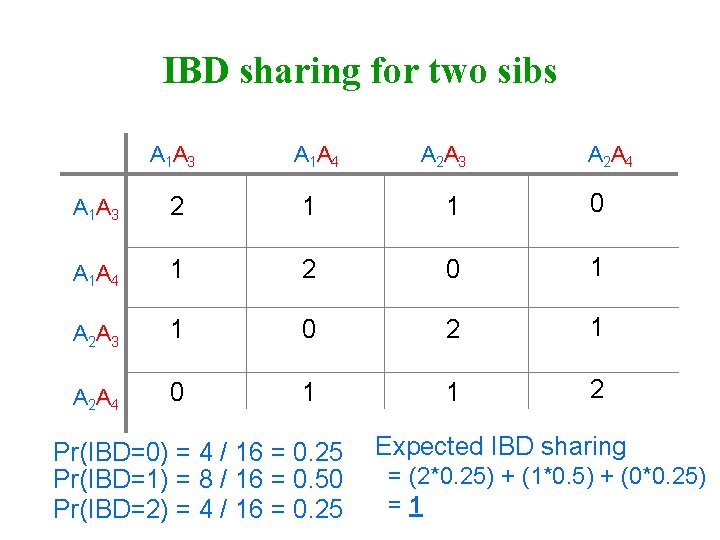
IBD sharing for two sibs A 1 A 3 A 1 A 4 A 2 A 3 A 2 A 4 A 1 A 3 2 1 1 0 A 1 A 4 1 2 0 1 A 2 A 3 1 0 2 1 A 2 A 4 0 1 1 2 Pr(IBD=0) = 4 / 16 = 0. 25 Pr(IBD=1) = 8 / 16 = 0. 50 Pr(IBD=2) = 4 / 16 = 0. 25 Expected IBD sharing = (2*0. 25) + (1*0. 5) + (0*0. 25) =1

Average correlation in QTL effects Sib pairs P(IBD 0) =¼ P(IBD 1) =½ P(IBD 2) =¼ Average correlation Additive component = ¼× 0 + ½×½ + ¼× 1 =½ Dominance component = ¼× 0+ ½× 0 + ¼× 1 =¼

Summary of shared genetic variance MZ = VA + VD P-O = ½ VA U =0 DZ, FS = ¼(VA+VD) + ½(VA/2) + ¼ (0) = ½VA + ¼VD • These single locus results can be summed over all loci to give total genetic variance, heritability.

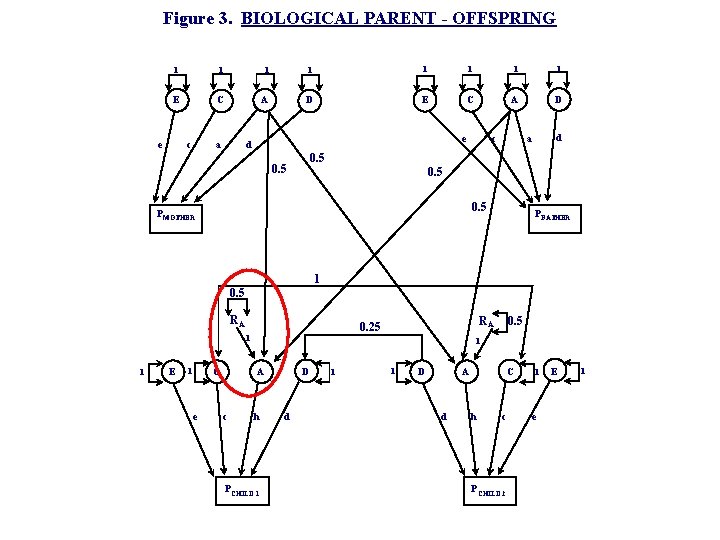
Figure 3. BIOLOGICAL PARENT - OFFSPRING 1 1 1 1 E C A D e a c d a c e d 0. 5 P MOTHER P FATHER 1 0. 5 RA 1 1 E C 1 e 0. 5 1 A c RA 0. 25 h PCHILD 1 D d 1 1 D A d C h c P CHILD 2 1 e E 1

Segregation variance • If both parents transmit 1 allele AO = AP/2 + AM/2 • which means Var(AO) = ¼ Var(AP) + ¼ Var(AM) • But… this violates Var(AO)=Var(AP)=Var(AM) • So this explains the extra path in the P-O model: as Var(A)=1, arbitrarily, then Var(AO) = ¼ Var(AP) + ¼ Var(AM) + ½

Segregation variance • Segregation variance (SV) represents the random variation among the gametes of an individual – i. e. an Aa parent could transmit either A or a AO= AP/2 + AM/2 + SP + SM Var (AO) = Var(AP)/4 + Var(AM)/4 + Var(SP) + Var(SM) • • For homozygous locus, Var(S) = 0 For heterozygous locus, Var(S) = 2/4 Given random mating, average SV = 2 pq 2/4 = VA/4 As VA is fixed to 1 in the path model, SV = ¼ + ¼ = ½
- Slides: 65