Biomedical Informatics The Lexical Grid Project Lex Grid

Biomedical Informatics The Lexical Grid Project: Lex. Grid Christopher G. Chute, MD Dr. PH Professor and Chair, Biomedical Informatics Mayo Clinic College of Medicine Rochester, Minnesota Ontolog Forum 14 December 2006 © 2006 Mayo Clinic College of Medicine

Biomedical Informatics The Lexical Grid Project: Lex. Grid Acknowledgements: Harold Solbrig James Buntrock Thomas Johnson Dan Armbrust © 2006 Mayo Clinic College of Medicine

Biomedical Informatics Outline - Lex. Grid • Overview • Functional Features • Problem Framing • Lex. Grid History • Present Status • Implementations • Future © 2006 Mayo Clinic College of Medicine 3

Biomedical Informatics Overview • The Lex. Grid package represents a comprehensive set of software and services to load, publish, and access vocabulary or ontological resources. • The package is based upon an open standard • HL 7 CTS (CTS II intended as more complete) • Reference implementations as open source © 2006 Mayo Clinic College of Medicine 4

Biomedical Informatics Lex. Grid Interlocking Components • Standards - access methods (programming APIs) and formats need to be published and openly available. • Tools - standards based tools must be readily available. • Content - commonly used vocabularies and © 2006 Mayo Clinic College of Medicine 5

Biomedical Informatics The Lexical Grid • Terminology as a commodity resource • Accessible online • under a common model • through a set of common API's • in web-space on web-time • cross-linked • loosely coupled • published individually, when ready • exportable • locally extendable • globally revised • open source tooling to browse, edit, © 2006 Mayo Clinic College of Medicine 6

Biomedical Informatics Overview purposes • Provides a single information model flexible enough to represent yesterday’s, today’s and tomorrow’s terminological or ontological resources • Allows resources to be published online, cross-linked, and indexed on demand • Provides standardized building blocks and tools that allow applications and users to take advantage of the content where and when it is needed • Provide consistency and standardization required to support large-scale © 2006 Mayo Clinic College of Medicine 7

Biomedical Informatics Lex. Grid Features • Accommodation of multiple vocabulary and ontology distribution formats. • Support of multiple data stores to accommodate federated vocabulary distribution. • Consistent and standardized access across multiple vocabularies. • Rich API for supporting lexical and graph search and traversal. • Fully compatible with HL 7 -CTS implementation. • Support for programmatic access via Java, . NET, and web services. • Open source tooling and code to facilitate adoption and use. © 2006 Mayo Clinic College of Medicine 8

Biomedical Informatics Lex. Grid Users • Vocabulary service providers. Describes organizations currently supporting externalized API-level interfaces to vocabulary content. • Vocabulary integrators. Describes organizations that desire to integrate new vocabulary content or relations to be served locally. • Vocabulary users. Describes persons and organizations desiring common, consistent access to vocabulary content © 2006 Mayo Clinic College of Medicine 9
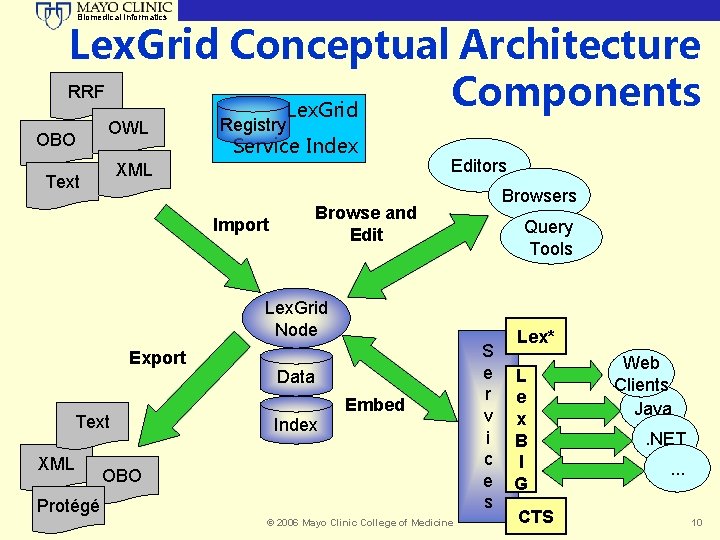
Biomedical Informatics Lex. Grid Conceptual Architecture RRF Components Lex. Grid OBO OWL Registry Service Index XML Text Editors Browse and Edit Import Query Tools Lex. Grid Node Export Text XML Data Index Embed OBO Protégé © 2006 Mayo Clinic College of Medicine S e r v i c e s Lex* L e x B I G CTS Web Clients Java. NET. . . 10

Biomedical Informatics Lex. Grid Node • The logical persistence layer for storing and managing vocabulary content. • The Lex. Grid node utilizes relational database management systems for management of data and indexing functions. • Lex. Grid nodes have been successful installed and tested using My. SQL, Postgres, UDB/DB 2, Oracle, Hypersonic, and LDAP/BDB. © 2006 Mayo Clinic College of Medicine 11

Biomedical Informatics The Import Toolkit(s) • Provides an API and a set of administration tools to load, index, publish, and manage vocabulary content for the vocabulary server. • Standard formats and models that have been developed include: • Rich Release Format (RRF) • Ontology Web Language (OWL) • Lex. Grid XML • Text Delimited • Ontylog XML (Apelon) format • Open Biomedical Ontology (OBO) © 2006 Mayo Clinic College of Medicine 12

Biomedical Informatics The Export Toolkit(s) • Provides an API and set of • administration tools to export content in a standard format from a Lex. Grid node. Standard formats provided for export include: • Lex. Grid XML • OWL © 2006 Mayo Clinic College of Medicine 13

Biomedical Informatics The Lex. Grid Editor • A light weight editor for creating, • • modifying, and changing vocabulary content. The Lex. Grid Editor is an Eclipse Based application that supports multi vocabulary query and browsing, interactive views, and logging and auditing. Recent enhancements have provided extensions to accommodate value set creation and management. © 2006 Mayo Clinic College of Medicine 14

Biomedical Informatics Lex. Grid Principles • Lex. Grid software is based on a model driven architecture. • The Lex. Grid model is maintained in XML Schema format • Represents a core component of design. • The Lex. BIG API • Java-based API to Lex. Grid content is formally modeled • Accommodates registration of additional load, index, and search functions • Provides a conscious separation of service and data classes in order to support deferred query resolution and © 2006 Mayo Clinic College of Medicine 15

Biomedical Informatics Lex. Grid Model • Lexical Semantics • Names • (Textual) Definitions • Comments • Other non-classification property • Context • Languages and dialects • Communities and specialties • Localizations • Logical Semantics • Roles and Relations © 2006 Mayo Clinic College of Medicine 16

Biomedical Informatics Lex. Grid Model • Proposal for standard storage of controlled vocabularies and ontologies • Flexible enough to accurately represent a wide variety of vocabularies and other lexicallybased resources • Defines • How vocabularies should be formatted and represented programmatically • Several different server storage © 2006 Mayo Clinic College of Medicine 17
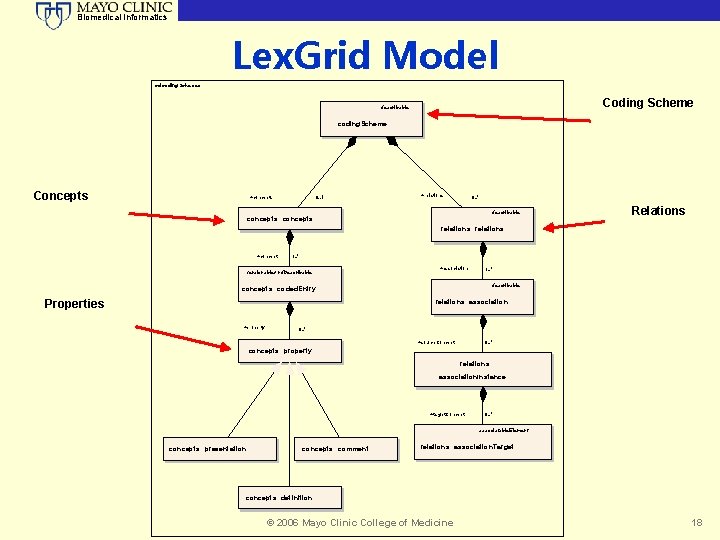
Biomedical Informatics Lex. Grid Model cd coding. Schemes Coding Scheme describable coding. Scheme Concepts +concepts 0. . 1 +relations 0. . * describable concepts: : concepts Relations relations: : relations +concept 1. . * versionable. And. Describable +association describable concepts: : coded. Entry Properties 1. . * relations: : association +property 0. . * +source. Concept 0. . * concepts: : property relations: : association. Instance +target. Concept 0. . * associatable. Element concepts: : presentation concepts: : comment relations: : association. Target concepts: : definition © 2006 Mayo Clinic College of Medicine 18

Biomedical Informatics Model: Code Systems • Each service defined to the Lex. Grid model can encapsulate the definition of one or more vocabularies. • Each vocabulary is modeled as an individual code system, known as a coding. Scheme. • Each scheme tracks information used to uniquely identify the code system, along with relevant metadata. • The collection of all code systems © 2006 Mayo Clinic College of Medicine 19

Biomedical Informatics Model: Concepts • A code system may define zero or more coded concepts, encapsulated within a single container. • A concept represents a coded entity (identified in the model as a coded. Entry) within a particular domain of discourse. • Each concept is unique within the code system that defines it. • Must be qualified by at least one term or designation, represented in the model as a property. • Each property is an attribute, facet, or some other characteristic that may represent or help define the intended meaning of the encapsulating coded. Entry. • A concept may be the source for and/or © 2006 Mayo Clinic College of Medicine 20

Biomedical Informatics Model: Relations • Each code system may define or more containers to encapsulate relationships between concepts. • Each named relationship (e. g. “has. Subtype” or “has. Part”) is represented as an association within the Lex. Grid model. • Each relations container must define or more association. • May also further define the nature of the relationship in terms of transitivity, symmetry, reflexivity, forward and inverse names, etc. • Multiple instances of each association can be defined, each of which provide a directed relationship between one source and one or more target concepts. • Source and target concepts may be contained in the same code system as the association or © 2006 Mayo Clinic College of Medicine 21

Biomedical Informatics Available Representations of the Lex. Grid Model • The master representation of the Lex. Grid model is provided in XML Schema Definition (XSD) format. • Conversions to other formal representations are available, including XML Metadata Interchange (XMI) and Unified Modeling Language (UML). • Implementation or technology-specific renderings of the model also exist. • Relational database schema • (My. SQL, Postgre. SQL, DB 2, Oracle, etc) • Lightweight Directory Access Protocol (LDAP) schema • Programming interfaces generated from the formal representation include Java bean © 2006 Mayo Clinic College of Medicine 22

Biomedical Informatics Disease Understanding Constrained by Knowledge • Carolus Linnaeus Carl von Linné • Genera Morborum (1763) • Underscored Content Difficulty • Pathophysiology vs Manifestation e. g. Rabies as psychiatric disease © 2006 Mayo Clinic College of Medicine 23

Biomedical Informatics The Genomic Era • The genomic transformation of medicine far exceeds the introduction of antibiotics and aseptic surgery • The binding of genomic biology and clinical medicine will accelerate • The implications for shared semantics across the basic science and clinical communities are unprecedented • The implications for Public Health surveillance and inference are © 2006 Mayo Clinic College of Medicine 24

Biomedical Informatics From Practice-based Evidence to Evidence-based Practice Clinical Databases Data Shared Semantics Patient Encounters Registries et al. Inference Ontology Medical Knowledge Vocabularies & Terminologies Decision Expert support Systems Clinical Knowledge Guidelines Managemen © 2006 Mayo Clinic College of Medicine 25

Biomedical Informatics The Historical Center of the Health Data Universe Clinical Data Billable Diagnoses © 2006 Mayo Clinic College of Medicine 26

Biomedical Informatics Copernican Health Data Universe (Niklas Koppernigk) Clinical Data Guidelines Billable Diagnoses Genomic Characteri Scientific Literature Medical Literatu Clinical Data © 2006 Mayo Clinic College of Medicine 27

Biomedical Informatics Continuum from Nomenclature to Classification • Patient Data is Highly Detailed • Modifiers: Anatomy, Stage, Severity, Extent • Qualifiers: Probability, Temporal Status • Aggregate Uses Require Categorization • Granularity of Classifiers • Focused Groups and Strata for CQI/Outcomes • Broad Statistical/Fiscal Groups © 2006 Mayo Clinic College of Medicine 28

Biomedical Informatics Familiar Points Along Continuum Modern Health Vocabularies • Nomenclature – Highly Detailed Descriptions (SNOMED) • Classification – Organized Aggregation of Descriptions into a Rubric (ICDs) • Groupings – High Level Categories of Rubrics (DRGs) Nomenclature Detailed Classification Groups Grouped © 2006 Mayo Clinic College of Medicine 29

Biomedical Informatics Blois, 1988 Medicine and the nature of vertical reasoning • Molecular: receptors, enzymes, vitamins, drugs • Genes, SNPs, gene regulation • Physiologic pathways, regulatory changes • Cellular metabolism, interaction, meiosis, … • Tissue function, integrity • Organ function, pathology © 2006 Mayo Clinic College of Medicine 30
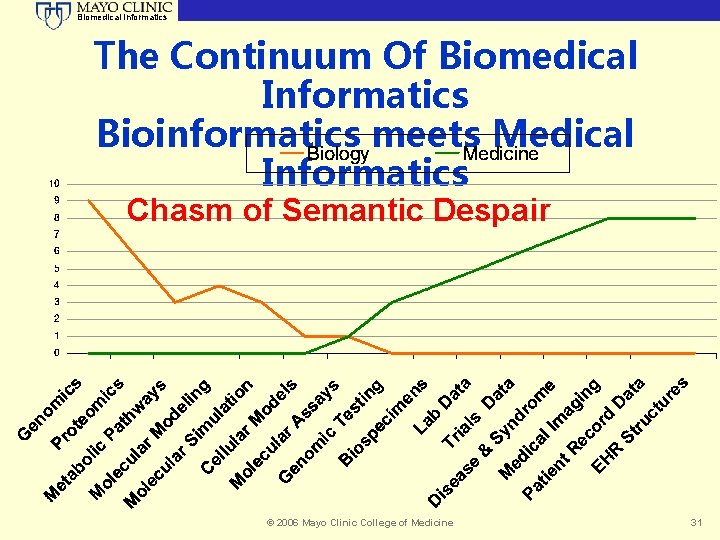
Biomedical Informatics The Continuum Of Biomedical Informatics Bioinformatics meets Medical Informatics Chasm of Semantic Despair © 2006 Mayo Clinic College of Medicine 31

Biomedical Informatics Feudal Cognition Intellectual Semantic Baronies • Genetic variation – Genomics • Haplotypes – Statistical Genomics • Molecular – Metabolomics, Proteomics • Binding – Molecular simulation • Pathways – Physiology and Systems Biology • Symptoms – Consumer Health • Rx and Px – Clinical Medicine © 2006 Mayo Clinic College of Medicine 32
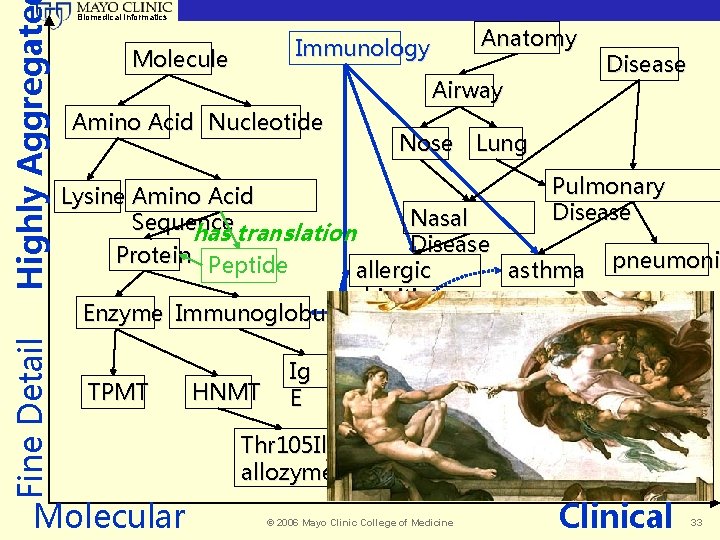
Highly Aggregate Fine Detail Biomedical Informatics Anatomy Immunology Molecule Airway Amino Acid Nucleotide Disease Nose Lung Pulmonary Lysine Amino Acid Disease Nasal Sequence has translation Disease Protein Peptide pneumoni allergic asthma pneumonia rhinitis Enzyme Immunoglobulin TPMT Molecular HNMT Ig E ? Thr 105 Ile allozyme © 2006 Mayo Clinic College of Medicine Clinical 33
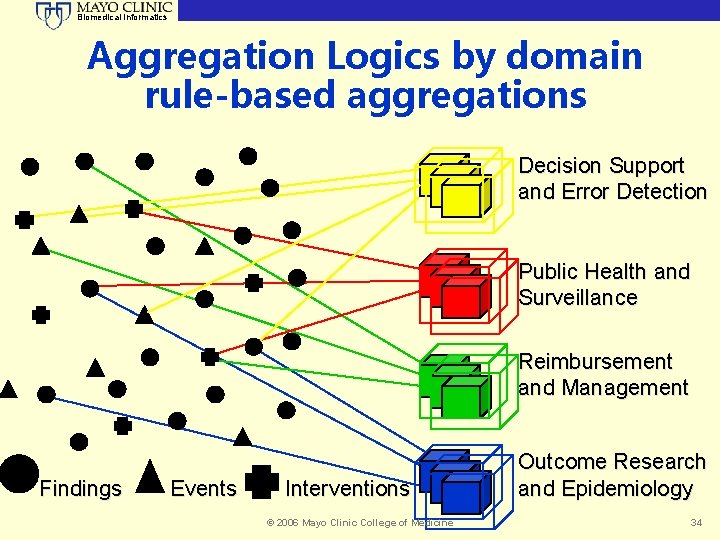
Biomedical Informatics Aggregation Logics by domain rule-based aggregations Decision Support and Error Detection Public Health and Surveillance Reimbursement and Management Findings Events Interventions © 2006 Mayo Clinic College of Medicine Outcome Research and Epidemiology 34
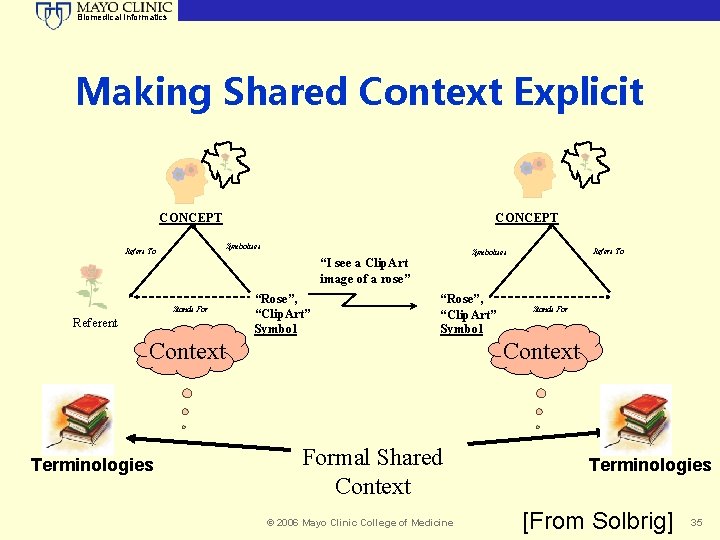
Biomedical Informatics Making Shared Context Explicit CONCEPT Symbolises Refers To Stands For Referent “Rose”, “Clip. Art” Symbol Context Terminologies Refers To Symbolises “I see a Clip. Art image of a rose” Stands For Context Formal Shared Context © 2006 Mayo Clinic College of Medicine Terminologies [From Solbrig] 35

Biomedical Informatics Proliferation of Content “Have it your way” Vocabulary Models • Major ontologies • SNOMED CT; Gene Ontology; LOINC; NDF-RT • UMLS Metathesaurus; NCI Thesaurus • HL 7 RIM and Vocabulary; DICOM Rad. Lex • CDC bioterrorism PHIN standards • ca. BIG DSR / CDEs (Common Data Elements) • All created with differing formats and models © 2006 Mayo Clinic College of Medicine 36

Biomedical Informatics History of Terminology Services in the US • YATN: yet another terminology service 1996 • Mayo, Kaiser, Lexical Technology • Meta. Phrase – Lexical Technology 1998 • LQS: Lexicon Querry Services; 3 M 1998 • Mayo Autocoder: UI to YATN suite 2000 • CTS: Common Terminology Services 2003 © 2006 Mayo Clinic College of Medicine 37

Biomedical Informatics Mayo’s Work with Problem List Interface Design • Premise upon Terminology Server • Meta. Phrase Prototypes on the Network • Iterative Usability Lab Evaluations • Mock-ups in VB, Delphi, Java, … • Evolve Toward Subset of Functional Needs • Problem List Specific • Drive Specification and Operation of T Server © 2006 Mayo Clinic College of Medicine 38
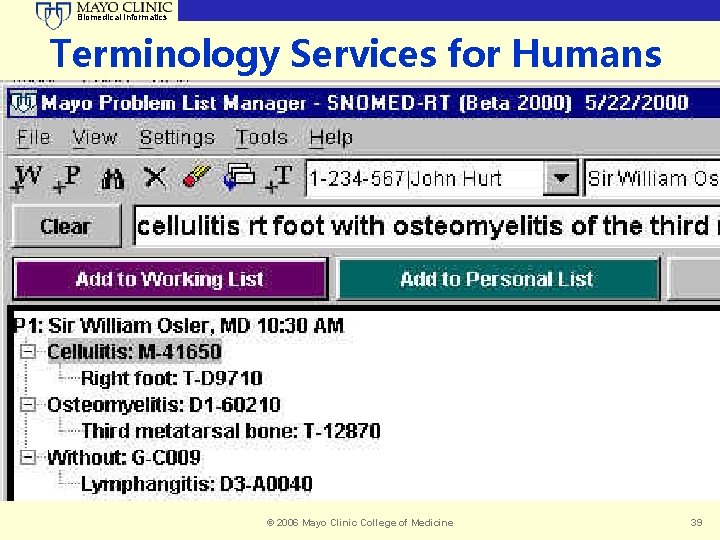
Biomedical Informatics Terminology Services for Humans © 2006 Mayo Clinic College of Medicine 39

Biomedical Informatics Common Terminology Services (CTS) • An HL 7 ANSI standard • Defines the minimum set of requirements for interoperability across disparate healthcare applications • A specification for accessing terminology content • The CTS identifies the minimum set of functional characteristics a terminology resource must possess for use in HL 7. • A functional model • Defining the functional characteristics of vocabulary as a set of Application © 2006 Mayo Clinic College of Medicine 40

Biomedical Informatics CTS APIs • Define the necessary functions for healthcare terminology • Decouples terminology from the terminology service. • Technology independent • Legacy database • Institutional infrastructure • Provide common interface and reference model • I know what you mean by • Code System • Coded Concept © 2006 Mayo Clinic College of Medicine 41

Biomedical Informatics Mayo Lex. Grid Project Services • Ontology HL 7 ANSI Standard • ISO Standard • Open specification • Provide consistency and standardization required to support large-scale vocabulary adoption and use • Common model, tools, formats, and interfaces • Standard terminology model (Excel to OWL) © 2006 Mayo Clinic College of Medicine 42

Biomedical Informatics Examples and Proof of Concept • NIH Road. Map: Nat. Center Biomedical Ontologies • Mayo Lex. Grid project [MLG] • Clinical and basic science (Gene Ontology) communities • NCI ca. BIG – Bioinformatics Grid [MLG] • HHS/ONC NHIN National Health Information Network • IBM Data Coordination project • NLM/HL 7 Coordination project; [MLG] • CDC PHIN Public Health Information © 2006 Mayo Clinic College of Medicine 43

Biomedical Informatics Lex. Grid Applications at Mayo for Semantic Annotation and Integration • Basis for NLP (Natural Language Processing) entity annotation – clinical notes • Harmonize data elements, values sets • Getting the data right • Information retrieval and navigation • Getting the right data • Grounding for data governance • Foundation for semantic © 2006 Mayo Clinic College of Medicine 44

Biomedical Informatics Cancer Biomedical Informatics Grid (ca. BIG) • Coordinated infrastructure for Cancer Research • Clinical Trials, Integrative Cancer Research, Tissue Banking and Pathology Tools • Vocabulary, Common Data Elements, Architecture © 2006 Mayo Clinic College of Medicine 45
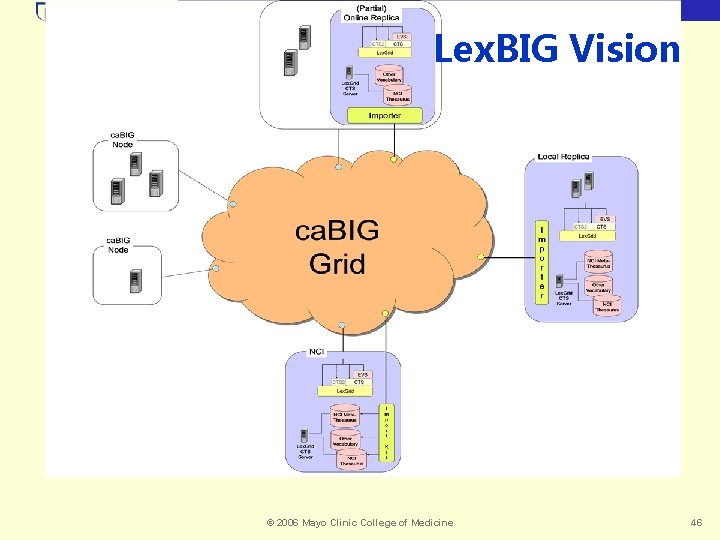
Biomedical Informatics Lex. BIG Vision © 2006 Mayo Clinic College of Medicine 46

Biomedical Informatics © 2006 Mayo Clinic College of Medicine 47

Biomedical Informatics Lex. PHIN CDC Public Health Informatics Network • Adoption of the Lex. Grid Model • Replace PHIN Vocabulary Services (VS) • Addresses genomic characterization of disease • Span semantic chasm with Gene Ontology • Organized Value Sets • Outbreak Management System • Biosurveillance and Biosense © 2006 Mayo Clinic College of Medicine 48
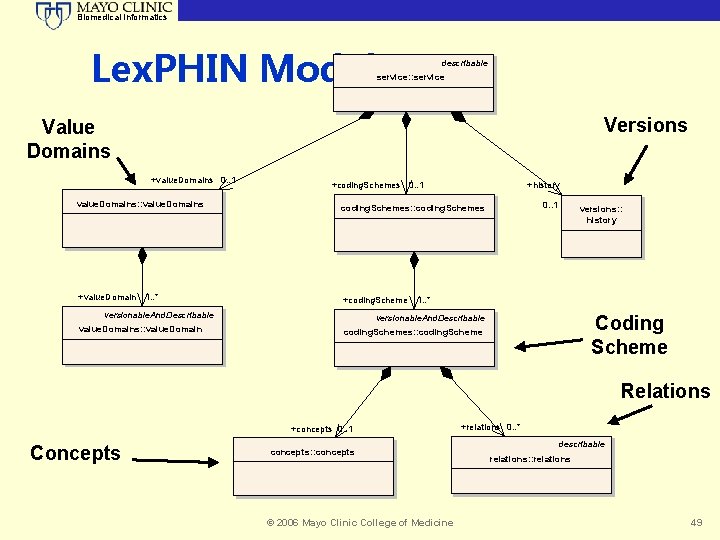
Biomedical Informatics Lex. PHIN Model describable service: : service Versions Value Domains +value. Domains 0. . 1 +coding. Schemes 0. . 1 +history value. Domains: : value. Domains coding. Schemes: : coding. Schemes +value. Domain 1. . * +coding. Scheme 1. . * versionable. And. Describable value. Domains: : value. Domain 0. . 1 versions: : history Coding Scheme versionable. And. Describable coding. Schemes: : coding. Scheme Relations +concepts 0. . 1 Concepts concepts: : concepts © 2006 Mayo Clinic College of Medicine +relations 0. . * describable relations: : relations 49

Biomedical Informatics Health Level Seven (HL 7) • Vocabulary and value domain management • Tooling for vocabulary submissions • Includes change events for HL 7 governance process © 2006 Mayo Clinic College of Medicine 50
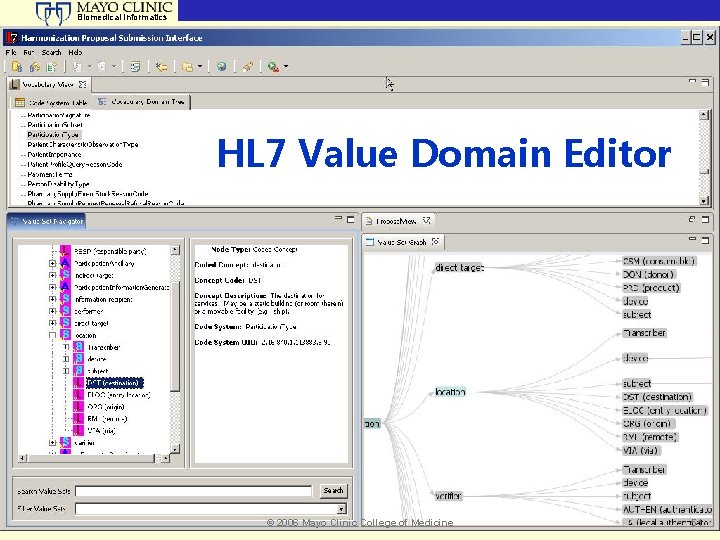
Biomedical Informatics HL 7 Value Domain Editor © 2006 Mayo Clinic College of Medicine 51

Biomedical Informatics NCBO – A Bridge Across the Chasm © 2006 Mayo Clinic College of Medicine 52
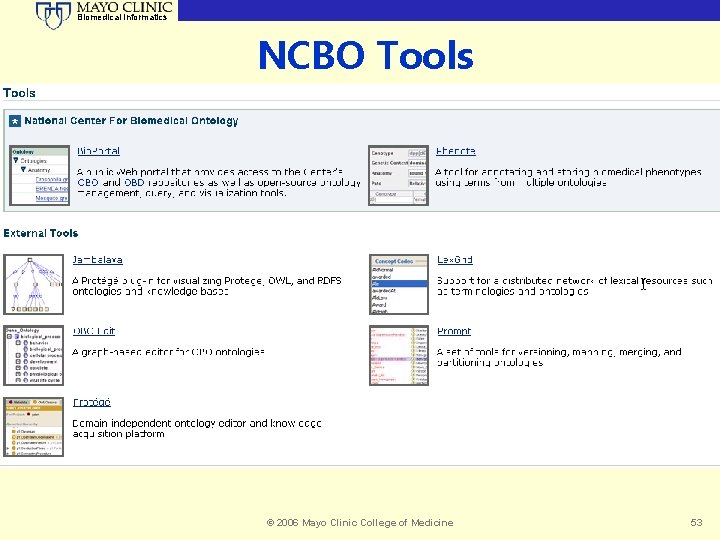
Biomedical Informatics NCBO Tools © 2006 Mayo Clinic College of Medicine 53
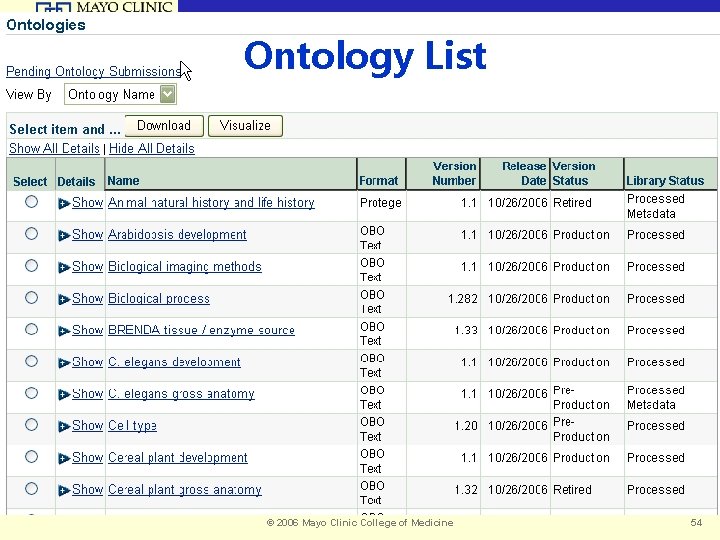
Biomedical Informatics Ontology List © 2006 Mayo Clinic College of Medicine 54

Biomedical Informatics Ontology Counts Total Number of Ontologies 52 NCBO Library 45 Remote 7 Number of Classes 175296* *ontologies which have been parsed and indexed © 2006 Mayo Clinic College of Medicine 55
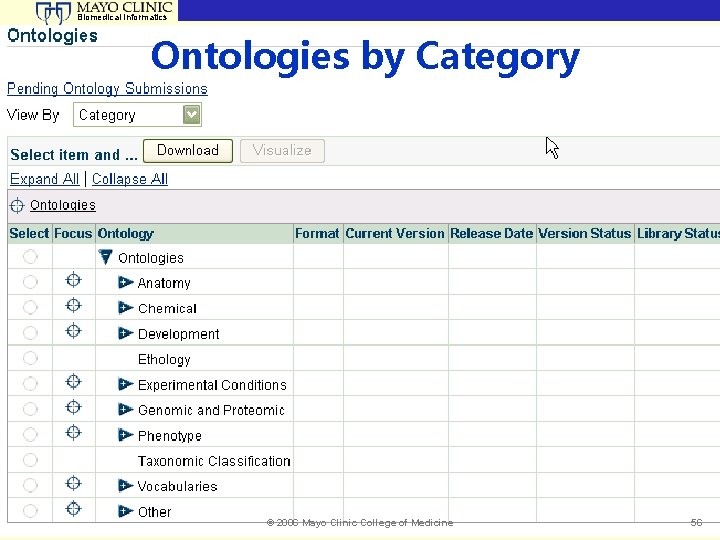
Biomedical Informatics Ontologies by Category © 2006 Mayo Clinic College of Medicine 56
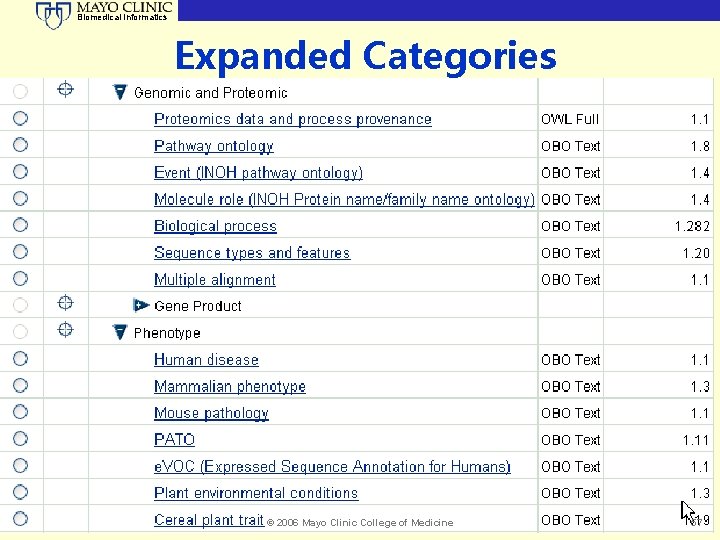
Biomedical Informatics Expanded Categories © 2006 Mayo Clinic College of Medicine 57
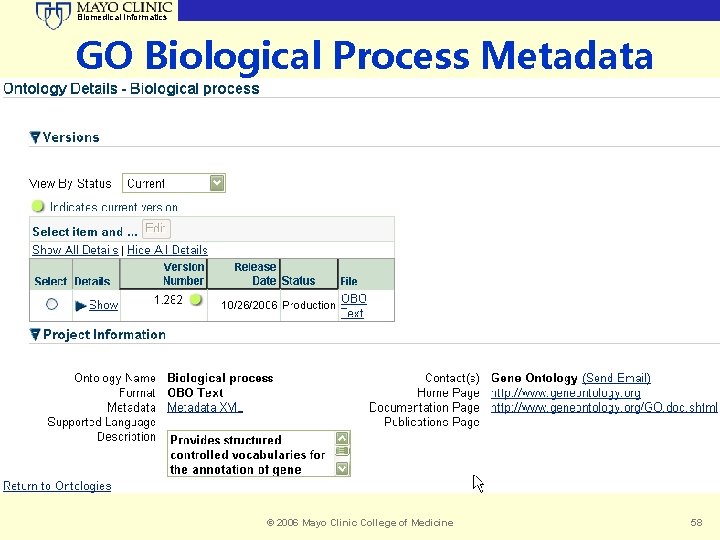
Biomedical Informatics GO Biological Process Metadata © 2006 Mayo Clinic College of Medicine 58
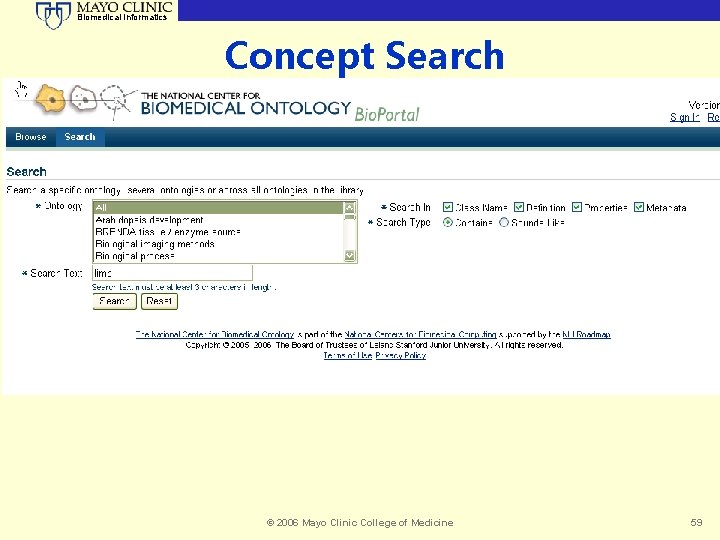
Biomedical Informatics Concept Search © 2006 Mayo Clinic College of Medicine 59
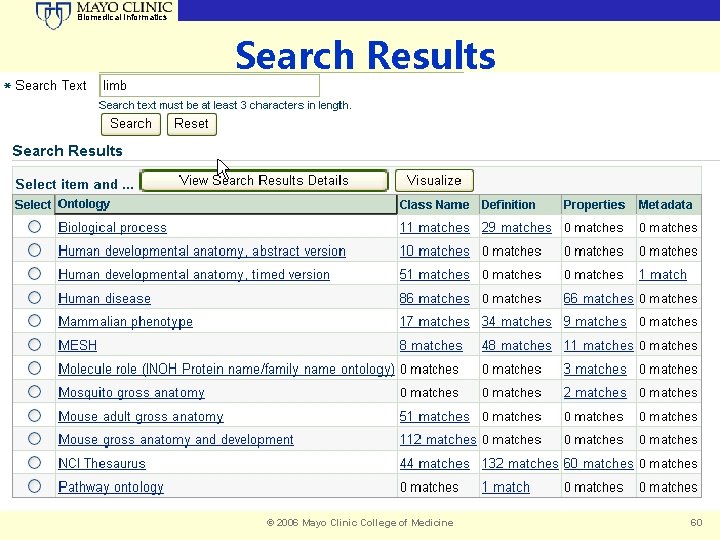
Biomedical Informatics Search Results © 2006 Mayo Clinic College of Medicine 60
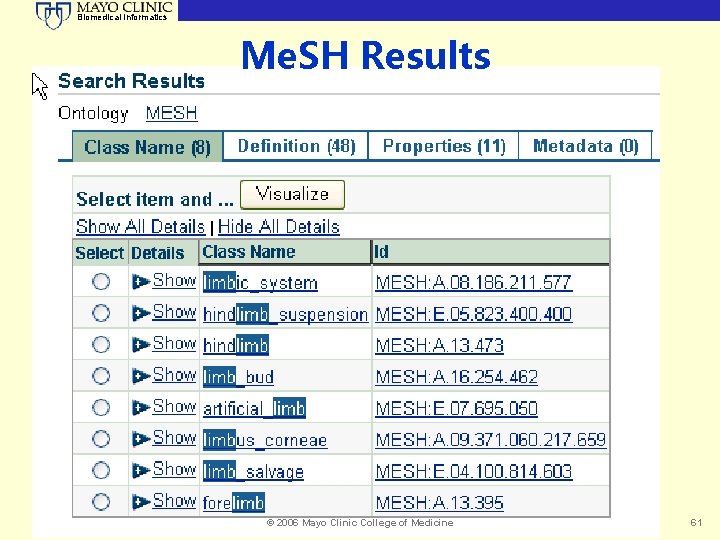
Biomedical Informatics Me. SH Results © 2006 Mayo Clinic College of Medicine 61
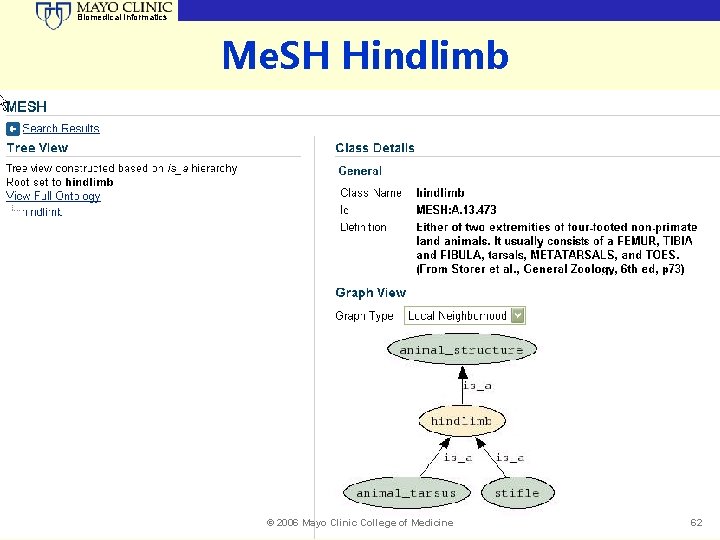
Biomedical Informatics Me. SH Hindlimb © 2006 Mayo Clinic College of Medicine 62
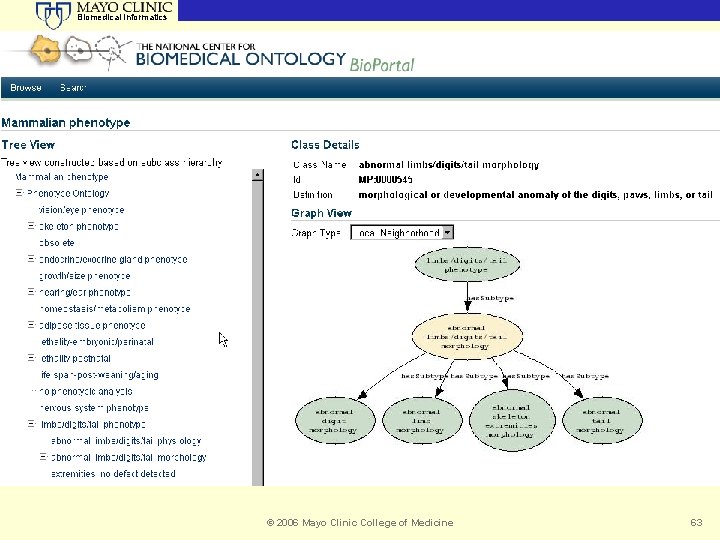
Biomedical Informatics Bio. Portal Stanford University Archana Vembakam and Lynn Murphy © 2006 Mayo Clinic College of Medicine 63

Biomedical Informatics Lex. Grid Future Issues • Federated vocabulary node synchronization and registration/discovery. • API extensions to support local vocabulary extensions and provider suggestions. • API extensions to support HL 7/CTSII API (currently being defined). • API extensions to support submission of vocabulary change requests. • API extensions to load and map between additional vocabulary formats. • ISO 11179 and Lex. Grid integration • Provide additional index services • Synonymy and normalized search • Reasoner or classifier adaptation • Automated coding of medical records • Provide a light-weight Representational State Transfer (REST) service implementation. © 2006 Mayo Clinic College of Medicine 64

Biomedical Informatics Conclusion • Biomedicine concepts have become complex and intertwined • Big science model of future research • 21 st Century Medicine will require comparable and consistent data (Clinical and Genomic) • Ontologies as formal models of concepts provide great opportunity • Tools, content, and resources are becoming increasingly available • Lex. Grid is emerging as an © 2006 Mayo Clinic College of Medicine 65

Biomedical Informatics Resources Lex. Grid Project http: //informatics. mayo. edu/Lex. Gri d Lex. BIG Forge Site http: //gforge. nci. nih. gov/projects/l exbig ca. BIG Lex. Grid CVS http: //cabigcvs. nci. nih. gov/viewcvs. cgi/lexgrid NCBO Project © 2006 Mayo Clinic College of Medicine 66
- Slides: 66