BIOMAT 2017 17 th International Symposium Mathematical BiologyBiological

BIOMAT 2017 – 17 th International Symposium Mathematical Biology/Biological Physics, Moscow, Russia Federation October 30 – November 3, 2017 MATHEMATICAL MODELING OF THE NEURAL ACTIVITY Galina V. Muratova, Vadim V. Bavin Southern Federal University Rostov-on-Don, Russia BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 1

Outline 1. 2. 3. 4. Computational neuroscience Neuron Models Multi-segment model. Сable equation Phenomenological models of neural networks. Izhikevich Model 5. Conclusions BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 2

Computational neuroscience • Bert Sakmann: «I know all the details: cell types, the properties of their activity, connectivity cell excitability of dendrites, synaptic dynamics. . . but I can not understand it (as a whole), I have to model this» . • Bert Sakmann, (born June 12, 1942, Stuttgart, Ger. ), German medical doctor and research scientist who in 1991, together with German physicist Erwin Neher, won the Nobel Prize for Physiology or Medicine for research into basic cell function and for their development of the patch-clamp technique—a laboratory method widely used in cell biology and neuroscience to detect electrical currents as small as a trillionth of an ampere through cell membranes. BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 3

Computational neuroscience Problems that we should be able to solve in the next few decades: - What is consciousness? - How do circuits of neurons compute? - What causes psychiatric and neurological illness? - How do learning and memory work? - Why do we sleep and dream? -How is time represented in the brain? - How do we make decisions? BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 4

Neuron models • Point neuron model Model Hodgkin- Huxley (electrogenesis study on the drug giant axon of the squid) • Multi-segment model, modeling neuron with processes. Сable equation • Phenomenological models of neurons. Izhikevich model BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 5
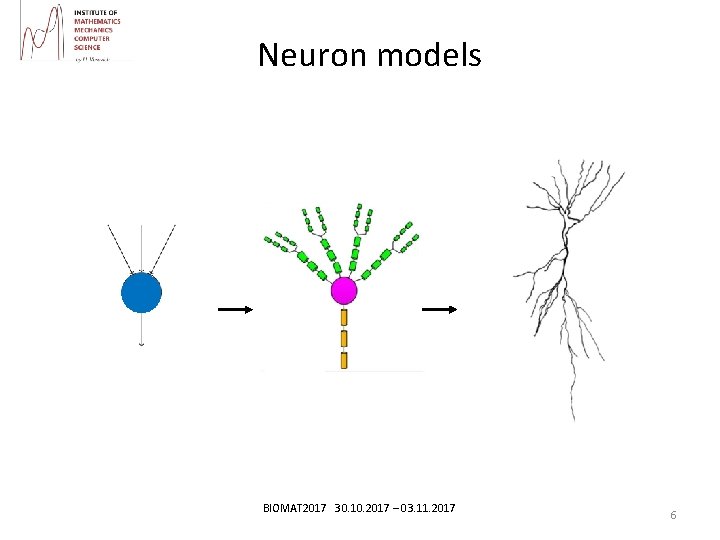
Neuron models BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 6
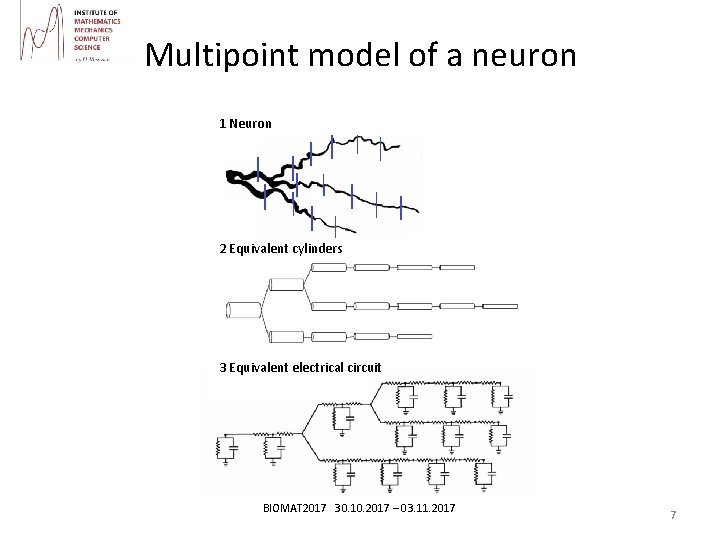
Multipoint model of a neuron 1 Neuron 2 Equivalent cylinders 3 Equivalent electrical circuit BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 7

Cable equation for node BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 8
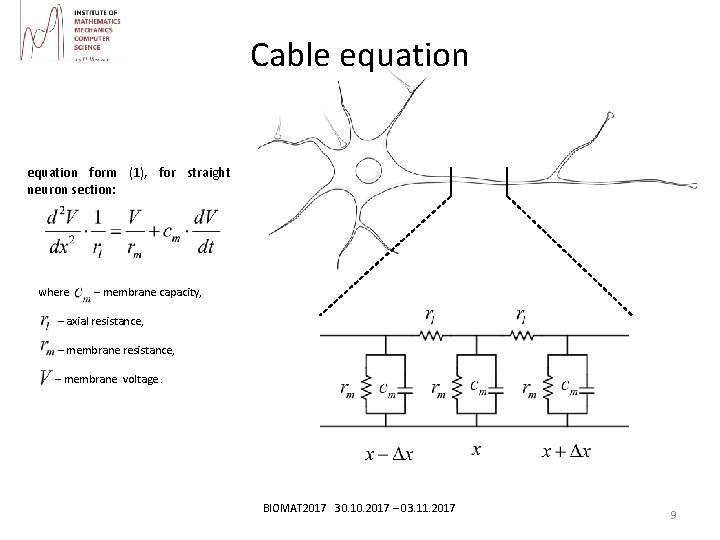
Cable equation form (1), for straight neuron section: where – membrane capacity, – axial resistance, – membrane voltage. BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 9

Cable equation the form of equation (1), for a branched part of the cable, includes an additional axial current between segments [ j, 0] и [ j+1, 1]: the form of equation (1), for a section of a cable with a synapse, includes a synaptic current: where – connection conductivity, – connection recovery potential. BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 10

Cable equation the boundary condition in case of termination of nerve fibers, axial current from the end is missing : the boundary condition in the case of connection segment with soma of neuron, the potential value of segment is fixed - the value of the soma potential: BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 11

Multigrid - history • The general idea of MGM was suggested by R. P. Fedorenko R. P. , A relaxation method for solving elliptic difference equations, Russian J. Comput. Math. Phys. 1 (1961) p. p. 1092 - 1096. • Prof. Fedorenko is an alumnus of Rostov State University 11. 03. 1930 -13. 09. 2009 BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 12

Multigrid method linear system: pre smoothing: after smoothing: correcting : residual : interpolation : coarsening : solving : BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 13

Algebraic Multigrid. Algebraic smooth Error where 1. error equation : 2. smooth error equation: : 3. if is the combination of eigenvectors then the eigenvalue quadratic form for error : if is big then the and can not differ greatly. coarsening in the direction of strong negative coefficient of A. BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 14
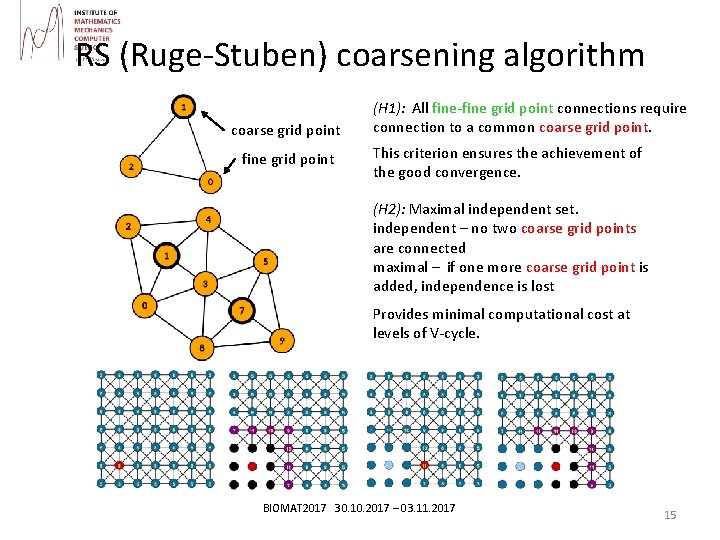
RS (Ruge-Stuben) coarsening algorithm coarse grid point fine grid point (H 1): All fine-fine grid point connections require connection to a common coarse grid point. This criterion ensures the achievement of the good convergence. (H 2): Maximal independent set. independent – no two coarse grid points are connected maximal – if one more coarse grid point is added, independence is lost Provides minimal computational cost at levels of V-cycle. BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 15
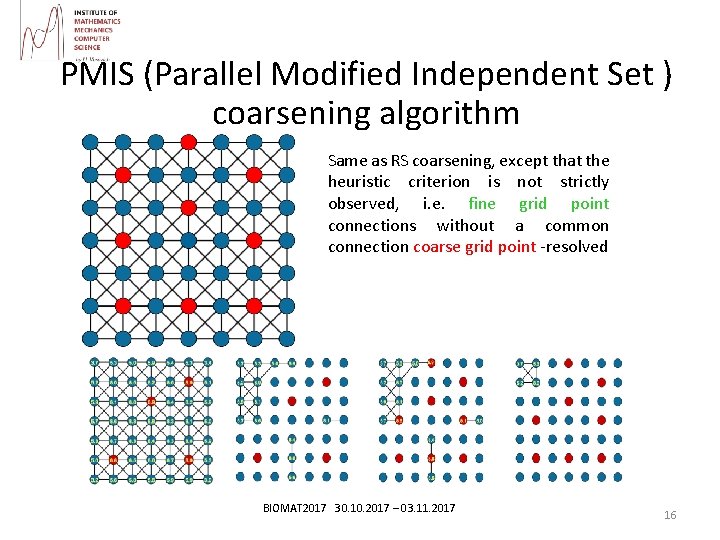
PMIS (Parallel Modified Independent Set ) coarsening algorithm Same as RS coarsening, except that the heuristic criterion is not strictly observed, i. e. fine grid point connections without a common connection coarse grid point -resolved BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 16
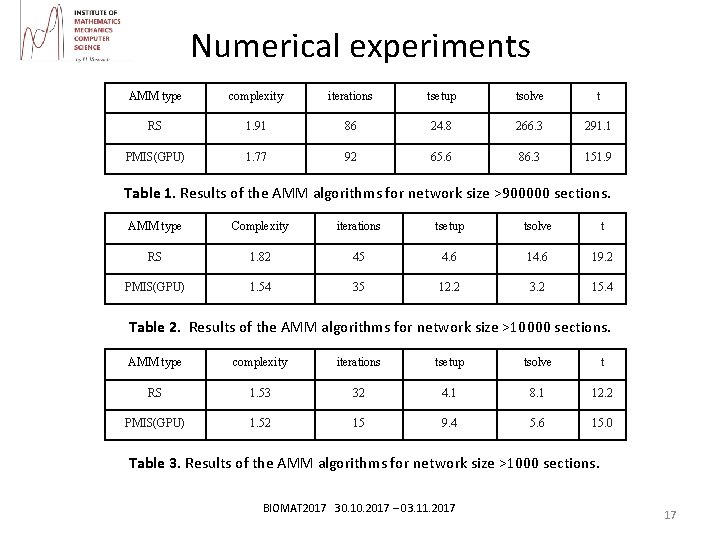
Numerical experiments AMM type complexity iterations tsetup tsolve t RS 1. 91 86 24. 8 266. 3 291. 1 PMIS(GPU) 1. 77 92 65. 6 86. 3 151. 9 Table 1. Results of the AMM algorithms for network size >900000 sections. AMM type Complexity iterations tsetup tsolve t RS 1. 82 45 4. 6 19. 2 PMIS(GPU) 1. 54 35 12. 2 3. 2 15. 4 Table 2. Results of the AMM algorithms for network size >10000 sections. AMM type complexity iterations tsetup tsolve t RS 1. 53 32 4. 1 8. 1 12. 2 PMIS(GPU) 1. 52 15 9. 4 5. 6 15. 0 Table 3. Results of the AMM algorithms for network size >1000 sections. BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 17
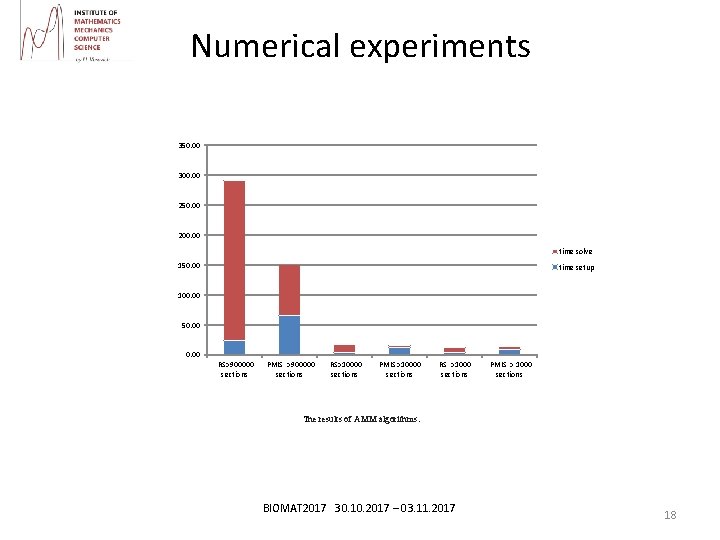
Numerical experiments 350. 00 300. 00 250. 00 200. 00 time solve 150. 00 time setup 100. 00 50. 00 RS>900000 sections PMIS >900000 sections RS>10000 sections PMIS >10000 sections RS >1000 sections PMIS > 1000 sections The results of AMM algorithms. BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 18
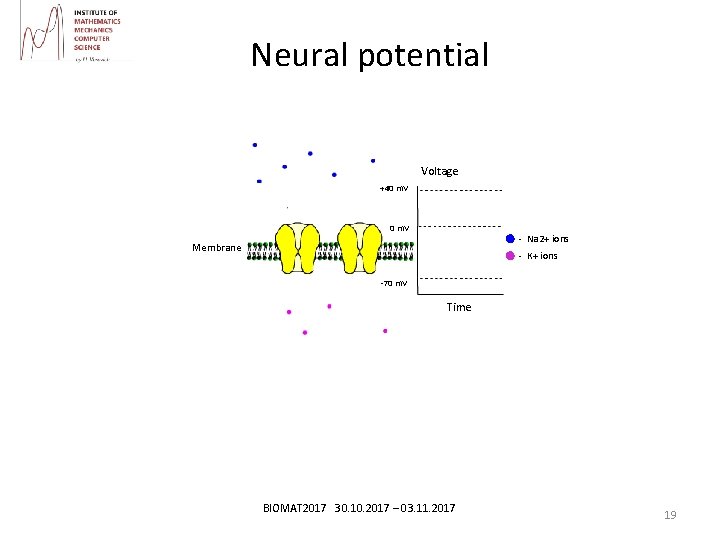
Neural potential Brain model Voltage +40 m. V - Na 2+ ions Membrane - K+ ions -70 m. V Time BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 19

Phenomenological models of neurons The problem in the modeling of neural networks: choice of an optimum approach, the scope for modeling The model reproduces the dynamics of the membrane potential as a phenomenon BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 20

Phenomenological models of neurons • Model integrating neuron (IN) (integrate -and -fire) • Model Fitz Hugh Nagumo (two-segment model, the basic element - differential equation with the polynomial of the 3 rd degree in the right-hand side) • Izhikevicha Model (modified FHN model, square polynomial on the right side of the membrane potential) BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 21

Izhikevich neuron model where – membrane capacity, – resting potential, – minimal potential for generation of action potential, – the peak potential of the spike, phase plane membrane voltage action potential – the sensitivity of U to the subthreshold fluctuations, – the value of the potential after spike, – the magnitude of increase of U after spike, – inverse of membrane resistance coefficient, – incoming synaptic current time BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 22
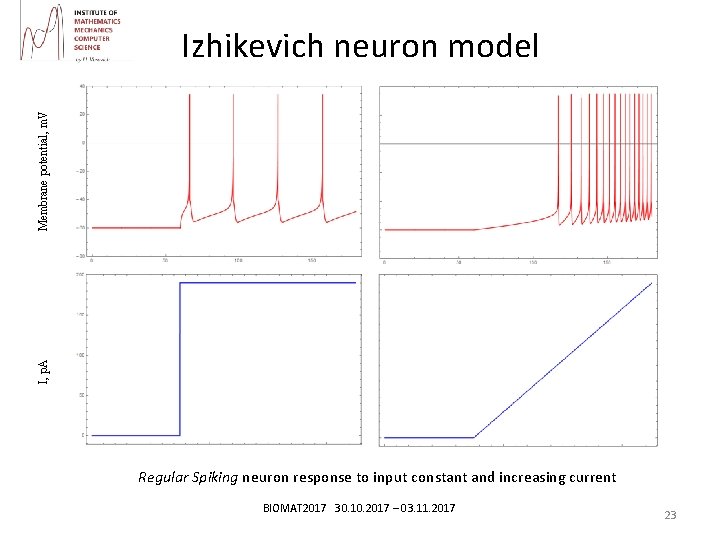
I, p. A Membrane potential, m. V Izhikevich neuron model Regular Spiking neuron response to input constant and increasing current BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 23
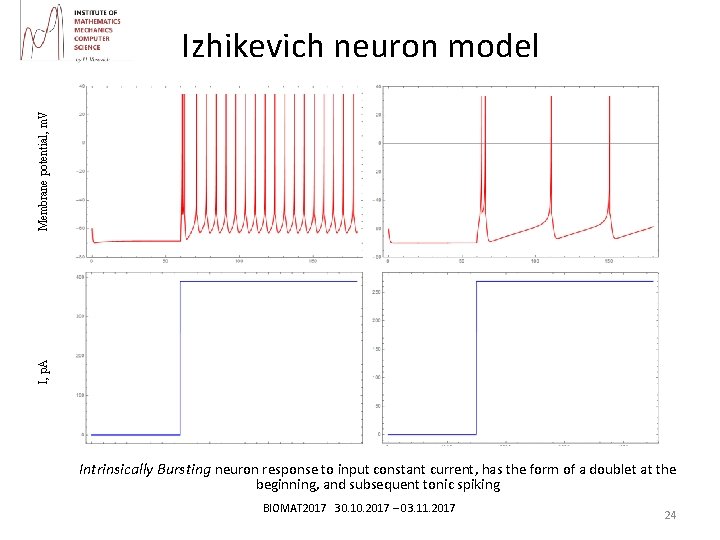
I, p. A Membrane potential, m. V Izhikevich neuron model Intrinsically Bursting neuron response to input constant current, has the form of a doublet at the beginning, and subsequent tonic spiking BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 24
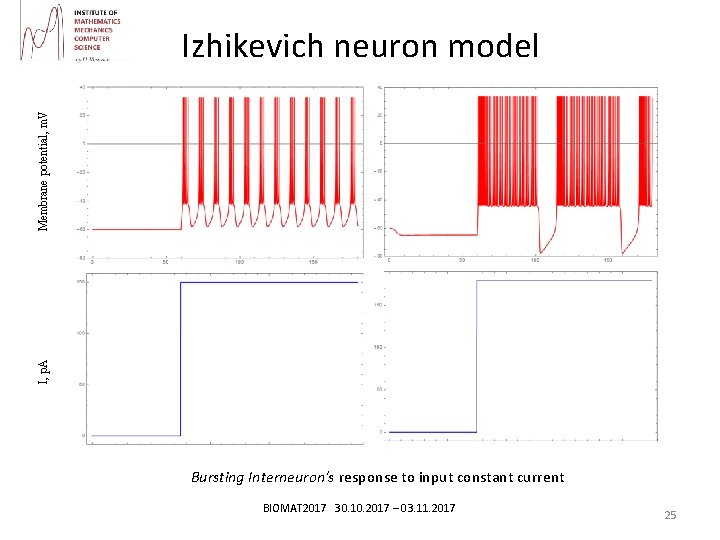
I, p. A Membrane potential, m. V Izhikevich neuron model Bursting Interneuron's response to input constant current BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 25

Conclusion • For cabel equation: – Efficiency of the algebraic multigrid method for solving the cable equation on non-uniform grids is demonstrated – RS, PMIS algorithms are presented • The model of the neural electrical activity based on the Izhikevich model is constructed BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 26

Thank you for your attention! Best wishes from Rostov on Don! muratova@sfedu. ru BIOMAT 2017 30. 10. 2017 – 03. 11. 2017 27

1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. References Fedorenko R. P. , A relaxation method for solving elliptic difference equations, Russian J. Comput. Math. Phys. 1 (1961) p. p. 1092 - 1096. Hodgkin A. L. , Huxley A. F. A quantitative description of ion currents and its applications to conduction and excitation in nerve membranes. Journal of Neurophysiology. 1952. V. 117. № 4. P. 500− 544. Izhikevich E. M. Dynamical systems in neuroscience: the geometry of excitability and bursting. Cambridge: MIT Press, 2007. 464 p. Hale J. , Koçak H. Dynamics and Bifurcations. New York: Springer, 1991. 567 p. Koch C. , Segev I. Methods in neuronal modeling: from ions to networks. Cambridge: MIT Press, 1999. 687 p. Nernst Equation. Calculating the cell potential under non-standard condition. URL: http: //www. roanestate. edu/faculty/chemistryslideshows/Nernst. pdf. (дата обращения: 05. 2015). Abbott L. F. In: Lapicque’s introduction of the integrate-and-fire model neuron. Amsterdam: Elsevier Science, 1999. P. 303− 304. Мareev V. V. , Stankova E. N. Multigrid methods. An introduction to the standard methods. Publishing house S. Peterb. unv-ty, 2012. P. 6 – 7. ( In Russian). Volkov K. N. , Deryugin YU. N. , Emelyanov V. N. Methods of acceleration hydrodynamic computations on unstructured grids. ISBN: 978 -5 -9221 -1542 -1 Fizmatlit. 2013. P. 344– 355. ( In Russian). R. D. Falgout. An Introduction to Algebraic Multigrid. Computing in Science and Engineering, 2006, P. 3 − 5. U. M. Yang. Parallel Algebraic Multigrid Methods - High Performance Preconditioners. Center for Applied Scientific Computing, Lawrence Livermore National Laboratory, 2006. P. 6 − 12. Koch C. , Segev I. Methods in Neuronal Modeling: From Ions to Networks. Cambridge: MIT Press, 1998. P. 28 − 35. L. N. Olson. Algebraic Multigrid Methods. Department of Computer Science University of Illinois at Urbana. Champaign Urbana. 2014. P. 4 − 5. W. Hackbusch Multi-Grid Method and Application Springer-Verlag, Berlin, 185. P. 18 − 20. Gee MW, Siefert ML 5. 0 Smoothed Aggregation Users Guide. 2007. P. 7 − 8. G. Karypis and V. Kumar, METIS: Unstructured graph partitining and sparse matrix ordering system, tech. rep. , 28 University of Minnesota, Department of Computer Science, 1998. P. 1 − 2.
- Slides: 28