BIOLOGY A Global Approach TENTH EDITION Global Edition

BIOLOGY A Global Approach TENTH EDITION Global Edition Campbell • Reece • Urry • Cain • Wasserman • Minorsky • Jackson 22 Phylogenetic Reconstruction Lecturer Dorothee Huchon Steinhardt Museum 527 036409817 Lecture Presentation by Nicole Tunbridge and huchond@tauex. tau. ac. il Kathleen Fitzpatrick © 2015 Pearson Education Ltd

Today’s subjects • 1 - Introduction – History of science (Linnaeus, Darwin) • 2 - Phylogenetic trees • 3 - How to reconstruct phylogenetic trees with morphological characters – The data matrix – The cladistics method – Molecular characters 2
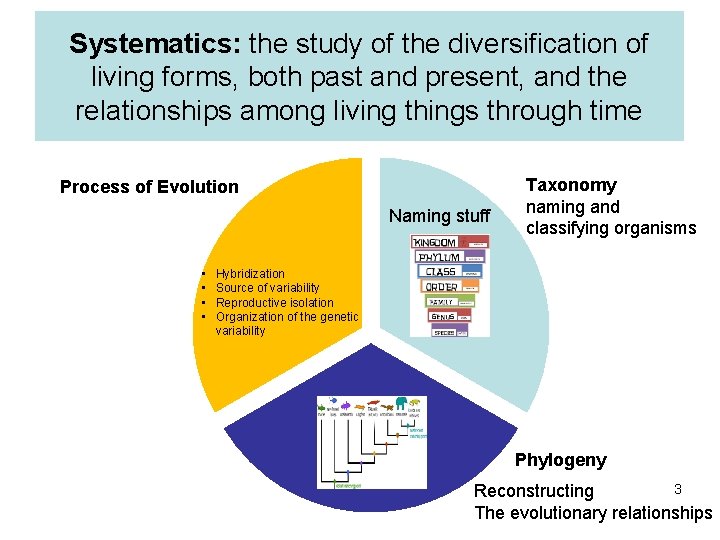
Systematics: the study of the diversification of living forms, both past and present, and the relationships among living things through time Process of Evolution Naming stuff • • Taxonomy naming and classifying organisms Hybridization Source of variability Reproductive isolation Organization of the genetic variability Phylogeny 3 Reconstructing The evolutionary relationships

Introduction • Phylogeny: – Phylum: a taxonomic division of animals below Kingdom and above Class; equivalent to “Division" in Botany ; represents a basic body plan. – Genesis: origin. • Phylogenetics: discipline that studies the evolutionary relationships among organisms or genes. 4

• Taxonomy is the science of naming and classifying organisms in regard to their natural relationships. • It is one of the oldest professions! Macrolepiota procera : tasty Lepiota cristata : highly poisonous 5
There are many classifications 6

There are many classifications 7

• Taxonomy is the science of naming and classifying organisms in regard to their natural relationships. 8

Carl Linneaus (1707 -1778). 4, 400 species of animals and 7, 700 species of plants - The first edition published in 1735 contained 11 very large pages - The last (12 th) edition published in 1766– 1768, contained 2, 400 pages 9

Before Linneaus After Linneaus binomial • Apis pubescens, thorace subgriseo, abdomine fusco, pedibus posticus glabis, untrinque margine ciliatus • Apis mellifera genus Specific epithet 10
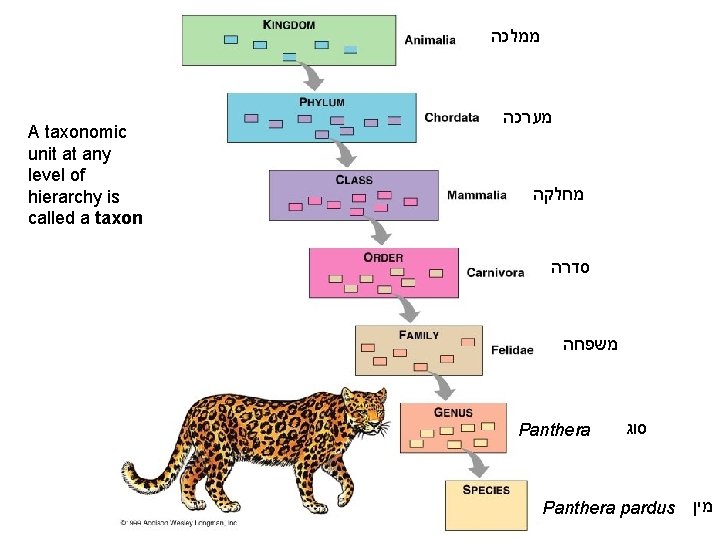
ממלכה A taxonomic unit at any level of hierarchy is called a taxon מערכה מחלקה סדרה משפחה Panthera סוג Panthera pardus מין

• Domain (or Superkingdom) – Kingdom • Subkingdom – Branch – Infrakingdom • Superphylum – Phylum (or Division) • Subphylum – Infraphylum » Microphylum • Superclass – Class • Subclass – Infraclass » Parvclass … 12
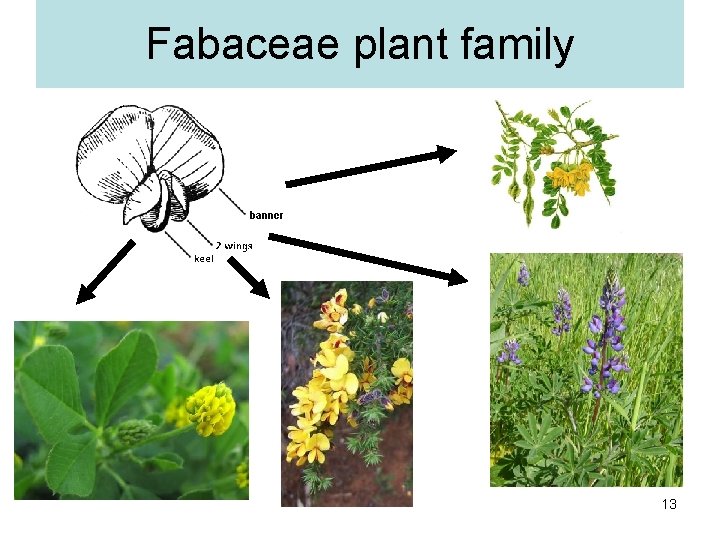
Fabaceae plant family 13

Darwinian revolution (Origin of species 1859) – If organisms have similar organs it is because they have a common ancestor. – Finding similar organs is finding evolutionary relationships. – Classification became evolutionary classification. 14

Today’s subjects • 1 - Introduction – History of science • 2 - Phylogenetic trees • 3 - How to reconstruct phylogenetic trees with morphological characters – The data matrix – The cladistics method – Molecular characters 15

Phylogenetic trees • The evolutionary history of a group of organisms can be represented in a branching phylogenetic tree. 16
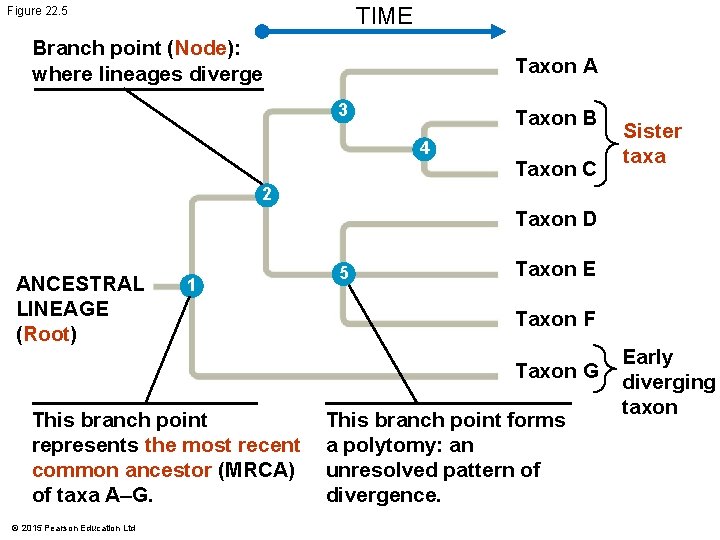
TIME Figure 22. 5 Branch point (Node): where lineages diverge Taxon A 3 Taxon B 4 Taxon C Sister taxa 2 Taxon D ANCESTRAL LINEAGE (Root) 1 5 Taxon E Taxon F Taxon G This branch point represents the most recent common ancestor (MRCA) of taxa A–G. © 2015 Pearson Education Ltd This branch point forms a polytomy: an unresolved pattern of divergence. Early diverging taxon
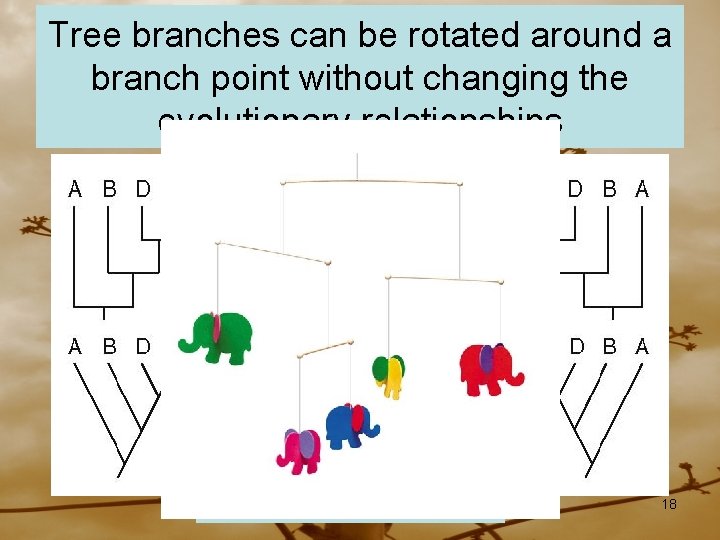
Tree branches can be rotated around a branch point without changing the evolutionary relationships All the same tree 18
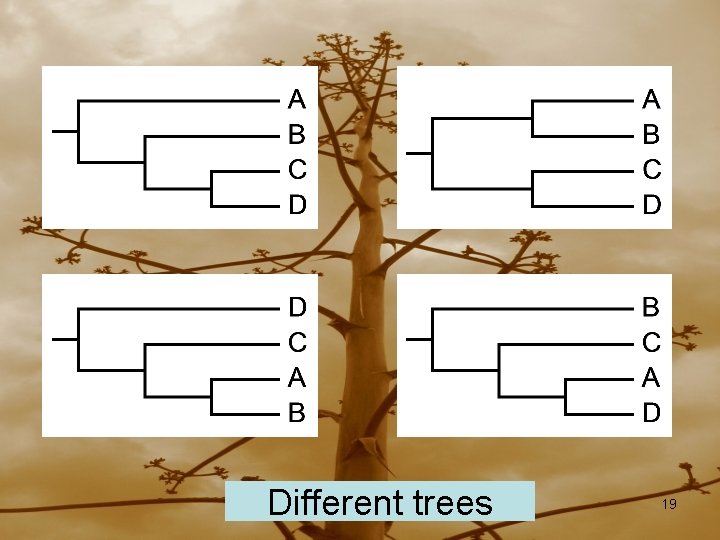
Different trees 19
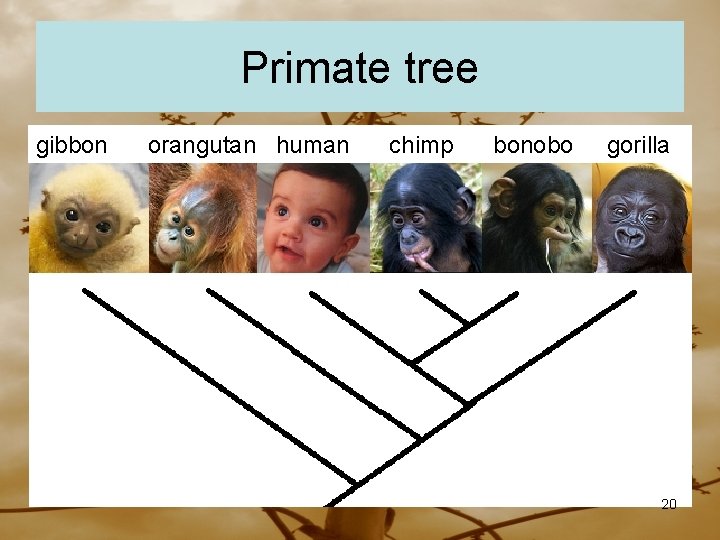
Primate tree gibbon orangutan human chimp bonobo gorilla 20

What We Can and Cannot Learn from Phylogenetic Trees • Phylogenetic trees show patterns of descent, not phenotypic similarity • Phylogenetic trees do not indicate when species evolved or how much change occurred in a lineage • It should not be assumed that a taxon evolved from the taxon next to it 21

What We Can and Cannot Learn from Phylogenetic Trees • A phylogenetic tree represents a hypothesis about evolutionary relationships. • Trees can be compared and alternative hypotheses tested. 22
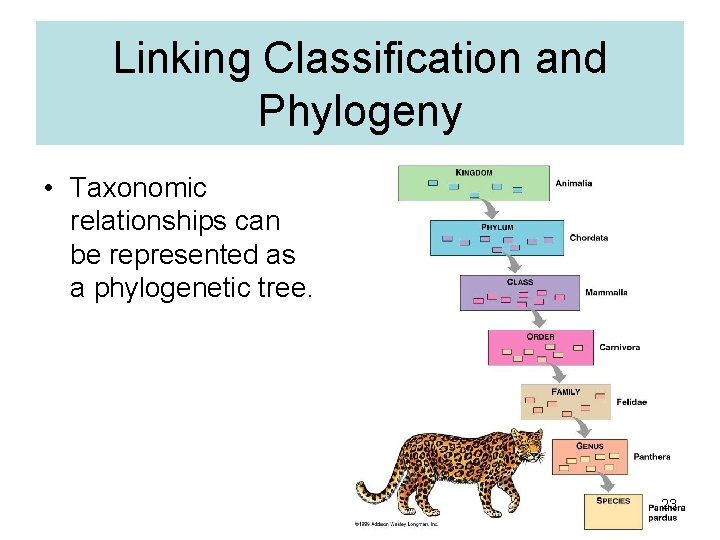
Linking Classification and Phylogeny • Taxonomic relationships can be represented as a phylogenetic tree. 23
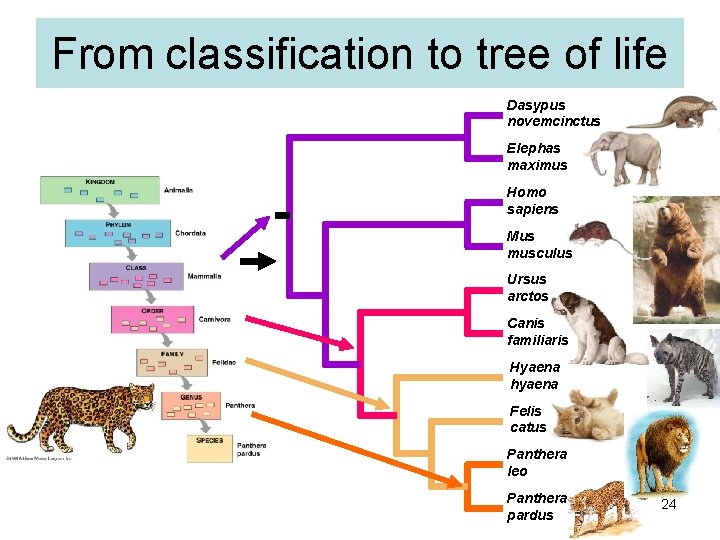
From classification to tree of life Dasypus novemcinctus Elephas maximus Homo sapiens Mus musculus Ursus arctos Canis familiaris Hyaena hyaena Felis catus Panthera leo Panthera pardus 24
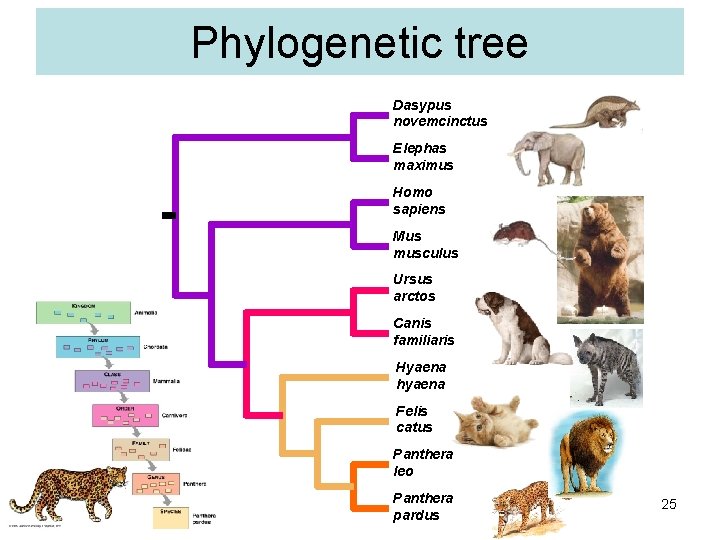
Phylogenetic tree Dasypus novemcinctus Elephas maximus Homo sapiens Mus musculus Ursus arctos Canis familiaris Hyaena hyaena Felis catus Panthera leo Panthera pardus 25
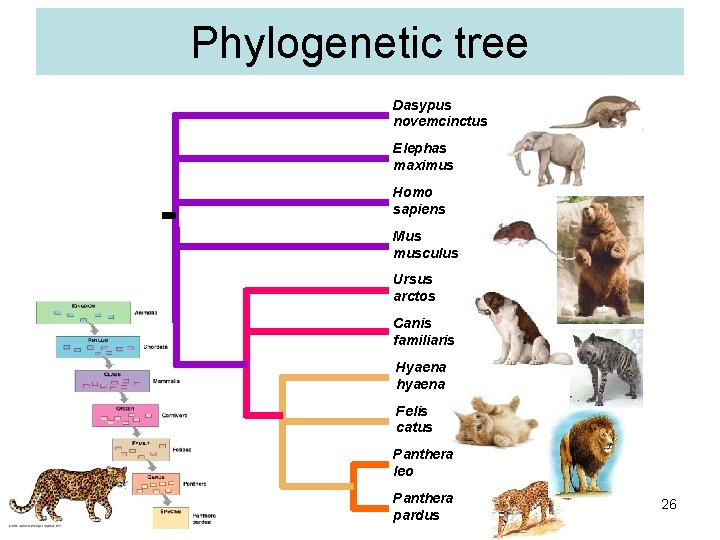
Phylogenetic tree Dasypus novemcinctus Elephas maximus Homo sapiens Mus musculus Ursus arctos Canis familiaris Hyaena hyaena Felis catus Panthera leo Panthera pardus 26

Today’s subjects • 1 - Introduction – History of science • 2 - Phylogenetic trees • 3 - How to reconstruct phylogenetic trees with morphological characters – The data matrix – The cladistics method – Molecular characters 27

1. Data 28

Five animals Cow Man Fish Frog Coral 29

Characters • Taxonomic characters: attributes of a taxon that allow its differentiation or potential differentiation from others. • Characters can be morphological, physiological, molecular, ecological, reproductive or behavioral. • Characters must be homologous. 30
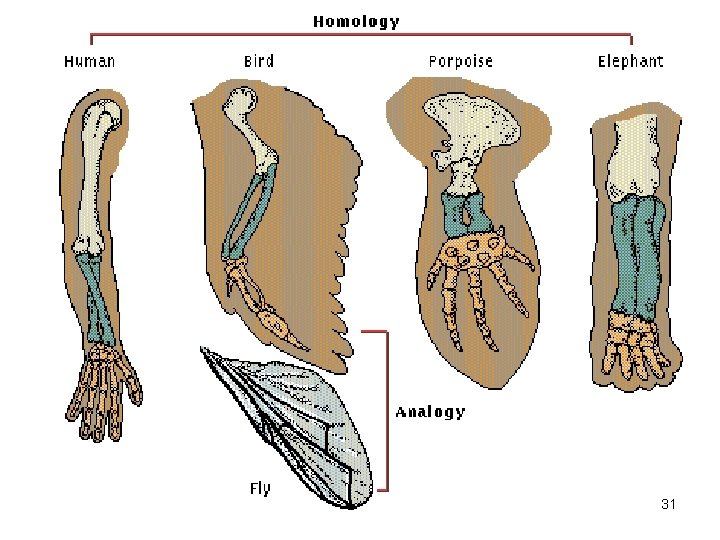
31

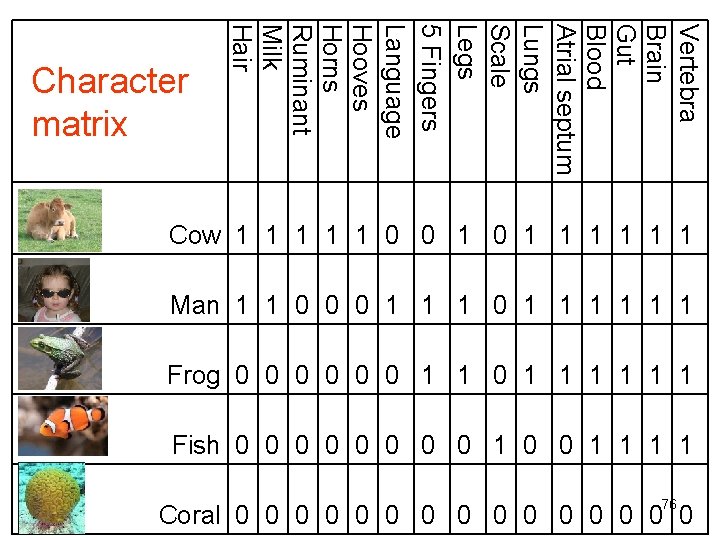
Character table • Characters are coded in a character table (or matrix). • A 0 usually indicates that a character is absent. • A 1 indicates that a character is present. 32
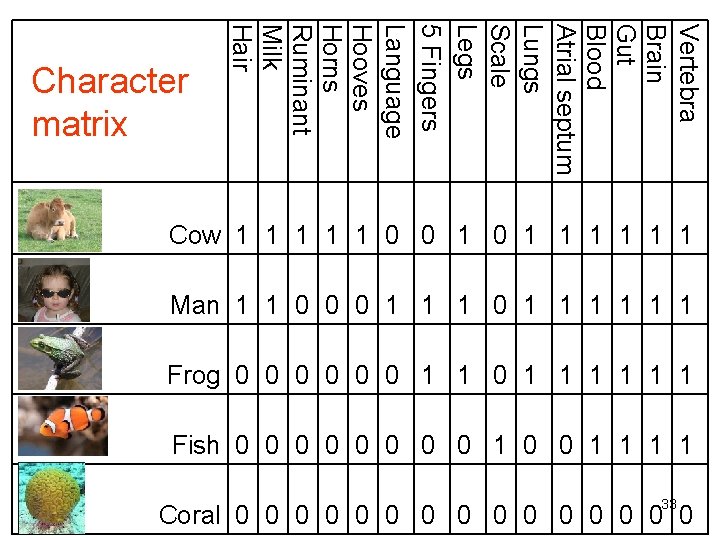
Vertebra Brain Gut Blood Atrial septum Lungs Scale Legs 5 Fingers Language Hooves Horns Ruminant Milk Hair Character matrix Cow 1 1 1 0 0 1 1 1 1 Man 1 1 0 0 0 1 1 1 Frog 0 0 0 1 1 1 Fish 0 0 0 0 1 1 33 Coral 0 0 0 0
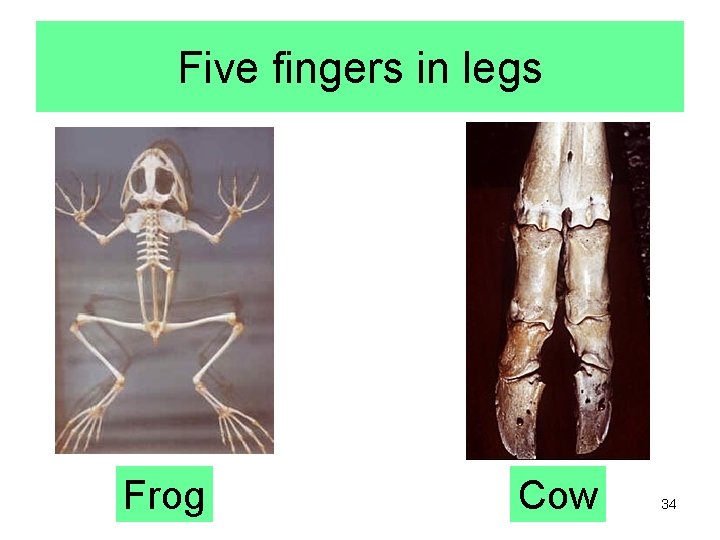
Five fingers in legs Frog Cow 34
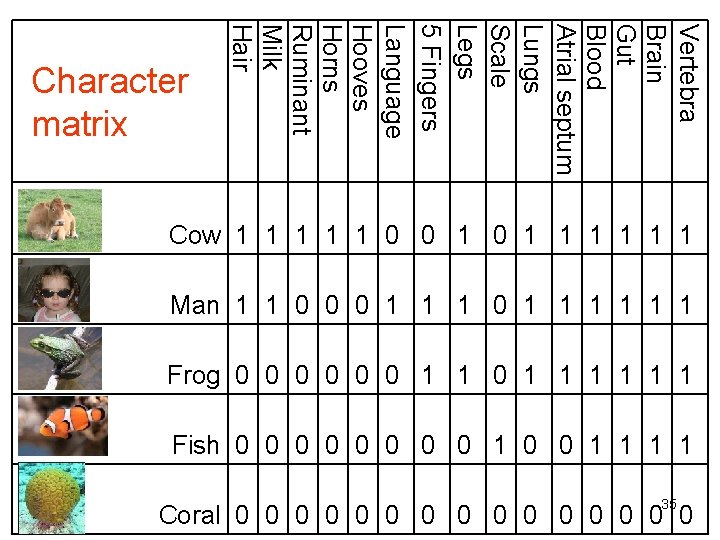
Vertebra Brain Gut Blood Atrial septum Lungs Scale Legs 5 Fingers Language Hooves Horns Ruminant Milk Hair Character matrix Cow 1 1 1 0 0 1 1 1 1 Man 1 1 0 0 0 1 1 1 Frog 0 0 0 1 1 1 Fish 0 0 0 0 1 1 35 Coral 0 0 0 0
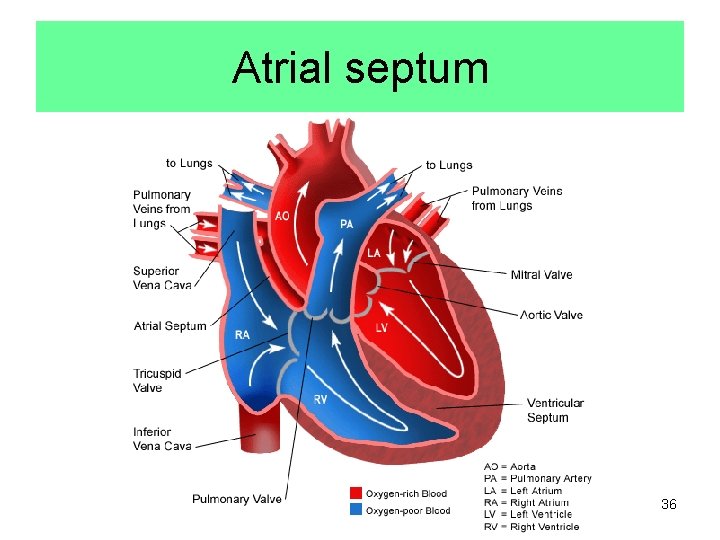
Atrial septum 36
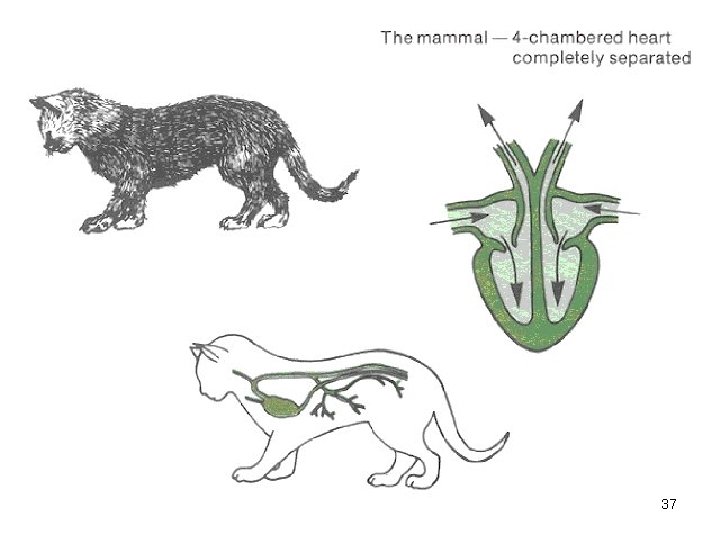
Heart vertebrate Frog Cow 37

38
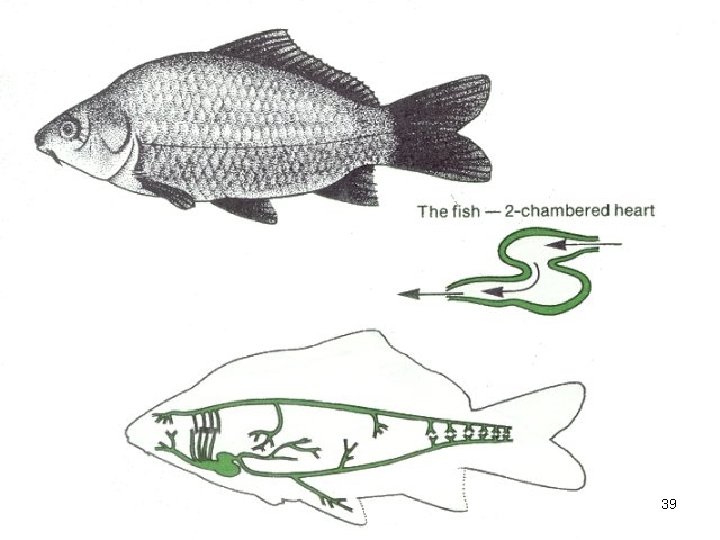
Heart vertebrate 39
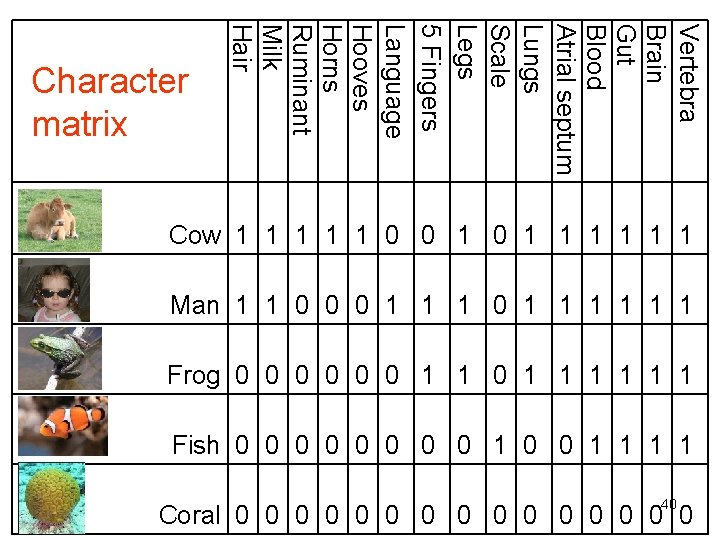
Vertebra Brain Gut Blood Atrial septum Lungs Scale Legs 5 Fingers Language Hooves Horns Ruminant Milk Hair Character matrix Cow 1 1 1 0 0 1 1 1 1 Man 1 1 0 0 0 1 1 1 Frog 0 0 0 1 1 1 Fish 0 0 0 0 1 1 40 Coral 0 0 0 0

2. Data->Tree 41

Today’s subjects • 1 - Introduction – History of science (Linnaeus, Darwin) • 2 - Phylogenetic trees • 3 - How to reconstruct phylogenetic trees with morphological characters – The data matrix – The cladistics method (monophyly / synapomorphy) – Molecular characters 42

Cladistic • Method developed by Willy Henning. • Willy Henning (1913 -1976) was a German entomologist. • Famous book in 1950 in German “Grundzüge einer Theorie der Phylogenetischen Systematik”. • Translated to English in 1966 “Phylogenetic Systematics”. Used only for morphological characters 43

Cladistic principles 1) Cladistics groups organisms strictly by common descent 44

A bit of vocabulary 45
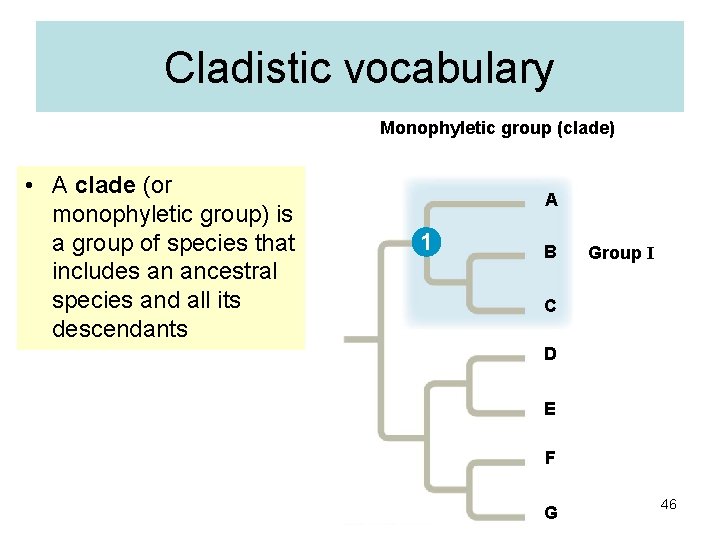
Cladistic vocabulary Monophyletic group (clade) • A clade (or monophyletic group) is a group of species that includes an ancestral species and all its descendants A 1 B Group I C D E F G 46
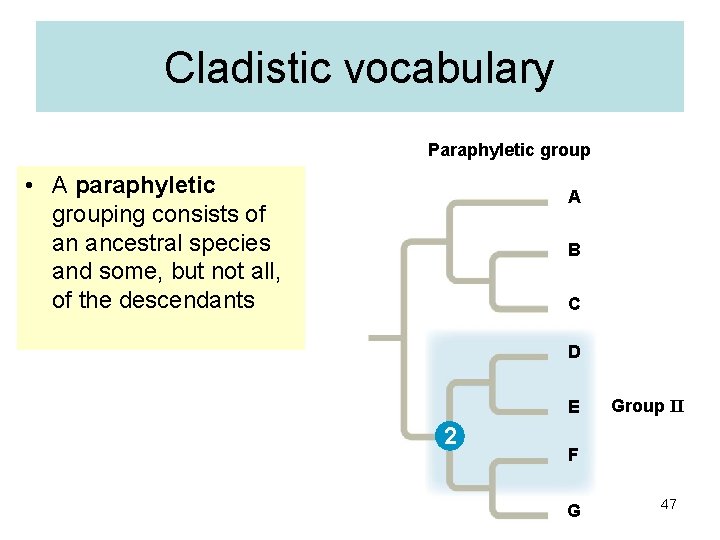
Cladistic vocabulary Paraphyletic group • A paraphyletic grouping consists of an ancestral species and some, but not all, of the descendants A B C D E 2 Group II F G 47
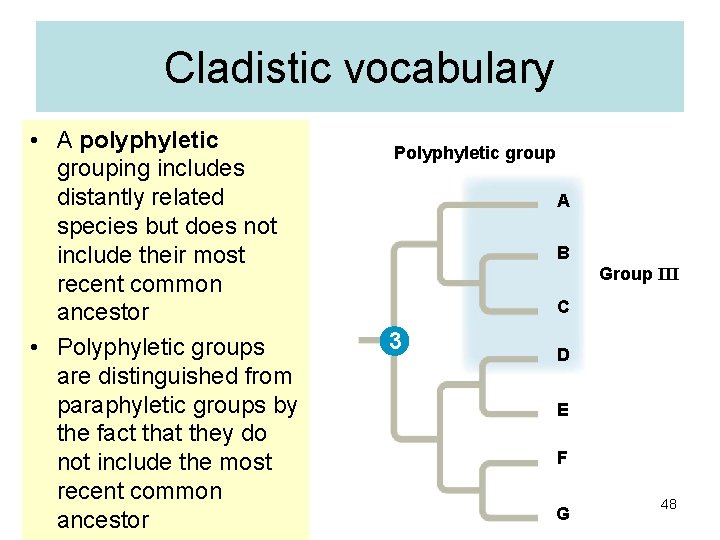
Cladistic vocabulary • A polyphyletic grouping includes distantly related species but does not include their most recent common ancestor • Polyphyletic groups are distinguished from paraphyletic groups by the fact that they do not include the most recent common ancestor Polyphyletic group A B Group III C 3 D E F G 48
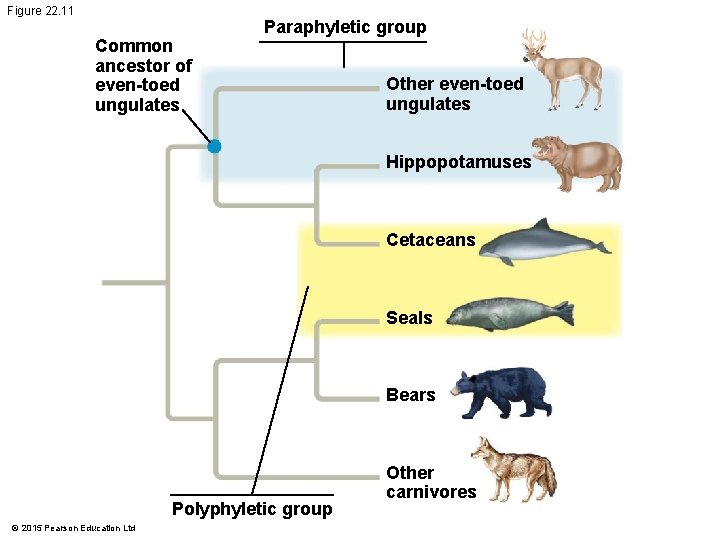
Figure 22. 11 Common ancestor of even-toed ungulates Paraphyletic group Other even-toed ungulates Hippopotamuses Cetaceans Seals Bears Polyphyletic group © 2015 Pearson Education Ltd Other carnivores
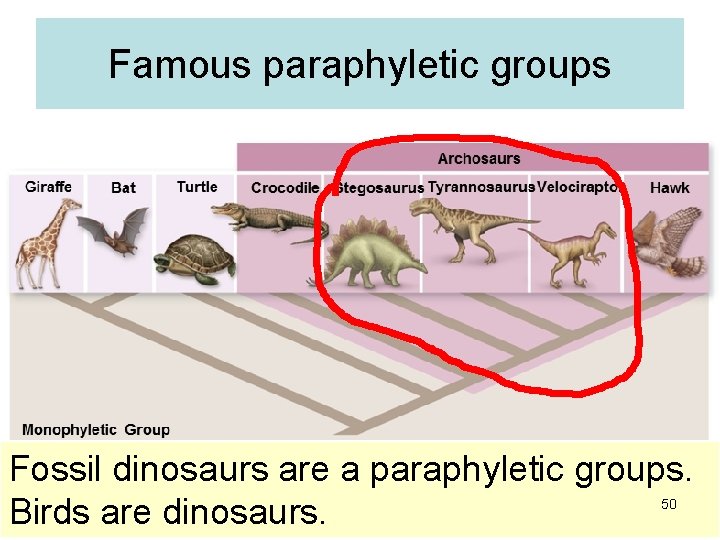
Famous paraphyletic groups Fossil dinosaurs are a paraphyletic groups. Birds are dinosaurs. 50

Velociraptors 51

52
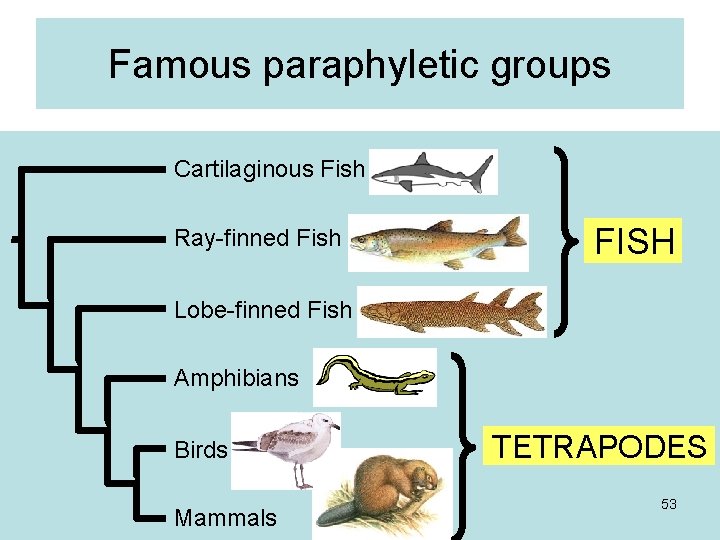
Famous paraphyletic groups Cartilaginous Fish Ray-finned Fish FISH Lobe-finned Fish Amphibians Birds Mammals TETRAPODES 53
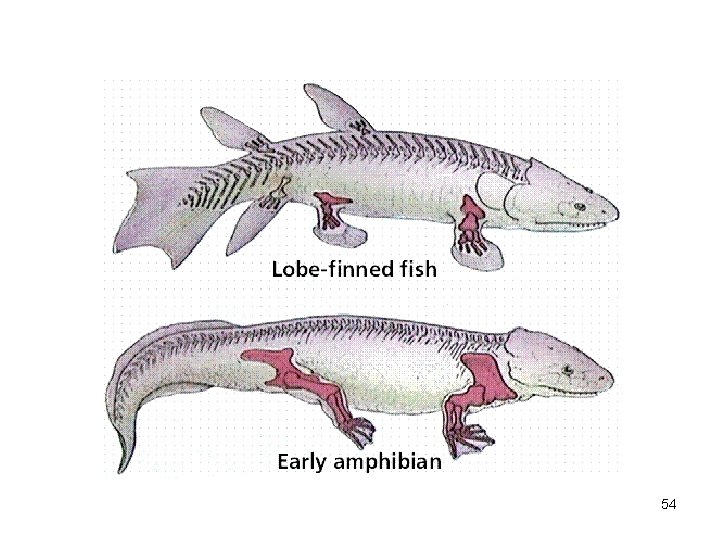
54
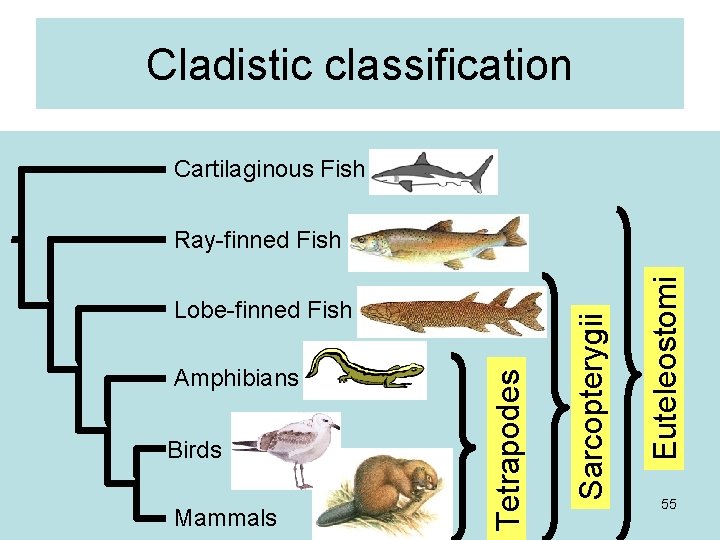
Cladistic classification Cartilaginous Fish Birds Mammals Euteleostomi Amphibians Tetrapodes Lobe-finned Fish Sarcopterygii Ray-finned Fish 55

Cladistic principles • (1) Organisms are grouped based on their common descent. • (2) Synapomorphies provide the only evidence for identifying relative recency of common ancestry. • Synapomorphies are understood to be the shared-derived (evolved, modified) features of organisms. 56

More vocabulary 57
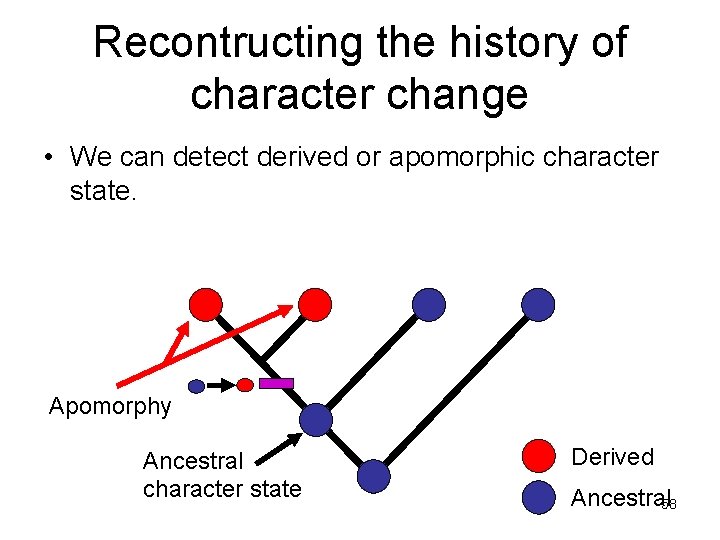
Recontructing the history of character change • We can detect derived or apomorphic character state. Apomorphy Ancestral character state Derived Ancestral 58
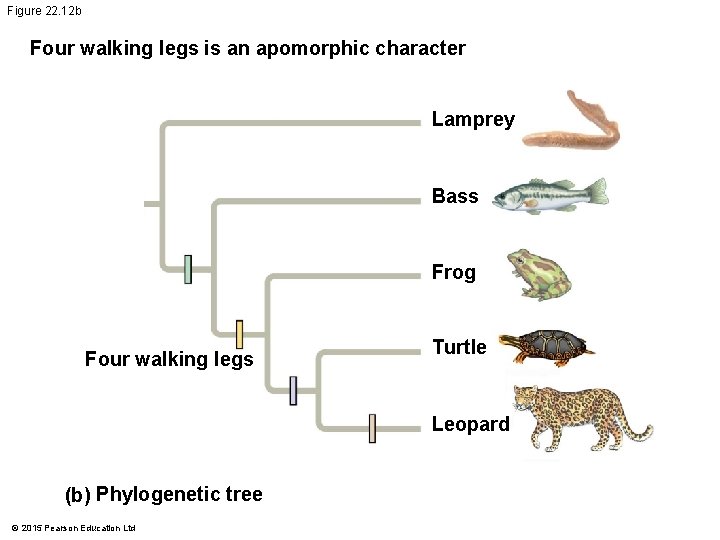
Figure 22. 12 b Four walking legs is an apomorphic character Lamprey Bass Frog Four walking legs Turtle Leopard (b) Phylogenetic tree © 2015 Pearson Education Ltd
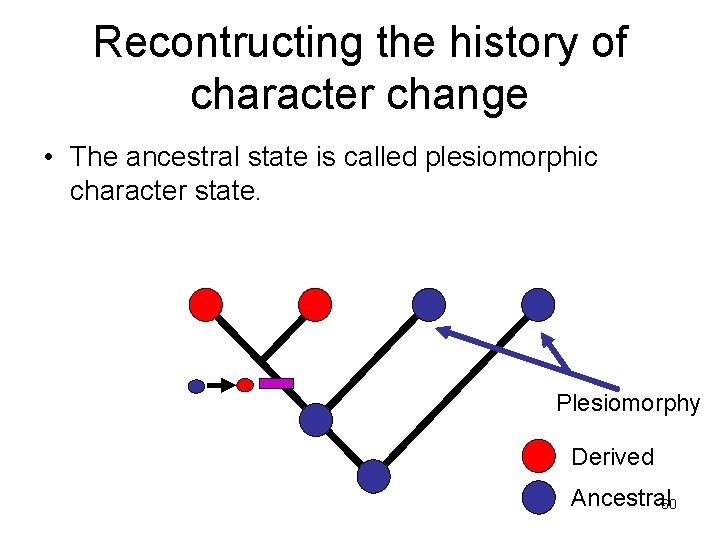
Recontructing the history of character change • The ancestral state is called plesiomorphic character state. Plesiomorphy Derived Ancestral 60
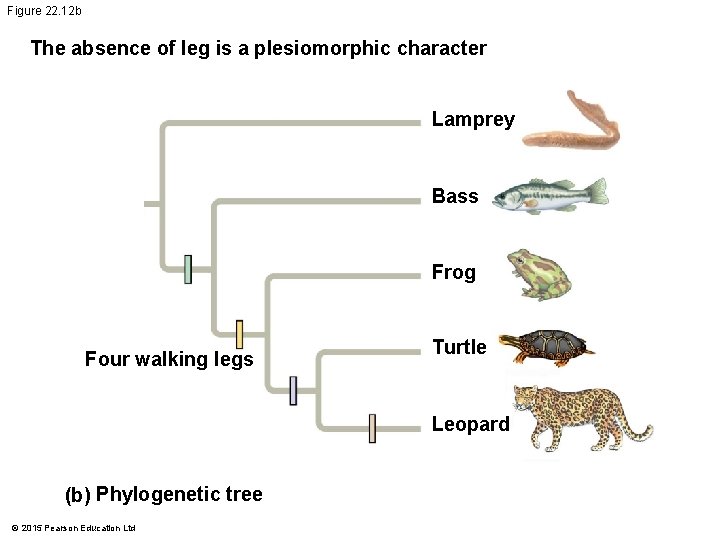
Figure 22. 12 b The absence of leg is a plesiomorphic character Lamprey Bass Frog Four walking legs Turtle Leopard (b) Phylogenetic tree © 2015 Pearson Education Ltd
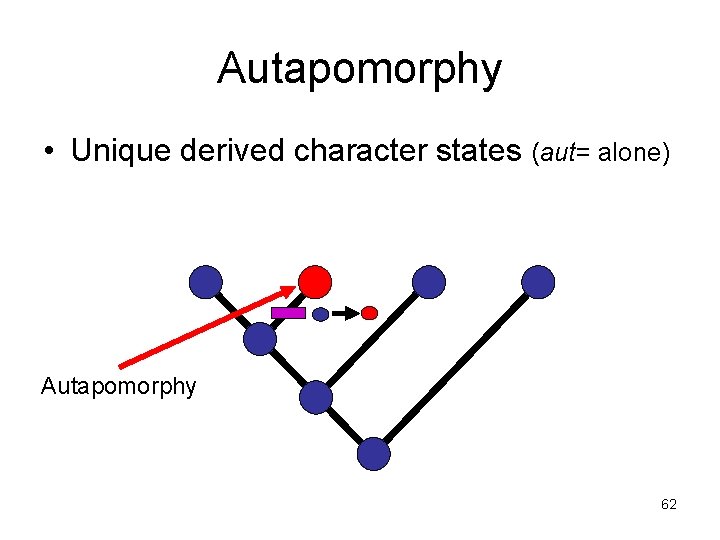
Autapomorphy • Unique derived character states (aut= alone) Autapomorphy 62
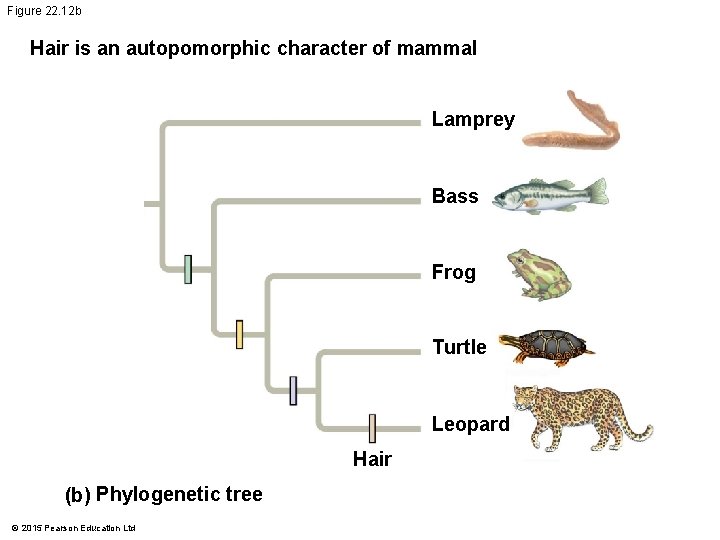
Figure 22. 12 b Hair is an autopomorphic character of mammal Lamprey Bass Frog Turtle Leopard Hair (b) Phylogenetic tree © 2015 Pearson Education Ltd
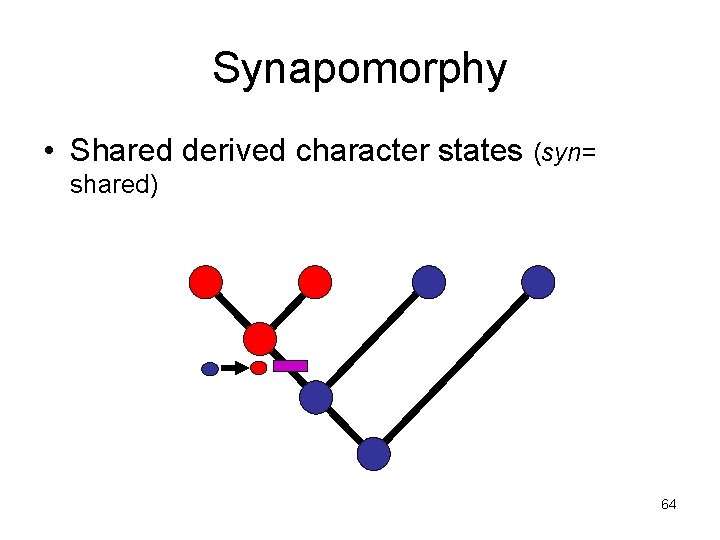
Synapomorphy • Shared derived character states (syn= shared) 64
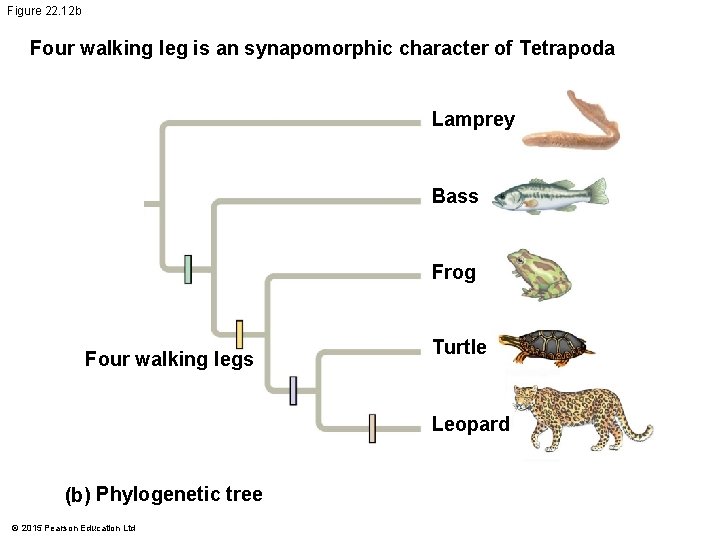
Figure 22. 12 b Four walking leg is an synapomorphic character of Tetrapoda Lamprey Bass Frog Four walking legs Turtle Leopard (b) Phylogenetic tree © 2015 Pearson Education Ltd
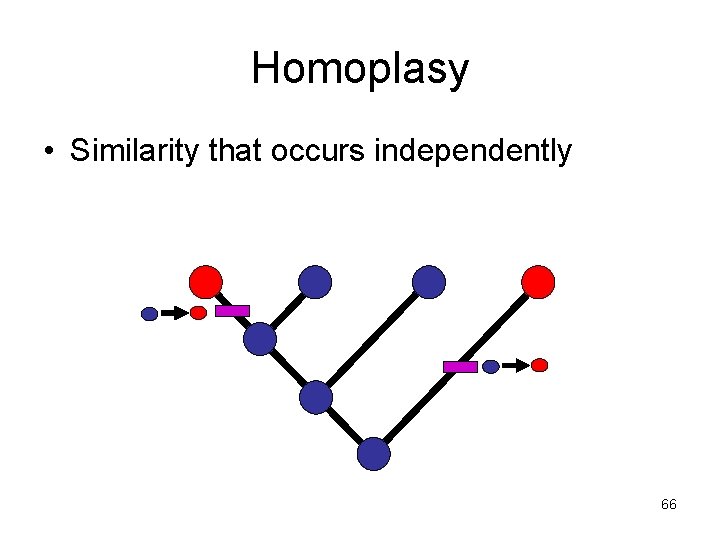
Homoplasy • Similarity that occurs independently 66

Warm blood is an homoplastic character (or "an instance of homoplasy“) that developed independently in mammals and birds Warm blood 67 Warm blood
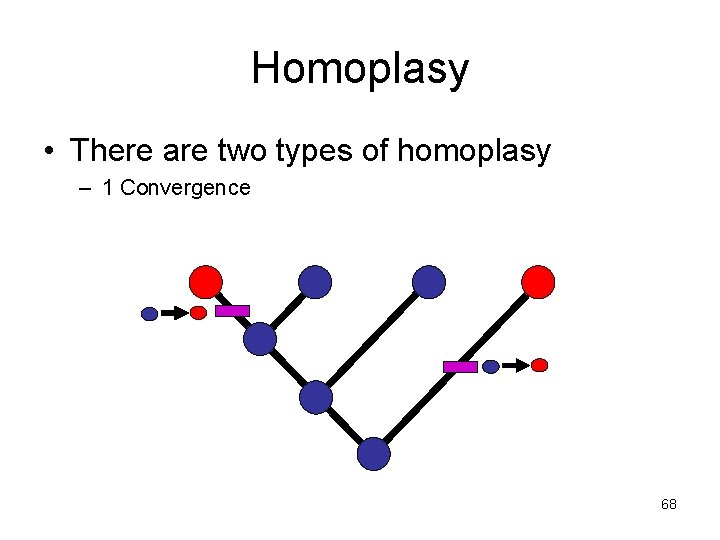
Homoplasy • There are two types of homoplasy – 1 Convergence 68
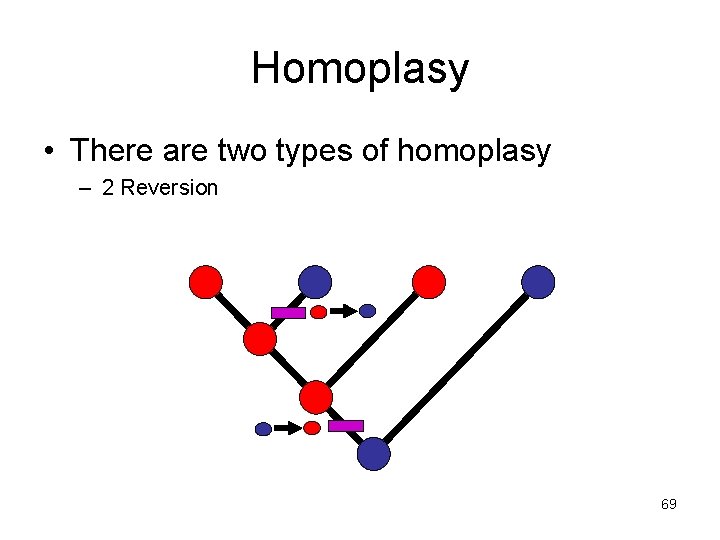
Homoplasy • There are two types of homoplasy – 2 Reversion 69
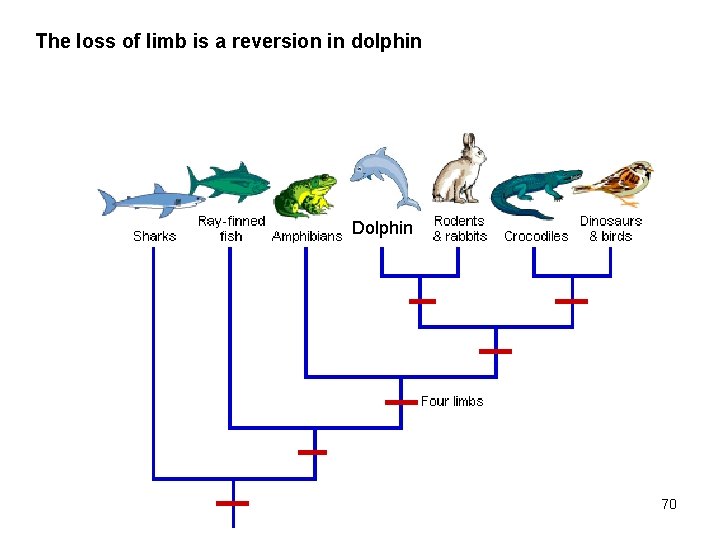
The loss of limb is a reversion in dolphin Dolphin 70

Cladistic principles • (1) organisms are grouped based on their common descent. • (2) Synapomorphies provide the only evidence for identifying relative recency of common ancestry. 71
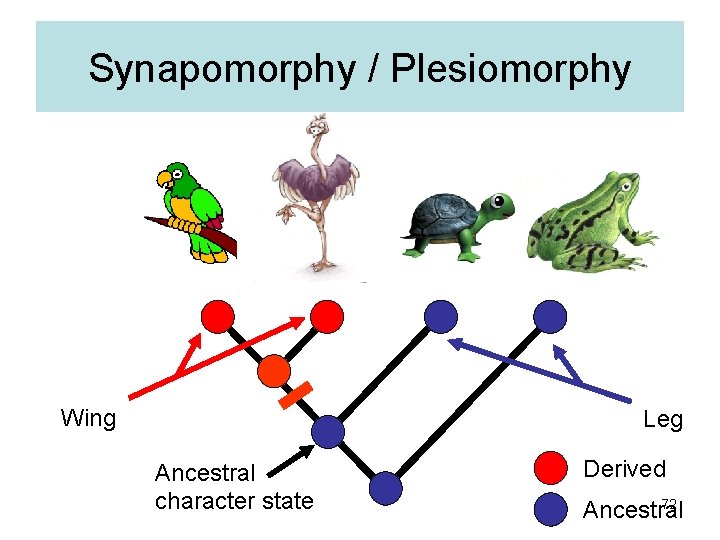
Synapomorphy / Plesiomorphy Wing Leg Ancestral character state Derived 72 Ancestral
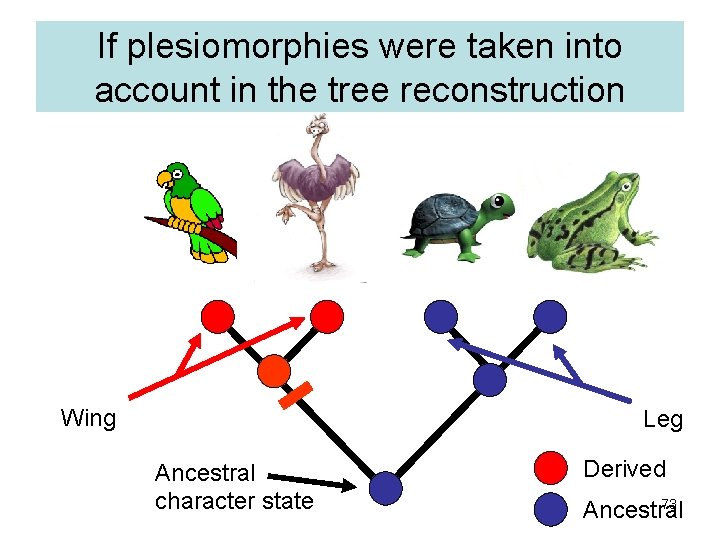
If plesiomorphies were taken into account in the tree reconstruction Wing Leg Ancestral character state Derived 73 Ancestral
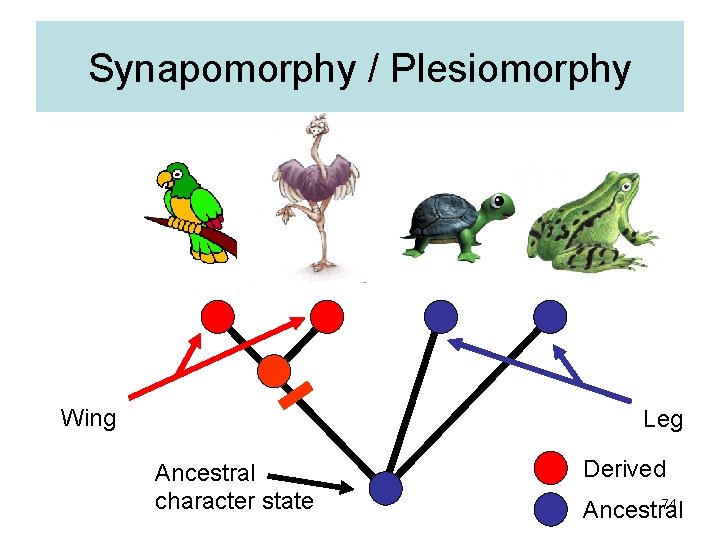
Synapomorphy / Plesiomorphy Wing Leg Ancestral character state Derived 74 Ancestral
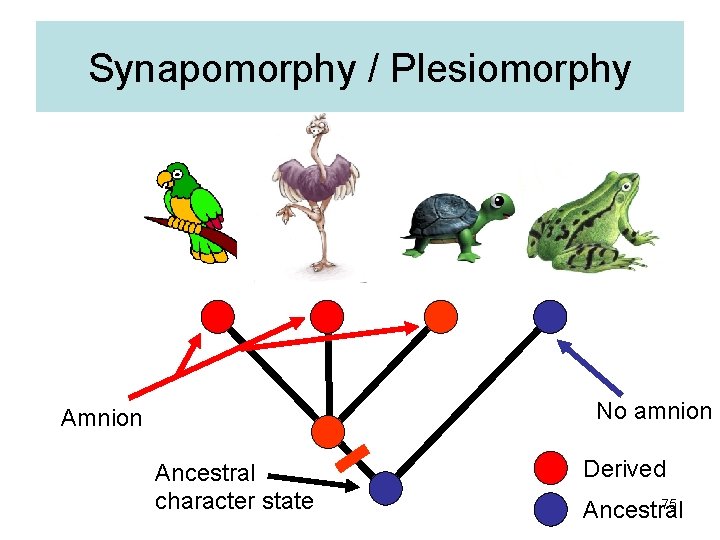
Synapomorphy / Plesiomorphy No amnion Ancestral character state Derived 75 Ancestral
Vertebra Brain Gut Blood Atrial septum Lungs Scale Legs 5 Fingers Language Hooves Horns Ruminant Milk Hair Character matrix Cow 1 1 1 0 0 1 1 1 1 Man 1 1 0 0 0 1 1 1 Frog 0 0 0 1 1 1 Fish 0 0 0 0 1 1 76 Coral 0 0 0 0
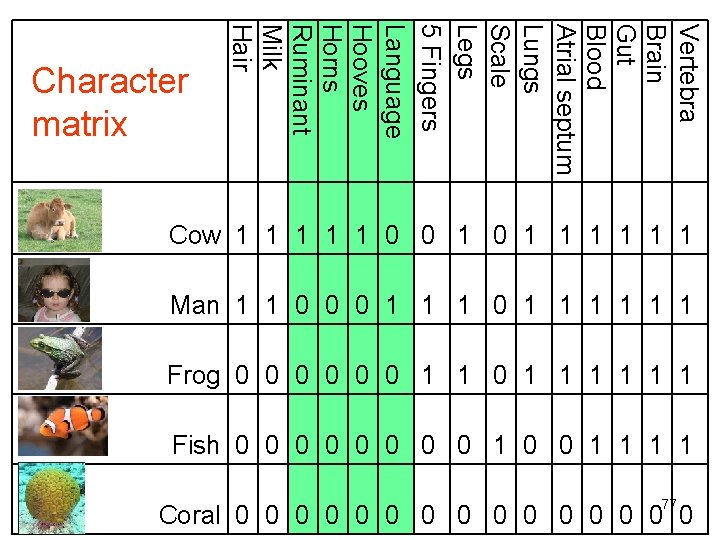
Vertebra Brain Gut Blood Atrial septum Lungs Scale Legs 5 Fingers Language Hooves Horns Ruminant Milk Hair Character matrix Cow 1 1 1 0 0 1 1 1 1 Man 1 1 0 0 0 1 1 1 Frog 0 0 0 1 1 1 Fish 0 0 0 0 1 1 77 Coral 0 0 0 0
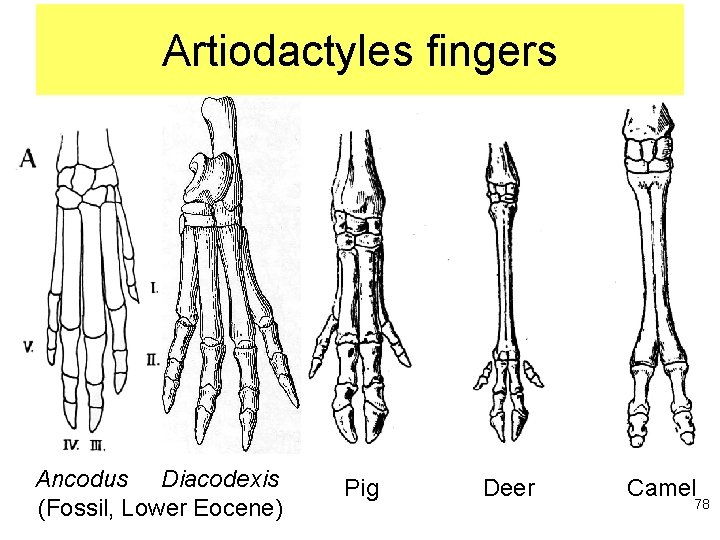
Artiodactyles fingers Ancodus Diacodexis (Fossil, Lower Eocene) Pig Deer Camel 78
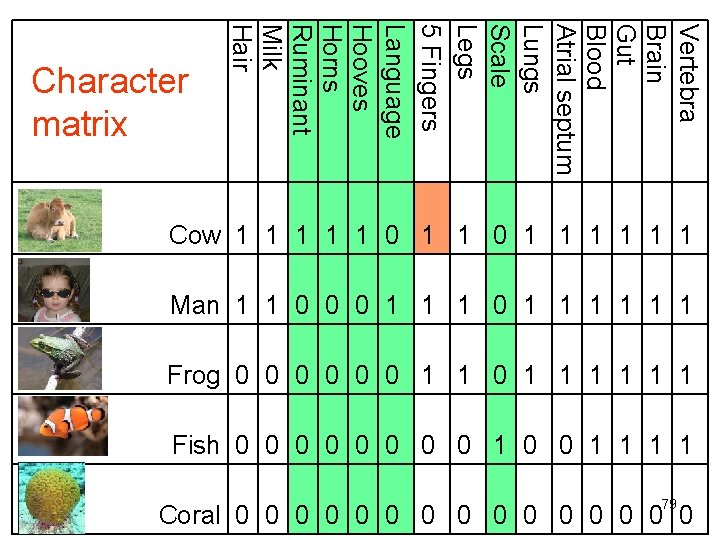
Vertebra Brain Gut Blood Atrial septum Lungs Scale Legs 5 Fingers Language Hooves Horns Ruminant Milk Hair Character matrix Cow 1 1 1 0 1 1 1 1 Man 1 1 0 0 0 1 1 1 Frog 0 0 0 1 1 1 Fish 0 0 0 0 1 1 79 Coral 0 0 0 0
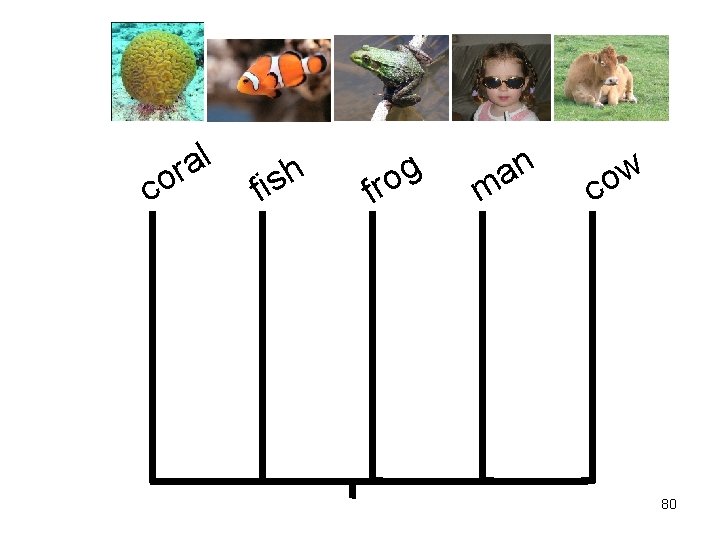
c l a or f h s i f g o r n a m w o c 80
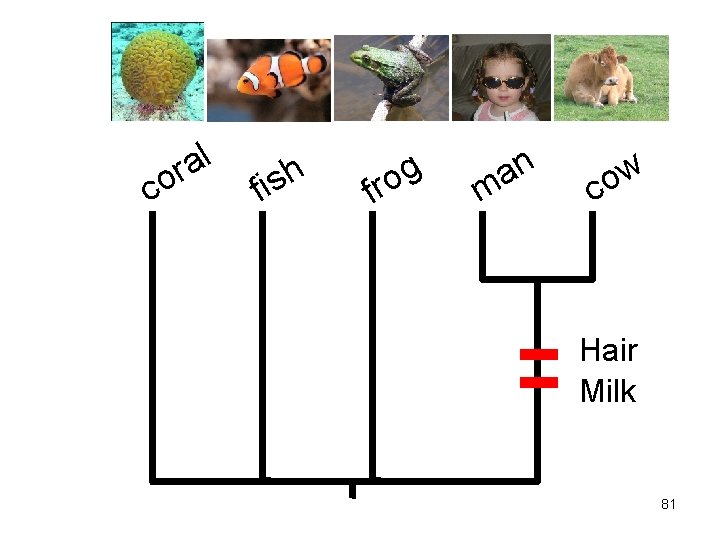
c l a or f h s i f g o r n a m w o c Hair Milk 81
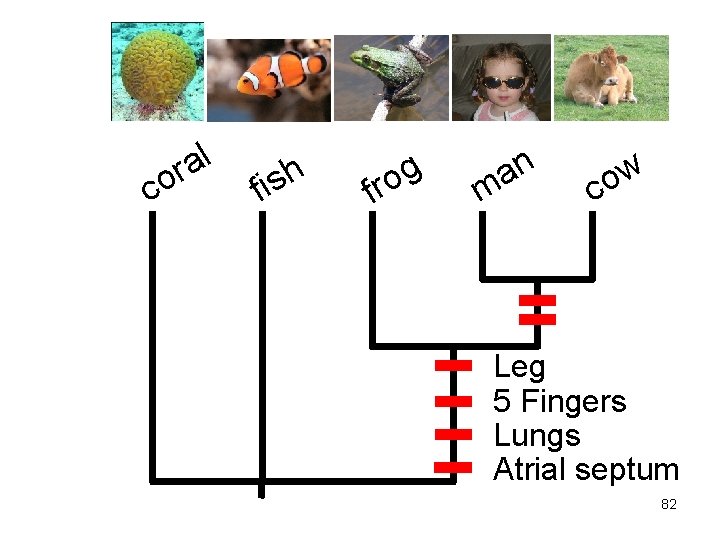
c l a or f h s i f g o r n a m w o c Leg 5 Fingers Lungs Atrial septum 82
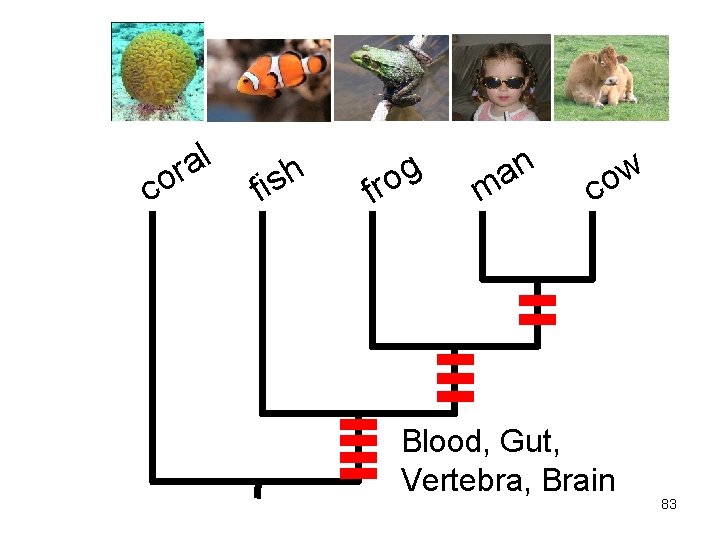
c l a or f h s i f g o r n a m w o c Blood, Gut, Vertebra, Brain 83
Cladistic steps • (1) Identify homologous characters. • (2) Differentiate ancestral character states from derived character states (identify polarity of characters). • (3) Identify convergent evolution. 84
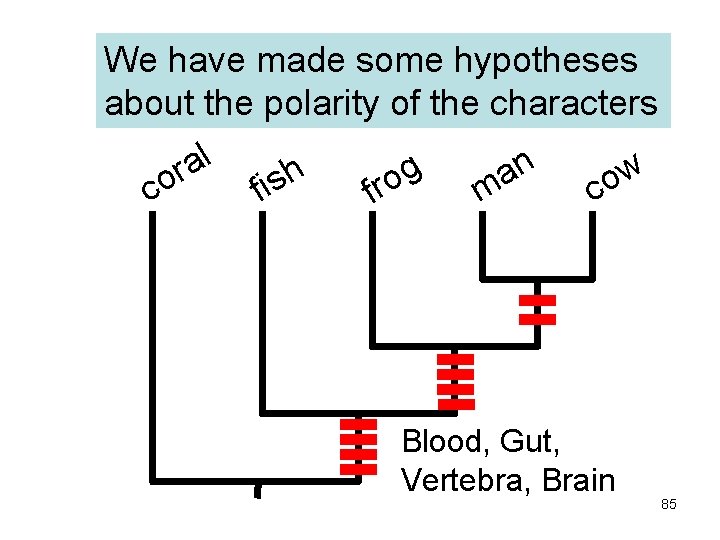
We have made some hypotheses about the polarity of the characters l a n g r w h a o o s o c fi m c fr Blood, Gut, Vertebra, Brain 85

How to identify character polarities • Outgroup comparison • Fossil record • Embryology (ontogeny recapitulates phylogeny) 86

Outgroup • A species or group of species from an evolutionary lineage that is known to have diverged before the lineage that includes the species we are studying (the ingroup). • A suitable outgroup can be determined based on evidence from morphology, paleontology, embryonic development, and gene sequences. 87

How to identify character polarities • Outgroup comparison • Fossil record • Embryology (ontogeny recapitulates phylogeny) 88
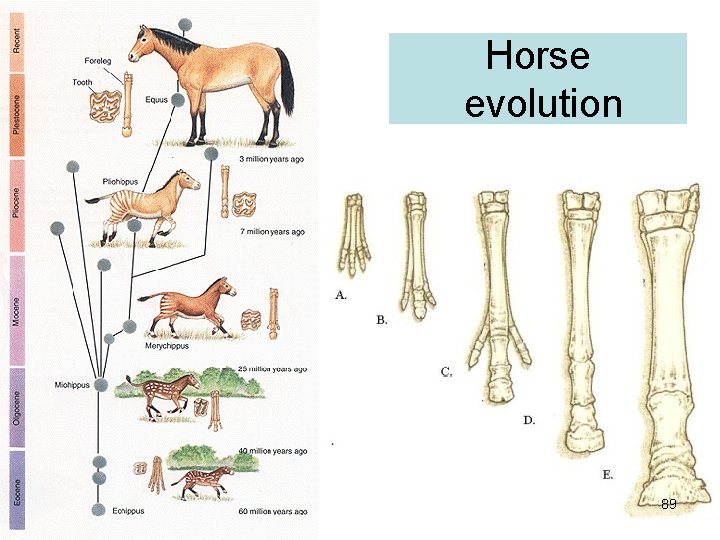
Horse evolution 89
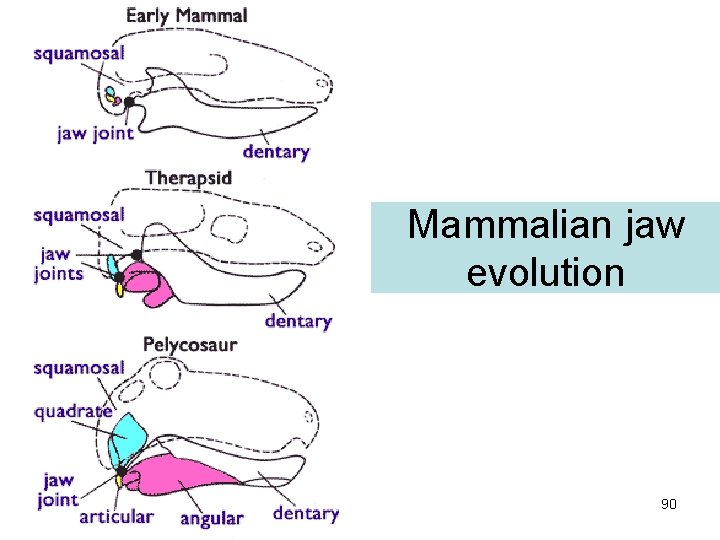
Mammalian jaw evolution 90
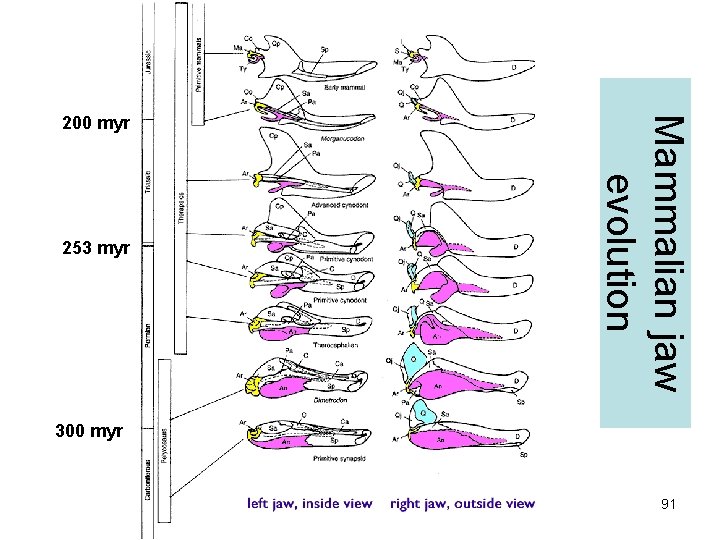
253 myr Mammalian jaw evolution 200 myr 300 myr 91

How to identify character polarities • Outgroup comparison • Fossil record • Embryology (ontogeny recapitulates phylogeny) 92
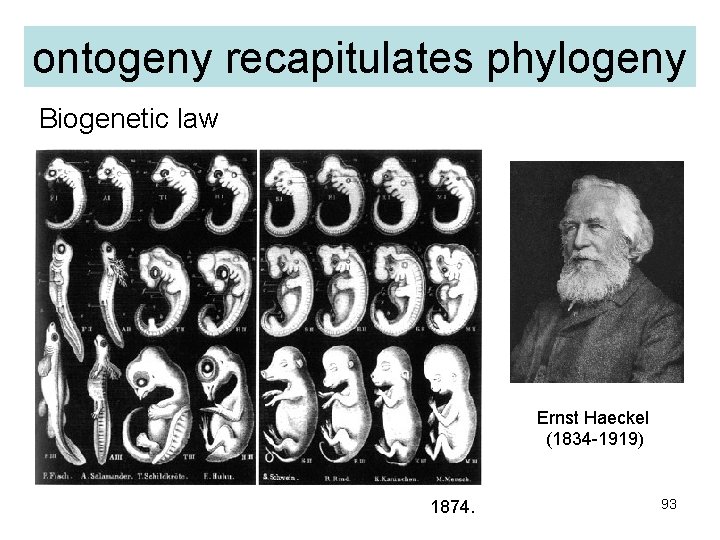
ontogeny recapitulates phylogeny Biogenetic law Ernst Haeckel (1834 -1919) 1874. 93
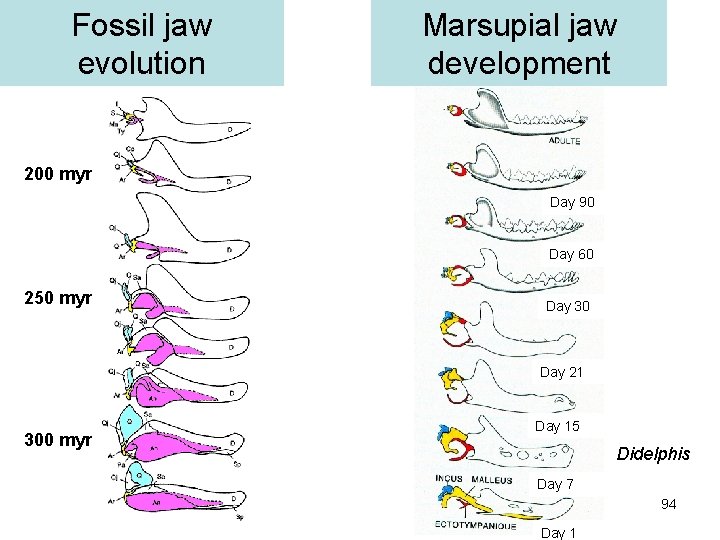
Fossil jaw evolution Marsupial jaw development 200 myr Day 90 Day 60 250 myr Day 30 Day 21 300 myr Day 15 Didelphis Day 7 94 Day 1

Advantage of cladistic • Every clades are clearly define by synapomorphy. • Cladistic is supposed to remove character conflicts. Theoretically, the conflict should be reduce to zero because real derived homologies cannot conflict. 95

Problem of cladistic • It might be difficult to identify character polarity. • It might be difficult to identify homoplasies. • Usually even after a cladistic analysis of the characters, some conflicts remain 96

Problem of cladistic • The cladistic method is not entirely objective. The systematist still has to make interpretations about homology and about how to weight and order characters for analysis. However, the cladistic paradigm does force the systematist to state explicitly why she believes characters are homologous or should be ordered in a certain way. This provides bases for falsifying hypotheses and promotes testing of alternatives. 97

Today’s subjects • 1 - Introduction – History of science • 2 - Phylogenetic trees • 3 - How to reconstruct phylogenetic trees with morphological characters – The data matrix – The cladistics method – Molecular characters 98
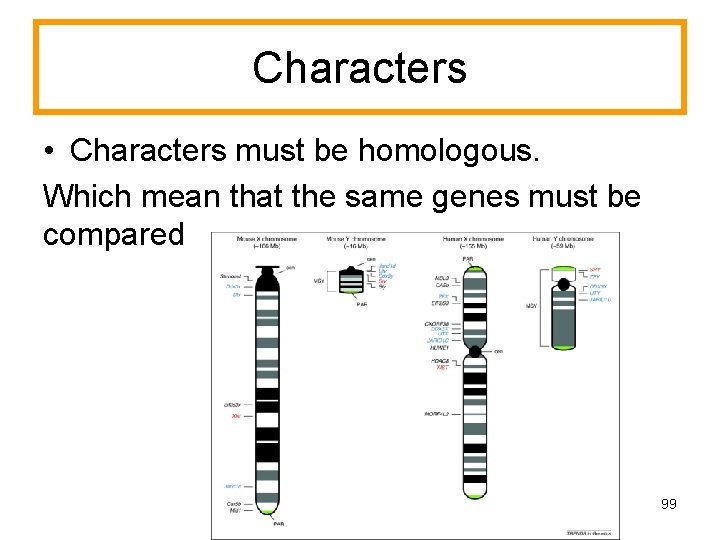
Characters • Characters must be homologous. Which mean that the same genes must be compared 99
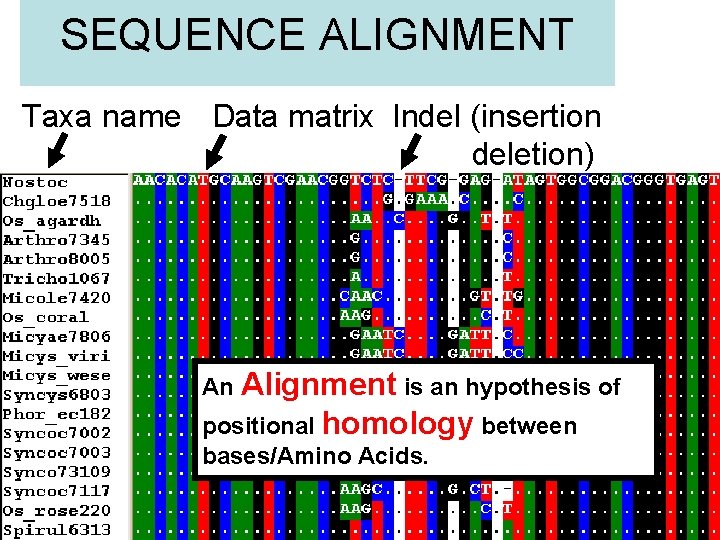
SEQUENCE ALIGNMENT Taxa name Data matrix Indel (insertion deletion) An Alignment is an hypothesis of positional homology between bases/Amino Acids.
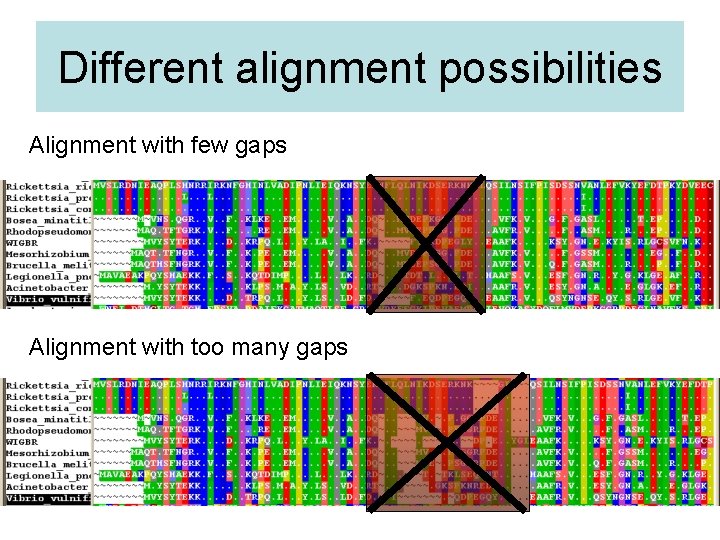
Different alignment possibilities Alignment with few gaps Alignment with too many gaps

Cladistic steps • (1) Identify homologous characters. s • (2) Differentiate ancestral character states r e t c a r a h c from derived character states (identify r a l u c e l o polarity of characters). m h t i w e l b i s s o p m i s i • T(3) his Identify convergent evolution. 102

The maximum likelihood method: A – Data matrix: B – Model of Sequence evolution: C – Tree topology: Species P= Tree a = 1 2 3 4 : : 1 A…TCT A…CTT A…CCT i G G A A n TAC…C CAC…C TAT…C Pdaa Pdac Pdag Pdat Pdca Pdcc Pdcg Pdct Pdga Pdgc Pdgg Pdtc Pdta Pdtc Pdtg Pdtt Species 1 Species 3 Species 2 Species 4

IMPORTANT TERMS AND CONCEPTS FROM THIS LECTURE l l l Phylogeny Linnaean classification Homology, Analogy What is a phylogenetic tree Monophyletic, Paraphyletic , Polyphyletic Sister taxa, sister clade 104

IMPORTANT TERMS AND CONCEPTS FROM THIS LECTURE l l Cladistic. Character polarity Synapomorphy, plesiomorphy, autapomorphy… Homoplasy, convergence, reversion… 105

In a comparison of birds and mammals, having four limbs is l l l a shared ancestral character. a character useful for sorting bird species. a shared derived character. an example of analogy rather than homology. a character useful for distinguishing birds from mammals. 106

In a comparison of birds and mammals, having four limbs is 1) 2) 3) 4) 5) a shared ancestral character. a character useful for sorting bird species. a shared derived character. an example of analogy rather than homology. a character useful for distinguishing birds from mammals. 107

Three living species X, Y, and Z share a common ancestor T, as do extinct species U and V. A grouping that consists of species T, X, Y, and Z (but not U or V) makes up 1) 2) 3) 4) 5) a monophyletic clade. a paraphyletic group. a polyphyletic group. an ingroup, with species U as the outgroup. a valid taxon. 108

Three living species X, Y, and Z share a common ancestor T, as do extinct species U and V. A grouping that consists of species T, X, Y, and Z (but not U or V) makes up 1) 2) 3) 4) 5) a monophyletic clade. a paraphyletic group. a polyphyletic group. an ingroup, with species U as the outgroup. a valid taxon. 109

Based on this tree, which statement is not correct? 1) 2) 3) 4) Salamanders are a sister group to the group containing lizards, goats, and humans. The salamander lineage is a basal taxon. Salamanders are as closely related to goats as to humans. Lizards are more closely related to salamanders than to humans. 110

Based on this tree, which statement is not correct? 1) 2) 3) 4) Salamanders are a sister group to the group containing lizards, goats, and humans. The salamander lineage is a basal taxon. Salamanders are as closely related to goats as to humans. Lizards are more closely related to salamanders than to humans. 111
- Slides: 111