Biological Pathways Janick Mathys Biological Pathways Definition Biochemical

Biological Pathways Janick Mathys

Biological Pathways • Definition • Biochemical compounds • Biological interactions • Energy • Control interactions • Levels of abstraction • Types of biological pathways • Integration of pathways • Inference Issues

Biological Pathways Definition: A biological pathway is a sequence of interactions between biochemical compounds aimed at the maintenance and control of the flow of information, energy and biochemical compounds in the cell and the ability of the cell to change its behaviour in response to stimuli.

Biological Pathways Definition: A biological pathway is a sequence of interactions between biochemical compounds aimed at the maintenance and control of the flow of information, energy and biochemical compounds in the cell. Main types of compounds in the context of pathways: - proteins and protein complexes - (part of) genes - metabolites

Biochemical compounds • (part of) genes • proteins and protein complexes • metabolites – – – – – Amino acids and peptides : A, C, F, G, H, S… Carbohydrates (sugars) Cell-structure components Cofactors, prosthetic groups and electron carriers: vitamins Fatty acids and lipids Nucleotides and nucleic acids: A, C, T, G Monocarbon compounds: CO, CH 4, CH 3 OH Essential elements: S, P, O 2, Fe, radicals Aromatic compounds: compounds with special stability and properties due to a closed loop of electrons (ring structure) – …
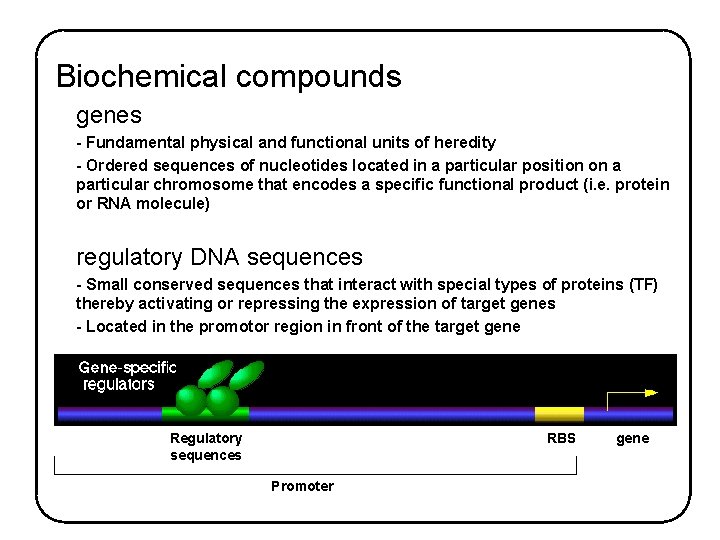
Biochemical compounds genes - Fundamental physical and functional units of heredity - Ordered sequences of nucleotides located in a particular position on a particular chromosome that encodes a specific functional product (i. e. protein or RNA molecule) regulatory DNA sequences - Small conserved sequences that interact with special types of proteins (TF) thereby activating or repressing the expression of target genes - Located in the promotor region in front of the target gene Regulatory sequences RBS Promoter gene
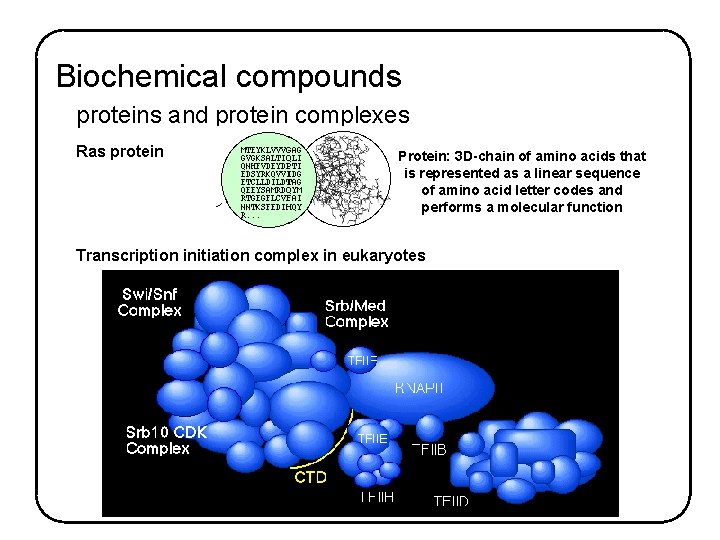
Biochemical compounds proteins and protein complexes Ras protein Protein: 3 D-chain of amino acids that is represented as a linear sequence of amino acid letter codes and performs a molecular function Transcription initiation complex in eukaryotes
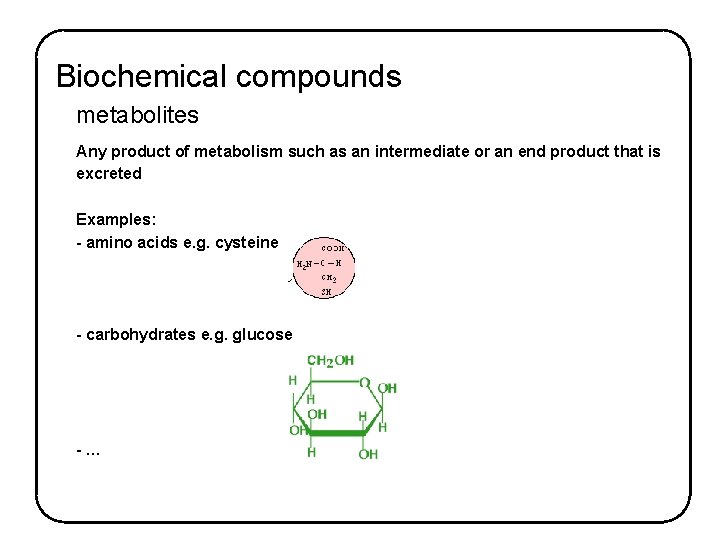
Biochemical compounds metabolites Any product of metabolism such as an intermediate or an end product that is excreted Examples: - amino acids e. g. cysteine - carbohydrates e. g. glucose -…
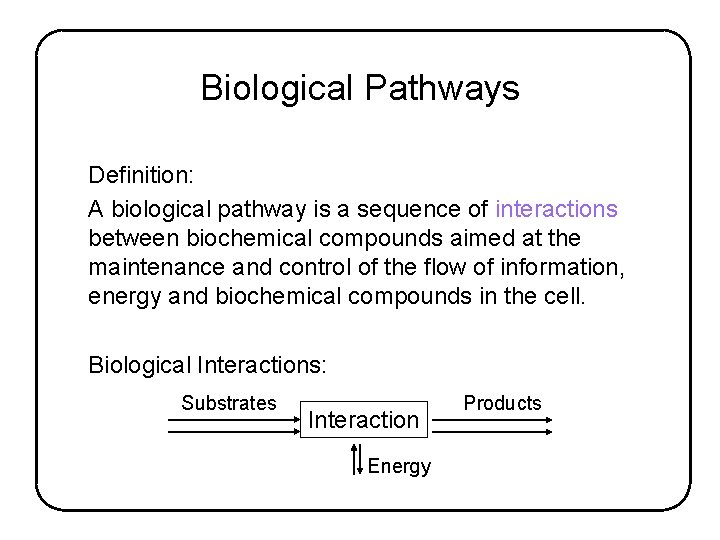
Biological Pathways Definition: A biological pathway is a sequence of interactions between biochemical compounds aimed at the maintenance and control of the flow of information, energy and biochemical compounds in the cell. Biological Interactions: Substrates Interaction Energy Products
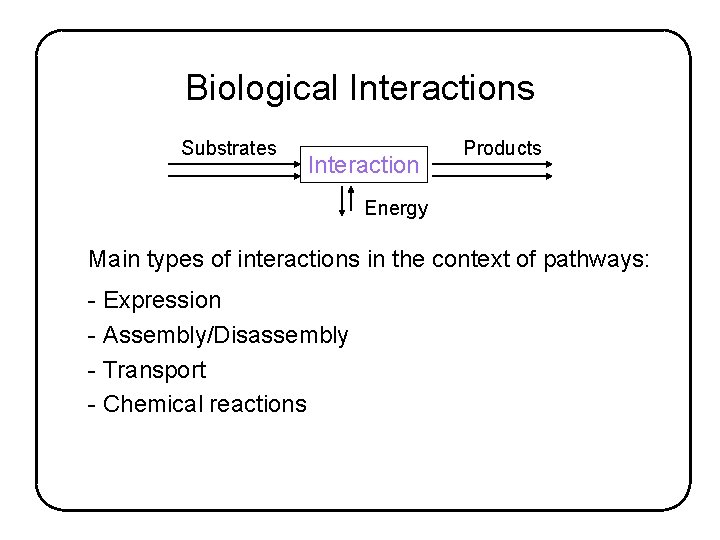
Biological Interactions Substrates Interaction Products Energy Main types of interactions in the context of pathways: - Expression - Assembly/Disassembly - Transport - Chemical reactions
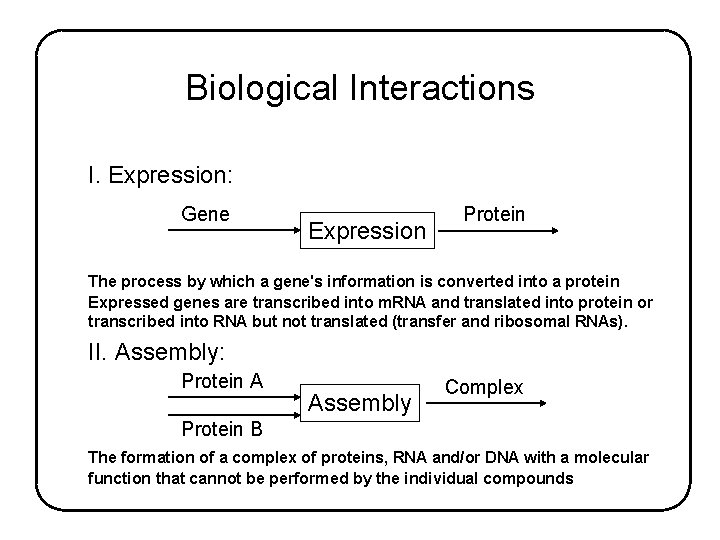
Biological Interactions I. Expression: Gene Expression Protein The process by which a gene's information is converted into a protein Expressed genes are transcribed into m. RNA and translated into protein or transcribed into RNA but not translated (transfer and ribosomal RNAs). II. Assembly: Protein A Assembly Complex Protein B The formation of a complex of proteins, RNA and/or DNA with a molecular function that cannot be performed by the individual compounds
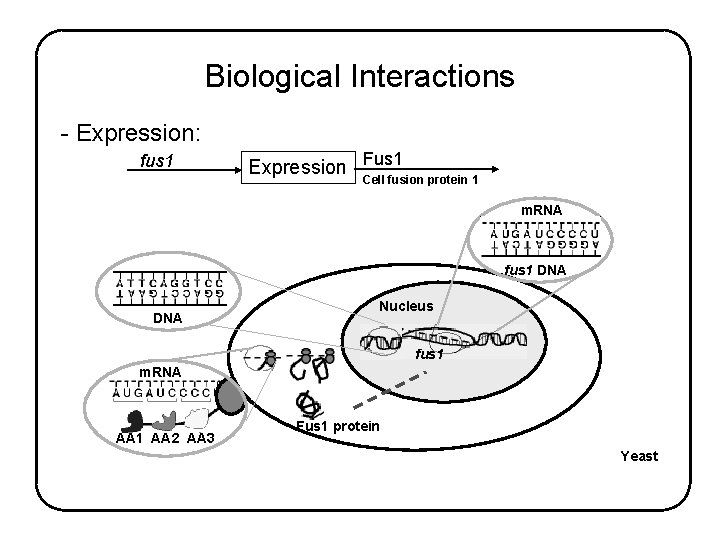
Biological Interactions - Expression: fus 1 Expression Fus 1 Cell fusion protein 1 m. RNA fus 1 DNA Nucleus fus 1 m. RNA AA 1 AA 2 AA 3 Fus 1 protein Yeast
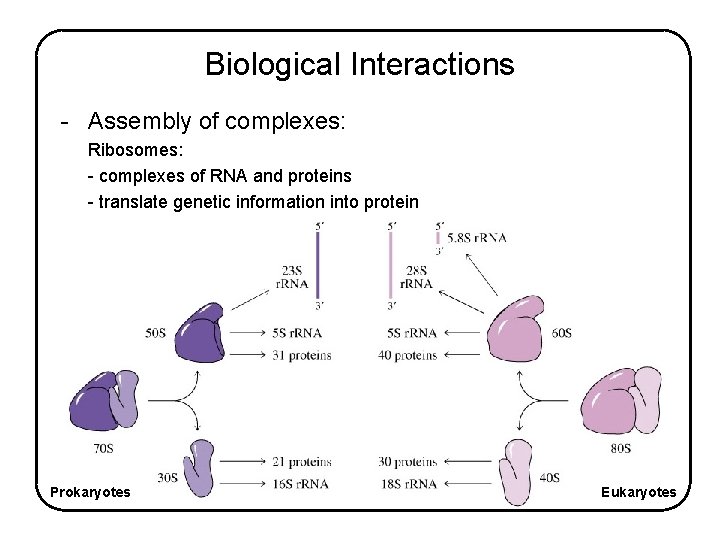
Biological Interactions - Assembly of complexes: Ribosomes: - complexes of RNA and proteins - translate genetic information into protein Prokaryotes Eukaryotes
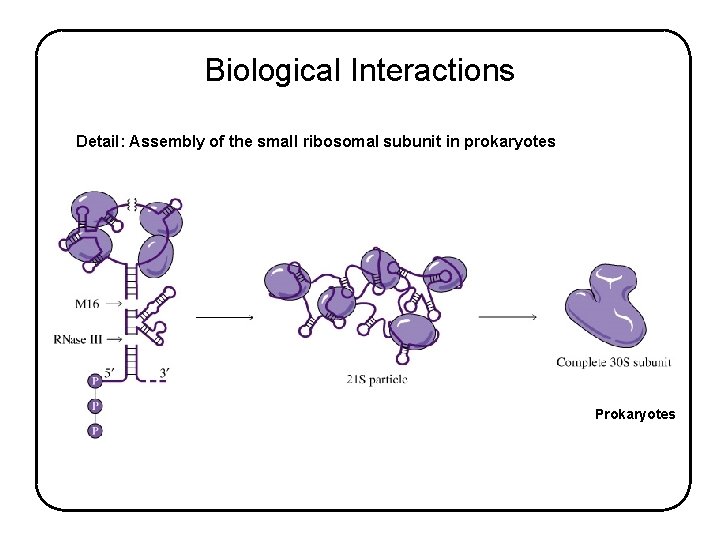
Biological Interactions Detail: Assembly of the small ribosomal subunit in prokaryotes Prokaryotes
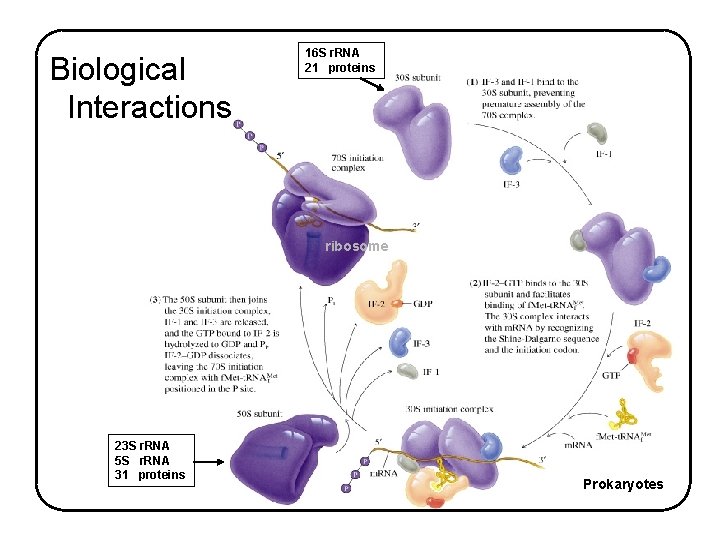
Biological Interactions 16 S r. RNA 21 proteins ribosome 23 S r. RNA 5 S r. RNA 31 proteins Prokaryotes
Biological Interactions III. Transport: Compound A at location 1 Transport Compound A at location 2 Change of location of compounds IV. Chemical reaction: Compound A Reaction Compound B

Biological Interactions - Transport: a. Transport of nascent proteins through plasma membrane of the ER: Ribosome - nascent protein complex Transport in cytoplasm Nascent protein in lumen of ER b. Transport of glucose from the lumen of the intestine into the blood: Glucose in intestine Transport Glucose in epithelial cell Transport Glucose in blood
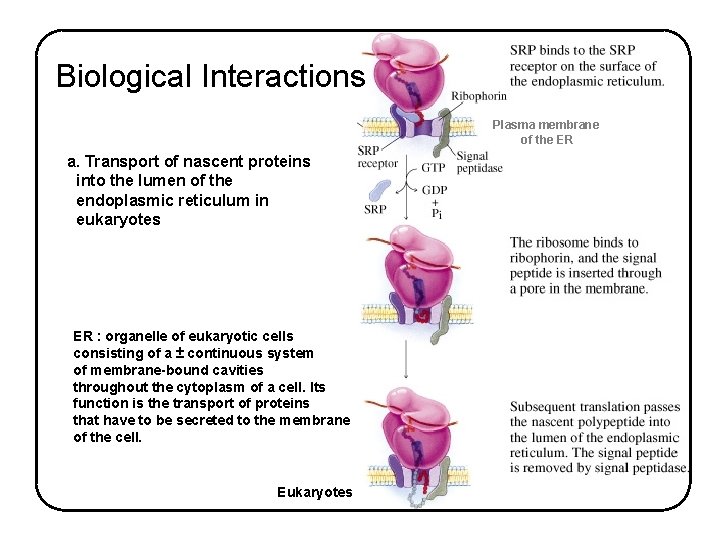
Biological Interactions Plasma membrane of the ER a. Transport of nascent proteins into the lumen of the endoplasmic reticulum in eukaryotes ER : organelle of eukaryotic cells consisting of a ± continuous system of membrane-bound cavities throughout the cytoplasm of a cell. Its function is the transport of proteins that have to be secreted to the membrane of the cell. Eukaryotes
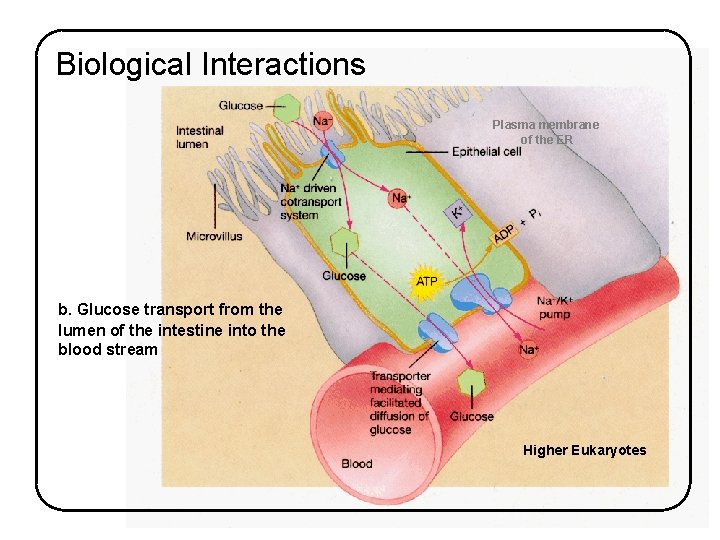
Biological Interactions Plasma membrane of the ER b. Glucose transport from the lumen of the intestine into the blood stream Higher Eukaryotes

Biological Interactions - Chemical reactions: 1. Redox reactions: Oxidation - Reduction (Photosynthesis): transfer of e- from electron donors to electron acceptors 2. Phosphorylation - Dephosphorylation (Signal transduction): addition/removal of phosphate groups 3. Hydrolysis: breakdown of bonds in compounds through the addition of water 4. Splitting or forming of a C-C bond 5. Isomerisation: Change of geometry or structure of a compound 6. Polymerisation 7. …
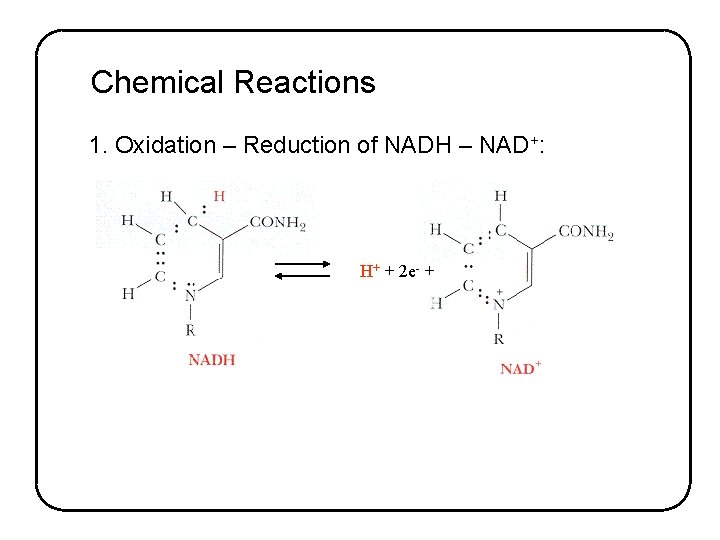
Chemical Reactions 1. Oxidation – Reduction of NADH – NAD+: H+ + 2 e- +
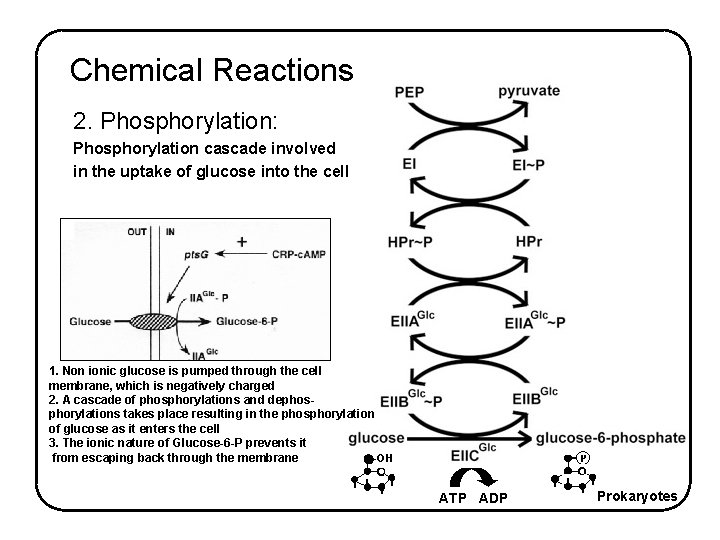
Chemical Reactions 2. Phosphorylation: Phosphorylation cascade involved in the uptake of glucose into the cell 1. Non ionic glucose is pumped through the cell membrane, which is negatively charged 2. A cascade of phosphorylations and dephosphorylations takes place resulting in the phosphorylation of glucose as it enters the cell 3. The ionic nature of Glucose-6 -P prevents it -OH from escaping back through the membrane ATP ADP Prokaryotes
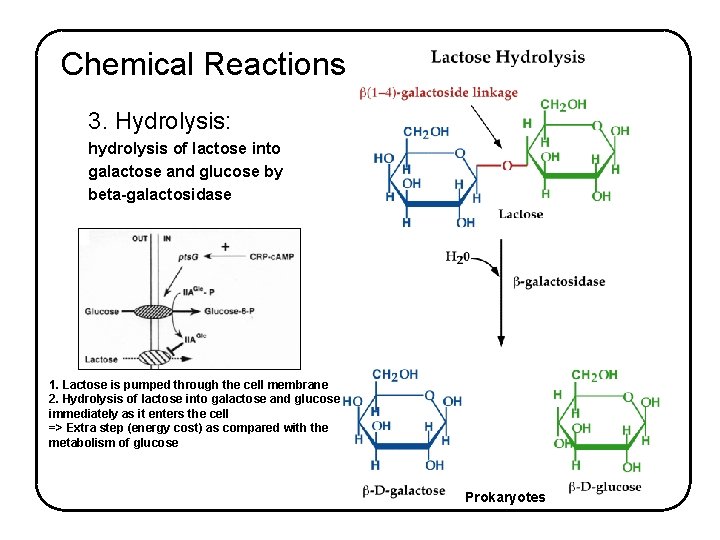
Chemical Reactions 3. Hydrolysis: hydrolysis of lactose into galactose and glucose by beta-galactosidase 1. Lactose is pumped through the cell membrane 2. Hydrolysis of lactose into galactose and glucose immediately as it enters the cell => Extra step (energy cost) as compared with the metabolism of glucose Prokaryotes
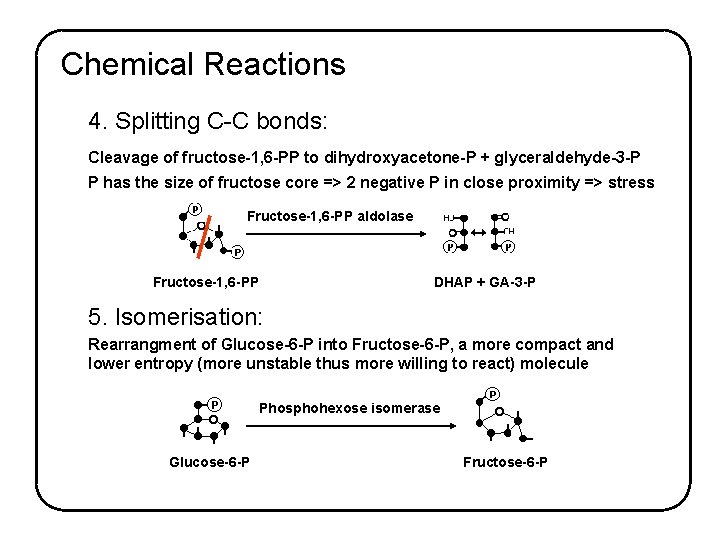
Chemical Reactions 4. Splitting C-C bonds: Cleavage of fructose-1, 6 -PP to dihydroxyacetone-P + glyceraldehyde-3 -P P has the size of fructose core => 2 negative P in close proximity => stress Fructose-1, 6 -PP aldolase Fructose-1, 6 -PP DHAP + GA-3 -P 5. Isomerisation: Rearrangment of Glucose-6 -P into Fructose-6 -P, a more compact and lower entropy (more unstable thus more willing to react) molecule Phosphohexose isomerase Glucose-6 -P Fructose-6 -P

Biological Interactions Substrate Interaction Product Energy • Energy is always required to form chemical bonds • Energy is sometimes released by the breaking of chemical bonds • For biological interactions the cell uses 3 energy sources: - ATP: Adenosine Tri. Phosphate - GTP: Guanine Tri. Phosphate - Creatine phosphate • ATP is generated by electron transfer in mitochondria: - Electron carriers pick up H+ and e- released by the breakdown of nutrients (e. g. glucose) - Electron carriers transfer H+ and e- to electron carriers in the mitochondrial membrane - Transfer of e- through the mitochondrial membrane down to O 2 releases energy - This energy is used to transport the H+ across the mitochondrial membrane - Rush of H+ releases energy used for phosphorylation of ADP to generate ATP
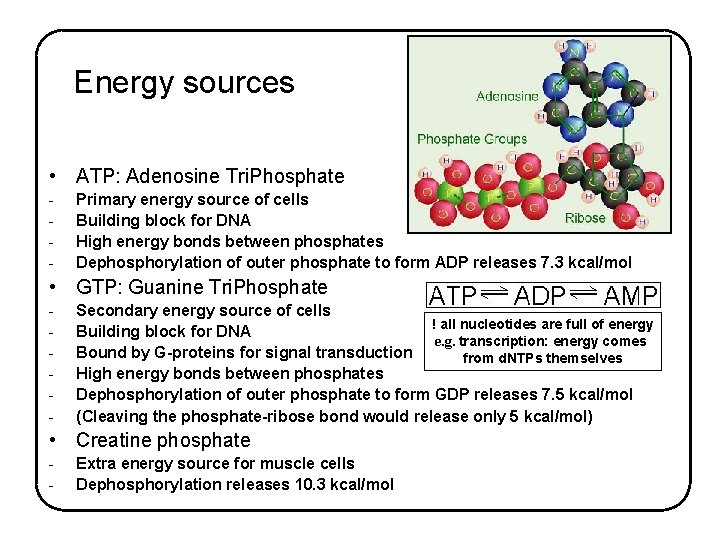
Energy sources • ATP: Adenosine Tri. Phosphate - Primary energy source of cells Building block for DNA High energy bonds between phosphates Dephosphorylation of outer phosphate to form ADP releases 7. 3 kcal/mol • GTP: Guanine Tri. Phosphate - Secondary energy source of cells ! all nucleotides are full of energy Building block for DNA e. g. transcription: energy comes Bound by G-proteins for signal transduction from d. NTPs themselves High energy bonds between phosphates Dephosphorylation of outer phosphate to form GDP releases 7. 5 kcal/mol (Cleaving the phosphate-ribose bond would release only 5 kcal/mol) • Creatine phosphate - Extra energy source for muscle cells Dephosphorylation releases 10. 3 kcal/mol
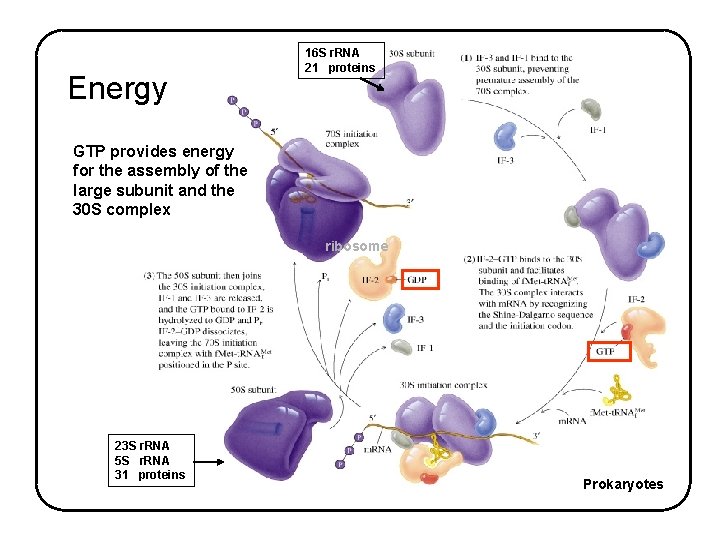
Energy 16 S r. RNA 21 proteins GTP provides energy for the assembly of the large subunit and the 30 S complex ribosome 23 S r. RNA 5 S r. RNA 31 proteins Prokaryotes
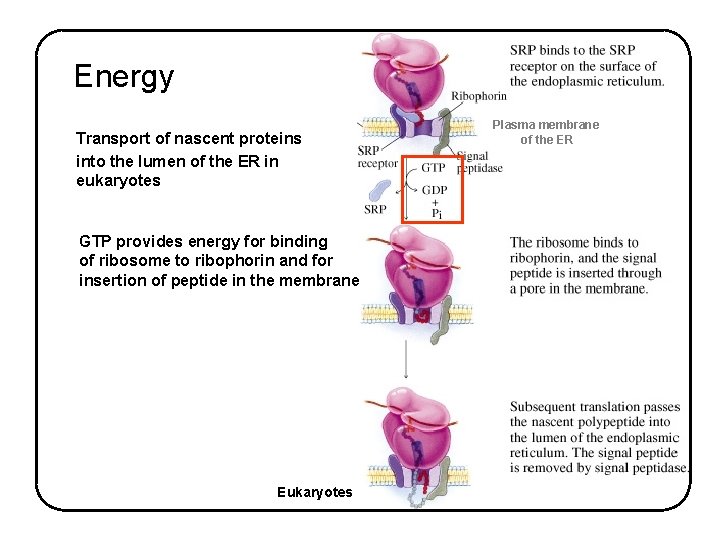
Energy Transport of nascent proteins into the lumen of the ER in eukaryotes GTP provides energy for binding of ribosome to ribophorin and for insertion of peptide in the membrane Eukaryotes Plasma membrane of the ER
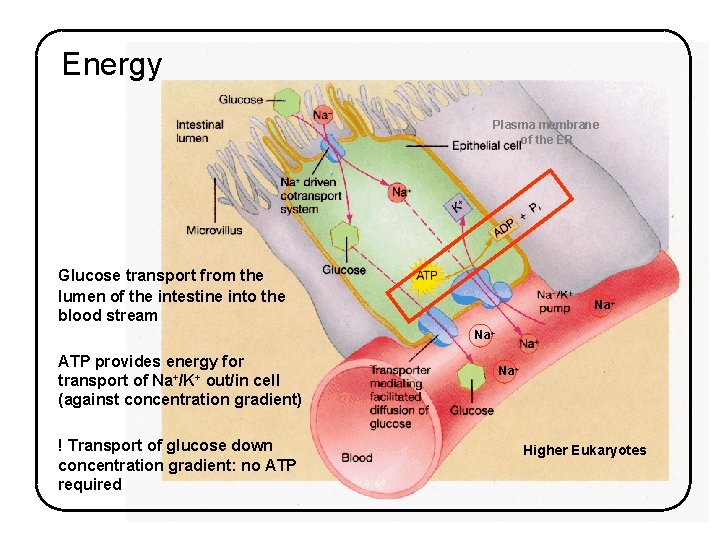
Energy Plasma membrane of the ER Glucose transport from the lumen of the intestine into the blood stream Na+ ATP provides energy for transport of Na+/K+ out/in cell (against concentration gradient) ! Transport of glucose down concentration gradient: no ATP required Na+ Higher Eukaryotes
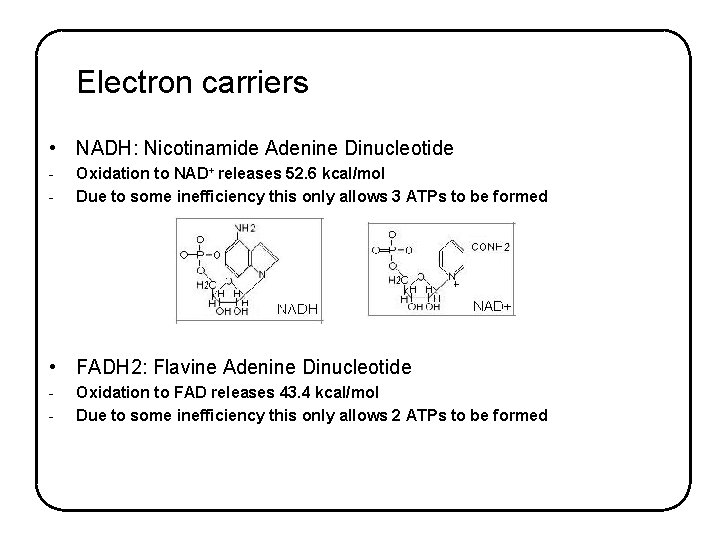
Electron carriers • NADH: Nicotinamide Adenine Dinucleotide - Oxidation to NAD+ releases 52. 6 kcal/mol Due to some inefficiency this only allows 3 ATPs to be formed • FADH 2: Flavine Adenine Dinucleotide - Oxidation to FAD releases 43. 4 kcal/mol Due to some inefficiency this only allows 2 ATPs to be formed
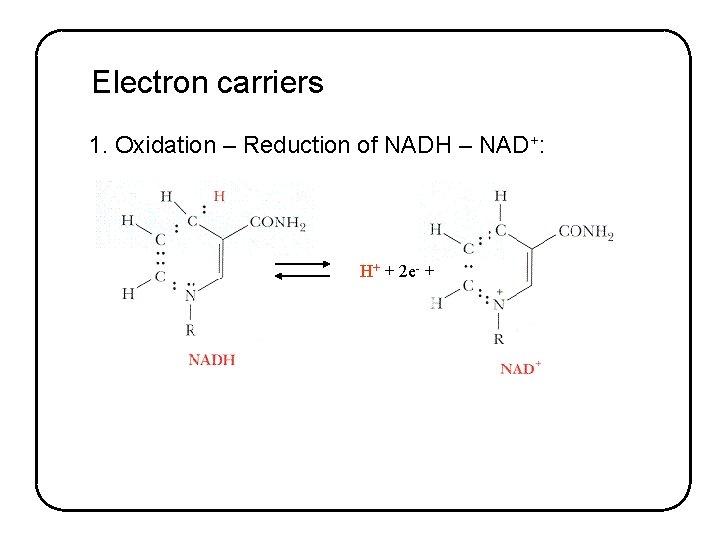
Electron carriers 1. Oxidation – Reduction of NADH – NAD+: H+ + 2 e- +

Biological Interactions Control Interactions: Enhance or repress other interactions + - Main types of control interactions: - transport facilitation - enzymatic catalysis - inhibition - activation of gene expression - repression of gene expression
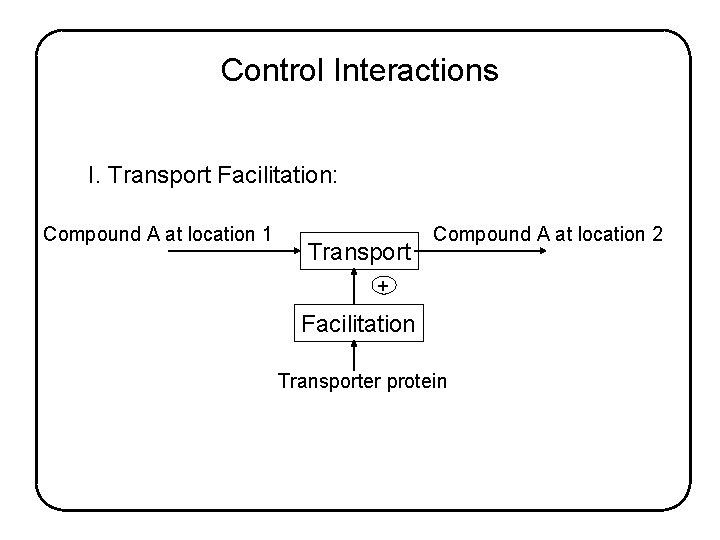
Control Interactions I. Transport Facilitation: Compound A at location 1 Transport Compound A at location 2 + Facilitation Transporter protein
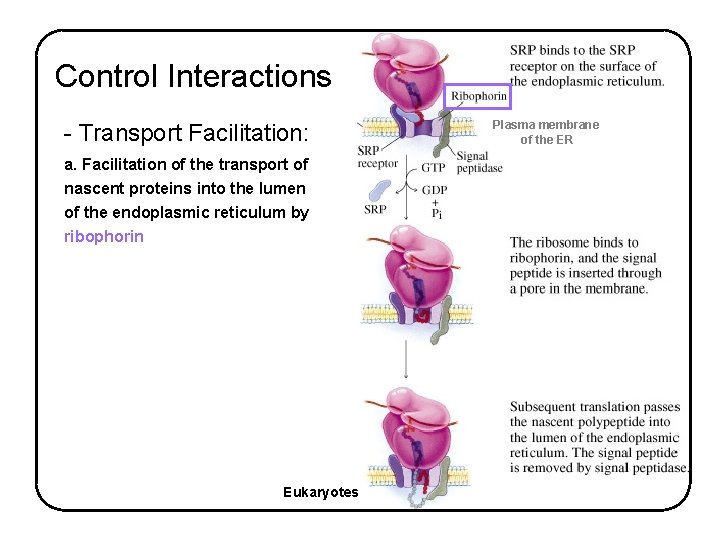
Control Interactions - Transport Facilitation: a. Facilitation of the transport of nascent proteins into the lumen of the endoplasmic reticulum by ribophorin Eukaryotes Plasma membrane of the ER
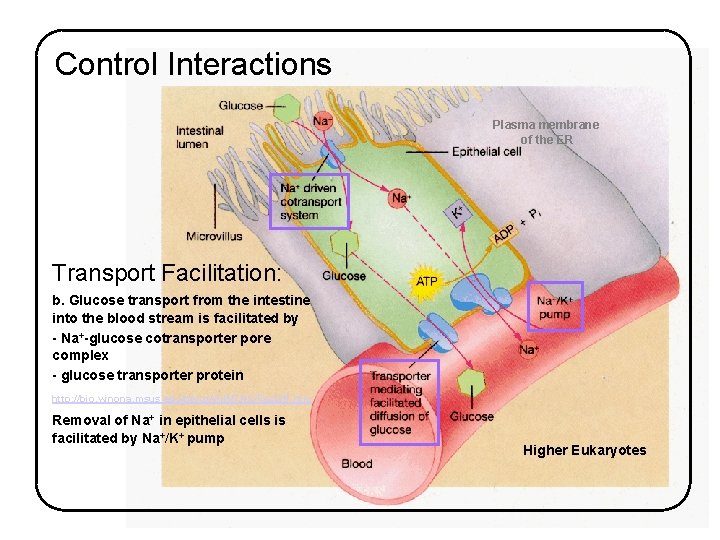
Control Interactions Plasma membrane of the ER Transport Facilitation: b. Glucose transport from the intestine into the blood stream is facilitated by - Na+-glucose cotransporter pore complex - glucose transporter protein http: //bio. winona. msus. edu/berg/ANIMTNS/Fac. Diff. htm Removal of Na+ in epithelial cells is facilitated by Na+/K+ pump Higher Eukaryotes
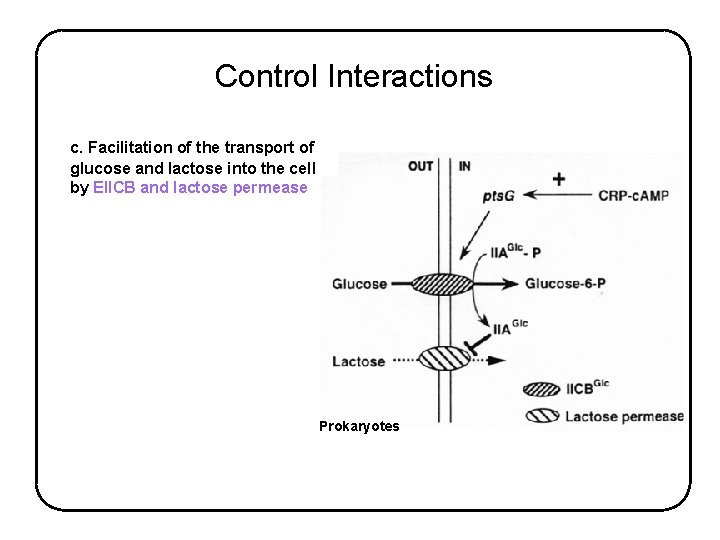
Control Interactions c. Facilitation of the transport of glucose and lactose into the cell by EIICB and lactose permease Prokaryotes
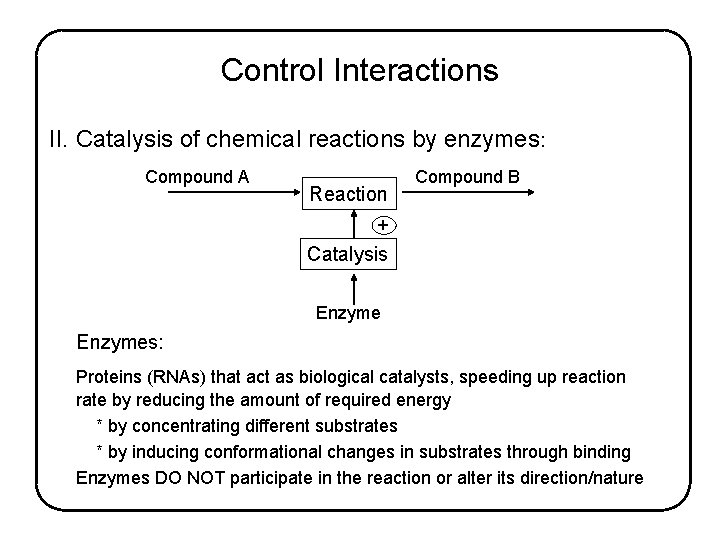
Control Interactions II. Catalysis of chemical reactions by enzymes: Compound A Reaction Compound B + Catalysis Enzymes: Proteins (RNAs) that act as biological catalysts, speeding up reaction rate by reducing the amount of required energy * by concentrating different substrates * by inducing conformational changes in substrates through binding Enzymes DO NOT participate in the reaction or alter its direction/nature

Control Interactions - Catalytic Enzymes : 1. Redox reactions: oxidase, dehydrogenase transfer of e- from electron donors to electron acceptors 2. Phosphorylation - Dephosphorylation: kinase, phosphatase addition - removal of phosphate groups 3. Hydrolysis: hydrolase breakdown of bonds through the addition (- removal) of water 4. Transfer of a side group: transferases 5. Splitting or forming a C-C bond: desmolase, aldolase 6. Changing geometry or structure of a compound: isomerase, gyrase 7. Joining two compounds through hydrolysis of ATP: ligase 8. Polymerisation: polymerase
Control Interactions III. Inhibition: Compound A Interaction Compound B Inhibition Compound Control interactions form a means of using compounds to introduce feedback !
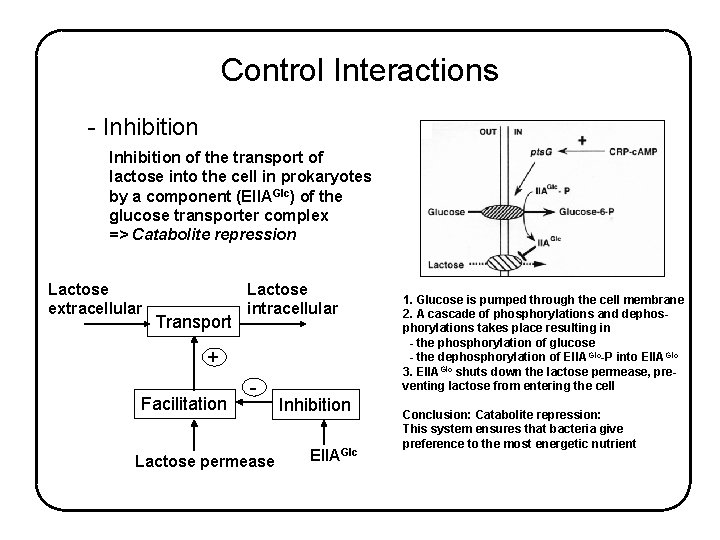
Control Interactions - Inhibition of the transport of lactose into the cell in prokaryotes by a component (EIIAGlc) of the glucose transporter complex => Catabolite repression Lactose extracellular Transport Lactose intracellular + Facilitation - Lactose permease Inhibition EIIAGlc 1. Glucose is pumped through the cell membrane 2. A cascade of phosphorylations and dephosphorylations takes place resulting in - the phosphorylation of glucose - the dephosphorylation of EIIA Glc-P into EIIAGlc 3. EIIAGlc shuts down the lactose permease, preventing lactose from entering the cell Conclusion: Catabolite repression: This system ensures that bacteria give preference to the most energetic nutrient
Control Interactions IV. Activation of gene expression: Gene Expression Protein + Activation Transcription factor V. Repression of gene expression: Gene Expression repression Transcription factor Protein
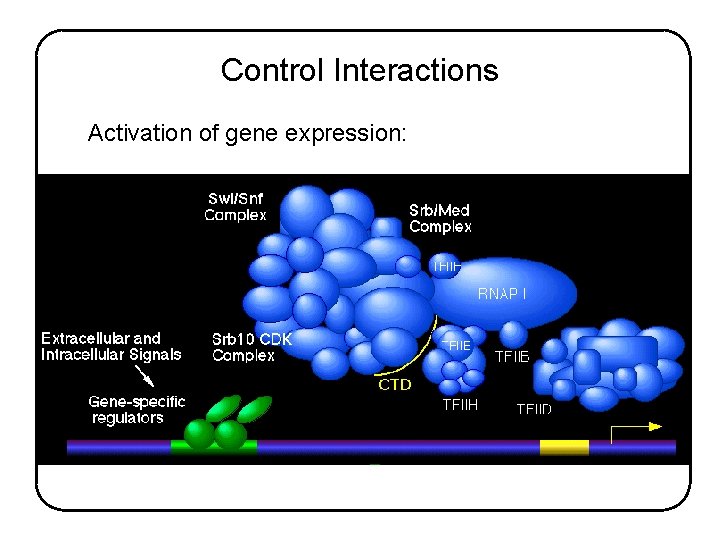
Control Interactions Activation of gene expression:
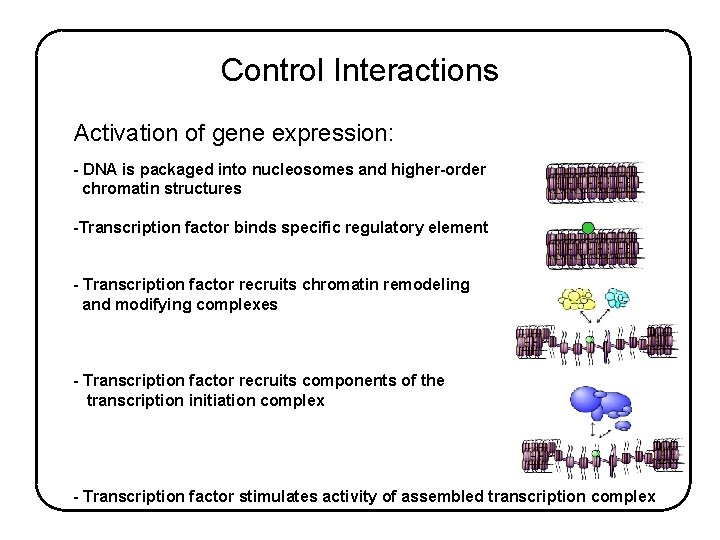
Control Interactions Activation of gene expression: - DNA is packaged into nucleosomes and higher-order chromatin structures -Transcription factor binds specific regulatory element - Transcription factor recruits chromatin remodeling and modifying complexes - Transcription factor recruits components of the transcription initiation complex - Transcription factor stimulates activity of assembled transcription complex

Control Interactions Transcription factor (complexes): • • • Proteins that bind to specific regulatory sequences in the DNA Regulate the level of expression of target gene(s) by controlling whether and how vigorously the gene is transcribed into RNA The on/off switches and rheostats of a (group of) target gene(s) Regulatory DNA sequences • • Every gene has its own cis-acting regulatory sequences Vary greatly in complexity among genes and organisms When active transcription factors associate with the regulatory sequences of their target genes, they can function to repress (down-regulate) or induce (up-regulate) transcription of the corresponding RNA
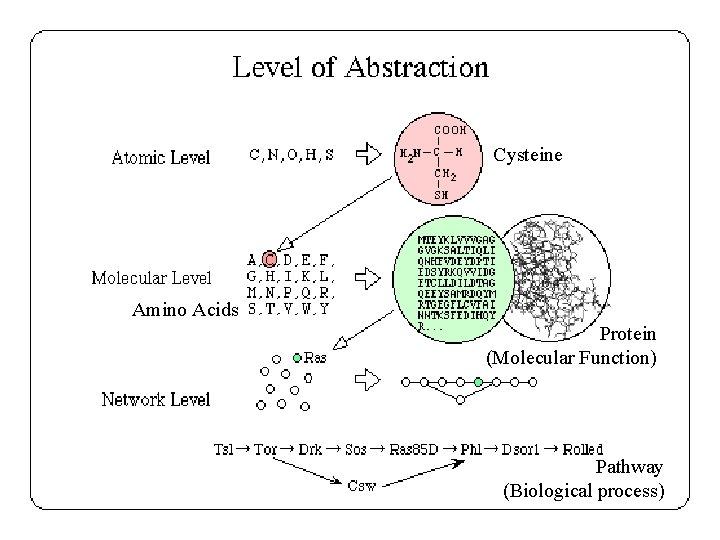
Cysteine Amino Acids Protein (Molecular Function) Pathway (Biological process)
Biological Pathways • • Metabolic pathways Developmental pathways Signal-transduction pathways Genetic regulatory circuits = genetic networks • Pathways interact • Pathways overlap => Biochemical compounds are involved in different pathways

Metabolic Pathways • Metabolism: The sum of all chemical reactions that take place within a cell providing energy for vital processes and for synthesizing new organic material • Eco. Cyc/Hin. Cyc/Meta. Cyc: Encyclopedia of Escherichia coli Genes and Metabolism Encyclopedia of Haemophilus influenzae Genes and Metabolism • EMP: Enzymes and Metabolic Pathways database • KEGG: Kyoto Encyclopedia of Genes and Genomes • …

Metabolic Pathways • Biosynthesis = Anabolism: Sequences of enzyme-catalyzed chemical reactions by which complex molecules are formed in living cells from building blocks with simple structures • Degradation = Catabolism: Sequences of enzyme-catalyzed chemical reactions by which large molecules in living cells are broken down or degraded into building blocks • Transport: Sequences of transport (facilitation) interactions by which compounds are transported from one location to another. • Energy Metabolism: Sequences of enzyme-catalyzed chemical reactions by which chemical energy obtained from the environment by degradation of nutrients or by capturing solar energy (plants) is transformed into energy-rich compounds that are required for metabolic processes

Metabolic Pathways • Metabolism of: – – – – – Amino acids, peptides, proteins and derivatives Carbohydrates (sugars) Cell-structure components Cofactors, prosthetic groups and electron carriers Fatty acids and lipids Nucleotides and nucleic acids Monocarbon compounds Essential elements Aromatic compounds
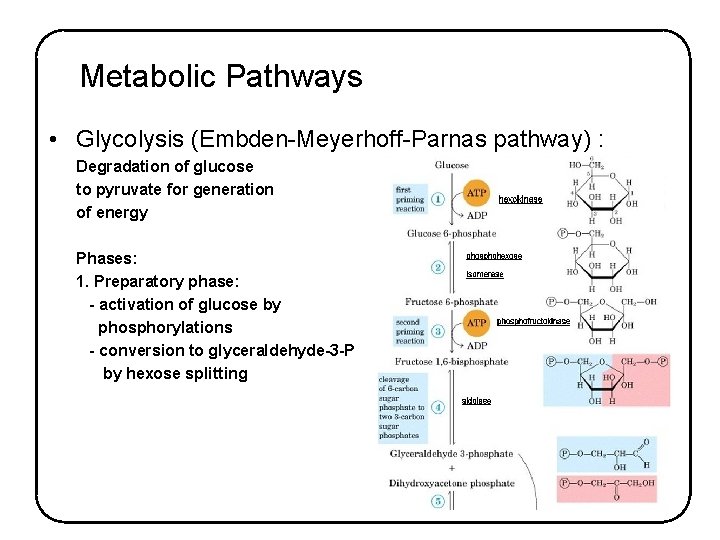
Metabolic Pathways • Glycolysis (Embden-Meyerhoff-Parnas pathway) : Degradation of glucose to pyruvate for generation of energy Phases: 1. Preparatory phase: - activation of glucose by phosphorylations - conversion to glyceraldehyde-3 -P by hexose splitting
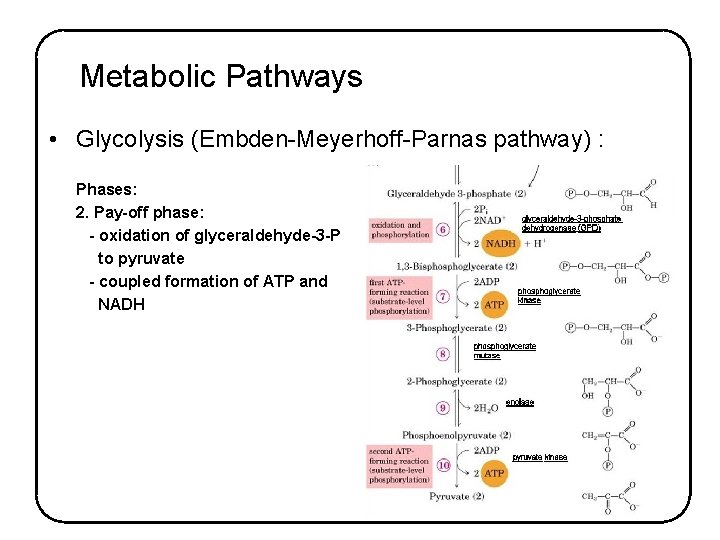
Metabolic Pathways • Glycolysis (Embden-Meyerhoff-Parnas pathway) : Phases: 2. Pay-off phase: - oxidation of glyceraldehyde-3 -P to pyruvate - coupled formation of ATP and NADH
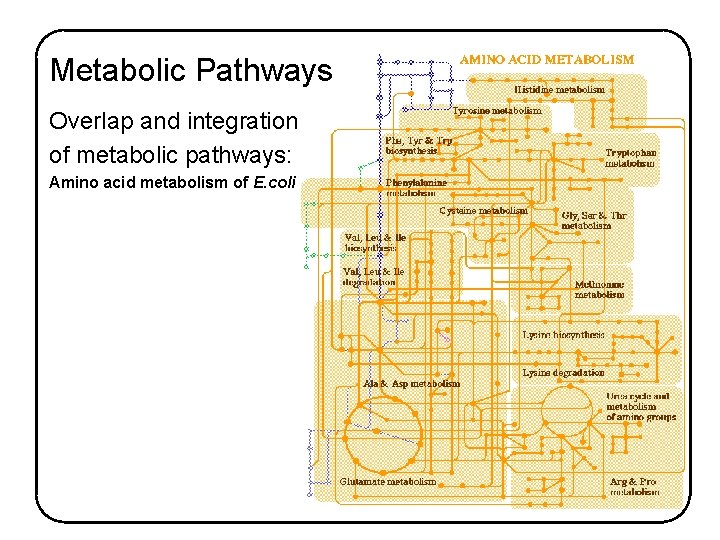
Metabolic Pathways Overlap and integration of metabolic pathways: Amino acid metabolism of E. coli
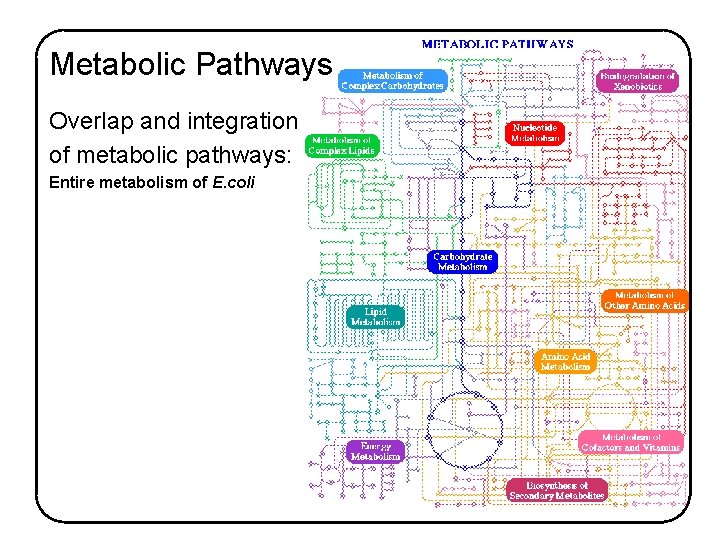
Metabolic Pathways Overlap and integration of metabolic pathways: Entire metabolism of E. coli
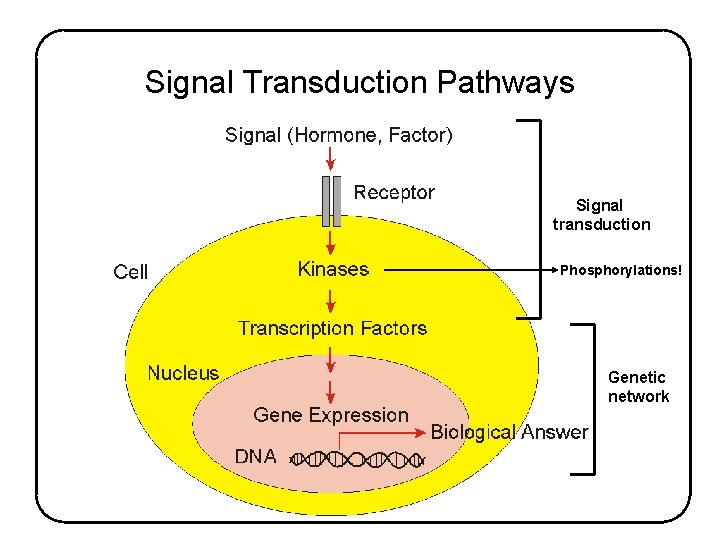
Signal Transduction Pathways Signal transduction Phosphorylations! Genetic network
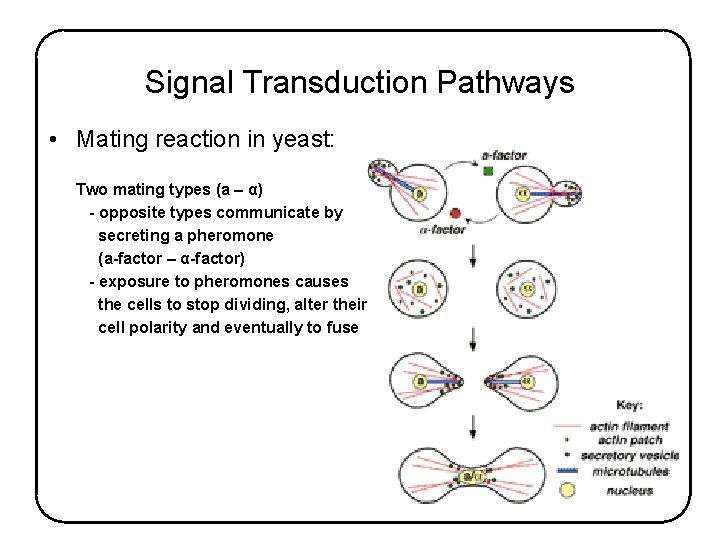
Signal Transduction Pathways • Mating reaction in yeast: Two mating types (a – α) - opposite types communicate by secreting a pheromone (a-factor – α-factor) - exposure to pheromones causes the cells to stop dividing, alter their cell polarity and eventually to fuse
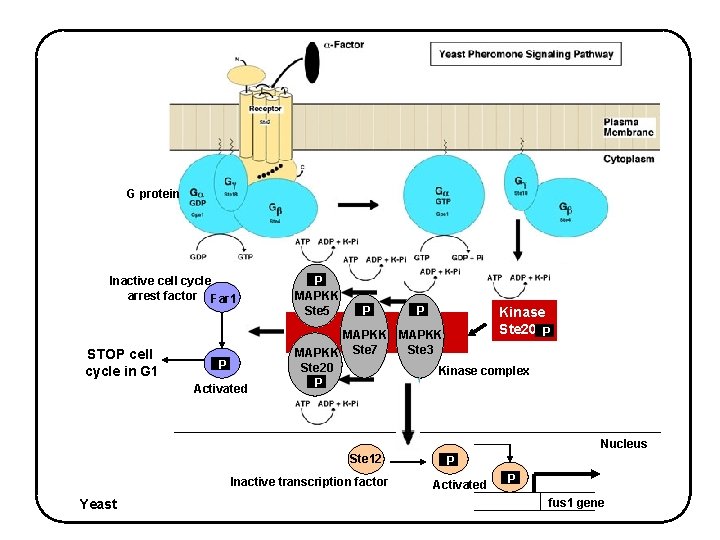
G protein Inactive cell cycle arrest factor Far 1 STOP cell cycle in G 1 P Activated P MAPKK Ste 5 P P Kinase Ste 20 P MAPKK Ste 3 MAPKK Ste 7 Ste 20 Kinase complex P Nucleus Ste 12 Inactive transcription factor Yeast P Activated P fus 1 gene
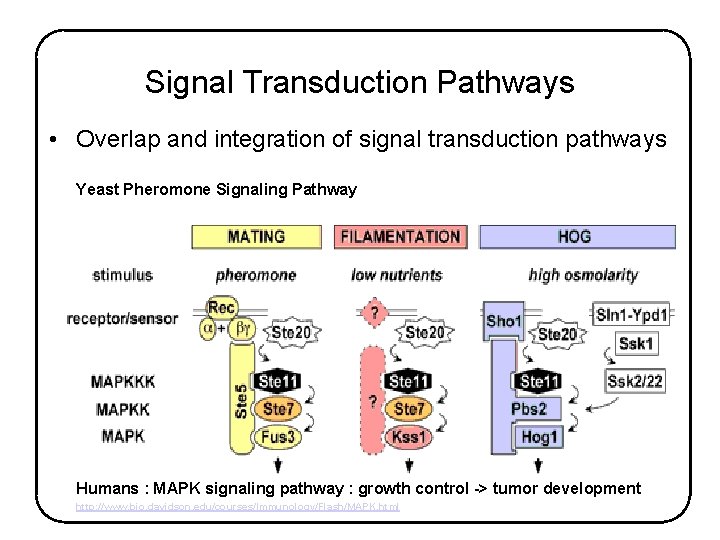
Signal Transduction Pathways • Overlap and integration of signal transduction pathways Yeast Pheromone Signaling Pathway Humans : MAPK signaling pathway : growth control -> tumor development http: //www. bio. davidson. edu/courses/Immunology/Flash/MAPK. html
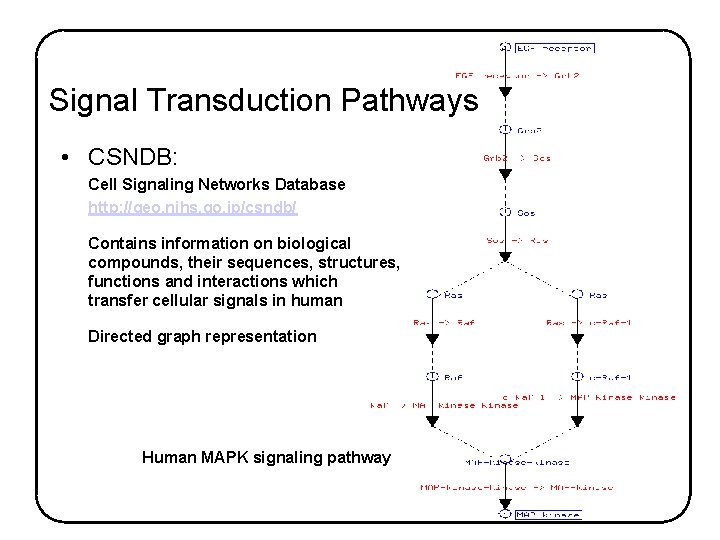
Signal Transduction Pathways • CSNDB: Cell Signaling Networks Database http: //geo. nihs. go. jp/csndb/ Contains information on biological compounds, their sequences, structures, functions and interactions which transfer cellular signals in human Directed graph representation Human MAPK signaling pathway
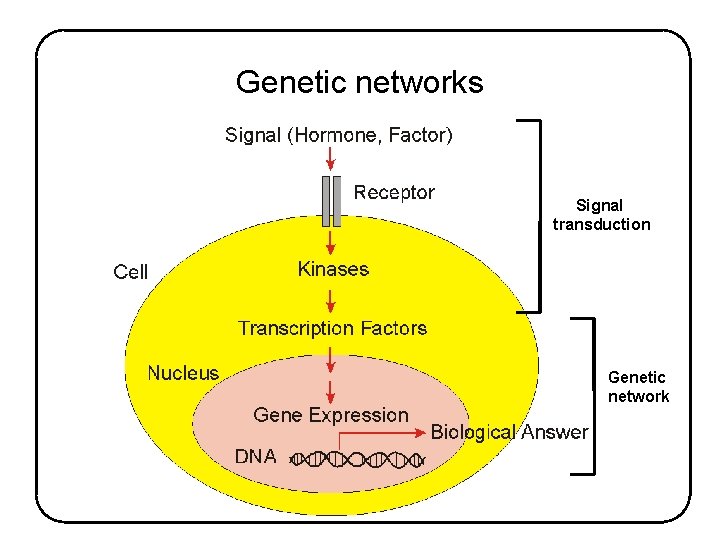
Genetic networks Signal transduction Genetic network

Genetic networks • • Biochemical computers controlling the on/off switches and rheostats of a cell at the gene level Dynamically orchestrate the expression level for each gene in the genome by controlling whether and how vigorously that gene is transcribed Essential interacting components: - Activated transcription factor complex - Regulatory DNA sequences Output : RNA and proteins Some of these proteins are the actuators of inhibition and repression => main feedback loops Co-regulated target genes often code for proteins that act together to build a specific cell structure or to effect a concerted change in cell function Often multiple waves of regulation with first wave products regulating expression of another group of genes and so on
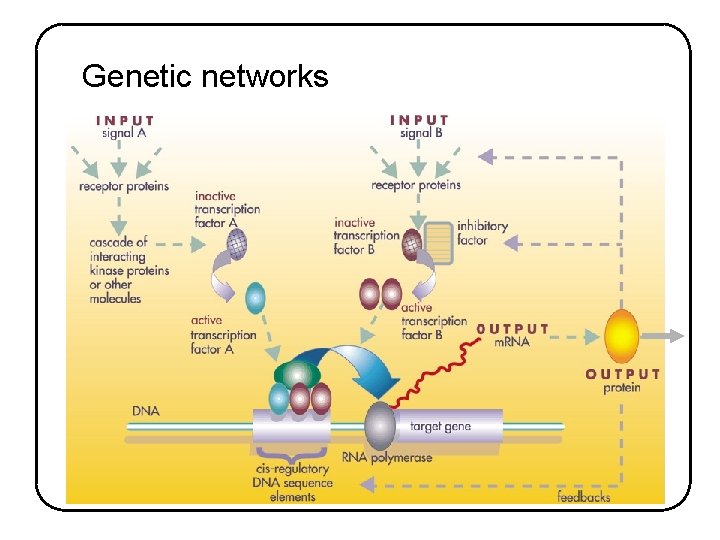
Genetic networks
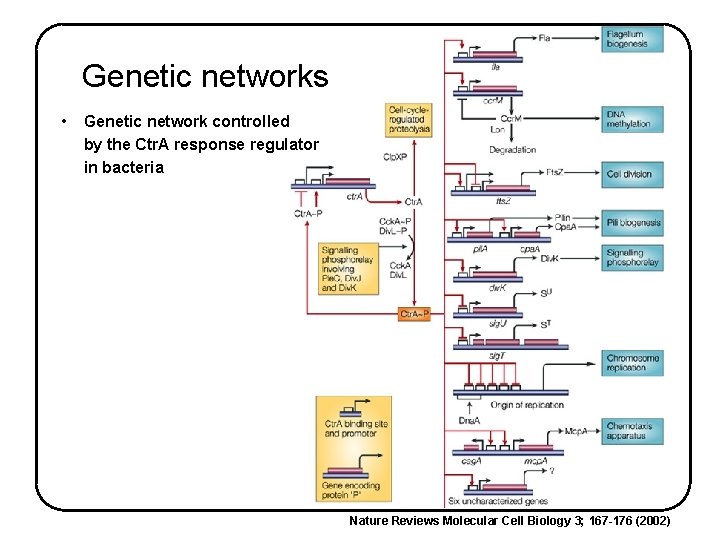
Genetic networks • Genetic network controlled by the Ctr. A response regulator in bacteria Nature Reviews Molecular Cell Biology 3; 167 -176 (2002)
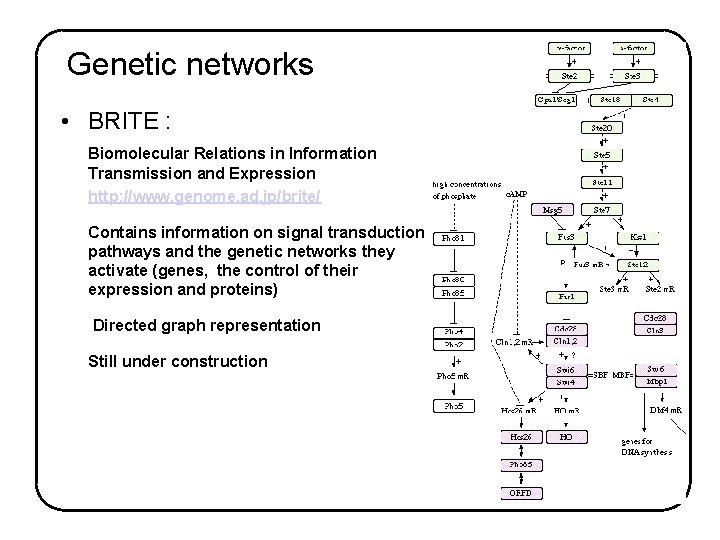
Genetic networks • BRITE : Biomolecular Relations in Information Transmission and Expression http: //www. genome. ad. jp/brite/ Contains information on signal transduction pathways and the genetic networks they activate (genes, the control of their expression and proteins) Directed graph representation Still under construction
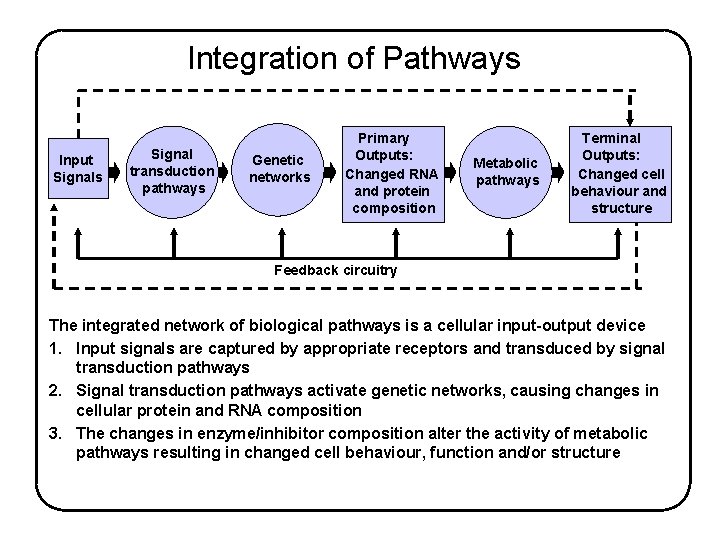
Integration of Pathways Input Signals Signal transduction pathways Genetic networks Primary Outputs: Changed RNA and protein composition Metabolic pathways Terminal Outputs: Changed cell behaviour and structure Feedback circuitry The integrated network of biological pathways is a cellular input-output device 1. Input signals are captured by appropriate receptors and transduced by signal transduction pathways 2. Signal transduction pathways activate genetic networks, causing changes in cellular protein and RNA composition 3. The changes in enzyme/inhibitor composition alter the activity of metabolic pathways resulting in changed cell behaviour, function and/or structure

Biological Pathways • Inference Issues: - Complete network of integrated biological pathways is blueprint of life - Genome is only an archiving system of building blocks - Regulatory DNA motifs are bar codes to retrieve the building blocks • Combination of high-throughput experiments, prior knowledge and bayesian network inference is necessary Experiments: - Microarrays - Proteome analysis - Yeast two-hybrid - Phosphorylations Expression is result of underlying network
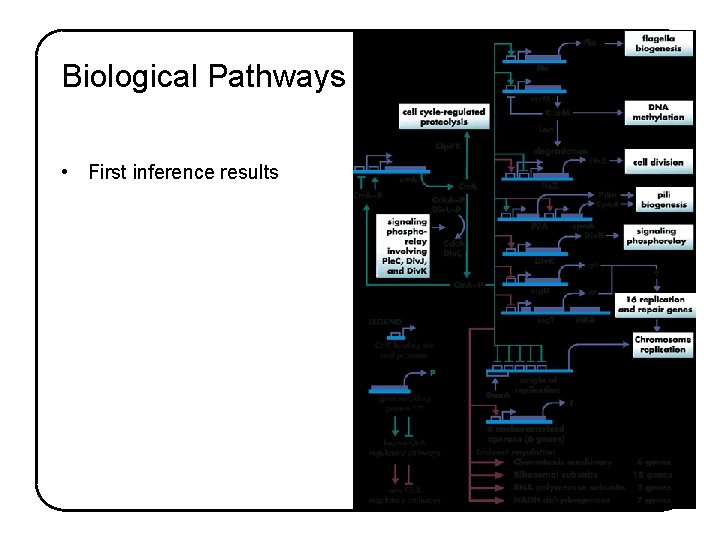
Biological Pathways • First inference results
- Slides: 67