BIOLOGICAL NETWORKS Woochang Hwang BIOLOGICAL NETWORKS Introduction p

BIOLOGICAL NETWORKS Woochang Hwang

BIOLOGICAL NETWORKS Introduction p Biological Networks p n n n Protein-Protein Interaction Networks Signaling & Metabolic Pathway Networks Expression Networks Biological Networks’ Properties p Databases p Discussion p STM Clustering Model p

Introduction

Bioinformatics p Informatics Its carrier is a set of digital codes and a language. In its manifestation in the space-time continuum, it has utility (e. g. to decrease entropy of an open system). p Bioinformatics The essence of life is information (i. e. from digital code to emerging properties of biosystems. ) Bioinformatics is the study of information content of life
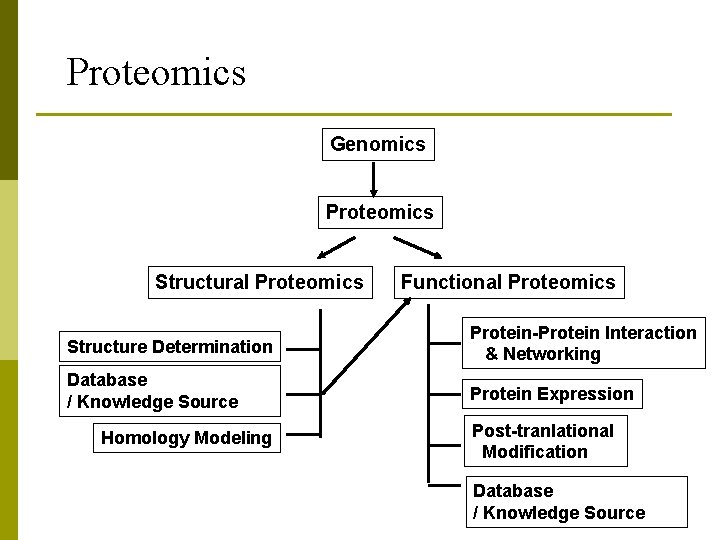
Proteomics Genomics Proteomics Structural Proteomics Functional Proteomics Structure Determination Protein-Protein Interaction & Networking Database / Knowledge Source Protein Expression Homology Modeling Post-tranlational Modification Database / Knowledge Source
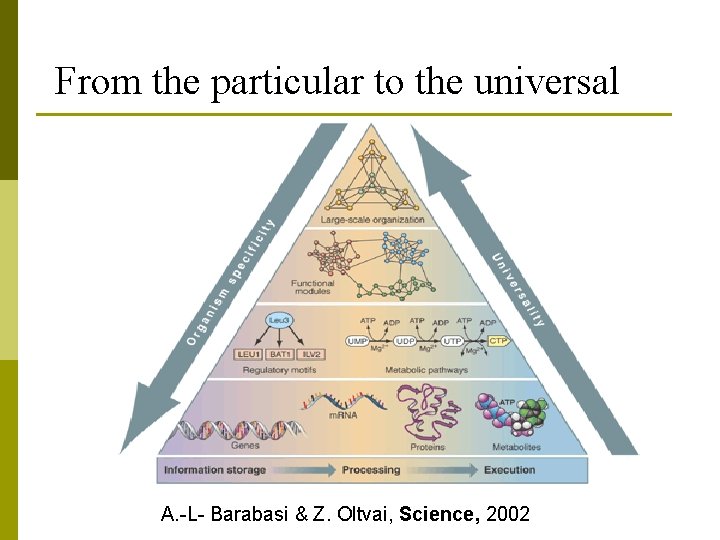
From the particular to the universal A. -L- Barabasi & Z. Oltvai, Science, 2002
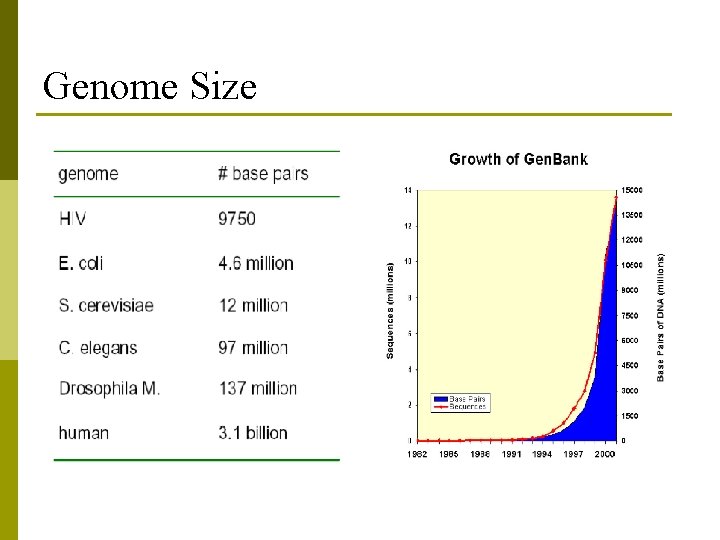
Genome Size
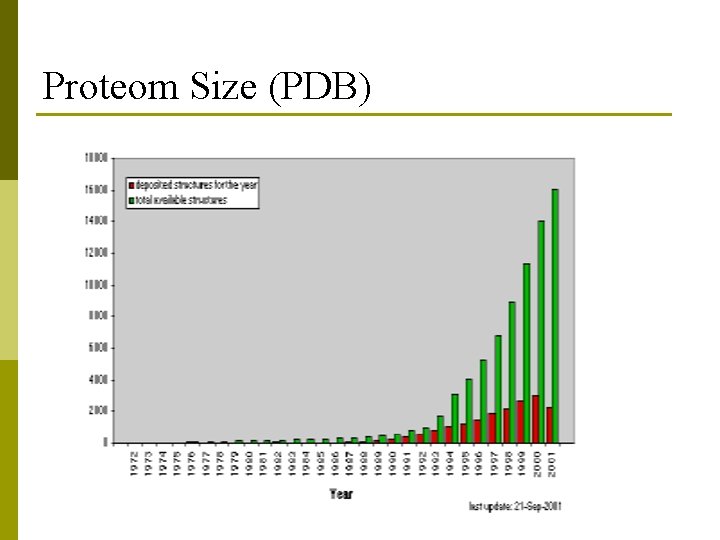
Proteom Size (PDB)

BIOLOGICAL NETWORK Networks are found in biological systems of varying scales: 1. Evolutionary tree of life 2. Ecological networks 3. Expression networks 4. Regulatory networks - genetic control networks of organisms 5. The protein interaction network in cells 6. The metabolic network in cells … more biological networks

Why Study Networks? It is increasingly recognized that complex systems cannot be described in a reductionist view. p Understanding the behavior of such systems starts with understanding the topology of the corresponding network. p Topological information is fundamental in constructing realistic models for the function of the network. p

Biological Network Model p Network n n A linked list of interconnected nodes. Node p § Protein, peptide, or non-protein biomolecules. Edges § Biological relationships, etc. , interactions, regulations, reactions, transformations, activation, inhibitions.
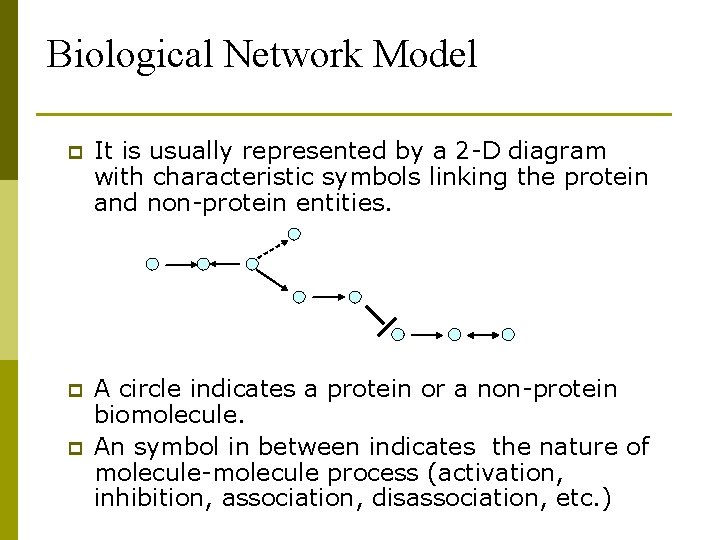
Biological Network Model p It is usually represented by a 2 -D diagram with characteristic symbols linking the protein and non-protein entities. p A circle indicates a protein or a non-protein biomolecule. An symbol in between indicates the nature of molecule-molecule process (activation, inhibition, association, disassociation, etc. ) p

Protein Interaction Network

Proteins in a cell p There are thousands of different active proteins in a cell acting as: n n n enzymes, catalysors to chemical reactions of the metabolism components of cellular machinery (e. g. ribosomes) regulators of gene expression Certain proteins play specific roles in special cellular compartments. Others move from one compartment to another as “signals”.

Protein Interactions Proteins perform a function as a complex rather as a single protein. p Knowing whether two proteins interact can help us discover unknown proteins’ functions: n If the function of one protein is known, the function of its binding partners are likely to be related- “guilt by association”. n Thus, having a good method for detecting interactions can allow us to use a small number of proteins with known function to characterize new proteins. p
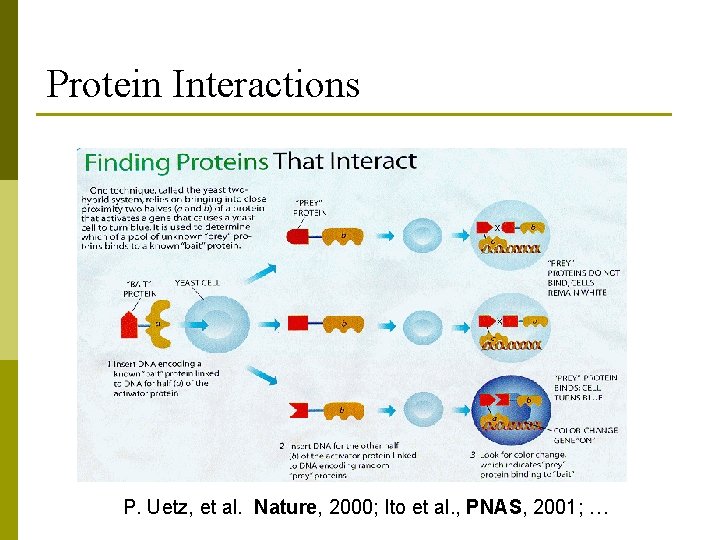
Protein Interactions P. Uetz, et al. Nature, 2000; Ito et al. , PNAS, 2001; …
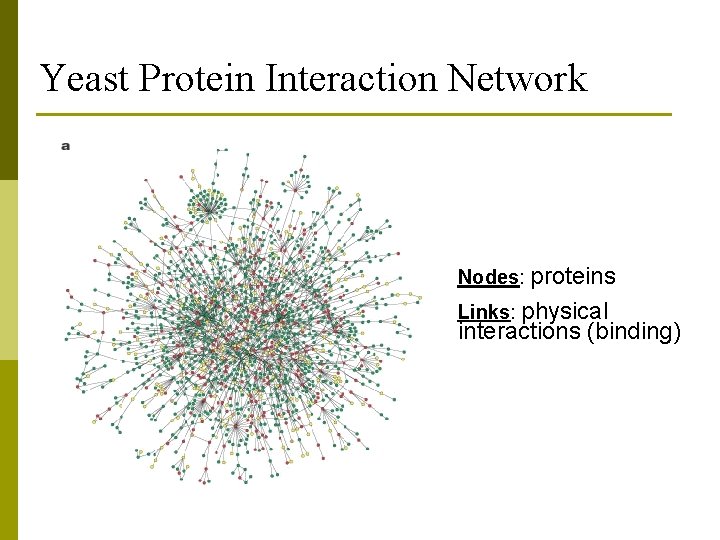
Yeast Protein Interaction Network Nodes: proteins Links: physical interactions (binding)

Pathway Networks

Signaling & Metabolic Pathway Network p p A Pathway can be defined as a modular unit of interacting molecules to fulfill a cellular function. Signaling Pathway Networks n n n p In biology a signal or biopotential is an electric quantity (voltage or current or field strength), caused by chemical reactions of charged ions. refer to any process by which a cell converts one kind of signal or stimulus into another. Another use of the term lies in describing the transfer of information between and within cells, as in signal transduction. Metabolic Pathway Networks n a series of chemical reactions occurring within a cell, catalyzed by enzymes, resulting in either the formation of a metabolic product to be used or stored by the cell, or the initiation of another metabolic pathway
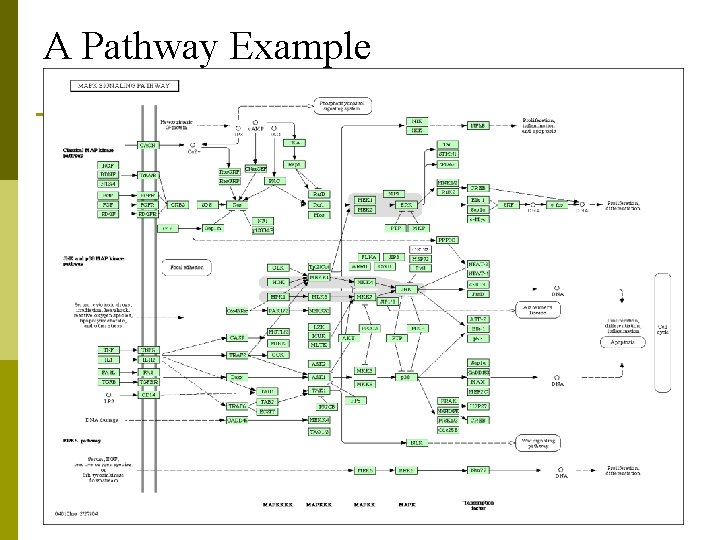
A Pathway Example
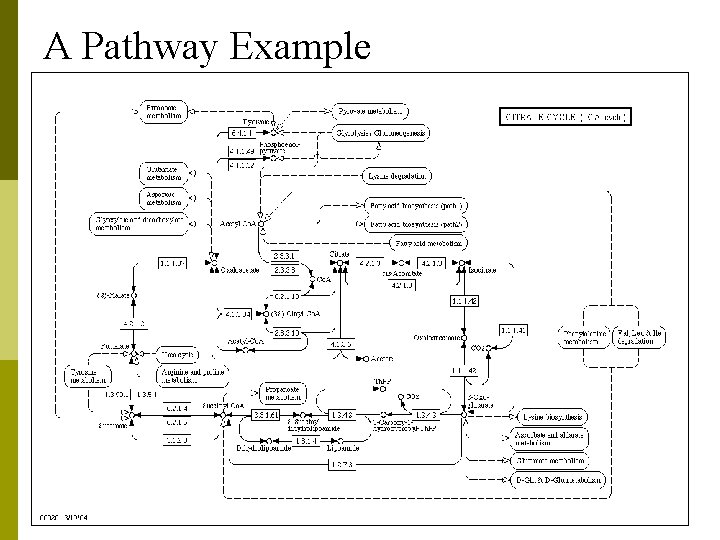
A Pathway Example
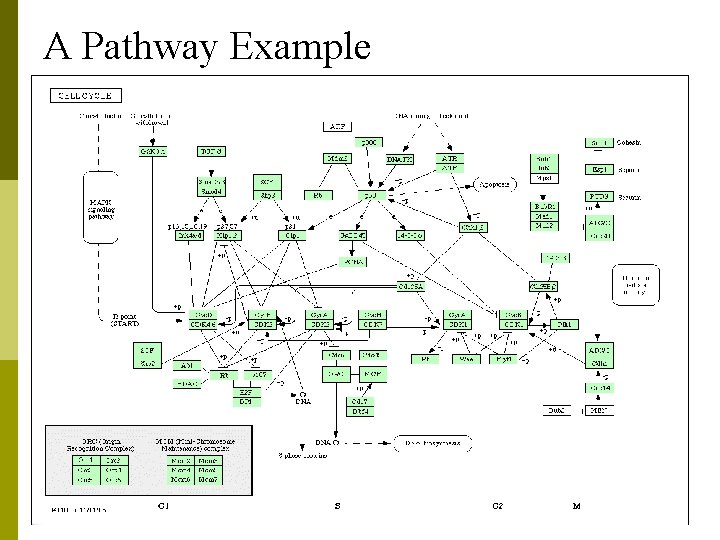
A Pathway Example

Regulatory Network p a collection of DNA segments (genes) in a cell which interact with each other and with other substances in the cell, thereby governing the rates at which genes in the network are transcribed into m. RNA.
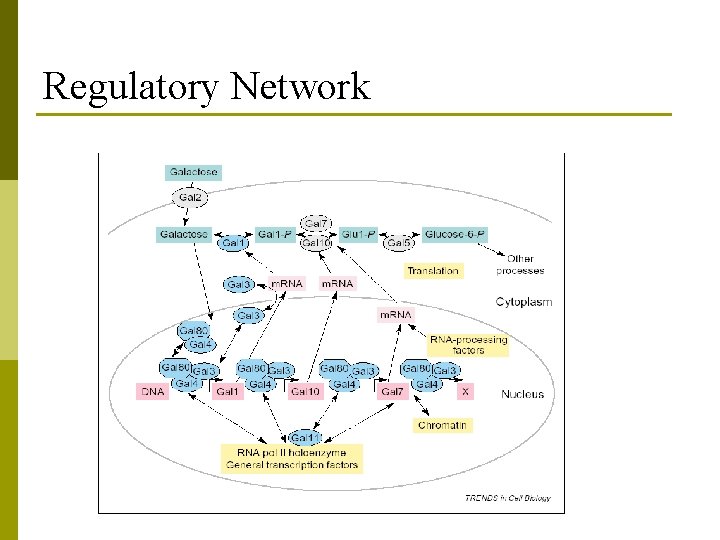
Regulatory Network
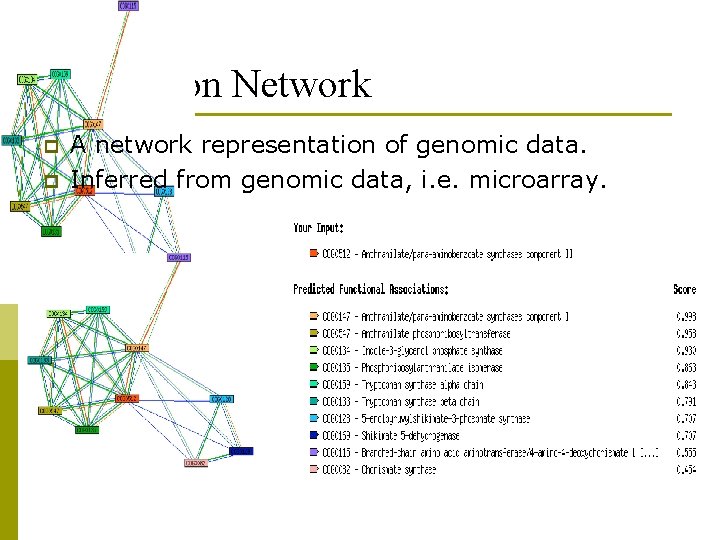
Expression Network p p A network representation of genomic data. Inferred from genomic data, i. e. microarray.

BIOLOGICAL NETWORK PROPERTY Interaction Network p Pathway Network p Regulatory Network p Expression Network p

Biological Networks Properties p p Power law degree distribution: Rich get richer Small World: A small average path length n Mean shortest node-to-node path Robustness: Resilient and have strong resistance to failure on random attacks and vulnerable to targeted attacks Hierarchical Modularity: A large clustering coefficient n How many of a node’s neighbors are connected to each other
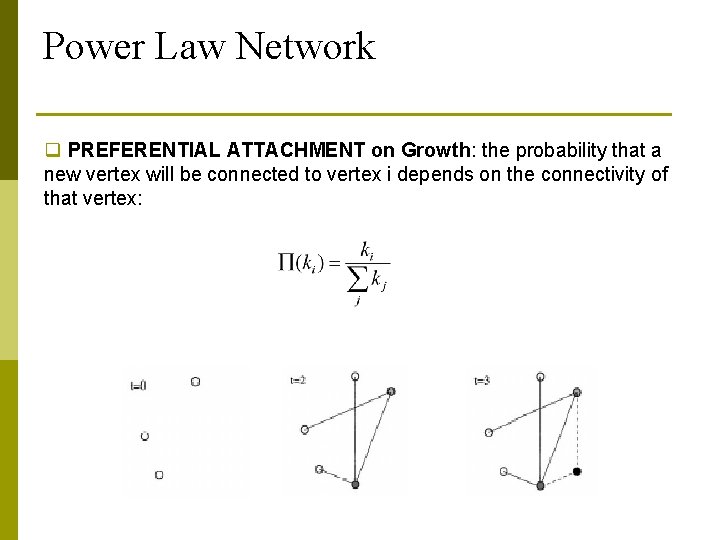
Power Law Network q PREFERENTIAL ATTACHMENT on Growth: the probability that a new vertex will be connected to vertex i depends on the connectivity of that vertex:
![The Barabási-Albert [BA] model ER Model WS Model (a) Random Networks Actors Power Grid The Barabási-Albert [BA] model ER Model WS Model (a) Random Networks Actors Power Grid](http://slidetodoc.com/presentation_image_h2/3b440fb7b2a434976936f9aae85d2fe2/image-29.jpg)
The Barabási-Albert [BA] model ER Model WS Model (a) Random Networks Actors Power Grid (b) Power law Networks Power Law Network (Scale Free) q The probability of finding a highly connected node decreases exponentially with k: www

Small World Property p p A small average path length Any node can be reached within a small number of edges, 4~5 hops.

Power Law Network p Power-law degree distribution & Small world phenomena also observed in: n n p communication networks web graphs research citation networks social networks Classical -Erdos-Renyi type random graphs do not exhibit these properties: n n n Links between pairs of fixed set of nodes picked uniformly: Maximum degree logarithmic with network size No hubs to make short connections between nodes
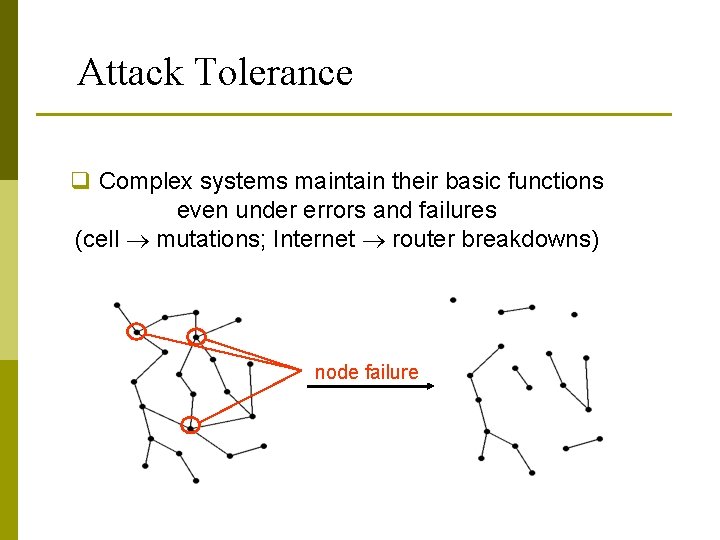
Attack Tolerance q Complex systems maintain their basic functions even under errors and failures (cell mutations; Internet router breakdowns) node failure
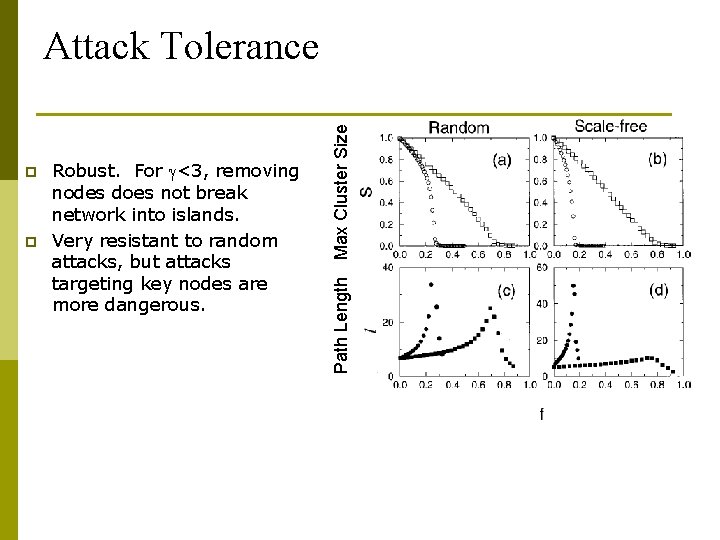
p p Robust. For <3, removing nodes does not break network into islands. Very resistant to random attacks, but attacks targeting key nodes are more dangerous. Path Length Max Cluster Size Attack Tolerance
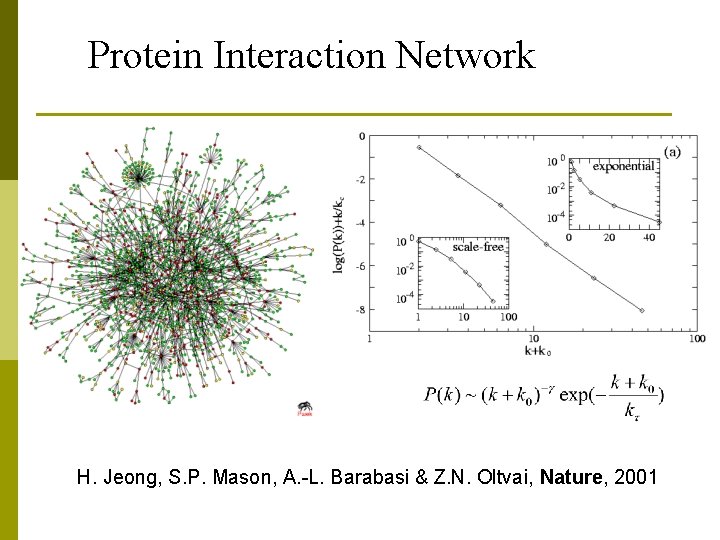
Protein Interaction Network H. Jeong, S. P. Mason, A. -L. Barabasi & Z. N. Oltvai, Nature, 2001

Protein Interaction Network p The yeast protein interaction network seems to reveal some basic graph theoretic properties: n n n The frequency of proteins having interactions with exactly k other proteins follows a power law. The network exhibits the small world phenomena: can reach any node within small number of hops, usually 4 or 5 hops Robustness: Resilient and have strong resistance to failure on random attacks and vulnerable to targeted attacks.
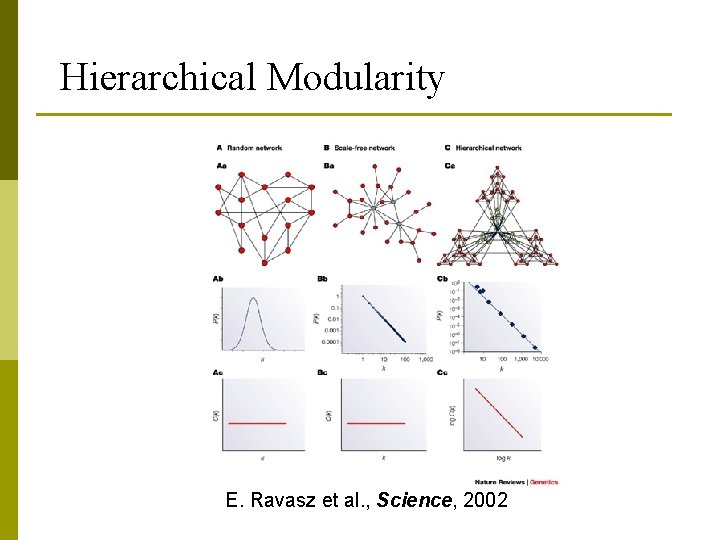
Hierarchical Modularity E. Ravasz et al. , Science, 2002
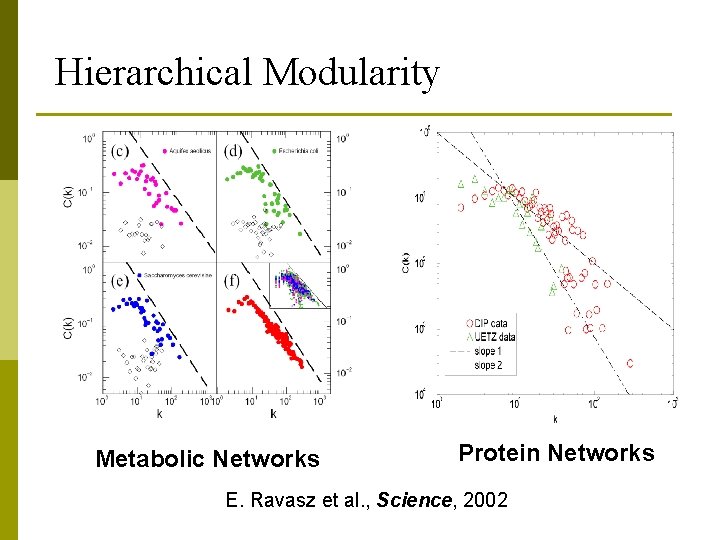
Hierarchical Modularity Metabolic Networks Protein Networks E. Ravasz et al. , Science, 2002

Implications From Observations p p p Biological complexity: # states ~2# of genes. Protein hubs critical for cells, 45%. Infections will target highly connected nodes. Cascading node failures could cause a critical problem. Development of drug and treatment with novel strategies like targeting effective nodes is indispensable.

Databases

Protein Databases q. Swiss-Prot (non-redundant database): Release 41. 0, 11/4/2003: 124, 464 entries. Release 41. 5, 23/4/2002: 125, 236 entries. q. Tr. EMBL (translations of EMBL nucleotide sequences not yet integrated into Swiss-Prot): Release 23. 7, 17/4/2003: 863, 248 entries q. This number keeps rapidly growing mainly due to large scale sequencing projects.

Protein Interaction Databases q Species-specific n n n Fly. Nets - Gene networks in the fruit fly MIPS - Yeast Genome Database Regulon. DB - A Data. Base On Transcriptional Regulation in E. Coli Soy. Base PIMdb - Drosophila Protein Interaction Map database q Function-specific n n n n Biocatalysis/Biodegradation Database BRITE - Biomolecular Relations in Information Transmission and Expression COPE - Cytokines Online Pathfinder Encyclopaedia Dynamic Signaling Maps EMP - The Enzymology Database FIMM - A Database of Functional Molecular Immunology CSNDB - Cell Signaling Networks Database

Protein Interaction Databases q Interaction type-specific n DIP - Database of Interacting Proteins n DPInteract - DNA-protein interactions n Inter-Chain Beta-Sheets (ICBS) - A database of protein interactions mediated by interchain beta-sheet formation n Interact - A Protein-Protein Interaction database n Gene. Net (Gene networks) q General n BIND - Biomolecular Interaction Network Database n Binding. DB - The Binding Database n MINT - a database of Molecular INTeractions n PATIKA - Pathway Analysis Tool for Integration and Knowledge Acquisition n PFBP - Protein Function and Biochemical Pathways Project n PIM (Protein Interaction Map)

Pathway Databases q KEGG (Kyoto Encyclopedia of Genes and Genomes) q q q http: //www. genome. ad. jp/kegg/ q Institute for Chemical Research, Kyoto University Path. DB q http: //www. ncgr. org/pathdb/index. html q National Center for Genomic Resources SPAD: Signaling PAthway Database q Graduate School of Genetic Resources Technology. Kyushu University. Cytokine Signaling Pathway DB. q Dept. of Biochemistry. Kumamoto Univ. Eco. Cyc and Meta. Cyc q Stanford Research Institute BIND (Biomolecular Interaction Network Database) q UBC, Univ. of Toronto

KEGG p Pathway Database: Computerize current knowledge of molecular and cellular biology in terms of the pathway of interacting molecules or genes. p Genes Database: Maintain gene catalogs of all sequenced organisms and link each gene product to a pathway component p Ligand Database: Organize a database of all chemical compounds in living cells and link each compound to a pathway component p Pathway Tools: Develop new bioinformatics technologies for functional genomics, such as pathway comparison, pathway reconstruction, and pathway design
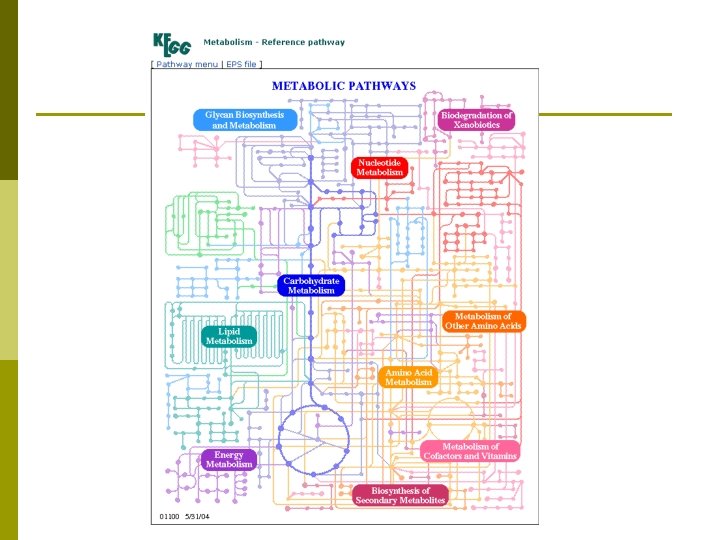
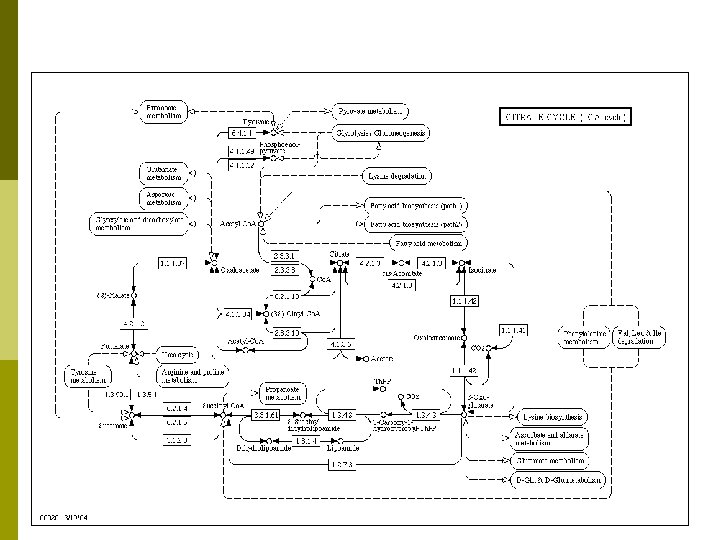

Discussion § Problems § Network Inference § Micro Array, Protein Chips, other high throughput assay methods § Function prediction § The function of 40 -50% of the new proteins is unknown § Understanding biological function is important for: § § § § Study of fundamental biological processes § Drug design § Genetic engineering Functional module detection § Cluster analysis Topological Analysis § Descriptive and Structural § Locality Analysis § Essential Component Analysis Dynamics Analysis § Signal Flow Analysis § Metabolic Flux Analysis § Steady State, Response, Fluctuation Analysis Evolution Analysis Biological Networks are very rich networks with very limited, noisy, and incomplete information. Discovering underlying principles is very challenging.

Signal Transduction Model Based Functional Module Detection Algorithm for Protein-Protein Interaction Networks Woochang Hwang 1 Young-Rae Cho 1 Aidong Zhang 1 Murali Ramanathan 2 1 Department of Computer Science and Engineering, State University of New York at Buffalo 2 Department of Pharmaceutical Sciences, State University of New York at Buffalo University at Buffalo The State University of New York

Contents p p Introduction Protein Interaction Networks Functional Categories Functional Module Detection Algorithm n p p p Signal Transduction Model (STM) Experimental Results Discussion Future Works

Introduction p Cellular Functions are coordinately carried out by groups of genes and gene products. p Detection of such functional modules in a complex molecular network is one of the most challenging problem. p Molecular networks: high data volume, high noise level, sparse connectivity, etc. p PPI data n S. Cerevisae full PPI data in DIP: over 4900 proteins and 18000 interactions. n PPI data provide us the good opportunity to analyze the underlying principles and the structure of large living systems.

Cluster Assessment p Clustering Coefficient: n p Betweeness Centrality: n p is the number of shortest paths from node s to t and (v) the number of shortest paths from s to t that pass through the node v. P-value: n n p N(v) is the set of the direct neighbors of node v and d(v) is the number of the direct neighbors of node v C is the size of the cluster containing k proteins with a given function; G is the size of the universal set of proteins of known proteins and contains n proteins with the function. The p-value is the probability that a cluster would be enriched with proteins with a particular function by chance alone. Density: n n is the number of proteins and e is the number of interactions in a sub graph s of a PPI network.
Protein-Protein Interaction (PPI) Data & MIPS Functional Category Data p DIP Yeast Protein Interaction core data n 2521 proteins, 5949 interactions n Average clustering coefficient: 0. 069 n Average path length: 5. 47 p MIPS Functional Category n 457 Hierarchical Functional Categories n Sub graphs of each functional categories are extracted from DIP core data. p p Average graph density: 0. 0025 Average diameter (longest path in a graph): 4. 23
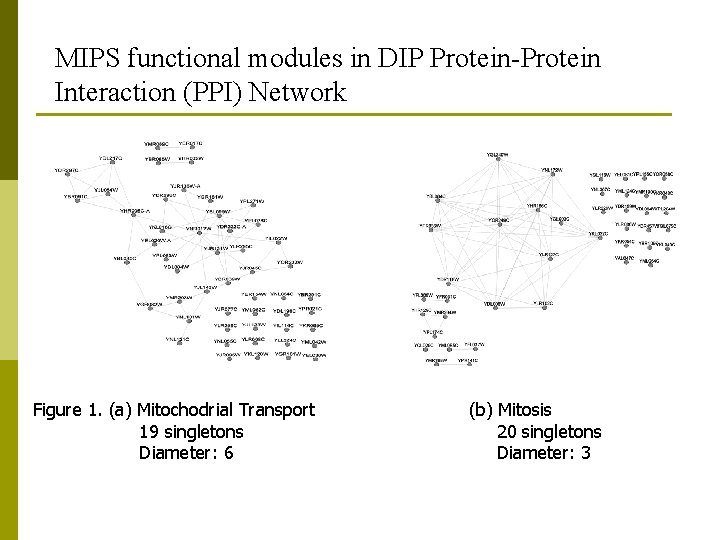
MIPS functional modules in DIP Protein-Protein Interaction (PPI) Network Figure 1. (a) Mitochodrial Transport 19 singletons Diameter: 6 (b) Mitosis 20 singletons Diameter: 3

Topological Properties of MIPS Functional Modules in DIP Protein Interaction Data p Sparse connectivity : low density, isolated sub graphs and singletons existence. p Longish shape: high diameter

Related works p Distance Based Approaches n Several distance metrics were introduced n Use traditional clustering algorithms p Graph Based Approaches n Density based approaches: Maximal Cliques, Quasi Cliques, RNSC, HCS, MCODE n Statistical approaches: MCL, Samantha

Related works p Suffered by their limited way of clustering. n identify only the clusters with specific shapes, e. g. , balanced round shapes, with high density. n But, the actual functional modules are not so densely connected as they expected. n Some members in functional categories do not have direct physical interaction with other members of the functional category they belong to. n Modules that have longish shapes are frequently observed. n The incompleteness of clustering is another distinct drawback of existing algorithms, which produce many clusters with small size and singletons.

Contribution Unexpected properties of functional categories and sparse connectivity in PPI networks. A relative excess of emphasis on density in the existing methods can be preferential for detecting clusters with relatively balanced round shapes, high discarding rate, and limit performance. § § STM Clustering Model § § Effective clustering should be able to detect clusters with arbitrary shape and density if the cluster members share biological and topological similarities. To take those unexpected properties of PPI networks and actual functional modules into consideration and to conquer the drawbacks of existing approaches effectively: § § STM clustering model utilizes a statistical signal transduction model to find the modules whose members share biological common feature even though they are sparsely connected. STM model also adopts the network’s topological properties into the model.

STM Clustering Model Process 1: Simulation of dynamic statistical signal transduction behavior in the network. § STM model simulates dynamic signal transduction behavior to find the most influential proteins on each protein in PPI network biologically and topologically. Process 2: Selection of the putative cluster representatives on each node. Process 3: Preliminary clusters formation. § Preliminary clusters will be formed by accumulating each node toward its chosen representatives. Process 4: Cluster merge. § § § So far, STM has considered only the biological features and topological connectivity of the network and its components, not similarity among preliminary clusters. Clusters that have significant interconnections between them should have substantial similarity. In process 4, STM will merge the clusters which has substantial similarity.

Statistical Signal Transduction Model p Signal transduction behavior of the network is modeled by the Erlang distribution, a special case of the Gamma distribution. (1) p p p where c > 0 is the shape parameter, b > 0 is the scale parameter, x >= 0 is the independent variable, usually time. The Erlang distribution with x/b = 1 is used and the value of c is set to the number of nodes between source protein node and the target protein Setting the value of x/b to unity assesses the perturbation at the target protein when the perturbation reaches 1/e of its initial value at the nearest neighbor of the source protein node.

Statistical Signal Transduction Model p p Statistically, the Erlang distribution represents the time required to carry out a sequence of c tasks whose durations are identical, exponential probability distributions. It represents the chance that the actual time to accomplish c tasks will be less than or equal to b. Figure 2. The pharmacodynamic signal transduction model whose bolus response is an Erlang distribution. The b is the time constant for signal transfer and c is the number of compartments.

Topologically Modified Signal Transduction Model p The Erlang distribution was further weighted to reflect network topology. (2) p p d(i) is the degree of node i, P(v, w) is the set of all visited nodes on the shortest path from node v to node w excluding the source node v and target node w, and F(c) is the signal transduction behavior function. The perturbation induced by the source protein node was assumed to be proportional to its degree and to follow the shortest path to the target protein node. n p Our choice of the shortest path is motivated by the finding that the majority of flux prefers the path of least resistance in many physicochemical and biological systems. During transduction to the target protein node, the perturbation was assumed to be dissipated at each intermediate node visited in proportion to the reciprocal of the degree of each intermediate node visited.
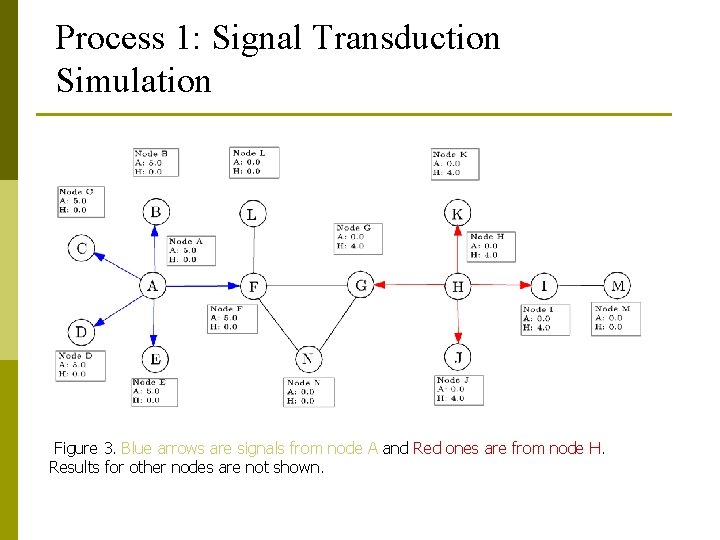
Process 1: Signal Transduction Simulation Figure 3. Blue arrows are signals from node A and Red ones are from node H. Results for other nodes are not shown.
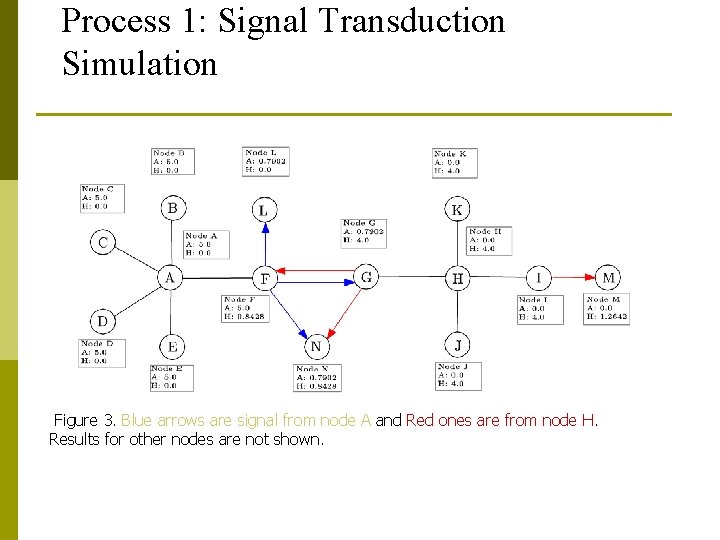
Process 1: Signal Transduction Simulation Figure 3. Blue arrows are signal from node A and Red ones are from node H. Results for other nodes are not shown.
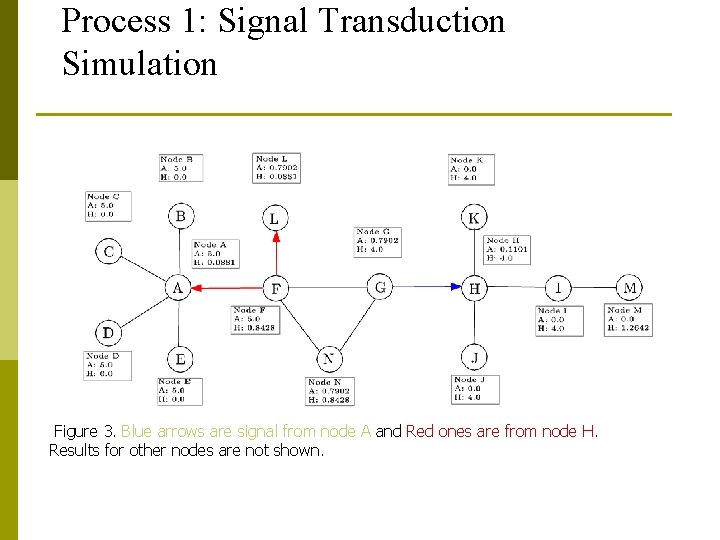
Process 1: Signal Transduction Simulation Figure 3. Blue arrows are signal from node A and Red ones are from node H. Results for other nodes are not shown.
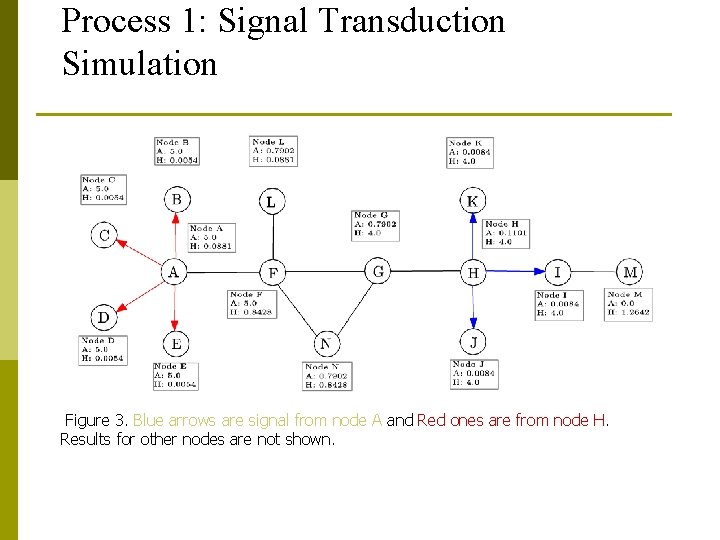
Process 1: Signal Transduction Simulation Figure 3. Blue arrows are signal from node A and Red ones are from node H. Results for other nodes are not shown.
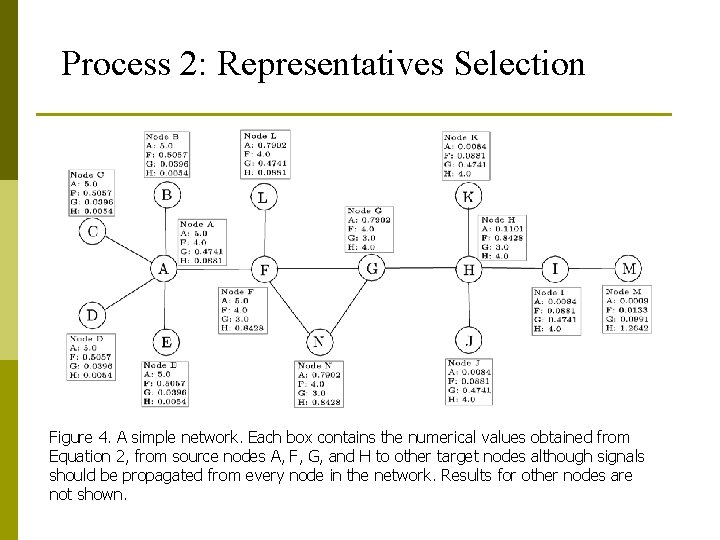
Process 2: Representatives Selection Figure 4. A simple network. Each box contains the numerical values obtained from Equation 2, from source nodes A, F, G, and H to other target nodes although signals should be propagated from every node in the network. Results for other nodes are not shown.
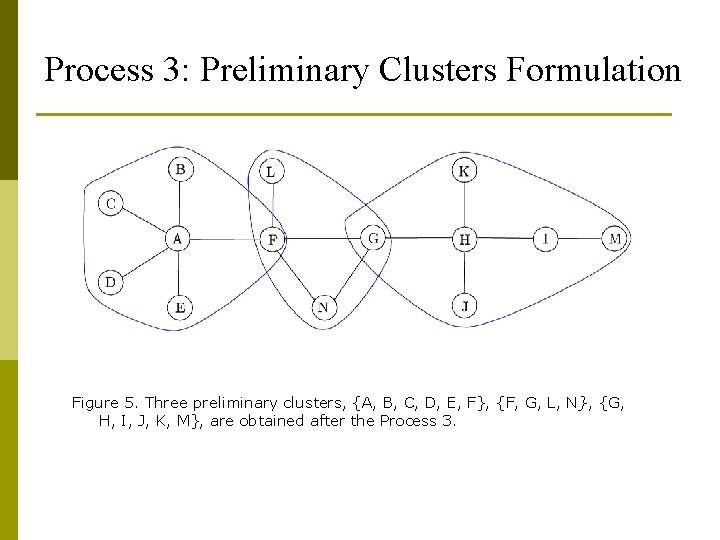
Process 3: Preliminary Clusters Formulation Figure 5. Three preliminary clusters, {A, B, C, D, E, F}, {F, G, L, N}, {G, H, I, J, K, M}, are obtained after the Process 3.

Cluster Merge p Similarity of two clusters i and j (3) p where interconnectivity(i, j) is the number of connections between clusters i and j, and minsize(i, j) is the size of the smaller cluster among clusters i and j. p The pair of clusters that have the highest similarity are merged in each iteration and the merge process iterates until the highest similarity of all cluster pairs is less than a given threshold. p We see when interconnectivity(i, j)>=minsize(i, j), clusters i and j have substantial interconnections.
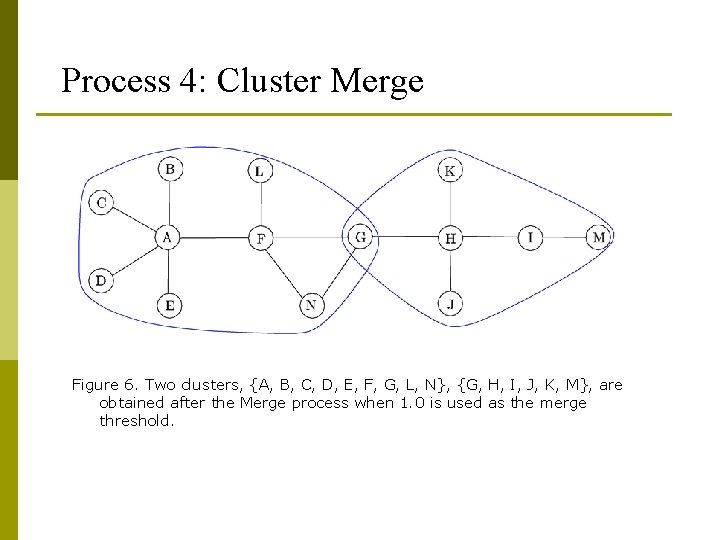
Process 4: Cluster Merge Figure 6. Two clusters, {A, B, C, D, E, F, G, L, N}, {G, H, I, J, K, M}, are obtained after the Merge process when 1. 0 is used as the merge threshold.
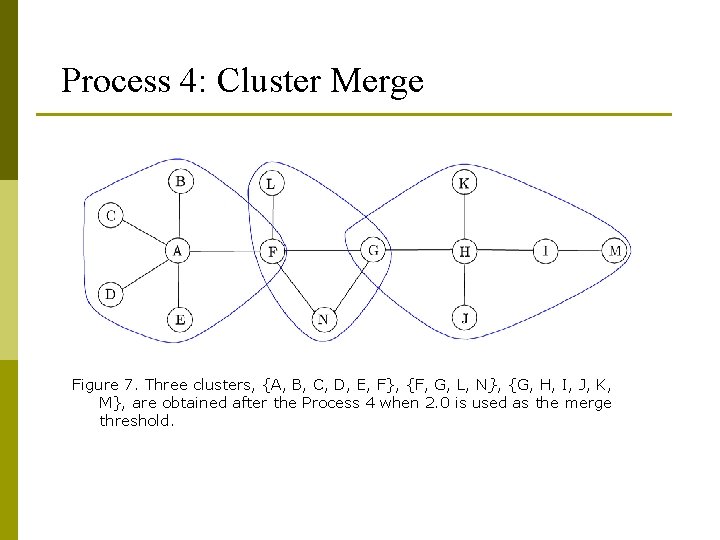
Process 4: Cluster Merge Figure 7. Three clusters, {A, B, C, D, E, F}, {F, G, L, N}, {G, H, I, J, K, M}, are obtained after the Process 4 when 2. 0 is used as the merge threshold.

Experimental Results p Protein Interaction Data n The core data of S. Cerevisiae was obtained from the DIP database. p n 2526 proteins and 5949 filtered reliable physical interactions. Species such as S. Cerevisae provide important test beds for the study of the PPI networks since it is a wellstudied organism for which most proteomics data is available for the organism, by virtue of the availability of a defined and relatively stable proteome, full genome clone libraries, established molecular biology experimental techniques and an assortment of well designed genomics databases.
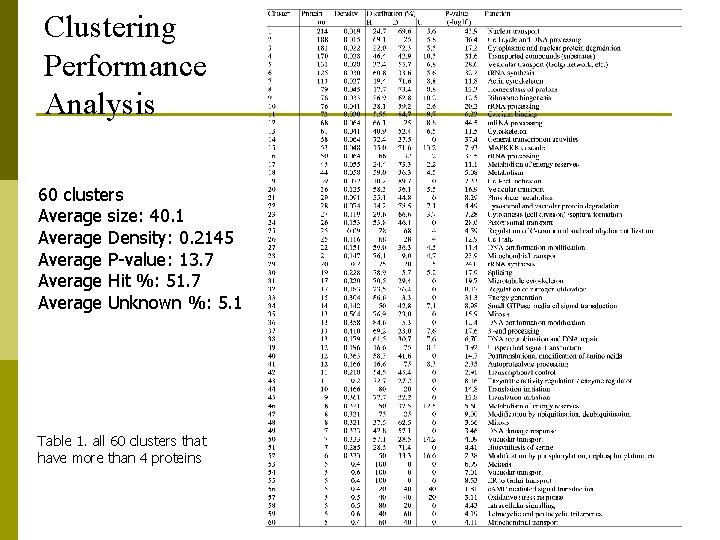
Clustering Performance Analysis 60 clusters Average size: 40. 1 Average Density: 0. 2145 Average P-value: 13. 7 Average Hit %: 51. 7 Average Unknown %: 5. 1 Table 1. all 60 clusters that have more than 4 proteins
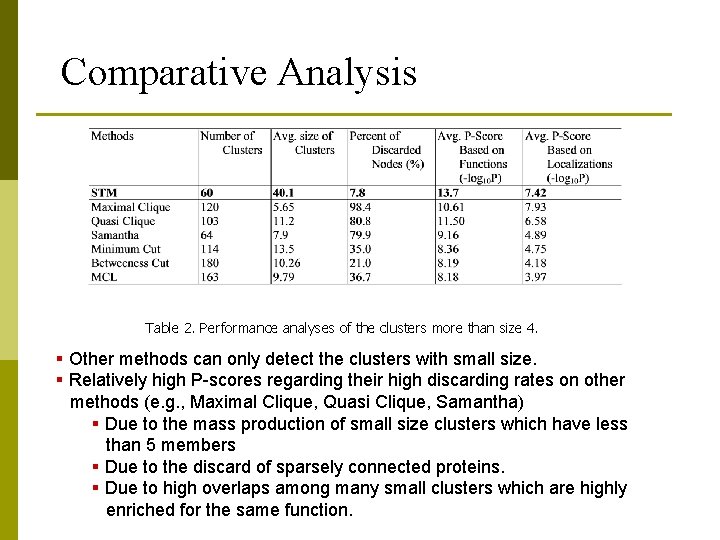
Comparative Analysis Table 2. Performance analyses of the clusters more than size 4. § Other methods can only detect the clusters with small size. § Relatively high P-scores regarding their high discarding rates on other methods (e. g. , Maximal Clique, Quasi Clique, Samantha) § Due to the mass production of small size clusters which have less than 5 members § Due to the discard of sparsely connected proteins. § Due to high overlaps among many small clusters which are highly enriched for the same function.

Computational Complexity p p Our signal transduction based model is fundamentally established on all pairs shortest path searching algorithm to measure the distance between all pairs of nodes: O(V 2 log. V+VE) where V is the number of nodes and E is the number of edges in a network. The time required to find the best cluster pair that has the most interconnections is O(k 2 logk) by using heapbased priority queue, where k is the number of preliminary clusters. But k is much smaller than V in sparse networks like the Yeast PPI network. So the total time complexity of our algorithm is bounded by the time consumed in measuring the distance between all pairs of nodes, which is O(V 2 log. V+VE).

Discussion p p p In head-to-head comparisons, our algorithm outperformed competing approaches and is capable of effectively detecting both dense and sparsely connected, biologically relevant functional modules with fewer discards. The clusters identified had p-values that are 2. 2 orders of magnitude or approximately 125 -fold lower than Quasi clique, the best performing alternative clustering method, on biological function. The incompleteness of clustering is another distinct drawback of existing algorithms, which produce many clusters with small size and singletons. Our method discarded only about 7. 8% of proteins which is tremendously lower than the other approaches did, 59% in average. In conclusion, our method has strong pharmacodynamics-based underpinnings and is an effective, versatile approach for analyzing protein interactions.

Thanks!
- Slides: 78