Biological Membranes Cell Membrane Membrane Structure I Two

Biological Membranes 薛雅薇 中央大學物理系
Cell Membrane

Membrane Structure (I) Two primary building blocks : - Protein - Lipid, or fat. Lipids form a bilayer. The glycocalyx carbohydrate network on the outside surface: - responsible for cell–cell recognition and adhesion to other cells.

Functions of cell membranes 1. Supporting and retaining the cytoplasm (inner cell) 2. Being a selective barrier: Get nutrients in and waste products out. non-polar molecules and some small polar molecules can cross. Most polar compounds, e. g. amino acids, must be specifically transported across the membrane by proteins (Transport). 3. Communication (signaling, via receptors) Information Flow Through the Plasma Membrane via a Membrane Receptor 4. Recognition Structure and function are related.
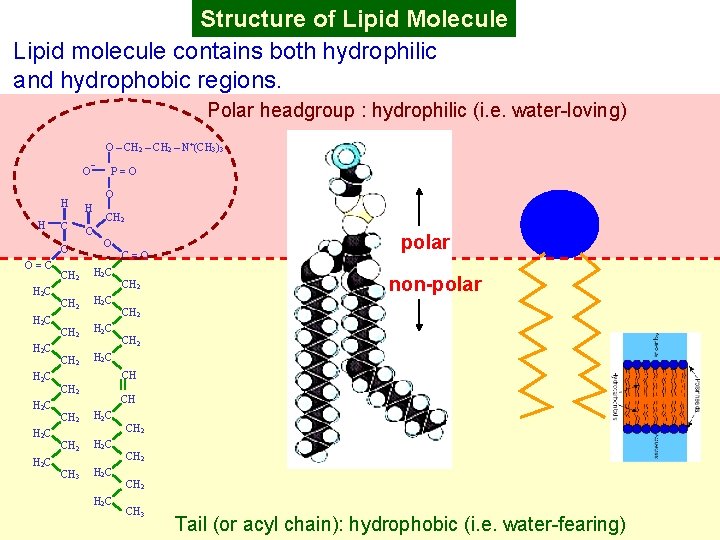
Structure of Lipid Molecule Lipid molecule contains both hydrophilic and hydrophobic regions. Polar headgroup : hydrophilic (i. e. water-loving) O – CH 2 – N+(CH 3)3 O¯ H O=C H 2 C H 2 C O H H C C O P=O CH 2 H 2 C CH 2 H 2 C C=O CH 2 polar non-polar CH 2 CH CH 2 H 2 C CH 3 H 2 C CH 2 CH 3 Tail (or acyl chain): hydrophobic (i. e. water-fearing)

60: 40 DPPC/Chol bilayers In liquid crystalline phase Lipid motion: - Lateral - Rotational - nodding -…

Membrane Structure (II) Consists of specialized domains of different compositions and properties. These specialized domains (called lipid rafts) are enriched in cholesterol (膽固醇) and sphingolipids (high melting lipids) Lipid rafts are postulated to be very important in signal transduction in cells (as signaling platforms, for example). Size of domain: 0 – 10 m It has found that many diseases, such as Alzheimer's, Huntington's, Parkinson's disease, AIDS. . etc are related to lipid rafts.

Fluorescence image (1: 1 brain. PC/brain. SPM)+25 mol%Chol AFM images (1: 1 DOPC/SPM)+25 mol%Chol Crane & Tamm 2004 Two bilayer thicknesses Kruijff et al. 2001

Probing Membrane’s structure & dynamics Use model membranes Nuclear magnetic resonance (NMR, 核磁共振) Differential scanning calorimetry (DSC) Fluorescence microscopy Atomic force microscopy (AFM, 原子力顯微鏡)
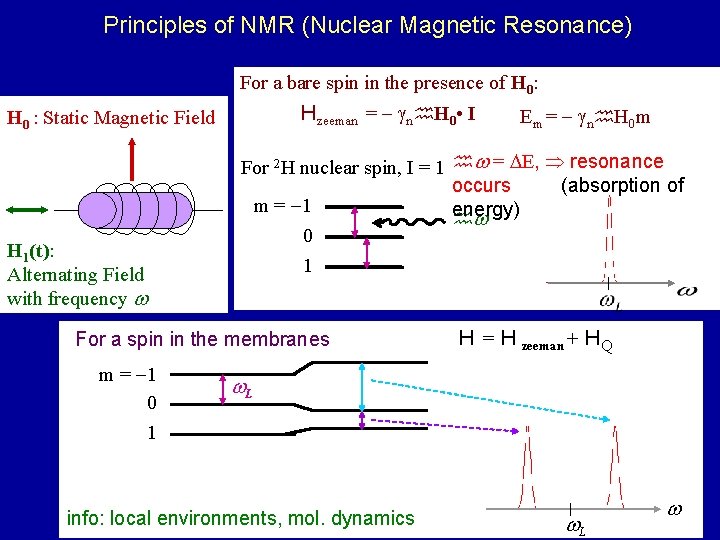
Principles of NMR (Nuclear Magnetic Resonance) For a bare spin in the presence of H 0: Hzeeman = n H 0 • I H 0 : Static Magnetic Field H 1(t): Alternating Field with frequency For 2 H nuclear spin, I = 1 = E, resonance occurs (absorption of m = 1 energy) 0 1 For a spin in the membranes m = 1 0 1 Em = n H 0 m H = H zeeman + HQ L info: local environments, mol. dynamics L

Solid-state NMR spectrometer (固態核磁共振儀)
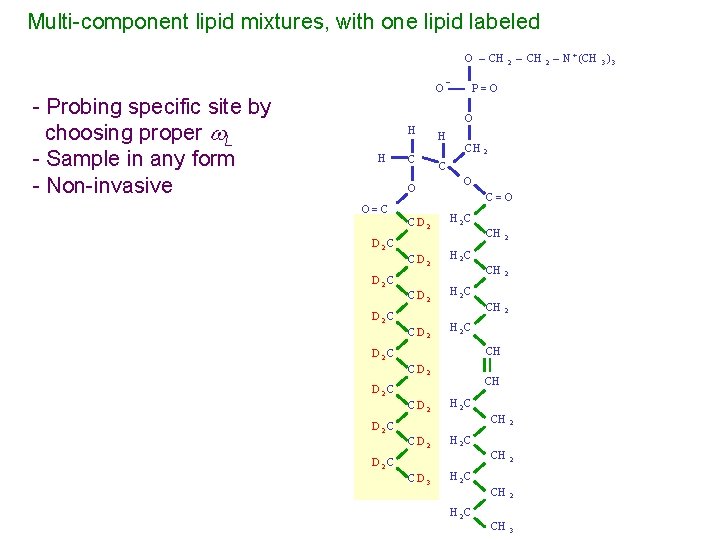
Multi-component lipid mixtures, with one lipid labeled O – CH - Probing specific site by choosing proper L - Sample in any form - Non-invasive O¯ H H C O O=C CD 2 D 2 C CD 2 2 P=O O H CH 2 C O C=O H 2 C CH 2 H 2 C CH D 2 C CD 2 D 2 C CD 3 CH H 2 C CH 2 CH 3 H 2 C – CH 2 – N + (CH 3 ) 3
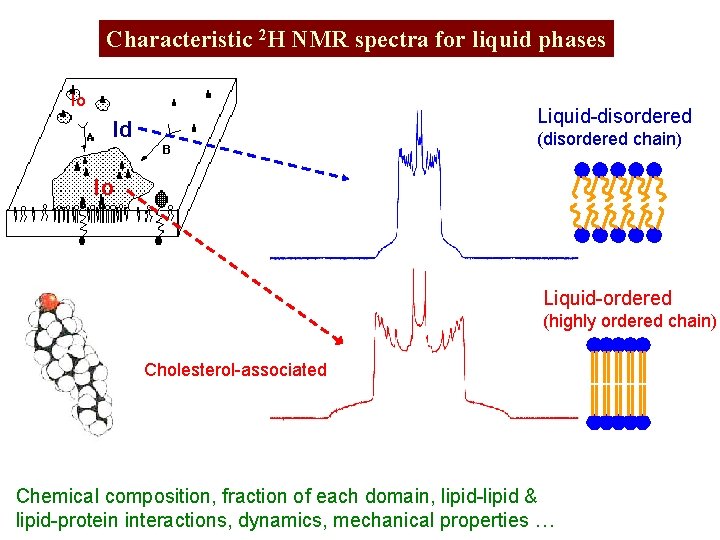
Characteristic 2 H NMR spectra for liquid phases lo Liquid-disordered ld (disordered chain) lo Liquid-ordered (highly ordered chain) Cholesterol-associated Chemical composition, fraction of each domain, lipid-lipid & lipid-protein interactions, dynamics, mechanical properties …
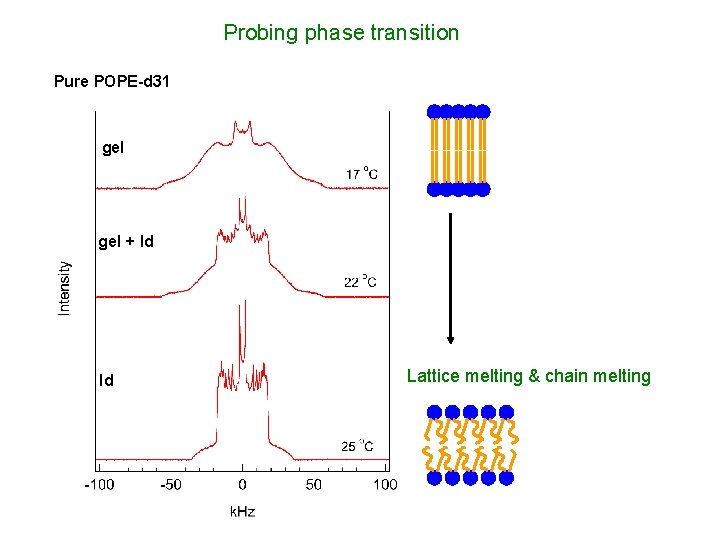
Probing phase transition Pure POPE-d 31 gel + ld ld Lattice melting & chain melting

Thank You
- Slides: 15