Biological Insights Gained from Genomic Prediction John B

Biological Insights Gained from Genomic Prediction John B. Cole Animal Improvement Programs Laboratory Agricultural Research Service, USDA Beltsville, MD 20705 -2350 john. cole@ars. usda. gov

What is whole-genome selection? • • Many markers used to track inheritance of chromosomal segments Impact of each segment estimated for each trait Estimates combined with traditional predicted transmitting ability (PTA) to produce genomic evaluation (GPTA) Animals can be selected shortly after birth University of Bern, November 23, 2009 (2) Cole

What is a SNP? • • Location on a chromosome at which nucleotides differ among animals Usually not a quantitative trait locus (QTL) With enough SNP, association between SNP and QTL alleles allows useful genetic evaluation SNP chosen to be distributed evenly and have both alleles well represented in the population University of Bern, November 23, 2009 (3) Cole

Source of genomic evaluations • • DNA is extracted from blood, hair, or semen 43, 385 SNP are evaluated SNP effect is difference in PTA from having 1 more A (BB v AB or AB v AA) Genomic evaluation combines SNP effect estimates with existing parent average (PA) or PTA University of Bern, November 23, 2009 (4) Cole

Preparation for genotyping • Nominate animal for genotyping • Collect hair, blood, or semen from animal • • • Blood may not be suitable for twins Send to laboratory for DNA extraction Laboratory transfers DNA to Bead. Chip (12 samples/chip) for 3 -day genotyping process University of Bern, November 23, 2009 (5) Cole

Genotyping • Read red/green intensities from chip • Call genotypes from intensity file • Check genotypes: • Duplicates • Parent-progeny conflicts • Wrong breed • Wrong sex University of Bern, November 23, 2009 (6) Cole
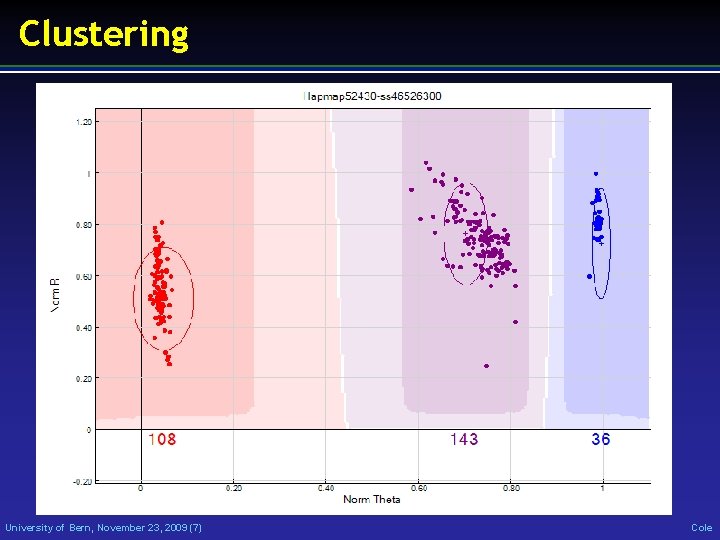
Clustering University of Bern, November 23, 2009 (7) Cole

Genotype for Elevation Chromosome 1 10001112200200121110111121111001121100020122002 2201111202101200211122110021112001111001011011010220 0110022011011200201101020222121122102010011100011220 22122211201202010020220200002110001120201122111022011110000212202000221012020002211220111012100 111211102112110020102100022010002011000022022112101121110122220012112122200200202020122211 002222222002212111121002111120011011200202220001 1120110102111212111020221002112012110011111021112110 2111220001011011102022002211101020111211110112021021 021211011022122001211012022011002220021100 011100211011100022200202212121100022201020022221 2122112111200201102020012222221122120212112101100121 1011020022000200100200011110110012110212121112010101 212022101010111110211221111112121112101101200111 110211110111112201212110102220202121122212022200 2121210201100111222121101 University of Bern, November 23, 2009 (8) Cole

Genotype for inbred bull (Megastar) Chromosome 24 1021222101021021011102110112112110 0220200022202000 202022000002200202222022020000200202222220000202222000 0022020000200200000022220000002022 00200022202022222000220222220000200220202220200020 00220000220222000000220020200022220020020202222022202000202202202222020202022002 22022002220000022020000200200200222220002222020200 22200222020000002222202020000200200222200020220 222200022220220022220202000220220222200220002 00220200000220222000022000222202002222000220 020020202202000222002220220220000022022002002002 02200022220220020220200222200000202200020 2002020200022000002202220020222020002200200022 00200200220222220022022000200002022002022 020020000222002002220000220220020022022 02020200022202000220200202202200002020200002 0202000222222000200220222200000202200202022 02202020002002202200 University of Bern, November 23, 2009 (9) Double grandson of Aerostar Cole

What can go wrong? • • • Inadequate DNA quality or quantity Genotype has many SNP that can’t be determined (90% call rate required) Conflicts between parents and progeny • Pedigree error • Sample ID error • Laboratory error • An unrelated animal qualifies as parent or progeny University of Bern, November 23, 2009 (10) Cole

Parent-progeny conflict • Parent 102120021012012110010201201 00 • Progeny 102020101002002210011201202 20 • The two genotypes are inconsistent because parents and offspring can’t be homozygous for different alleles at the same locus University of Bern, November 23, 2009 (11) Cole
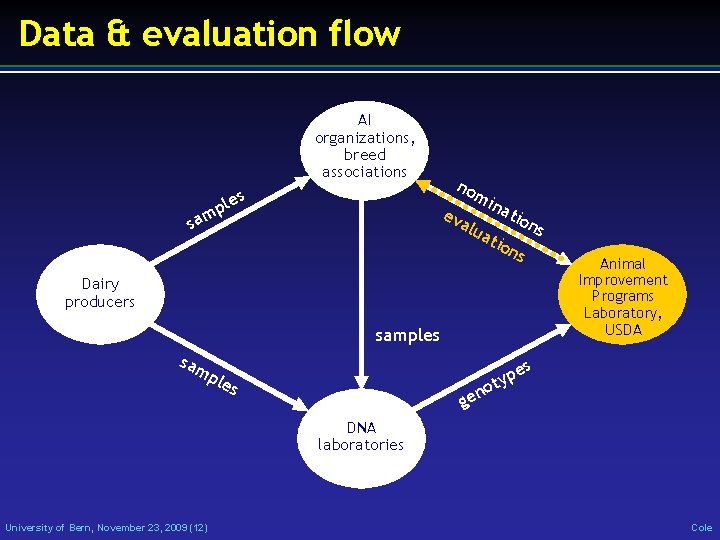
Data & evaluation flow AI organizations, breed associations les p m a s no ev mi na alu ati tio ns on s Dairy producers USDA samples sam Animal Improvement Programs Laboratory, es ple p oty s n ge DNA laboratories University of Bern, November 23, 2009 (12) Cole
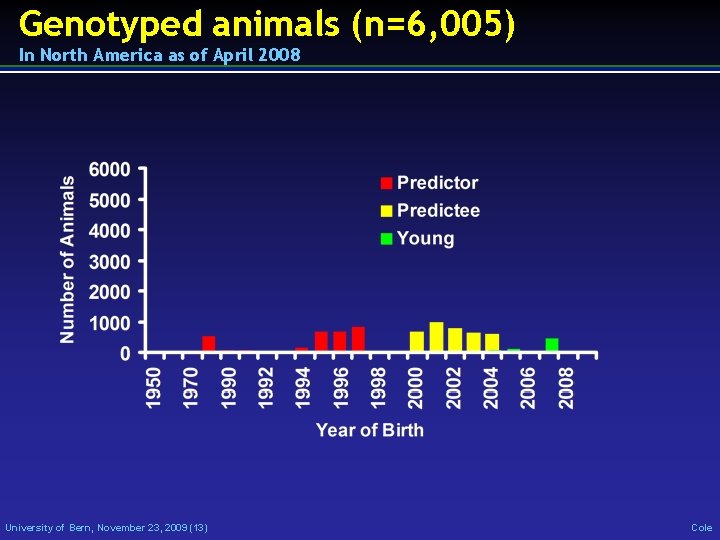
Genotyped animals (n=6, 005) In North America as of April 2008 University of Bern, November 23, 2009 (13) Cole
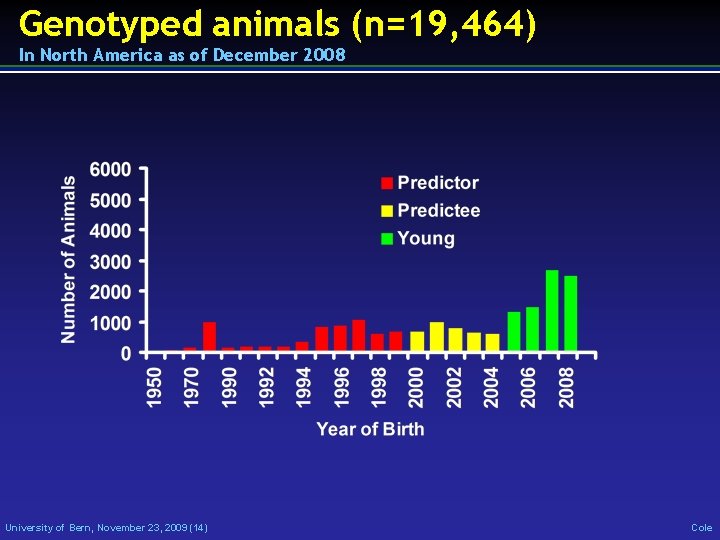
Genotyped animals (n=19, 464) In North America as of December 2008 University of Bern, November 23, 2009 (14) Cole
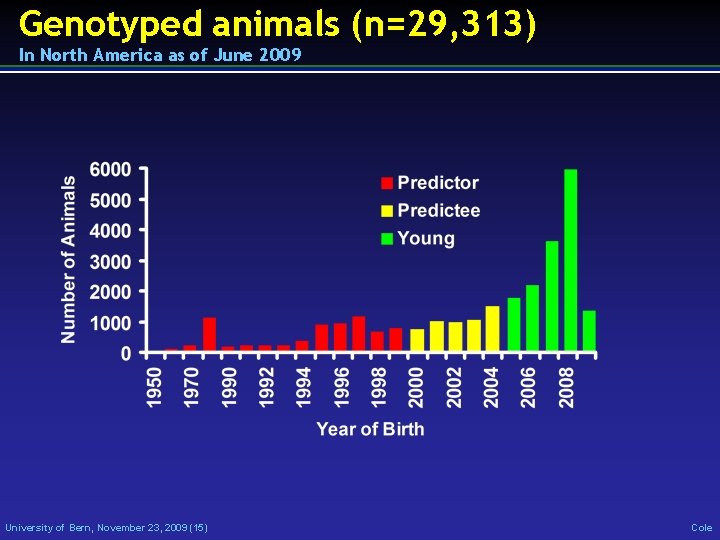
Genotyped animals (n=29, 313) In North America as of June 2009 University of Bern, November 23, 2009 (15) Cole
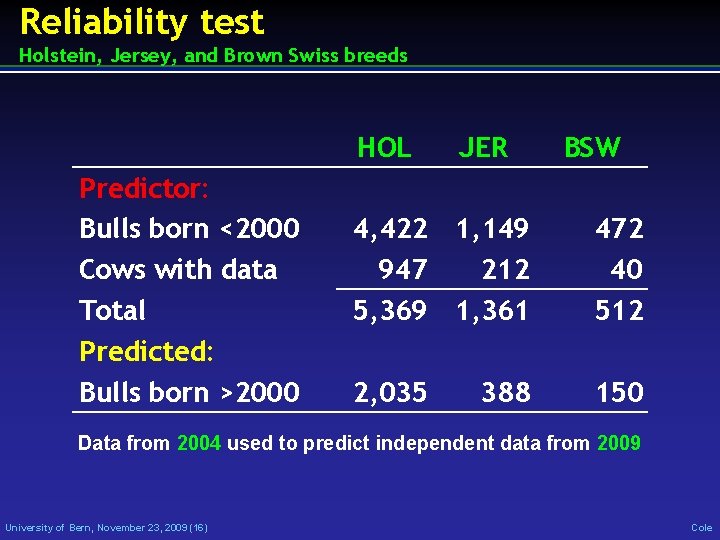
Reliability test Holstein, Jersey, and Brown Swiss breeds HOL Predictor: Bulls born <2000 Cows with data Total Predicted: Bulls born >2000 JER BSW 4, 422 1, 149 947 212 5, 369 1, 361 472 40 512 2, 035 150 388 Data from 2004 used to predict independent data from 2009 University of Bern, November 23, 2009 (16) Cole
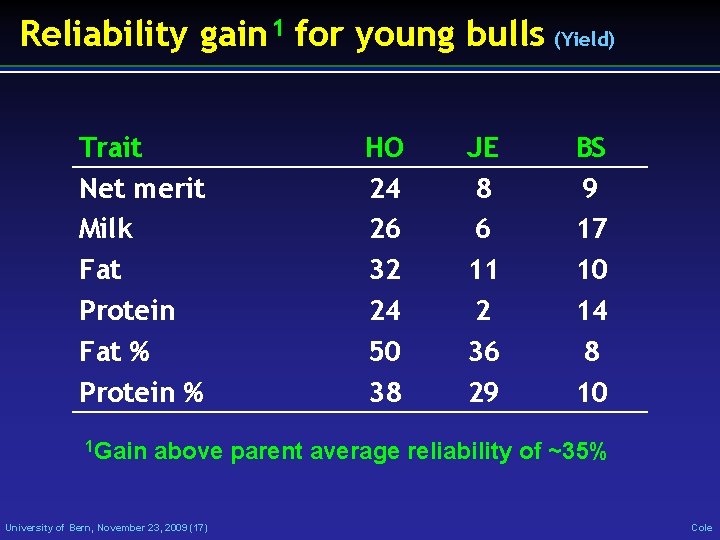
Reliability gain 1 for young bulls (Yield) Trait Net merit Milk Fat Protein Fat % Protein % 1 Gain HO 24 26 32 24 50 38 JE 8 6 11 2 36 29 BS 9 17 10 14 8 10 above parent average reliability of ~35% University of Bern, November 23, 2009 (17) Cole
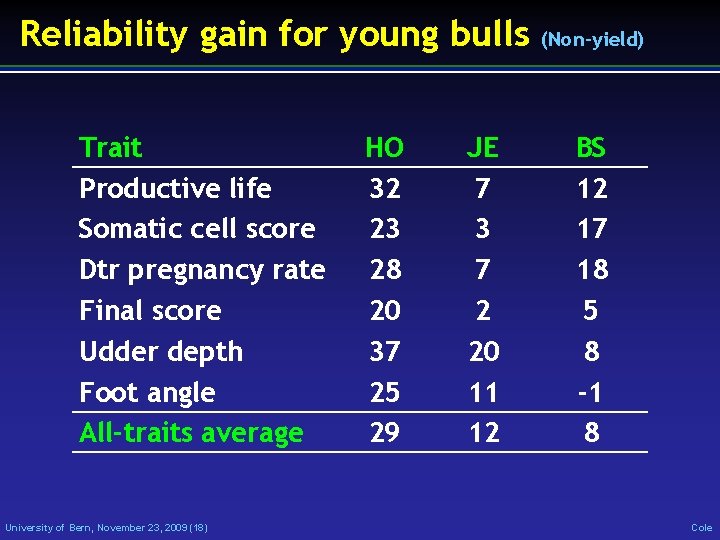
Reliability gain for young bulls Trait Productive life Somatic cell score Dtr pregnancy rate Final score Udder depth Foot angle All-traits average University of Bern, November 23, 2009 (18) HO 32 23 28 20 37 25 29 JE 7 3 7 2 20 11 12 (Non-yield) BS 12 17 18 5 8 -1 8 Cole
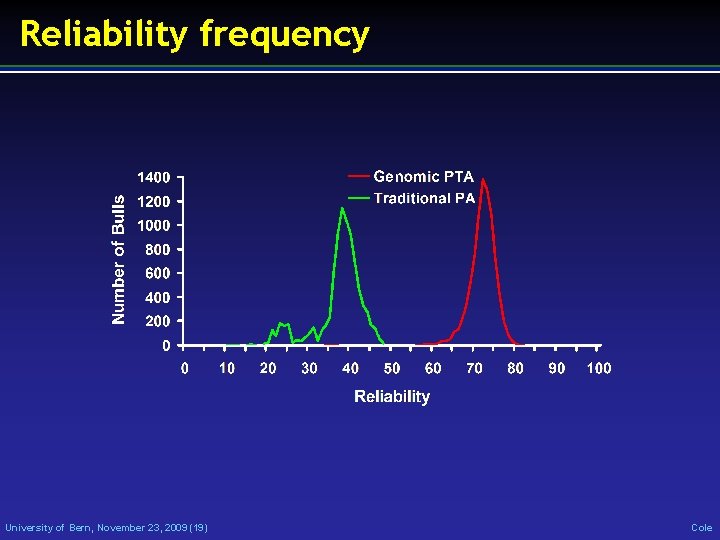
Reliability frequency University of Bern, November 23, 2009 (19) Cole
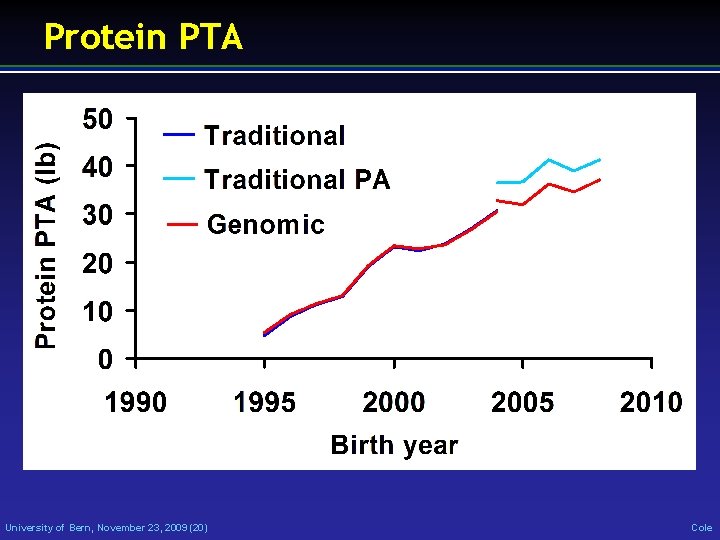
Protein PTA University of Bern, November 23, 2009 (20) Cole
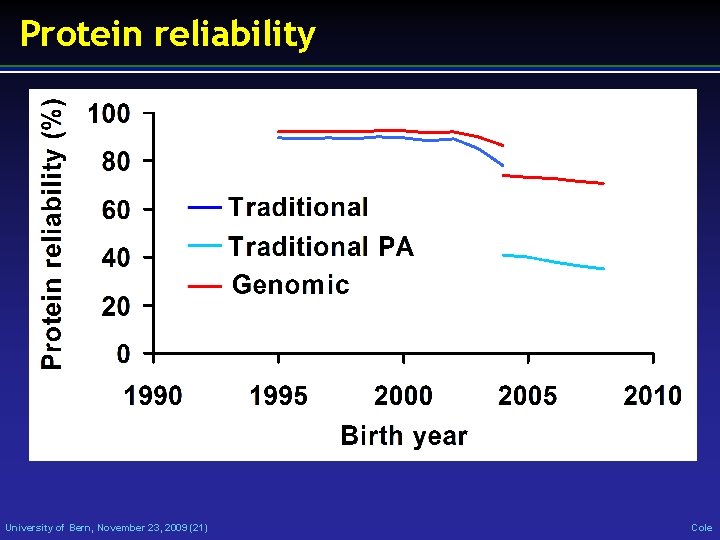
Protein reliability University of Bern, November 23, 2009 (21) Cole
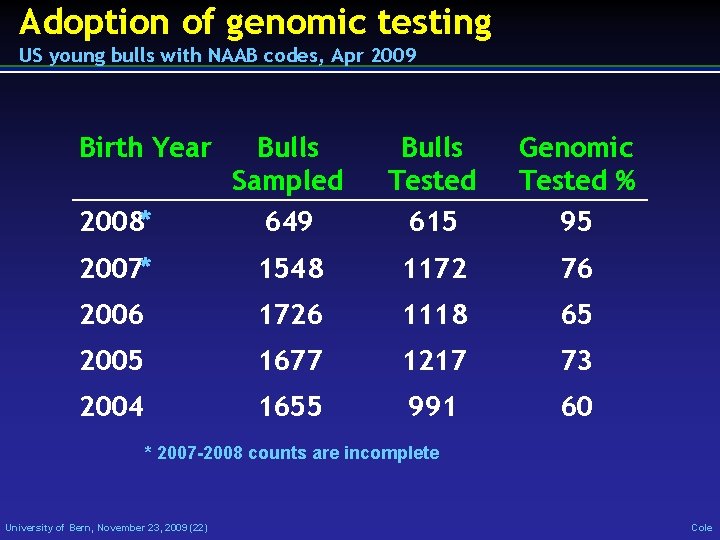
Adoption of genomic testing US young bulls with NAAB codes, Apr 2009 Birth Year 2008* Bulls Sampled 649 Bulls Tested 615 Genomic Tested % 95 2007* 1548 1172 76 2006 1726 1118 65 2005 1677 1217 73 2004 1655 991 60 * 2007 -2008 counts are incomplete University of Bern, November 23, 2009 (22) Cole

Actual results from 50 K chip • • • High correlation between genomic merit in November 2004 and August 2009 merit that includes performance data Bull with highest genomic net merit in November 2004 (Man O Man) now ranks 4 th of 1, 925 bulls Bull with highest genomic net merit in January 2009 (Freddie) now ranks 2 nd University of Bern, November 23, 2009 (23) Cole

Use of genomic evaluations • Determine which young bulls to bring into AI • Aid in selection of mating sires • Increasing impact on bull dam selection • Market semen from 2 -year-old bulls University of Bern, November 23, 2009 (24) Cole

Updates between official evaluations • • • Genomic evaluations calculated every 2 months Not released for animals that already have an official evaluation Evaluations of new animals distributed to owners • Females • Males • by breed associations by NAAB Usually 2, 000 new genotypes included University of Bern, November 23, 2009 (25) Cole

Impact on producers • • • Young-bull evaluations with accuracy of early 1 st-crop evaluations AI organizations marketing genomically evaluated 2 -year-olds Bull dams likely to be required to be genotyped Rate of genetic improvement likely to increase by up to 50% Progeny-test programs changing University of Bern, November 23, 2009 (26) Cole

International implications • • • All major dairy countries investigating genomic selection Interbull researching how to integrate genomic evaluations European collaboration to share genotypes Prediction accuracy continues to increase with increasing numbers of predictor animals Importing countries must change rules to allow for genomically evaluated young bulls University of Bern, November 23, 2009 (27) Cole

Possible selection of embryos • • In vitro fertilization of embryos from immature animals • Further reduces generation interval • Not yet feasible Frozen, genotyped embryo market • Cost of genotyping < cost of ET • Could replace AI if accuracy high and vitality not affected University of Bern, November 23, 2009 (28) Cole
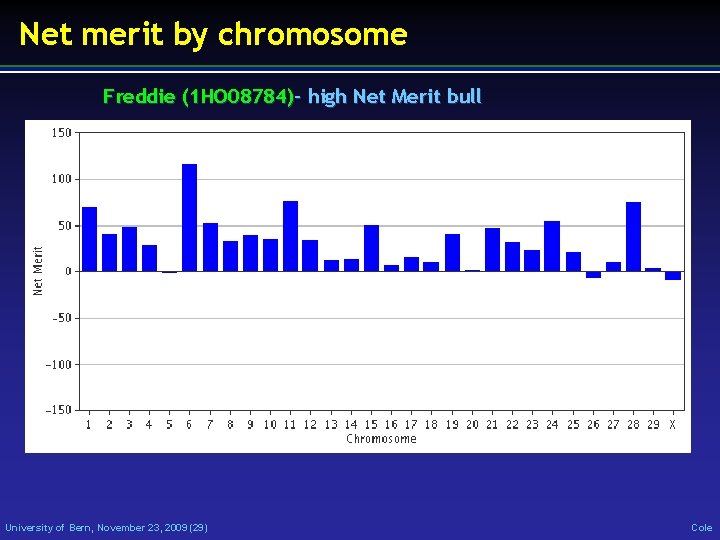
Net merit by chromosome Freddie (1 HO 08784)- high Net Merit bull University of Bern, November 23, 2009 (29) Cole
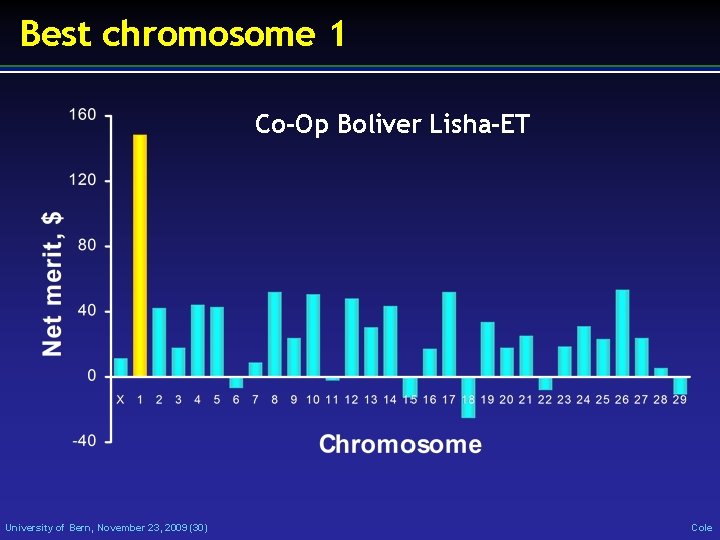
Best chromosome 1 Co-Op Boliver Lisha-ET University of Bern, November 23, 2009 (30) Cole
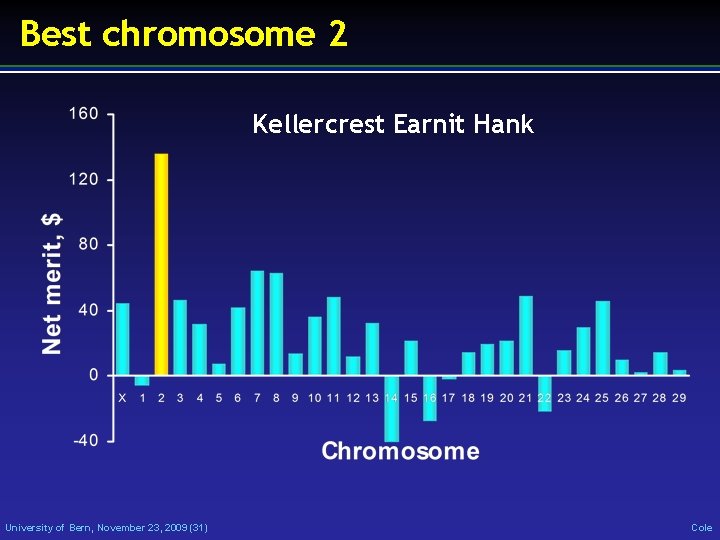
Best chromosome 2 Kellercrest Earnit Hank University of Bern, November 23, 2009 (31) Cole
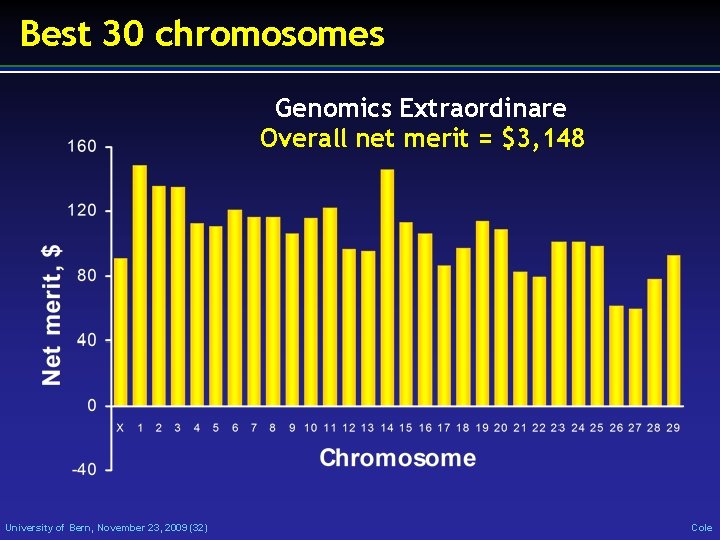
Best 30 chromosomes Genomics Extraordinare Overall net merit = $3, 148 University of Bern, November 23, 2009 (32) Cole
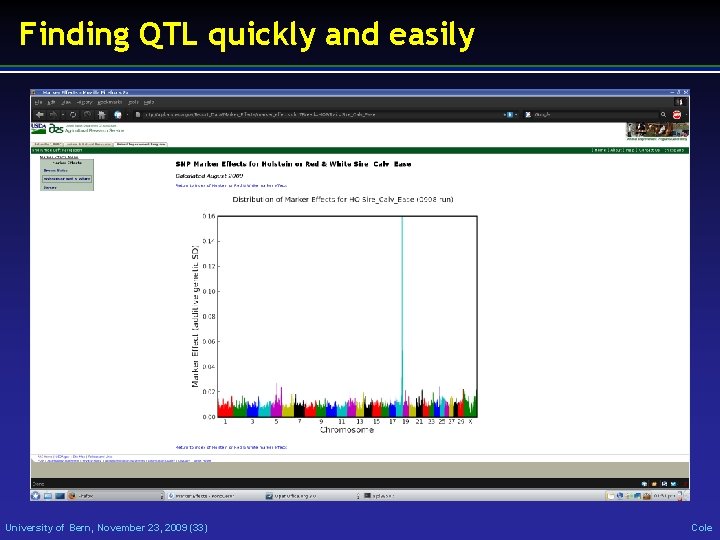
Finding QTL quickly and easily University of Bern, November 23, 2009 (33) Cole
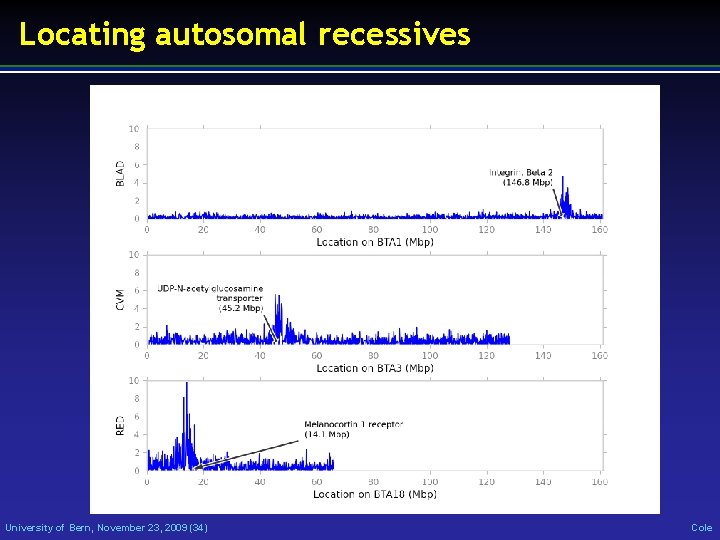
Locating autosomal recessives University of Bern, November 23, 2009 (34) Cole
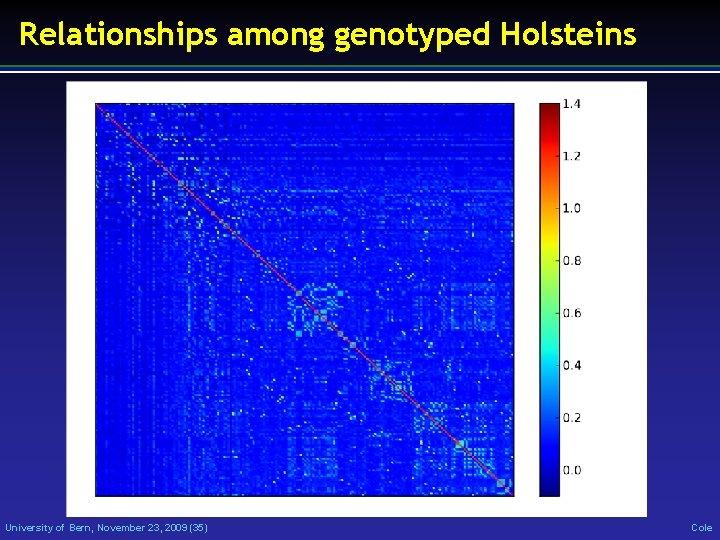
Relationships among genotyped Holsteins University of Bern, November 23, 2009 (35) Cole
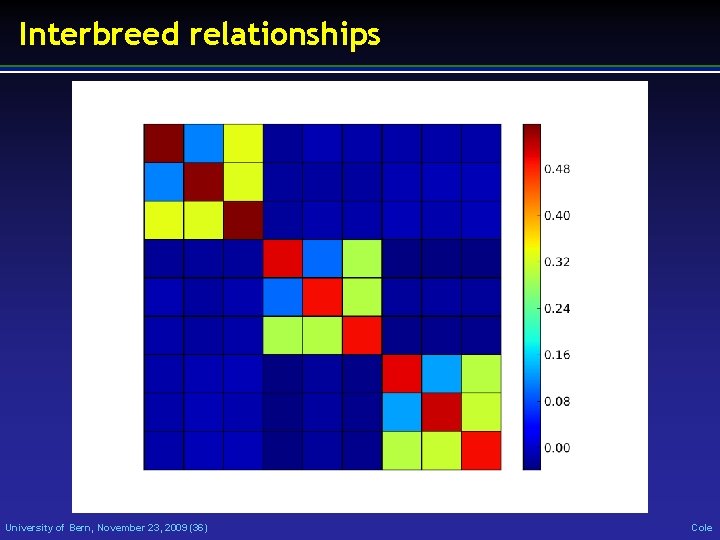
Interbreed relationships University of Bern, November 23, 2009 (36) Cole

Closing thoughts • • Extraordinarily rapid implementation of genomic evaluations Young-bull acquisition and marketing now based on genomic evaluations Genomic evaluations may allow more cows from commercial herds to be used as bull dams SNP data are valuable tools for basic research University of Bern, November 23, 2009 (37) Cole

Acknowledgments • Genotyping and DNA extraction: • • Computing: • • BFGL, U. Missouri, U. Alberta, Gene. Seek, Genetics & IVF Institute, Genetic Visions, and Illumina AIPL staff (Leigh Walton, Jay Megonigal) Funding: • • NRI grants 2006 -35205 -16888 and 200635205 -16701 Agriculture Research Service Holstein, Jersey and Brown Swiss breed associations Cooperative Dairy DNA Repository (CDDR) University of Bern, November 23, 2009 (38) Cole

CDDR contributors • National Association of Animal Breeders (Columbia, MO) • • ABS Global (De. Forest, WI) Accelerated Genetics (Baraboo, WI) Alta (Balzac, AB, Canada) Genex (Shawano, WI) New Generation Genetics (Fort Atkinson, WI) Select Sires (Plain City, OH) Semex Alliance (Guelph, ON, Canada) Taurus-Service (Mehoopany, PA) University of Bern, November 23, 2009 (39) Cole
- Slides: 39