Biological Databases What types of data are available

Biological Databases What types of data are available? What is a database? What are Genbank and Entrez? What does a typical entry look like? How does one use the database? BIO 520 Bioinformatics Jim Lund

NCBI Biological Databases Central Dogma-o-centric • • • Genomic DNA sequence m. RNA/c. DNA sequence Protein 3 D structure Literature (Function)

Biological Data • Genomic DNA sequence (complete) • m. RNA/c. DNA sequence • Gene expression data (NEW) – Microarrays, SAGE – Expression catalogs • Protein sequence – Protein interaction/complex data (NEW) • Protein 3 D structure • Literature (Function) – Organism databases (NEW) – Annotation and classification projects (NEW)

What is a Biological Database? An organized body of persistent data and associated computer software for updating, querying, and retrieving data records. • Collection of records and files • Organized for a particular purpose • The database is separate from the interface and can have several interfaces. – NCBI Protein can be searched by protein name or using BLAST (Basic Local Alignment Search Tool).

Common database features • Relational Databases – Tables – Relationships between tables • Version Control • Consistency enforcement • Multiauthor/multiuser with security
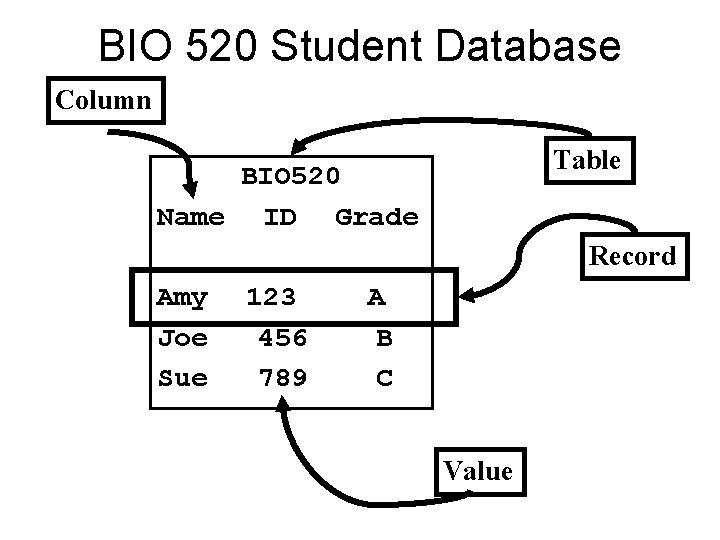
BIO 520 Student Database Column Table BIO 520. Name ID Grade Record Amy Joe Sue 123 456 789 A B C Value

Genbank Entry LOCUS BC 005255 495 bp m. RNA linear PRI 23 -JUN-2006 DEFINITION Homo sapiens insulin, m. RNA (c. DNA clone IMAGE: 3950204), complete cds. ACCESSION BC 005255 VERSION BC 005255. 1 GI: 13528923 KEYWORDS MGC. SOURCE Homo sapiens (human) ORGANISM Homo sapiens Eukaryota; Metazoa; Chordata; Craniata; Vertebrata; Euteleostomi; Mammalia; Eutheria; Euarchontoglires; Primates; Haplorrhini; Catarrhini; Hominidae; Homo. FEATURES Location/Qualifiers source 1. . 495 /organism="Homo sapiens" gene 1. . 495 /gene="INS" /db_xref="Gene. ID: 3630" CDS 60. . 392 /gene="INS" /translation="MALWMRLLPLLALLALWGPDPAAAFVNQHLCGSHLVEALYLVCG ERGFFYTPKTRREAEDLQVGQVELGGGPGAGSLQPLALEGSLQKRGIVEQCCTSICSL YQLENYCN" ORIGIN 1 agccctccag gacaggctgc atcagaagag gccatcaagc agatcactgt ccttctgcca … 421 ccgcctcctg caccgagaga gatggaataa agcccttgaa ccaacaaaaa 481 aaaaa //
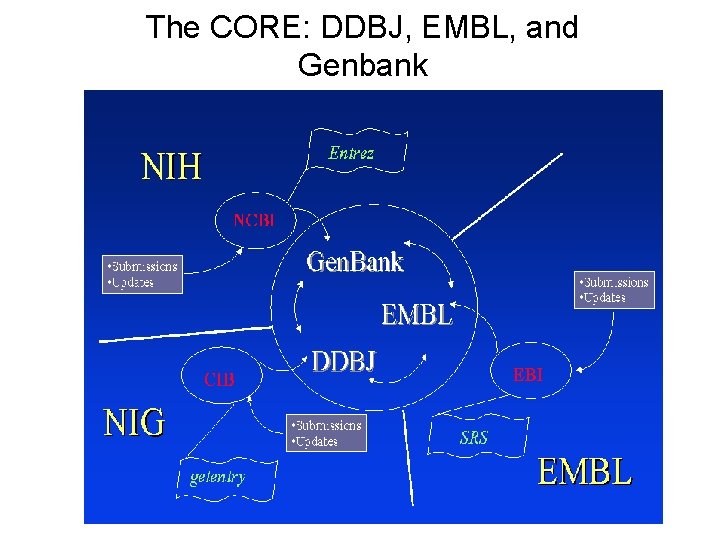
The CORE: DDBJ, EMBL, and Genbank

Genbank DNA Sequence Database • Genbank/EMBL/DDBJ mirror & exchange sequence records. • Primary vs. Secondary Databases – nr (non-redundant database) • Primary vs. secondary records – Sequence vs. inferred property (coding region)

Primary vs. Derivative Databases • Primary Databases – Original submissions by experimentalists – Content controlled by the submitter • Examples: Gen. Bank, SNP, GEO • Derivative Databases – Built from primary data – Content controlled by third party (NCBI) • Examples: Refseq, TPA, Ref. SNP, Uni. Gene, NCBI Protein, Structure, Conserved Domain

LOCUS DEFINITION AY 182241 1931 bp m. RNA linear PLN 04 -MAY-2004 Malus x domestica (E, E)-alpha-farnesene synthase (AFS 1) m. RNA, complete cds. ACCESSION AY 182241 VERSION AY 182241. 2 GI: 32265057 KEYWORDS. SOURCE Malus x domestica (cultivated apple) ORGANISM Malus x domestica Eukaryota ; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta; Spermatophyta; Magnoliophyta; eudicotyledons ; core eudicots; rosids; eurosids I; Rosales; Rosaceae; Maloideae; Malus. REFERENCE 1 (bases 1 to 1931) AUTHORS Pechous, S. W. and Whitaker, B. D. TITLE Cloning and functional expression of an (E, E) -alpha-farnesene synthase c. DNA from peel tissue of apple fruit JOURNAL Planta 219, 84 -94 (2004) REFERENCE 2 (bases 1 to 1931) AUTHORS Pechous, S. W. and Whitaker, B. D. TITLE Direct Submission JOURNAL Submitted (18 -NOV-2002) PSI-Produce Quality and Safety Lab, USDA-ARS, 10300 Baltimore Ave. Bldg. 002, Rm. 205, Beltsville, MD 20705, USA REFERENCE 3 (bases 1 to 1931) AUTHORS Pechous, S. W. and Whitaker, B. D. TITLE Direct Submission JOURNAL Submitted (25 -JUN-2003) PSI-Produce Quality and Safety Lab, USDA-ARS, 10300 Baltimore Ave. Bldg. 002, Rm. 205, Beltsville, MD 20705, USA REMARK Sequence update by submitter COMMENT On Jun 26, 2003 this sequence version replaced gi: 27804758. FEATURES Location/Qualifiers source 1. . 1931 /organism="Malus x domestica" /mol_type="m. RNA" /cultivar="'Law Rome'" /db_xref="taxon : 3750" /tissue_type="peel" gene 1. . 1931 /gene="AFS 1" CDS 54. . 1784 /gene="AFS 1" /note="terpene synthase" /codon_start=1 /product="(E, E) -alpha-farnesene synthase" /protein_id="AAO 22848. 2" /db_xref="GI: 32265058" /translation="MEFRVHLQADNEQKIFQNQMKPEPEASYLINQRRSANYKPNIWK NDFLDQSLISKYDGDEYRKLSEKLIEEVKIYISAETMDLVAKLELIDSVRKLGLANLF EKEIKEALDSIAAIESDNLGTRDDLYGTALHFKILRQHGYKVSQDIFGRFMDEKGTLE DFLHKNEDLLYNISLIVRLNNDLGTSAAEQERGDSPSSIVCYMREVNASEETARKNIK GMIDNAWKKVNGKCFTTNQVPFLSSFMNNATNMARVAHSLYKDGDGFGDQEKGPRTHI LSLLFQPLVN" ORIGIN 1 ttcttgtatc ccaaacatct cgagcttctt gtacaccaaa ttaggtattc actatggaat 61 tcagagttca cttgcaagct gataatgagc agaaaatttt tcaaaaccag atgaaacccg 121 aacctgaagc ctcttacttg attaatcaaa gacggtctgc aaattacaag ccaaatattt 181 ggaagaacga tttcctagat caatctctta tcagcaaata cgatggagat gagtatcgga 241 agctgtctga gaagttaata gaagaagtta agatttatat atctgctgaa acaatggatt // A Traditional Gen. Bank Record Header The Flatfile Format (formatted text) Feature Table Sequence

Genbank Entry LOCUS PCU 30791 -MAY-1996 1234 bp m. RNA PLN 31 DEFINITION Pneumocystis carinii form 6 guanine nucleotide binding protein alpha subunit (pcg 1) m. RNA, complete cds. ACCESSION U 30791 NID g 1345098 VERSION U 30791. 1 GI: 1345098 Unique ID Version Control

Content-Taxonomy SOURCE Pneumocystis carinii f. sp. carinii. ORGANISM Pneumocystis carinii f. sp. carinii Eukaryota; Fungi; Ascomycota; Archiascomycetes; Pneumocystidaceae; Pneumocystis.

Reference REFERENCE 1 (bases 1 to 1234) AUTHORS Smulian, A. G. , Ryan, M. , Staben, C. and Cushion, M. TITLE Signal transduction in Pneumocystis carinii: characterization of the genes (pcg 1) encoding the alpha subunit of the G protein (PCG 1) of Pneumocystis carinii and Pneumocystis carinii ratti JOURNAL Infect. Immun. 64 (3), 691 -701 (1996) PUBMED 96186460 • Unique cross reference • Can be >1 reference

Features FEATURES Location/Qualifiers source 1. . 1234 /organism="Pneumocystis carinii f. sp. carinii“ /strain="Form 6“ /note="450 kb chromosome" /db_xref="taxon: 38081“ 5'UTR 1. . 90 gene 91. . 1155 /gene="pcg 1"

CDS 91. . 1155 /gene="pcg 1” Related info in another database /note="G-protein alpha subunit" /codon_start=1 /product= "guanosine nucleotide binding protein alpha subunit" /protein_id="AAC 49295. 1" /db_xref="PID: g 1345099". INFERRED /db_xref="GI: 1345099" /translation="MGCCFSATYNQDTLRSKEI E SYLRQEQEHACHEAKILLLGAGES…

DNA BASE COUNT 421 a 171 c 195 g 447 t ORIGIN 1 tgaattctaa attttatatt … 1201 … tattttttta tgctccagat aaaa //

Genbank entries • Combination of required (LOCUS, SOURCE) and optional fields. – The entry is hierarchical, some fields contain subfields. • REFERENCE->AUTHORS • Some fields can appear multiple times (REFERENCE, /gene) • Some fields are numerical, other are text. Some fields contain free text, others use a controlled vocabulary or an database ID.

Other Genbank output formats • FASTA – Simple, little annotation information – Easy to use – Common denominator format • ASN 1 – Computer friendly, human unfriendly • XML, INSDSeq. XML, Tiny. Seq. XML • Graph (graphical map of seq features) …and more

DNA Sequence Files Common formats • Genbank (used by Vector. NTI) • FASTA • GCG – Accelrys GCG (Genetics Computer Group) package – formerly GCG Wisconsin Package Many others!

FASTA One annotation line only! >gi|1345098|gb|U 30791. 1|PCU 30791 TGAATTCTAAATTTTATATTTCTAATTGCATTTTATATT TTTGATAATACTAGATTTATTCCTGGAAACT TAAATTAGTTATTTTAAGTTATGGGATGTTGTTTTTCT GCTACATATAACCAAGATACACTTCGTTCCAA

Submitting sequences to Genbank • Sequin –Stand-alone sequence submission tool. • Bank. It –Web based sequence submission.

Genbank is an ARCHIVE • The literature and secondary databases are the knowledge sources. • There are many additional NCBI annotation databases

NCBI annotation databases! • Genbank -> Ref. Seq (Single sequence for each gene) • Entrez Gene (Gene-based links to annotation sources). • Homolo. Gene (Homologs) • OMIM • Conserved domains, 3 D domains • GEO (Gene expression datasets) • DNA, protein, 3 D structures • Interaction data • Links to other databases! • NCBI Genomes • NCBI Map viewer

Finding and editing DNA files • • Find DNA: Entrez Downloading files Format Conversion Sequence viewing/editing
Entrez • Database searching/browsing • Example: Pneumocystis Gproteins – PCR a c. DNA to express in E. coli – Read about it and related genes – Check similarity to • http: //www. ncbi. nlm. nih. gov/ Entrez/ related G-proteins – View the 3 D structure? ?
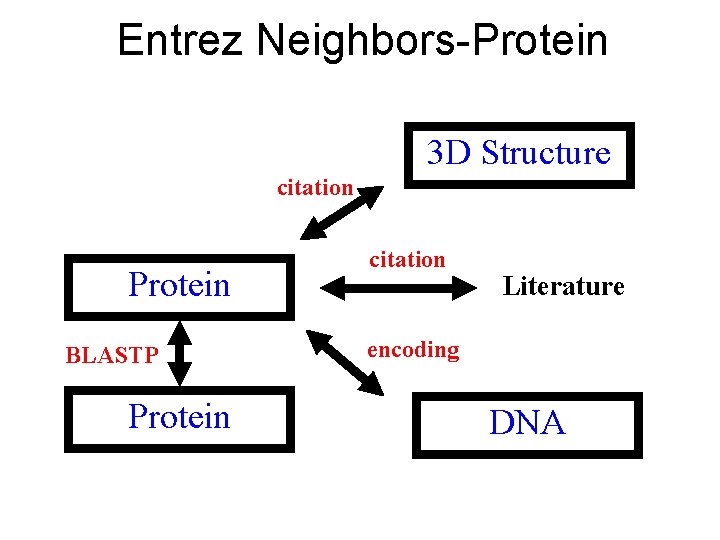
Entrez Neighbors-Protein 3 D Structure citation Protein BLASTP Protein citation Literature encoding DNA

Mapping the menagerie of biological databases

Nucleic Acid Manipulations • On the web: – Baylor Human Genome Center (BCM) http: //searchlauncher. bcm. tmc. edu/seq -util/seq-util. html – European Bioinformatics Institute (EBI) http: //www. ebi. ac. uk/Tools/misc. html

DNA/Protein sequence format conversion • Readseq – Download program: – http: //iubio. indiana. edu/soft/molbio/rea dseq – Use online: – http: //www. ebi. ac. uk/cgi-bin/readseq. cgi – http: //searchlauncher. bcm. tmc. edu/sequtil/readseq. html Beware Information Loss!

Reverse Complementing 5’-GAATCA-3’ 5’-TGATTC-3’ NOT 5’-ACTAAAG-3’

Sequence Statistics • • Nucleotide frequencies (di, tri…) UV Absorbance MW Tm

Restriction Map • Linear vs Circular • Enzyme sets – Which enzymes, where they cut. • Gel simulation – Gel-to-map MUCH harder!! • Useful for: – Cloning – Southern blots – Specialized mol bio techniques

Translation/ORFs • Translation table – Standard vs non-standard • Frame (1, 2, 3, 4, 5, 6) • Segmental translation (exon-intron) • Primary translation vs mature polypeptide

Sequence Annotation and Editing • Artemis • Sequin • NCBI’s Genbank entry creation/viewing tool • Text editor – Notepad – Word processor – vi MWGTCC Nonproportional fonts (courier, monospaced…) IIIIII MWGTCC IIIIII
Primer design program: Primer 3 http: //frodo. wi. mit. edu/cgi-bin/primer 3_www. cgi

Primary vs. Derivative Databases • Primary Databases – Original submissions by experimentalists – Content controlled by the submitter • Examples: Gen. Bank, SNP, GEO • Derivative Databases – Built from primary data – Content controlled by third party (NCBI) • Examples: Refseq, TPA, Ref. SNP, Uni. Gene, NCBI Protein, Structure, Conserved Domain

Other NCBI Databases • Structure: imported structures (PDB) Cn 3 D viewer, NCBI curation • CDD: conserved domain database Protein families (COGs and KOGs) Single domains (PFAM, SMART, CD) • db. SNP: • Gene: nucleotide polymorphism gene records Unifies Locus. Link and Microbial Genomes
Homologene Cluster
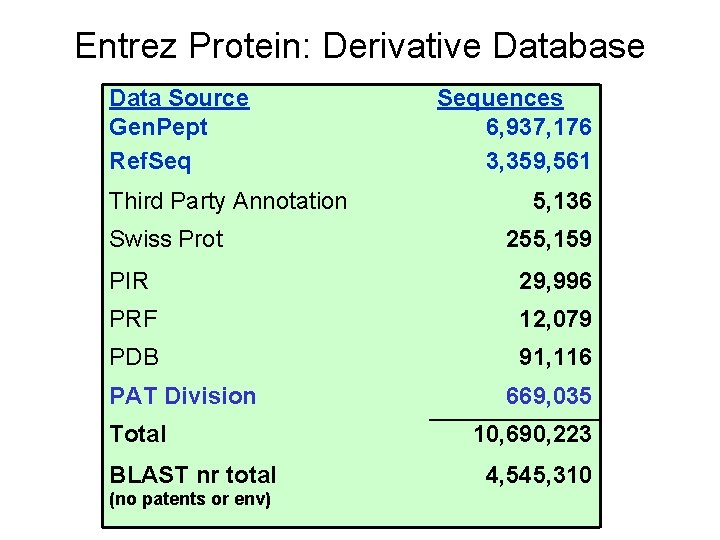
Entrez Protein: Derivative Database Data Source Gen. Pept Ref. Seq Third Party Annotation Swiss Prot Sequences 6, 937, 176 3, 359, 561 5, 136 255, 159 PIR 29, 996 PRF 12, 079 PDB 91, 116 PAT Division Total BLAST nr total (no patents or env) 669, 035 10, 690, 223 4, 545, 310
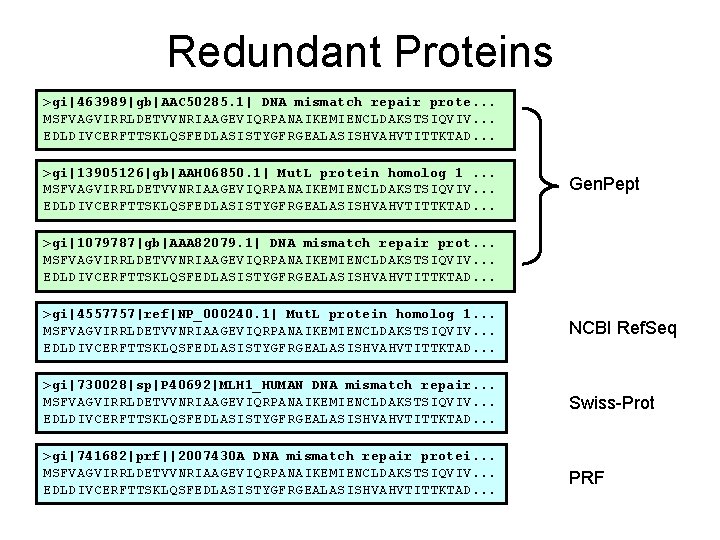
Redundant Proteins >gi|463989|gb|AAC 50285. 1| DNA mismatch repair prote. . . MSFVAGVIRRLDETVVNRIAAGEVIQRPANAIKEMIENCLDAKSTSIQVIV. . . EDLDIVCERFTTSKLQSFEDLASISTYGFRGEALASISHVAHVTITTKTAD. . . >gi|13905126|gb|AAH 06850. 1| Mut. L protein homolog 1. . . MSFVAGVIRRLDETVVNRIAAGEVIQRPANAIKEMIENCLDAKSTSIQVIV. . . EDLDIVCERFTTSKLQSFEDLASISTYGFRGEALASISHVAHVTITTKTAD. . . Gen. Pept >gi|1079787|gb|AAA 82079. 1| DNA mismatch repair prot. . . MSFVAGVIRRLDETVVNRIAAGEVIQRPANAIKEMIENCLDAKSTSIQVIV. . . EDLDIVCERFTTSKLQSFEDLASISTYGFRGEALASISHVAHVTITTKTAD. . . >gi|4557757|ref|NP_000240. 1| Mut. L protein homolog 1. . . MSFVAGVIRRLDETVVNRIAAGEVIQRPANAIKEMIENCLDAKSTSIQVIV. . . EDLDIVCERFTTSKLQSFEDLASISTYGFRGEALASISHVAHVTITTKTAD. . . NCBI Ref. Seq >gi|730028|sp|P 40692|MLH 1_HUMAN DNA mismatch repair. . . MSFVAGVIRRLDETVVNRIAAGEVIQRPANAIKEMIENCLDAKSTSIQVIV. . . EDLDIVCERFTTSKLQSFEDLASISTYGFRGEALASISHVAHVTITTKTAD. . . Swiss-Prot >gi|741682|prf||2007430 A DNA mismatch repair protei. . . MSFVAGVIRRLDETVVNRIAAGEVIQRPANAIKEMIENCLDAKSTSIQVIV. . . EDLDIVCERFTTSKLQSFEDLASISTYGFRGEALASISHVAHVTITTKTAD. . . PRF

Ref. Seq: NCBI’s Derivative Sequence Database • Curated transcripts and proteins – reviewed – human, mouse, rat, fruit fly, zebrafish, arabidopsis microbial genomes (proteins), and more • Model transcripts and proteins • Assembled Genomic Regions (contigs) – human – mouse – rat – chicken – honeybee – sea urchin – zebrafish – cow – dog – black poplar • Chromosome records – Human genome – microbial srcdb_refseq[Properties] – organelle ftp: //ftp. ncbi. nih. gov/refseq/release/

Ref. Seq Accession Numbers m. RNAs and Proteins NM_123456 NP_123456 NR_123456 XM_123456 XP_123456 XR_123456 Gene Records NG_123456 Chromosome NC_123455 Assemblies NT_123456 NW_123456 Curated m. RNA Curated Protein Curated non-coding RNA Predicted m. RNA Predicted Protein Predicted non-coding RNA Reference Genomic Sequence also Microbial replicons, organelles Contig WGS Supercontig http: //www. ncbi. nlm. nih. gov/Ref. Seq/key. html
- Slides: 43