BIOL 697 2017 Lesson 7 Correspondence Analysis Detrended

BIOL 697 2017 Lesson 7 Correspondence Analysis & Detrended Correspondence Analysis

Deadlines n Ordination exercise (100 pts) Due 30 Oct Questions? n Sorted table exercise (100 pts) Due 6 Nov n Final oral presentations (200 pts) Due 15 Nov n Final written papers (300 pts) First draft due 20 Nov Final papers due 9 Dec

Questions about ordination? Simple terms can be hard to define Ordination vs. classification Direct vs. indirect ordination Ordination space vs. joint plot Species matrix vs. environmental matrix Redundancy in a species matrix Plot ordination vs. species ordination Floristic similarity Polar ordination: Defining the endpoints of the axes, Orthogonal axes

From Michael Palmer’s web page: (http: //ordination. okstate. edu. htm) Eigenanalysis n n Central to the mathematical discipline of linear (matrix) algebra. A thorough understanding requires a training in matrix algebra. Eigenanalysis is a mathematical operation on a square, symmetric matrix (same number of rows as columns) and is based on the process of reciprocal averaging. Principal Components Analysis, Correspondence Analysis (Reciprocal Averaging), DCA (Detrended Correspondence Analysis) are all examples of eigenanalysis-based ordination methods. It is possible to perform a eigenanalyses analytically (that is, get exact results) only for very small matrices (e. g. three rows and columns). For large matrices, eigenanalysis requires an iterative approach which eventually "closes in" on the answer (in most cases).

Principal Components Analysis (PCA) n n n Available in most statistical packages. The basic idea of PCA is to eliminate the high intercorrelation between variables (minimize redundency) and reduce the number of variables to a few uncorrelated (orthogonal) “super-variables” or “components” that explain all or most of the variation in the data. PCA is a "rigid rotation" of the data matrix: it does not change the positions of points relative to each other; it just changes the coordinate systems. In PCA, axes (components) are created such that the perpendicular distance from each object to the ordination axes is minimized. Axes are linear combinations of species/variables represented by eigenvectors, which are the weightings of the variables on each component An eigenanalysis consists of a series of eigenvectors, eigenvalues, and component scores. Eigenvector: The weighting of a variable vector on the component axis. ¨ Eigenvalue: The total amount of variation explained by a component axis. A measure of the strength of an axis (component). The sum of the eigenvalues equals the sum of the variance of all variables. ¨ Component scores: Coordinates of the samples or variables along each of the axes. ¨ n n PCA is good for data that are not in the same units, in which case data must be standardized to zero mean and unit variance (this is known in PCA as the correlation matrix). PCA is useful for analyzing samples in environmental space, but not useful for analyzing samples in species space.

PCA reduces many highly correlated variables into a few uncorrelated (orthogonal) variables or components. 1 Plots N Species 1 1 Raw data matrix 1 Plots N Components Matrix 5 M Components represent ‘new” super variables made up of highly correlated combinations of the original M number of species. • In the original raw data matrix each variable (species in the left matrix) has a score for each plot. The new components matrix (right) has the same number of plots as the original matrix but fewer variables (the components), and each plot has a score for each component. These are called the component scores.

Variable vectors: In a matrix of plots (= quadrats) and species (= variables): A G The variable vectors represent combined scores for all quadrats for a give variable. (The variable vector for variable A is OA. ) The vectors are all normalized to unit length of 1. F C O D B E 7

Correlation between variables: The cosine of the angles between each pair of vectors is equivalent to the correlations between each pair of vectors (derived from the correlation matrix, below). A G F C O α D The cosine of α is equivalent to the correlation between variables C and D (0. 93). Correlation matrix using Pearson’s Product Moment Correlation showing correlations between variables 8 B E
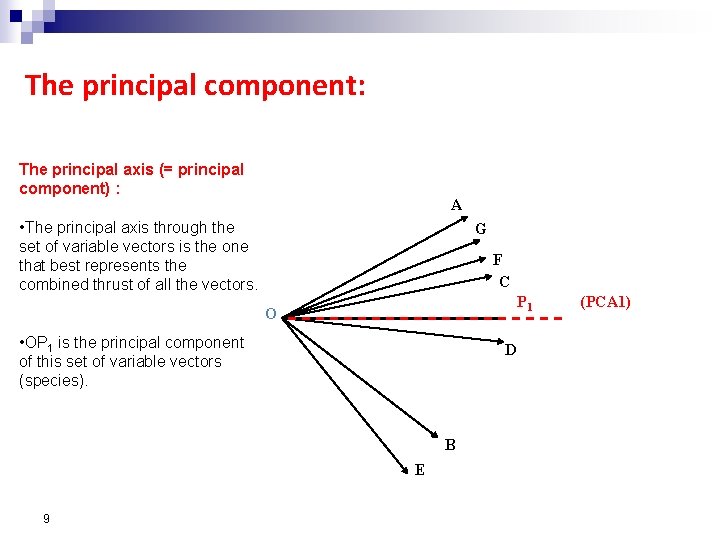
The principal component: The principal axis (= principal component) : A • The principal axis through the set of variable vectors is the one that best represents the combined thrust of all the vectors. G F C P 1 O • OP 1 is the principal component of this set of variable vectors (species). D B E 9 (PCA 1)

Eigenvectors for the first component (PCA 1) : • The projections of vectors (species) onto principal component (OP 1) are the eigenvectors (OA’, OB’, … OG’). • The lengths of the eigenvectors are the weighting of each vector (species) on the principal component axis. • Eigenvector scores close to 1 are strongly correlated with the Principal Component Axis 1 (PCA 1). Values O close to 0 are poorly correlated. A G F C A’ G’F’C’ E’ B’ D’ P 1 (PCA 1) D • The position of the principal axis OP 1 is found by finding the position at which the sum of the squared values of the all the eigenvectors is maximized. B E 10 EIGENVECTOR SCORES FOR PCA 1 OA’ =. 90 OB’ =. 71 OC’ =. 97 OD’ =. 99 OE’ =. 59 OF’ =. 95 OG’ =. 92 OP 1 = 1. 0

The eigenvalue of PCA 1: The eigenvalue = The sums of the squares of the eigenvectors Eigenvalue of Component 1 = (0. 90)2 + (0. 71)2 + (0. 97)2 + (0. 99)2 + (0. 59)2 + (0. 95)2 + (0. 92)2 = 5. 33 11 EIGENVECTORS FOR PCA 1 OA’ =. 90 OB’ =. 71 OC’ =. 97 OD’ =. 99 OE’ =. 59 OF’ =. 95 OG’ =. 92 OP 1 = 1. 0
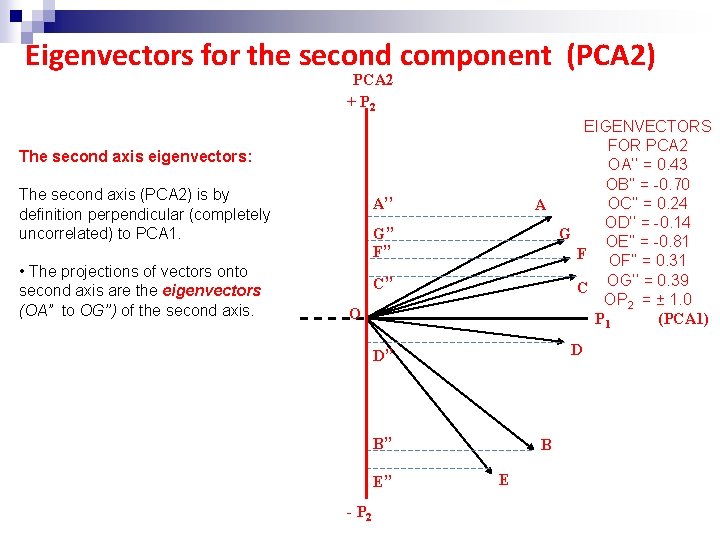
Eigenvectors for the second component (PCA 2) PCA 2 + P 2 EIGENVECTORS FOR PCA 2 OA’’ = 0. 43 OB’’ = -0. 70 OC’’ = 0. 24 A OD’’ = -0. 14 G OE’’ = -0. 81 F OF’’ = 0. 31 C OG’’ = 0. 39 OP 2 = ± 1. 0 P 1 (PCA 1) The second axis eigenvectors: The second axis (PCA 2) is by definition perpendicular (completely uncorrelated) to PCA 1. • The projections of vectors onto second axis are the eigenvectors (OA” to OG’’) of the second axis. A’’ G’’ F’’ C’’ O D D’’ B’’ E’’ - P 2 B E
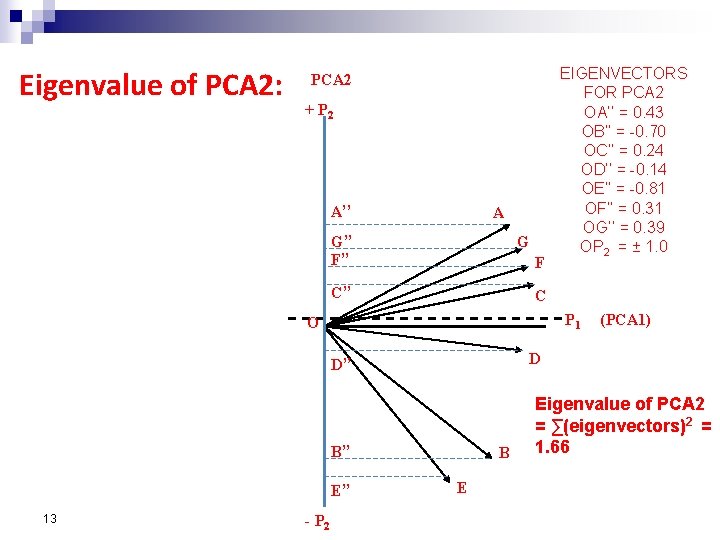
Eigenvalue of PCA 2: PCA 2 + P 2 A’’ A G’’ F’’ G F C’’ C P 1 O B’’ E’’ - P 2 (PCA 1) D D’’ 13 EIGENVECTORS FOR PCA 2 OA’’ = 0. 43 OB’’ = -0. 70 OC’’ = 0. 24 OD’’ = -0. 14 OE’’ = -0. 81 OF’’ = 0. 31 OG’’ = 0. 39 OP 2 = ± 1. 0 B E Eigenvalue of PCA 2 = ∑(eigenvectors)2 = 1. 66
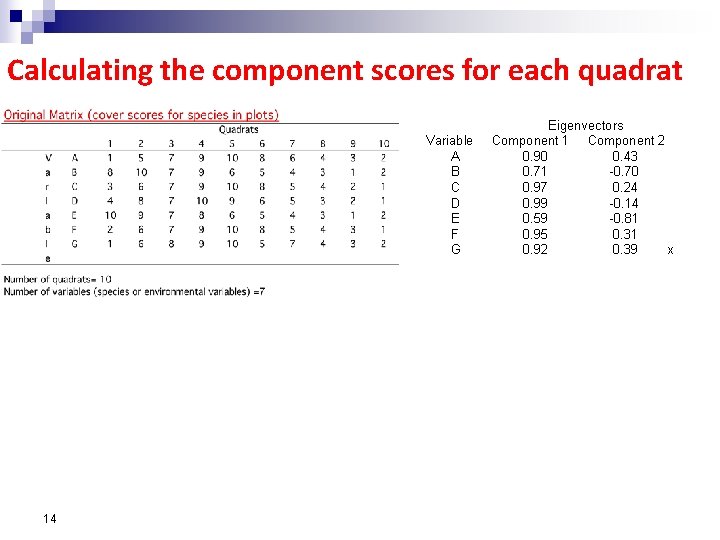
Calculating the component scores for each quadrat Variable A B C D E F G 14 Eigenvectors Component 1 Component 2 0. 90 0. 43 0. 71 -0. 70 0. 97 0. 24 0. 99 -0. 14 0. 59 -0. 81 0. 95 0. 31 0. 92 0. 39 x
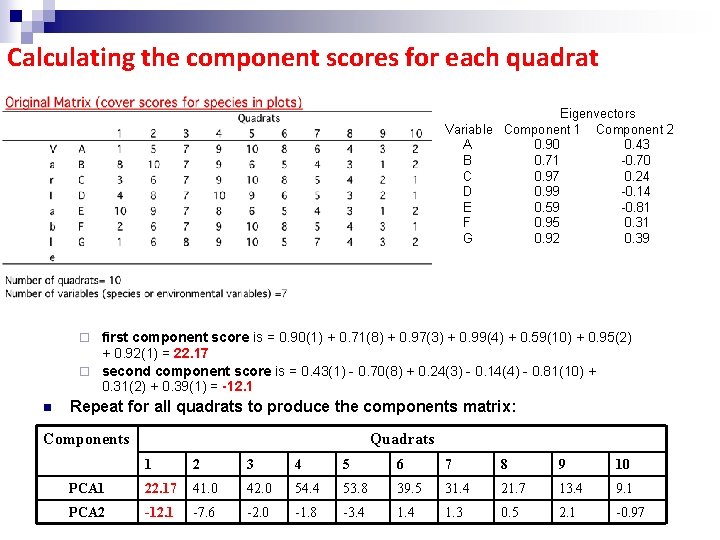
Calculating the component scores for each quadrat Eigenvectors Variable Component 1 Component 2 A 0. 90 0. 43 B 0. 71 -0. 70 C 0. 97 0. 24 D 0. 99 -0. 14 E 0. 59 -0. 81 F 0. 95 0. 31 G 0. 92 0. 39 n For quadrat 1: first component score is = 0. 90(1) + 0. 71(8) + 0. 97(3) + 0. 99(4) + 0. 59(10) + 0. 95(2) + 0. 92(1) = 22. 17 ¨ second component score is = 0. 43(1) - 0. 70(8) + 0. 24(3) - 0. 14(4) - 0. 81(10) + 0. 31(2) + 0. 39(1) = -12. 1 ¨ n Repeat for all quadrats to produce the components matrix: Components Quadrats 1 2 3 4 5 6 7 8 9 10 PCA 1 22. 17 41. 0 42. 0 54. 4 53. 8 39. 5 31. 4 21. 7 13. 4 9. 1 PCA 2 -12. 1 -7. 6 -2. 0 -1. 8 -3. 4 1. 3 0. 5 2. 1 -0. 97
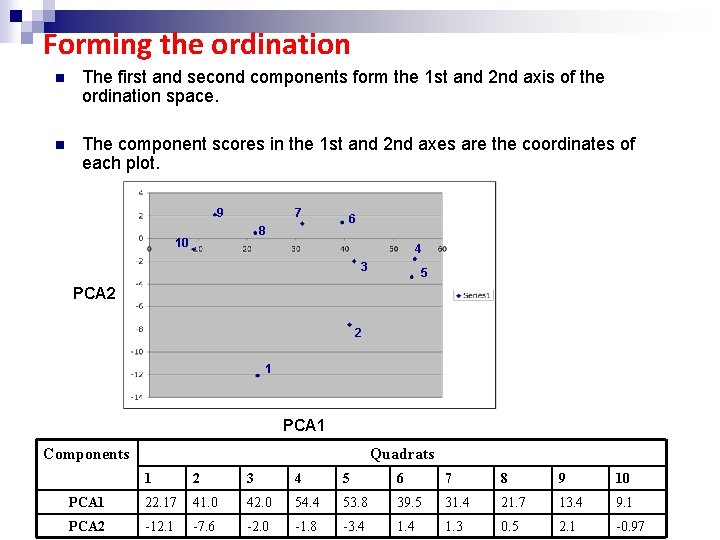
Forming the ordination n The first and second components form the 1 st and 2 nd axis of the ordination space. n The component scores in the 1 st and 2 nd axes are the coordinates of each plot. 9 7 8 10 6 4 3 5 PCA 2 2 1 PCA 1 Components Quadrats 1 2 3 4 5 6 7 8 9 10 PCA 1 22. 17 41. 0 42. 0 54. 4 53. 8 39. 5 31. 4 21. 7 13. 4 9. 1 PCA 2 -12. 1 -7. 6 -2. 0 -1. 8 -3. 4 1. 3 0. 5 2. 1 -0. 97

Beyond PCA: Why go further? n The linear model PCA usually doesn’t fit most species data. Species come and go across a gradient -- often with a Gausian or bellshaped distribution. If you sample across a broad enough gradient, this non-linear distribution patterns usually occurs. n Correspondence Analysis (CA) is another eigenvector-based technique, but it assumes a bell-shaped distribution, rather than a linear relationship among the variables. n The results are very similar to PCA, the big change is how the axes are derived.

Correspondence Analysis n Also known as Reciprocal Averaging n Basis for numerical classification, two-way indicator species analysis like TWINSPAN

Correspondence Analysis (Reciprocal averaging ) n Correspondence analysis works on much of the same principle as weighted averaging, but simultaneously orders both the columns and rows of a matrix. n Also, rather than forcing an external structure into the results by use of a weighting factor, it finds inherent structure within a data set. n Provides the basis for more advanced methods of ordinations developed after 1970. n Papers by Hill (1973, 1974) first made CA well known to ecologists.

Basic idea of Correspondence Analysis · To get a simultaneous ordination of the columns and rows (plots and species) · Like all ordination methods, it is based on the inherent structure of the data due to redundancy in the data, i. e. the co-occurrence of groups of species within the set of plots, and multiple plots showing the same groups of species. · The method of weighted averages is applied to a data matrix such that plot scores are derived from species scores and weightings. These are carried out successively using an iterative procedure. The scores eventually stabilize to get a set of scores for plots and a set of scores for species.

Calculation of the first axis 1. 2. 3. 4. 5. 6. Calculate the row and column totals Allocate weights to the species (can be arbitrary at first) Reciprocal averaging then commences The averaging process is then applied in reverse to give a new set of scores for the species using the sample scores To avoid calculation with very small numbers these new species scores are rescaled from 1 to 100 The species scores of the final iteration are the positions of the species along the first axis (0 to 100) of the ordination, and the plot scores are the positions of the plots along the first axis
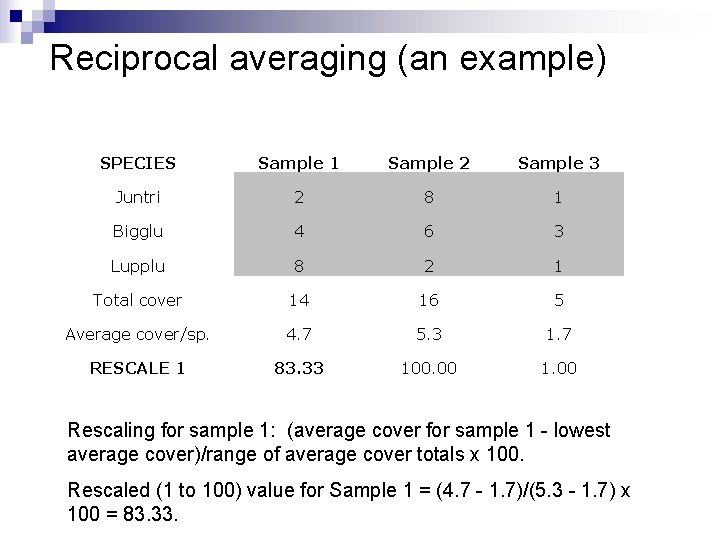
Reciprocal averaging (an example) SPECIES Sample 1 Sample 2 Sample 3 Juntri 2 8 1 Bigglu 4 6 3 Lupplu 8 2 1 Total cover 14 16 5 Average cover/sp. 4. 7 5. 3 1. 7 RESCALE 1 83. 33 100. 00 1. 00 Rescaling for sample 1: (average cover for sample 1 - lowest average cover)/range of average cover totals x 100. Rescaled (1 to 100) value for Sample 1 = (4. 7 - 1. 7)/(5. 3 - 1. 7) x 100 = 83. 33.
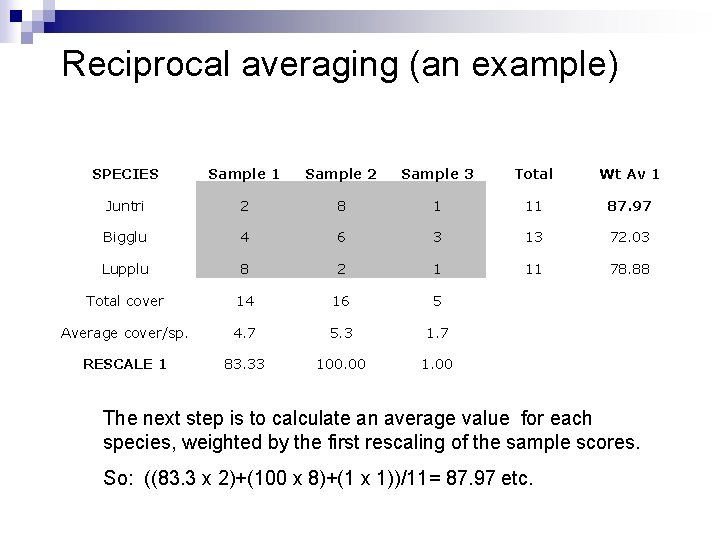
Reciprocal averaging (an example) SPECIES Sample 1 Sample 2 Sample 3 Total Wt Av 1 Juntri 2 8 1 11 87. 97 Bigglu 4 6 3 13 72. 03 Lupplu 8 2 1 11 78. 88 Total cover 14 16 5 Average cover/sp. 4. 7 5. 3 1. 7 RESCALE 1 83. 33 100. 00 1. 00 The next step is to calculate an average value for each species, weighted by the first rescaling of the sample scores. So: ((83. 3 x 2)+(100 x 8)+(1 x 1))/11= 87. 97 etc.
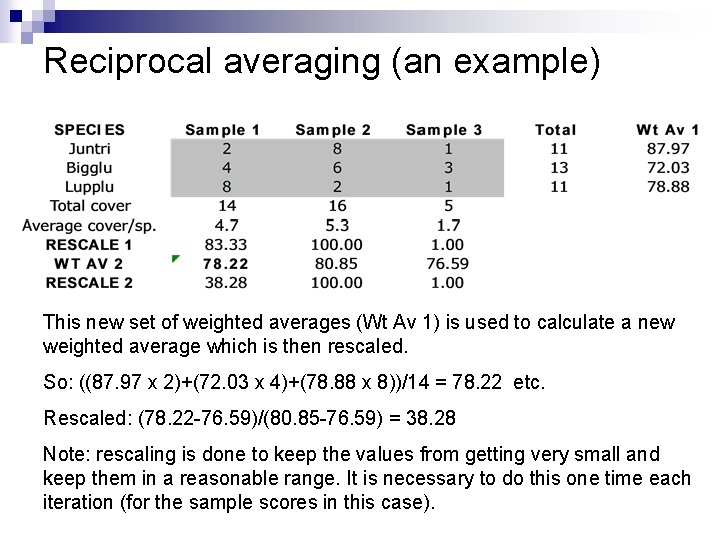
Reciprocal averaging (an example) This new set of weighted averages (Wt Av 1) is used to calculate a new weighted average which is then rescaled. So: ((87. 97 x 2)+(72. 03 x 4)+(78. 88 x 8))/14 = 78. 22 etc. Rescaled: (78. 22 -76. 59)/(80. 85 -76. 59) = 38. 28 Note: rescaling is done to keep the values from getting very small and keep them in a reasonable range. It is necessary to do this one time each iteration (for the sample scores in this case).
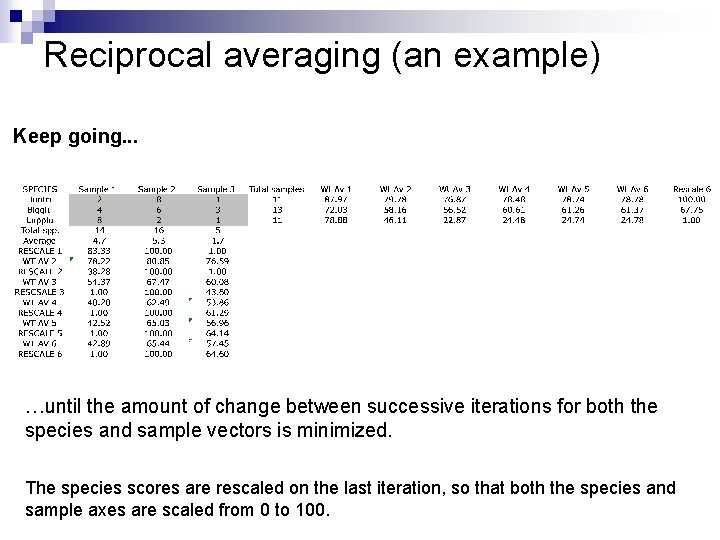
Reciprocal averaging (an example) Keep going. . . …until the amount of change between successive iterations for both the species and sample vectors is minimized. The species scores are rescaled on the last iteration, so that both the species and sample axes are scaled from 0 to 100.

Reciprocal averaging n If there is indeed a diagonal structure to the matrix (there is a major axis of variation in the data cloud) then the scores will stabilize. n Once they stabilize, you have the first axis!

Calculation of the eigenvalue n As with PCA, the eigenvalue can be thought of as a measure of the proportion of the total variation in the data explained by the axis. n It is obtained by taking the range of the unscaled scores in the final iteration and dividing by the range of the scaled scores. n In our example, the eigenvalue of the first axis = (65. 44 - 42. 89)/100 = 0. 236. n There is no specific eigenvalue cutoff for significant axes, but generally axes with eigenvalues >0. 25 should be discussed. In our trivial example with 3 plots and 3 species, the eigenvalue of the first axis is close to 0. 25, and no subsequent axis would be > 0. 25.

Calculation of the second axis · A second axis will be necessary if there are plots that lie close together in the first axis but which differ in species composition. · The second axis is extracted by the same iteration process, with one extra step in which the trial scores for the second axis are made uncorrelated with the first axis. In other words, the linear correlation with the first axis is removed. · This is done by taking the trial scores for the second axis and regressing them against the site scores for the first axis. The residuals from this regression are the new trial axis. This is done once for the sample scores at each iteration. · Similar procedure for subsequent axes
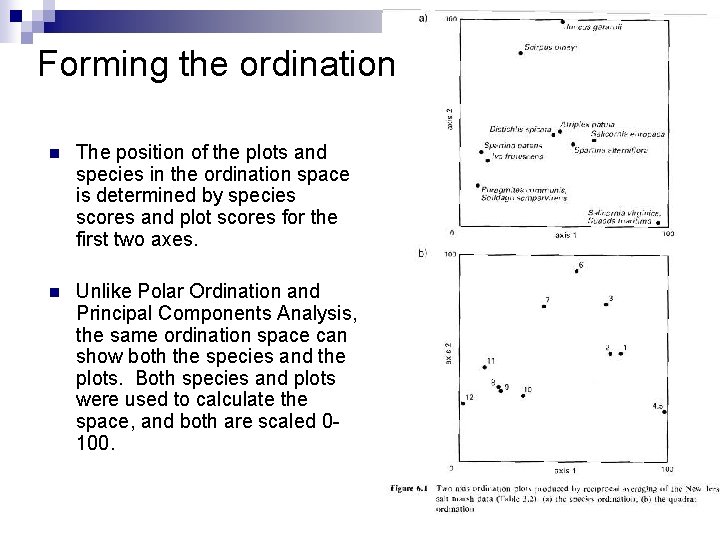
Forming the ordination n The position of the plots and species in the ordination space is determined by species scores and plot scores for the first two axes. n Unlike Polar Ordination and Principal Components Analysis, the same ordination space can show both the species and the plots. Both species and plots were used to calculate the space, and both are scaled 0100.

The math of Correspondence Analysis n In PC-ORD, correspondence analysis calculated using matrix algebra n Similar mathematically to a non-centered PCA standardized by species, but ¨ CA uses chi-square distances ¨ both plot and species weights are proportional to the plot or species total (double transformation)

Review of correspondence analysis (reciprocal averaging ) n Axis 1 ¨ ¨ ¨ n Assign arbitrary scores to each species Use these to calculate a weighted average for each plot Rescale these plot scores Use the plot scores to calculate a weighted average for each species Use the species scores to calculate a weighted average for each plot Continue until scores converge to a unique solution. Axis 2 Assign a new set of random scores to each species. ¨ Calculate a trial axis as above. ¨ Perform a multiple regression between the trial axis and the final axis obtained (above). ¨ Take the residual values as the new trial axis. ¨

Why does reciprocal averaging work? You start with meaningless numbers, then just average them in a fancy way, and expect to find a meaningful pattern! Well, it turns out that a meaningful pattern emerges because: n You get the same results no matter what your starting point of species scores (i. e. you are guaranteed to find "convergence") n The end result is that species scores and plot scores are maximally correlated with each other (that is, we could not hope for a better solution, given the data) n The eigenvalue is a measure of how well the species scores correspond with the plot scores (hence the name Correspondence Analysis) n This first axis usually turns out to be related to important environmental gradients http: //www. okstate. edu/artsci/botany/ordinate

Problems with Correspondence Analysis n THE “ARCH EFFECT” The second axis is a quadratic distortion of the first axis ¨ In data sets where there is no strong controlling gradient for the second axis, an arch effect is likely to occur ¨ In this case the second axis may have no ecological significance, but be simply a mathematical artifact ¨ Ecological effects may be found in the third or fourth axes ¨ n Best for data sets with long environmental gradients n COMPRESSION NEAR THE ENDS OF THE AXES ¨ Related to the arch effect and does not show the actual comings and goings of species along the first axis.
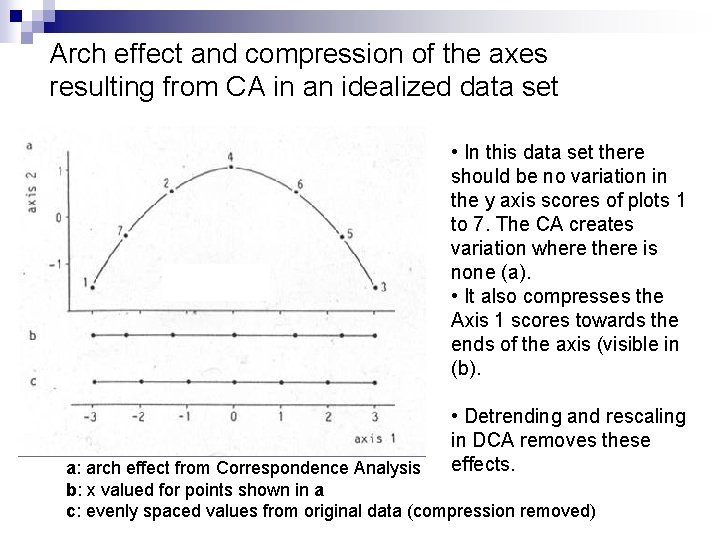
Arch effect and compression of the axes resulting from CA in an idealized data set • In this data set there should be no variation in the y axis scores of plots 1 to 7. The CA creates variation where there is none (a). • It also compresses the Axis 1 scores towards the ends of the axis (visible in (b). • Detrending and rescaling in DCA removes these effects. a: arch effect from Correspondence Analysis b: x valued for points shown in a c: evenly spaced values from original data (compression removed)
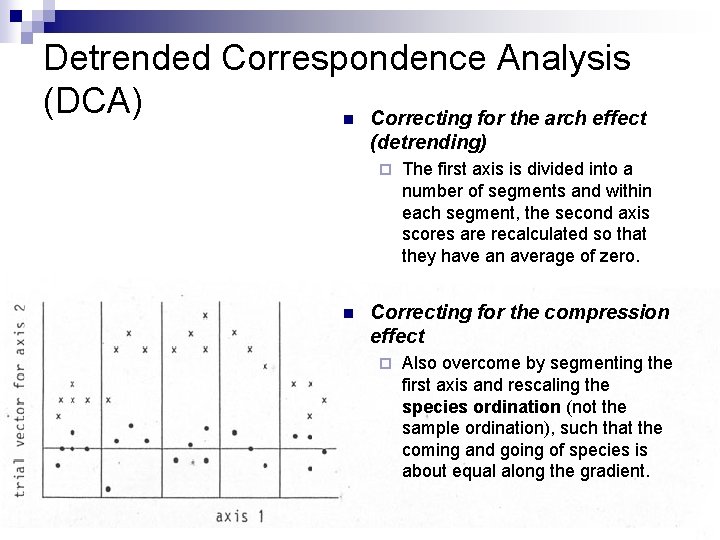
Detrended Correspondence Analysis (DCA) n Correcting for the arch effect (detrending) ¨ n The first axis is divided into a number of segments and within each segment, the second axis scores are recalculated so that they have an average of zero. Correcting for the compression effect ¨ Also overcome by segmenting the first axis and rescaling the species ordination (not the sample ordination), such that the coming and going of species is about equal along the gradient.

Detrended Correspondence Analysis (DCA): scaling of the axes n Scaling of the axes in sd units ¨ The axes in DCA are scaled into units that are the average standard deviation of species turnover (sd units). ¨ A 50% change in species composition occurs in a distance of about 1 SD unit. Species appear, rise to their modes, and disappear over a distance of about 4 SD units. ¨ The more SD units that occur along the axis the more change in species composition is shown. Thus, the sd units of DCA are a useful measure of beta diversity in the total data set.
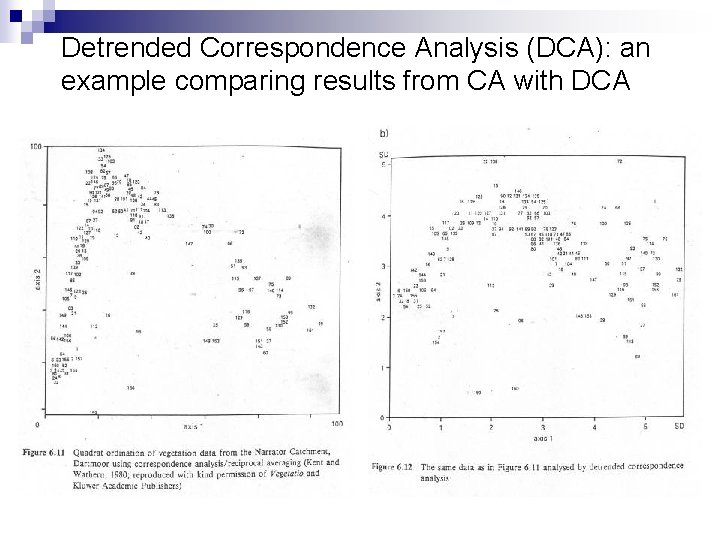
Detrended Correspondence Analysis (DCA): an example comparing results from CA with DCA

PC-ORD DCA output • List of residuals. Eigenvectors are found by iteration and the residuals indicate how close a given vector is to an eigenvector at each iteration. A vector is considered an eigenvector if it has a residual of less than 10 -7. • The eigenvalue is calculated at the final iteration. For the first axis it can be considered in the same way as in PCA, as an indicator of the amount of variance accounted for by the first axis. However, for higher axes, detrending in DCA destroys the correspondence between the eigenvalue and the structure along that axis. PC-ORD recommends using an after the fact coefficient of determination between Relative Euclidean distance in the unreduced species space and the Euclidean distance in the ordination space (Graph/ Ordination/ Statistics/ Percent of Variance).

PC-ORD DCA output (cont’) • Length of segments. These lengths indicate the degree of rescaling that occurred. For example, small values for the end segments suggests that a gradient was compressed at the ends before rescaling. • Length of the gradient. This is the total length of the axis before and after rescaling. The units are in sd units, a measure of beta-diversity. Four sd units is approximately equal to one complete turnover of species. Note: in PC-Ord 5, the length of the gradient is multiplied by 100.

PC-ORD DCA output (cont’) • At the end of the output are the species scores for the three axes, and their rankings along the first two axes. Along with the eigenvalue scores for each axis. • This followed by a similar table for the plot scores.

How do you tell if DCA is for you? n A general rule of thumb: if the axis is less than 2 units long, you should consider PCA, if it is more than 4 units long then DCA will likely be appropriate, and in between that… you will have to look harder at the data and make some “educated” choices. . .

Criticism of DCA n n n The method used for correcting the arch effect and compression have no empirical or theoretical basis (Wartenberg et. al 1987) The assumption that species turnover is constant or even along gradients is likely not true It removes any arch effect… even if there is really is an arch effect in your data set. Because it is done by arbitrarily dividing the axis into pieces, and then shifting those pieces up or down (to get a constant mean), the relationships within the segment are maintained, but other relationships can be “ripped” apart. Affected by outliers and gaps in gradient

DCA OVERALL n Despite these criticisms, DCA remains one of the most popular methods of indirect gradient analysis and is computationally very efficient. n It is the best method to to use when there are no environmental data. n The interpretation of results from DCA is best carried out with some knowledge of its limitations and comparison with other techniques. By doing this, one can get a better feel for patterns created by actual structure within the data set.
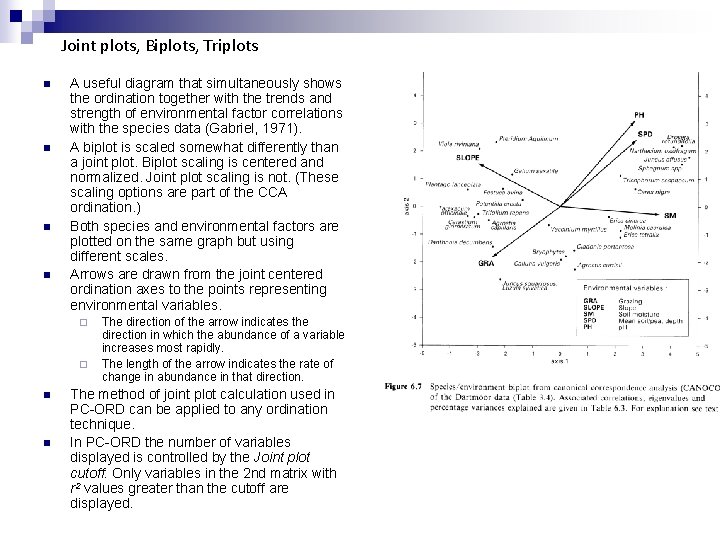
Joint plots, Biplots, Triplots n n A useful diagram that simultaneously shows the ordination together with the trends and strength of environmental factor correlations with the species data (Gabriel, 1971). A biplot is scaled somewhat differently than a joint plot. Biplot scaling is centered and normalized. Joint plot scaling is not. (These scaling options are part of the CCA ordination. ) Both species and environmental factors are plotted on the same graph but using different scales. Arrows are drawn from the joint centered ordination axes to the points representing environmental variables. ¨ ¨ n n The direction of the arrow indicates the direction in which the abundance of a variable increases most rapidly. The length of the arrow indicates the rate of change in abundance in that direction. The method of joint plot calculation used in PC-ORD can be applied to any ordination technique. In PC-ORD the number of variables displayed is controlled by the Joint plot cutoff. Only variables in the 2 nd matrix with r 2 values greater than the cutoff are displayed.
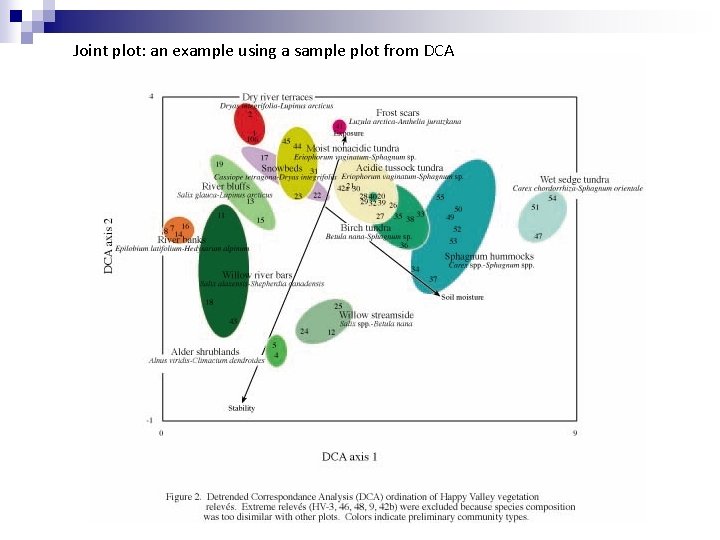
Joint plot: an example using a sample plot from DCA
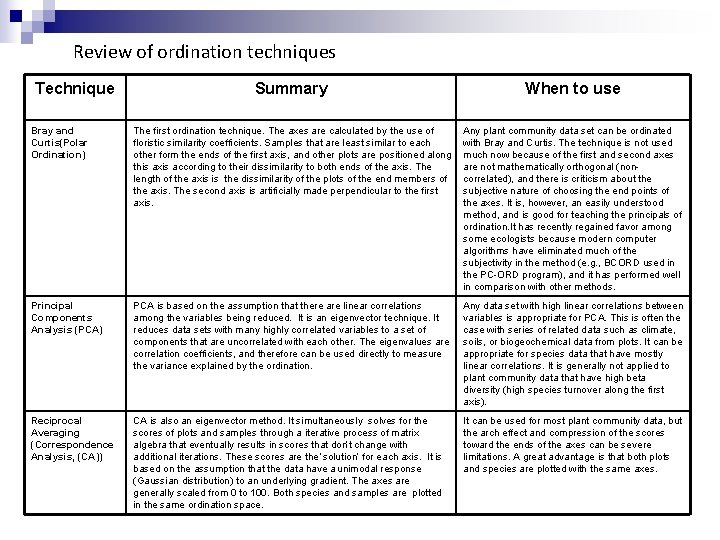
Review of ordination techniques Technique Summary When to use Bray and Curtis(Polar Ordination) The first ordination technique. The axes are calculated by the use of floristic similarity coefficients. Samples that are least similar to each other form the ends of the first axis, and other plots are positioned along this axis according to their dissimilarity to both ends of the axis. The length of the axis is the dissimilarity of the plots of the end members of the axis. The second axis is artificially made perpendicular to the first axis. Any plant community data set can be ordinated with Bray and Curtis. The technique is not used much now because of the first and second axes are not mathematically orthogonal (noncorrelated), and there is criticism about the subjective nature of choosing the end points of the axes. It is, however, an easily understood method, and is good for teaching the principals of ordination. It has recently regained favor among some ecologists because modern computer algorithms have eliminated much of the subjectivity in the method (e. g. , BCORD used in the PC-ORD program), and it has performed well in comparison with other methods. Principal Components Analysis (PCA) PCA is based on the assumption that there are linear correlations among the variables being reduced. It is an eigenvector technique. It reduces data sets with many highly correlated variables to a set of components that are uncorrelated with each other. The eigenvalues are correlation coefficients, and therefore can be used directly to measure the variance explained by the ordination. Any data set with high linear correlations between variables is appropriate for PCA. This is often the case with series of related data such as climate, soils, or biogeochemical data from plots. It can be appropriate for species data that have mostly linear correlations. It is generally not applied to plant community data that have high beta diversity (high species turnover along the first axis). Reciprocal Averaging (Correspondence Analysis, (CA)) CA is also an eigenvector method. It simultaneously solves for the scores of plots and samples through a iterative process of matrix algebra that eventually results in scores that don’t change with additional iterations. These scores are the ‘solution’ for each axis. It is based on the assumption that the data have a unimodal response (Gaussian distribution) to an underlying gradient. The axes are generally scaled from 0 to 100. Both species and samples are plotted in the same ordination space. It can be used for most plant community data, but the arch effect and compression of the scores toward the ends of the axes can be severe limitations. A great advantage is that both plots and species are plotted with the same axes.
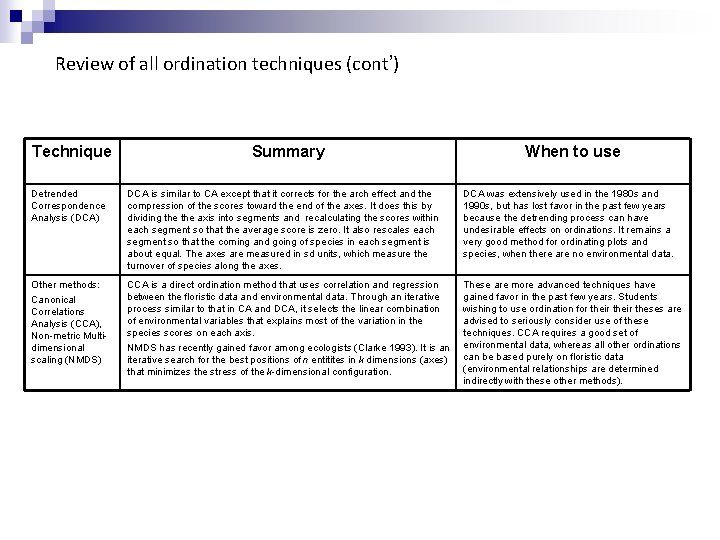
Review of all ordination techniques (cont’) Technique Summary When to use Detrended Correspondence Analysis (DCA) DCA is similar to CA except that it corrects for the arch effect and the compression of the scores toward the end of the axes. It does this by dividing the axis into segments and recalculating the scores within each segment so that the average score is zero. It also rescales each segment so that the coming and going of species in each segment is about equal. The axes are measured in sd units, which measure the turnover of species along the axes. DCA was extensively used in the 1980 s and 1990 s, but has lost favor in the past few years because the detrending process can have undesirable effects on ordinations. It remains a very good method for ordinating plots and species, when there are no environmental data. Other methods: Canonical Correlations Analysis (CCA), Non-metric Multidimensional scaling (NMDS) CCA is a direct ordination method that uses correlation and regression between the floristic data and environmental data. Through an iterative process similar to that in CA and DCA, it selects the linear combination of environmental variables that explains most of the variation in the species scores on each axis. NMDS has recently gained favor among ecologists (Clarke 1993). It is an iterative search for the best positions of n entitites in k dimensions (axes) that minimizes the stress of the k-dimensional configuration. These are more advanced techniques have gained favor in the past few years. Students wishing to use ordination for their theses are advised to seriously consider use of these techniques. CCA requires a good set of environmental data, whereas all other ordinations can be based purely on floristic data (environmental relationships are determined indirectly with these other methods).
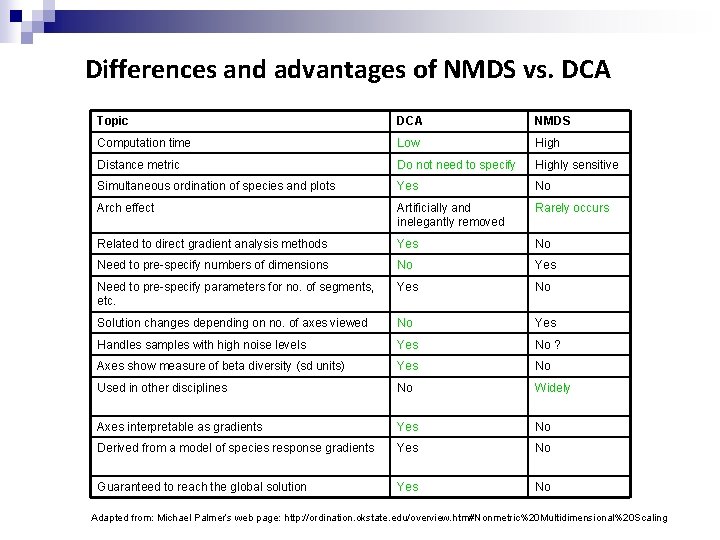
Differences and advantages of NMDS vs. DCA Topic DCA NMDS Computation time Low High Distance metric Do not need to specify Highly sensitive Simultaneous ordination of species and plots Yes No Arch effect Artificially and inelegantly removed Rarely occurs Related to direct gradient analysis methods Yes No Need to pre-specify numbers of dimensions No Yes Need to pre-specify parameters for no. of segments, etc. Yes No Solution changes depending on no. of axes viewed No Yes Handles samples with high noise levels Yes No ? Axes show measure of beta diversity (sd units) Yes No Used in other disciplines No Widely Axes interpretable as gradients Yes No Derived from a model of species response gradients Yes No Guaranteed to reach the global solution Yes No Adapted from: Michael Palmer’s web page: http: //ordination. okstate. edu/overview. htm#Nonmetric%20 Multidimensional%20 Scaling

Recommended reading Schickhoff, U. , M. D. Walker, and D. A. Walker. 2002. Riparian willow communities on the Arctic Slope of Alaska and their environmental relationships: A classification and ordination analysis. Phytocoenologia 32: 145 -204. http: //www. geobotany. org/library/pubs/Schickhoff. U 2002_pcoe_32_145. pdf
- Slides: 49