BIOL 2020 Chapter 4 The shape and structure

BIOL 2020 Chapter 4 The shape and structure of proteins How proteins work How Proteins are Controlled How Proteins are studied (labs 2 -3) Familiarize yourself with panels 4 -3, 4 -4 and 4 -5

Ch. 4 - is all about how proteins form, what they do, and how they are regulated. The big picture you need to understand is: 1 o structure contains all the info needed to get fully functional protein machines. How? - the order of aa (1 o structure), determines types of 2 o and 3 o structures that are possible. Genes tell cells order of aa’s in a protein, but this order also has all the information for the higher order interactions the proteins will make and therefore: 1 - how it will fold, 2 - what it will do, and 3 -where it will be located in the cell. Genes are DNA sequences selected for over millions of years that reliably fold into “same” stable structure every time. (keeping in mind that 2 o and 3 o structure is constantly variable depending on situation (inhibitors , bad p. H, phosphorylation, etc) It takes E to form original 1 o structure (peptide bonds of aa’s in specific order) but once that is done, all the other levels form spontaneously, due to noncovalent interactions (not energy requiring!) so self assembling

Theme #1 THE SHAPE AND STRUCTURE OF PROTEINS • The Shape of a Protein Is Specified by Its Amino Acid Sequence • Proteins fold into a conformation of lowest energy • protein folding landscape • Proteins come in a wide variety of complicated shapes • The Alpha Helix and the Beta Sheet are common 2 o structure folding patterns • Helices form readily in biological structures • Beta Sheets form rigid structures at the core of many proteins • Proteins have several levels of organization (1 o 2 o 3 o 4 o) • Few of the many possible polypeptide chains will be useful (only specific possibilities are selected for) • Proteins can be classified into Families • Large protein molecules often contain more than one polypeptide chain (4 o structure) • Proteins can assemble into Filaments, Sheets, or “Spheres“(globular) • Some types of proteins have elongated fibrous shapes • Extracellular proteins are often stabilized by covalent cross-linkages (disulfide bonds)

Proteins do (just about) everything! Proteins include a diversity of structures, resulting in a wide range of functions • FUNCTIONS: • used for structure and for metabolism (enzymes) • Proteins account for more than 50% of the dry mass of most cells – Protein functions include structural support, storage, transport, cellular communications, movement, defense against foreign substances, etc. Most cell functions depend on proteins! • We are built of proteins and require functioning proteins to live (enzymes, transport proteins, structural proteins, etc. ) This is how DNA controls cells: DNA codes for proteins! And proteins do pretty much everything else.

Proteins have numerous functions They are responsible for giving cells shape and structure, and for enabling them to do what they do.



General functions of proteins include: 1. Enzyme (catalysis and anabolic rxns) 2. Defense 3. Transport 4. Support 5. Motion 6. Regulation (ligands and receptors) 7. Storage Serve many functions since can have many shapes. What determines the shape of a specific protein? Each protein’s function is determined by its 3 D shape. (ie, its 3 o and 4 o structure)

Ch. 4 - is all about how proteins form, what they do, and how they are regulated. The big picture you need to understand is: 1 o structure contains all the info needed to get fully functional protein machines. How? - the order of aa (1 o structure), determines types of 2 o and 3 o structures that are possible. Genes tell cells order of aa’s in a protein, but this order also has all the information for the higher order interactions the proteins will make and therefore: 1 - how it will fold, 2 - what it will do, and 3 -where it will be located in the cell. Genes are DNA sequences selected for over millions of years that reliably fold into “same” stable structure every time. (keeping in mind that 2 o and 3 o structure is constantly variable depending on situation (inhibitors , bad p. H, phosphorylation, etc) It takes E to form original 1 o structure (peptide bonds of aa’s in specific order) but once that is done, all the other levels form spontaneously, due to noncovalent interactions (not energy requiring!) so self assembling

Proteins can have variable shapes How many active genes are needed to form functional hemoglobin? How many polypeptides are in one molecule of hemoglobin? What shape is hemoglobin? How does this affect its function? What level of protein structure does it exhibit? What determines the primary structure of each subunit? What causes subunits to interact with each other? How many cofactors are necessary for 1 hemoglobin to function? What emergent property does hemoglobin have that β hemoglobin lacks? What codes for the 3 D structure of hemoglobin? For its function? HOW? A globular protein made of 4 subunits Can be sheets, globular or filamentous; small or large, etc Can consist of one or multiple subunits (ie, 3 o or 4 o) Can require other molecules for function (cofactors) Specific shape and binding properties of each results in their specific function Protein shape, and thus function, is ultimately determined by its 1 o structure Hemoglobin
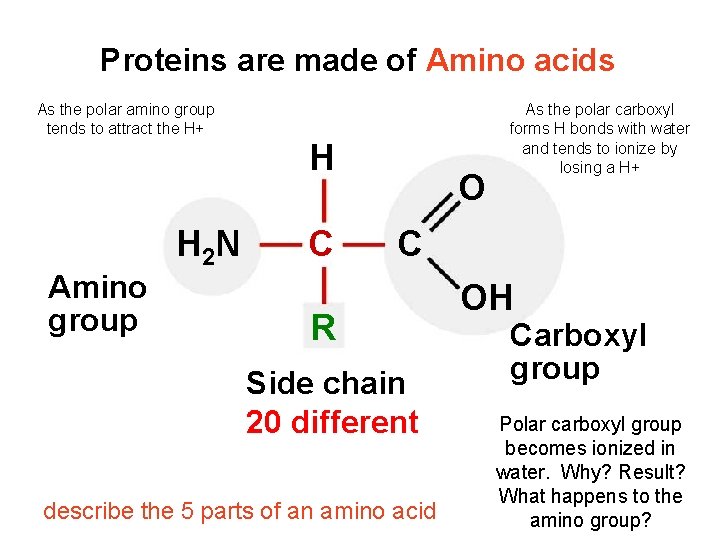
Proteins are made of Amino acids Proteins are made of Amino Acids As the polar amino group tends to attract the H+ Amino group H 2 N H C O As the polar carboxyl forms H bonds with water and tends to ionize by losing a H+ C R Side chain 20 different describe the 5 parts of an amino acid OH Carboxyl group Polar carboxyl group becomes ionized in water. Why? Result? What happens to the amino group?
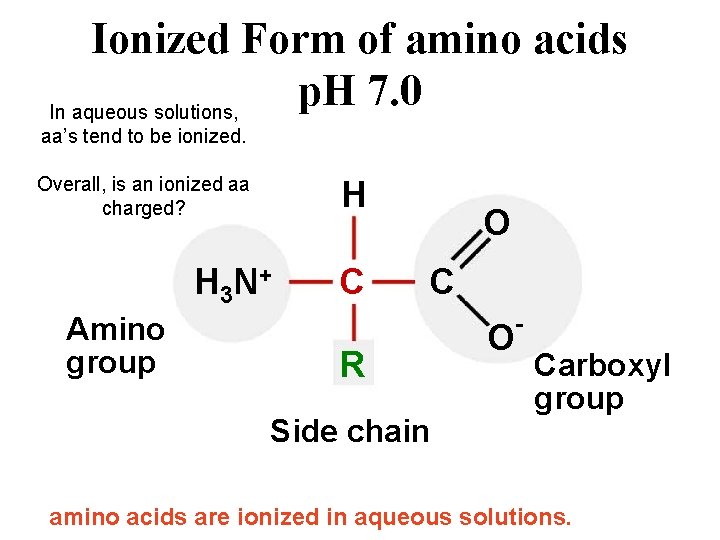
Ionized Form of amino acids p. H 7. 0 In aqueous solutions, aa’s tend to be ionized. Overall, is an ionized aa charged? H H 3 N + Amino group C O C R Side chain O - Carboxyl group amino acids are ionized in aqueous solutions.
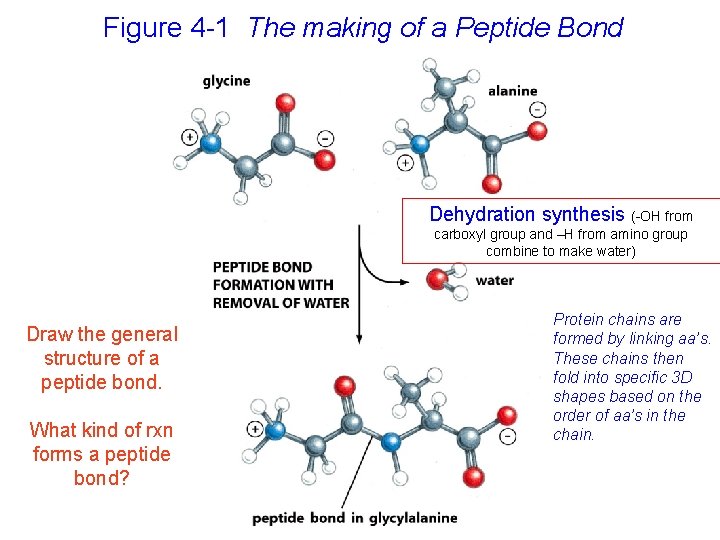
Figure 4 -1 The making of a Peptide Bond Dehydration synthesis (-OH from carboxyl group and –H from amino group combine to make water) Draw the general structure of a peptide bond. What kind of rxn forms a peptide bond? Protein chains are formed by linking aa’s. These chains then fold into specific 3 D shapes based on the order of aa’s in the chain.
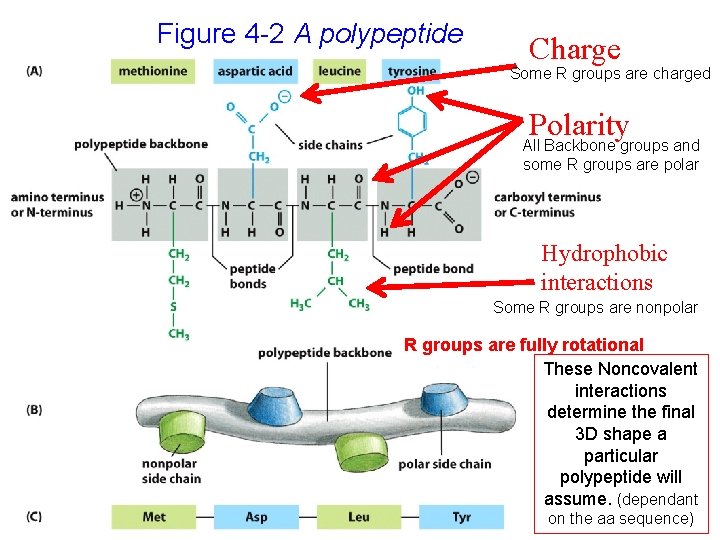
Figure 4 -2 A polypeptide Charge Some R groups are charged Polarity All Backbone groups and some R groups are polar Hydrophobic interactions Some R groups are nonpolar R groups are fully rotational These Noncovalent interactions determine the final 3 D shape a particular polypeptide will assume. (dependant on the aa sequence)
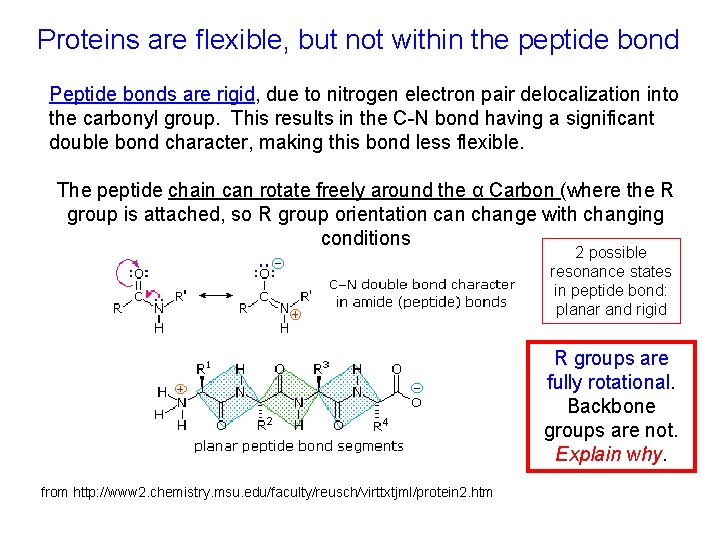
Proteins are flexible, but not within the peptide bond Peptide bonds are rigid, due to nitrogen electron pair delocalization into the carbonyl group. This results in the C-N bond having a significant double bond character, making this bond less flexible. The peptide chain can rotate freely around the α Carbon (where the R group is attached, so R group orientation can change with changing conditions 2 possible resonance states in peptide bond: planar and rigid R groups are fully rotational. Backbone groups are not. Explain why. from http: //www 2. chemistry. msu. edu/faculty/reusch/virttxtjml/protein 2. htm

How different amino acids interact with others in the chain, determines the final 3 D shape of the protein. 4 levels of protein structure: 1 - Primary structure: is the order of aa’s in the polypeptide chain. Ribosomes build the 1 o structure one aa at a time. It takes energy to form the peptide bonds in the chain. The growing polypeptide then spontaneously folds into its final 3 D shape, due to 2 o, 3 o and sometimes 4 o structure. 2 - Secondary structure: is due to hydrogen bonding between backbone groups (α helices and β sheets, turns) 3 - Tertiary structure: is due to numerous noncovalent interactions between R groups with each other, or with backbone groups. 3 o structure is also stabilized by disulfide bonds (which are covalent bonds). Final shape may be globular, filamentous, hollow, flat, etc. Some (but not all) proteins also have 4 - Quaternary structure: due to interaction of multiple polypeptide strands to form one functional protein (ex. Hemoglobin, collagen)
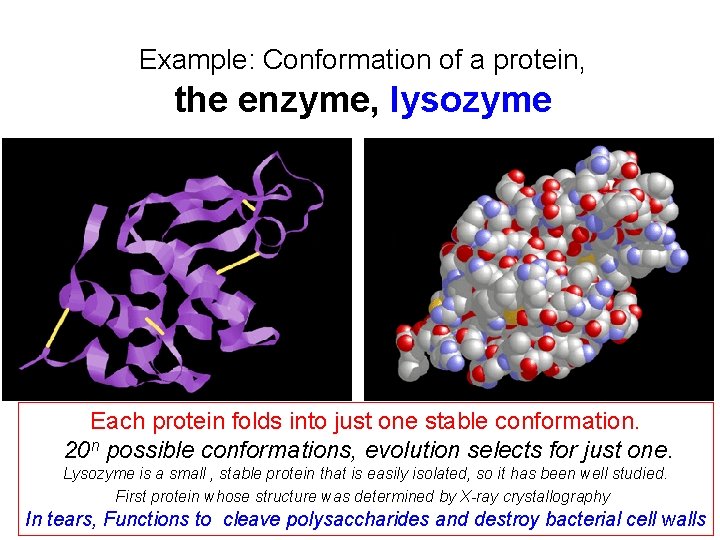
Example: Conformation of a protein, the enzyme, lysozyme Each protein folds into just one stable conformation. 20 n possible conformations, evolution selects for just one. Lysozyme is a small , stable protein that is easily isolated, so it has been well studied. First protein whose structure was determined by X-ray crystallography In tears, Functions to cleave polysaccharides and destroy bacterial cell walls
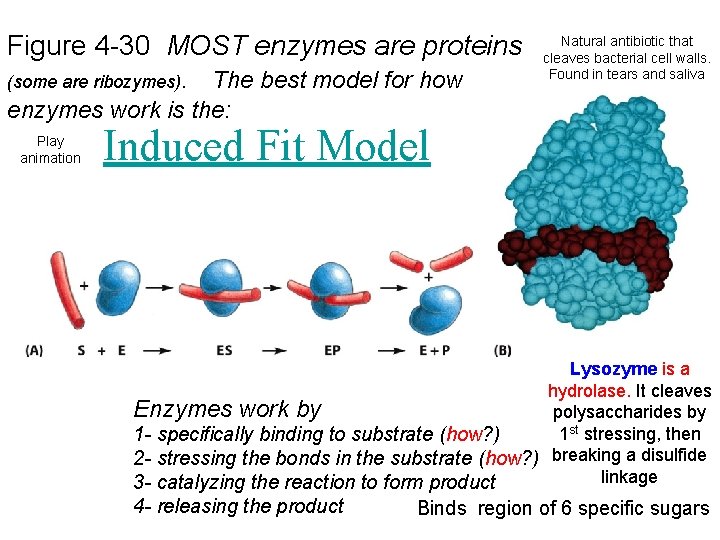
Figure 4 -30 MOST enzymes are proteins (some are ribozymes). The best model for how Natural antibiotic that cleaves bacterial cell walls. Found in tears and saliva enzymes work is the: Play animation Induced Fit Model Lysozyme is a hydrolase. It cleaves Enzymes work by polysaccharides by 1 st stressing, then 1 - specifically binding to substrate (how? ) 2 - stressing the bonds in the substrate (how? ) breaking a disulfide linkage 3 - catalyzing the reaction to form product 4 - releasing the product Binds region of 6 specific sugars
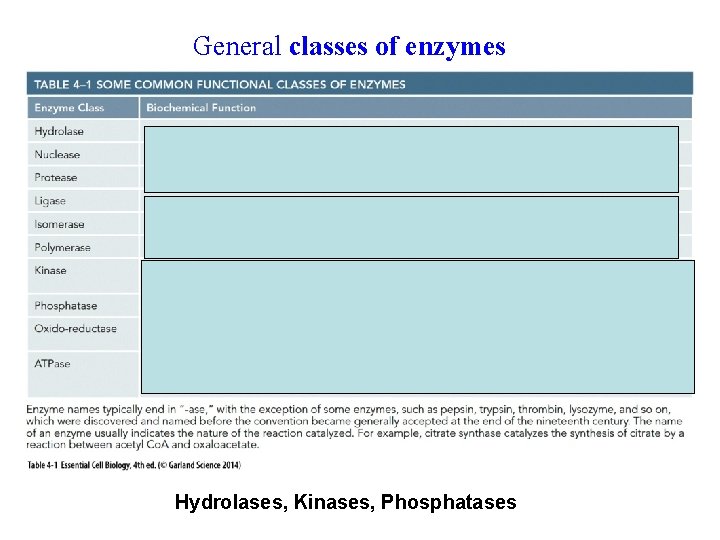
General classes of enzymes peroxidase Hydrolases, Kinases, Phosphatases
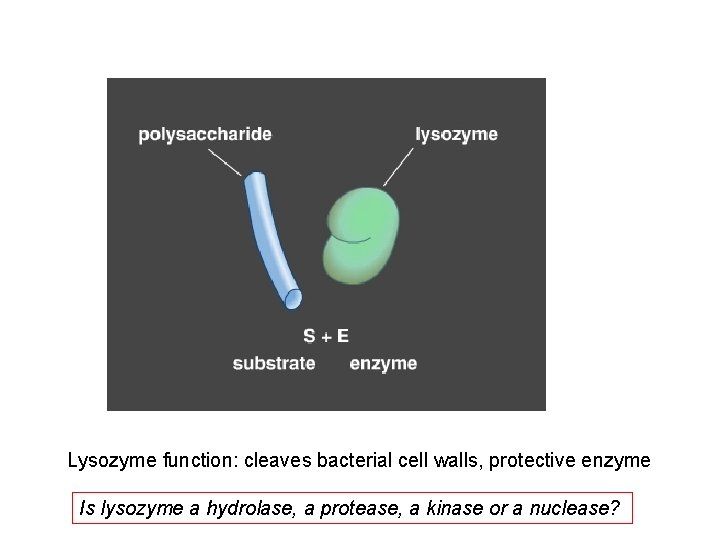
Lysozyme function: cleaves bacterial cell walls, protective enzyme Is lysozyme a hydrolase, a protease, a kinase or a nuclease?
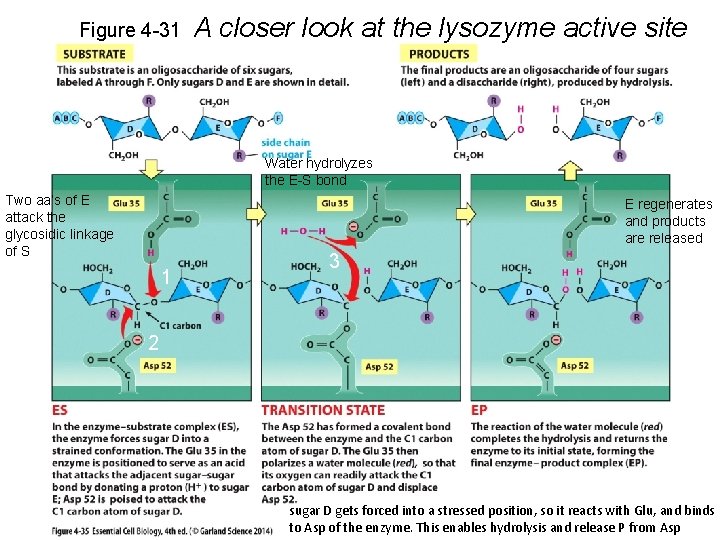
Figure 4 -31 A closer look at the lysozyme active site Water hydrolyzes the E-S bond Two aa’s of E attack the glycosidic linkage of S E regenerates and products are released 1 3 2 sugar D gets forced into a stressed position, so it reacts with Glu, and binds to Asp of the enzyme. This enables hydrolysis and release P from Asp
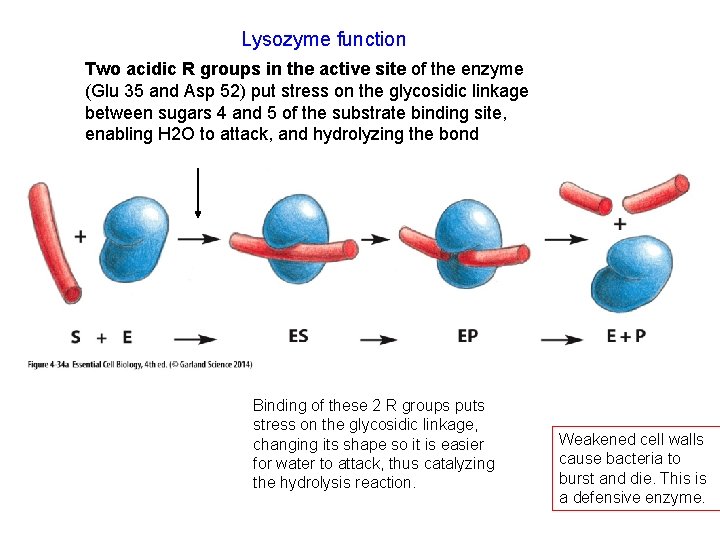
Lysozyme function Two acidic R groups in the active site of the enzyme (Glu 35 and Asp 52) put stress on the glycosidic linkage between sugars 4 and 5 of the substrate binding site, enabling H 2 O to attack, and hydrolyzing the bond Binding of these 2 R groups puts stress on the glycosidic linkage, changing its shape so it is easier for water to attack, thus catalyzing the hydrolysis reaction. Weakened cell walls cause bacteria to burst and die. This is a defensive enzyme.
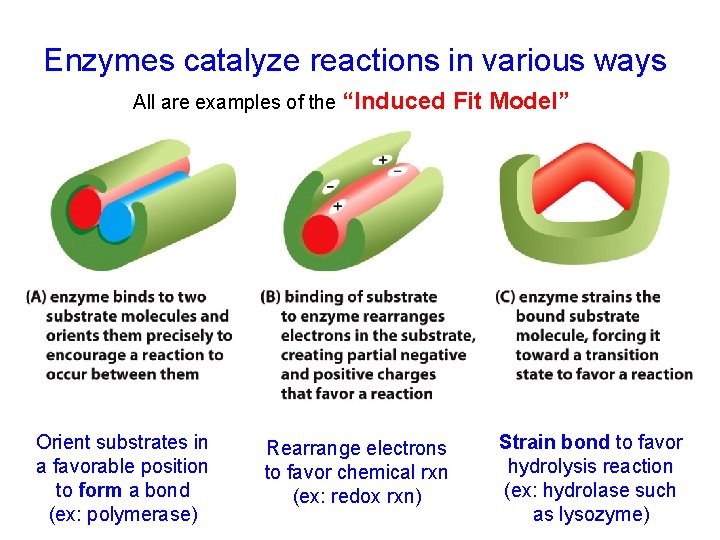
Enzymes catalyze reactions in various ways All are examples of the “Induced Orient substrates in a favorable position to form a bond (ex: polymerase) Rearrange electrons to favor chemical rxn (ex: redox rxn) Fit Model” Strain bond to favor hydrolysis reaction (ex: hydrolase such as lysozyme)

How do proteins get their specific 3 D shape, so that they perform their correct function? For example, explain how the active site of an enzyme forms? Why does an enzyme’s active site bind to a specific substrate?
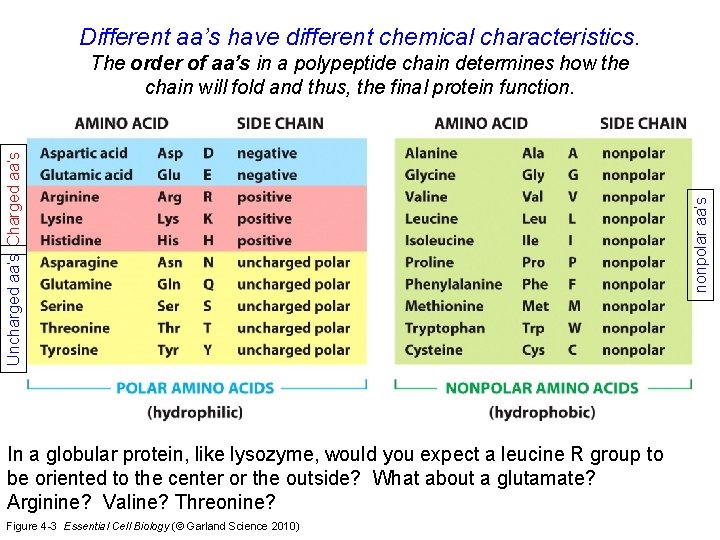
Different aa’s have different chemical characteristics. In a globular protein, like lysozyme, would you expect a leucine R group to be oriented to the center or the outside? What about a glutamate? Arginine? Valine? Threonine? Figure 4 -3 Essential Cell Biology (© Garland Science 2010) nonpolar aa’s Uncharged aa’s Charged aa’s The order of aa’s in a polypeptide chain determines how the chain will fold and thus, the final protein function.
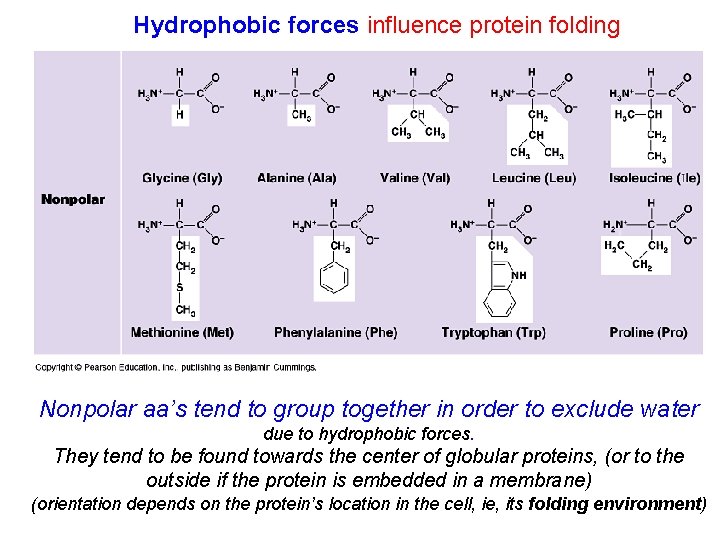
Hydrophobic forces influence protein folding Nonpolar aa’s tend to group together in order to exclude water due to hydrophobic forces. They tend to be found towards the center of globular proteins, (or to the outside if the protein is embedded in a membrane) (orientation depends on the protein’s location in the cell, ie, its folding environment)
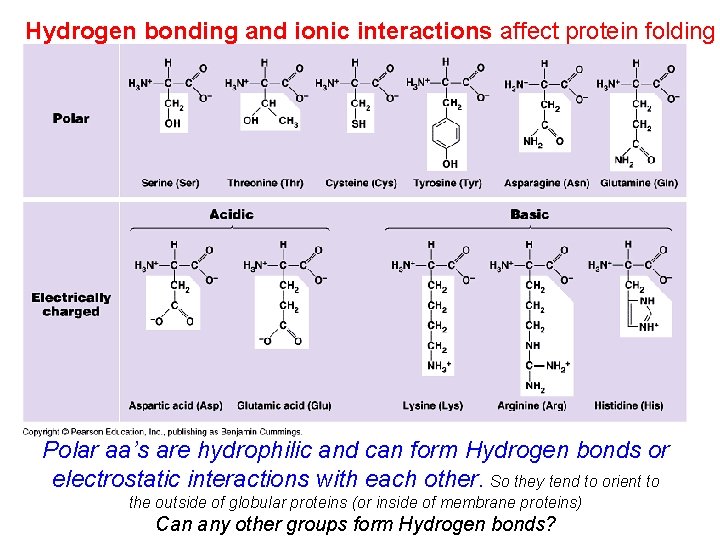
Hydrogen bonding and ionic interactions affect protein folding Polar aa’s are hydrophilic and can form Hydrogen bonds or electrostatic interactions with each other. So they tend to orient to the outside of globular proteins (or inside of membrane proteins) Can any other groups form Hydrogen bonds?

3 key ideas!!!! Figure 2 -31 1 - Shape is Everything Each protein folds into one stable, functional conformation. The amino acid sequence (1 o structure) contains all the information telling the protein how to fold into the correct shape (in various circumstances) Protein structure is unstable and changing when cellular conditions are not correct for the protein to be functional
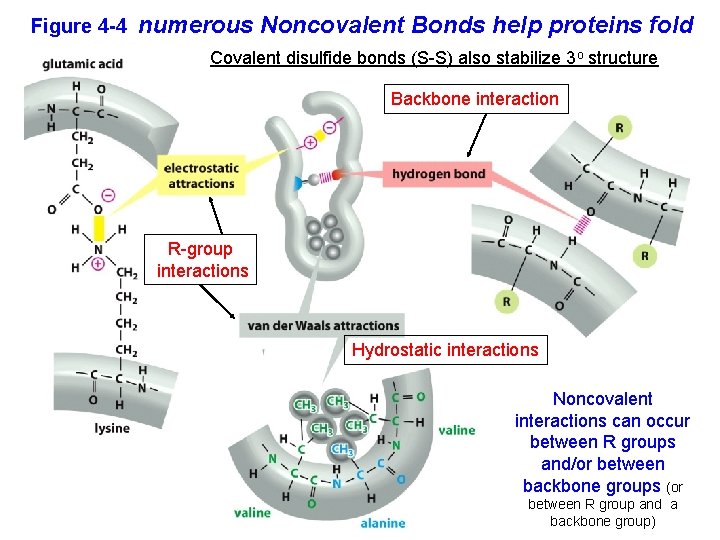
Figure 4 -4 numerous Noncovalent Bonds help proteins fold Covalent disulfide bonds (S-S) also stabilize 3 o structure Backbone interaction R-group interactions Hydrostatic interactions Noncovalent interactions can occur between R groups and/or between backbone groups (or between R group and a backbone group)
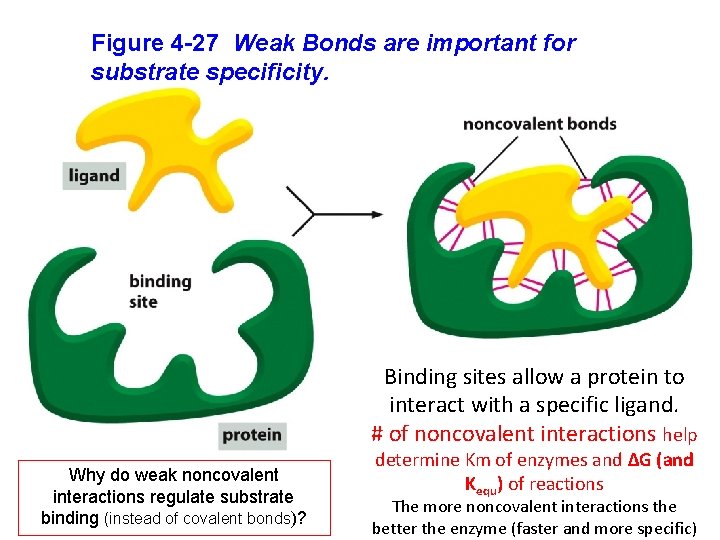
Figure 4 -27 Weak Bonds are important for substrate specificity. Binding sites allow a protein to interact with a specific ligand. # of noncovalent interactions help Why do weak noncovalent interactions regulate substrate binding (instead of covalent bonds)? determine Km of enzymes and ΔG (and Kequ) of reactions The more noncovalent interactions the better the enzyme (faster and more specific)
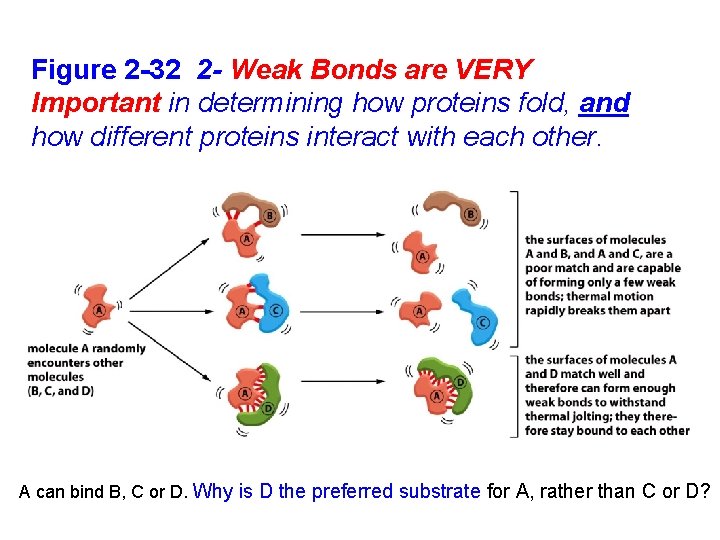
Figure 2 -32 2 - Weak Bonds are VERY Important in determining how proteins fold, and how different proteins interact with each other. A can bind B, C or D. Why is D the preferred substrate for A, rather than C or D?
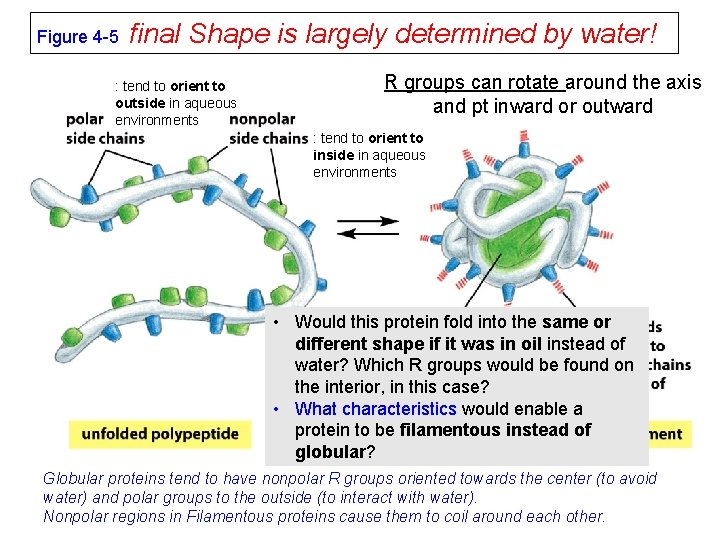
Figure 4 -5 final Shape is largely determined by water! : tend to orient to outside in aqueous environments R groups can rotate around the axis and pt inward or outward : tend to orient to inside in aqueous environments • Would this protein fold into the same or different shape if it was in oil instead of water? Which R groups would be found on the interior, in this case? • What characteristics would enable a protein to be filamentous instead of globular? Globular proteins tend to have nonpolar R groups oriented towards the center (to avoid water) and polar groups to the outside (to interact with water). Nonpolar regions in Filamentous proteins cause them to coil around each other.
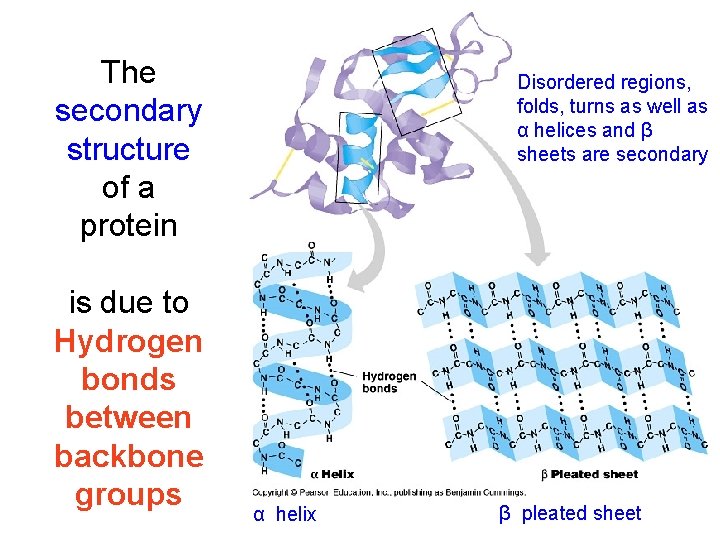
The secondary structure of a protein is due to Hydrogen bonds between backbone groups Disordered regions, folds, turns as well as α helices and β sheets are secondary α helix β pleated sheet

Movie 4. 2 alpha helix 2 o structures can occur anywhere in a protein, since they are due to interactions between backbone groups. (why) However, Which type of 2 o structures form, and where they form is codependent on the types of 3 o interactions forming between R groups. In other words, 2 o and 3 o structure form simultaneously, as the protein is being built, an thy can change as the protein grows, until they reach their final stable conformation [see: protein folding landscape]
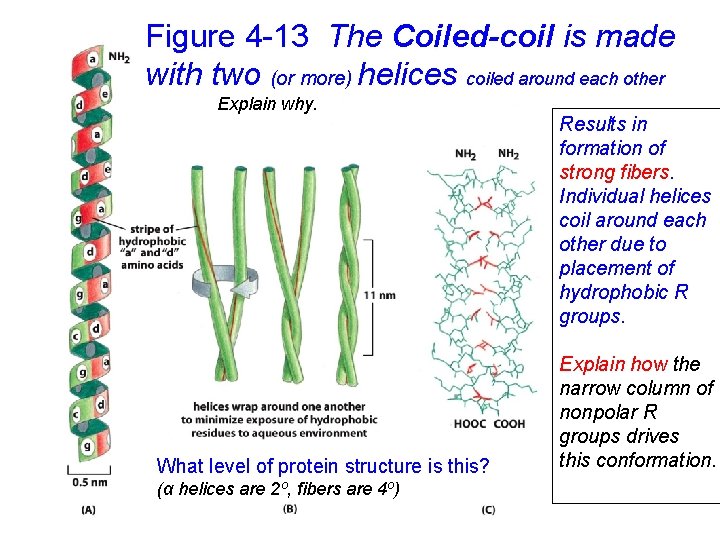
Figure 4 -13 The Coiled-coil is made with two (or more) helices coiled around each other Explain why. What level of protein structure is this? (α helices are 2 o, fibers are 4 o) Results in formation of strong fibers. Individual helices coil around each other due to placement of hydrophobic R groups. Explain how the narrow column of nonpolar R groups drives this conformation.
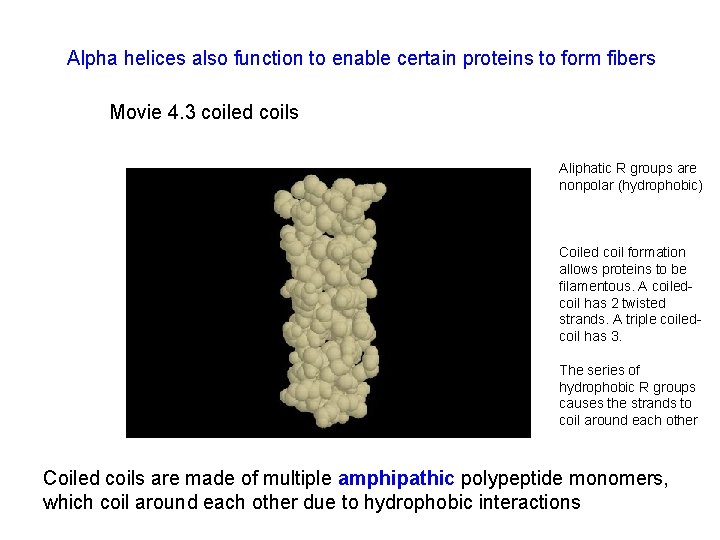
Alpha helices also function to enable certain proteins to form fibers Movie 4. 3 coiled coils Aliphatic R groups are nonpolar (hydrophobic) Coiled coil formation allows proteins to be filamentous. A coiledcoil has 2 twisted strands. A triple coiledcoil has 3. The series of hydrophobic R groups causes the strands to coil around each other Coiled coils are made of multiple amphipathic polypeptide monomers, which coil around each other due to hydrophobic interactions
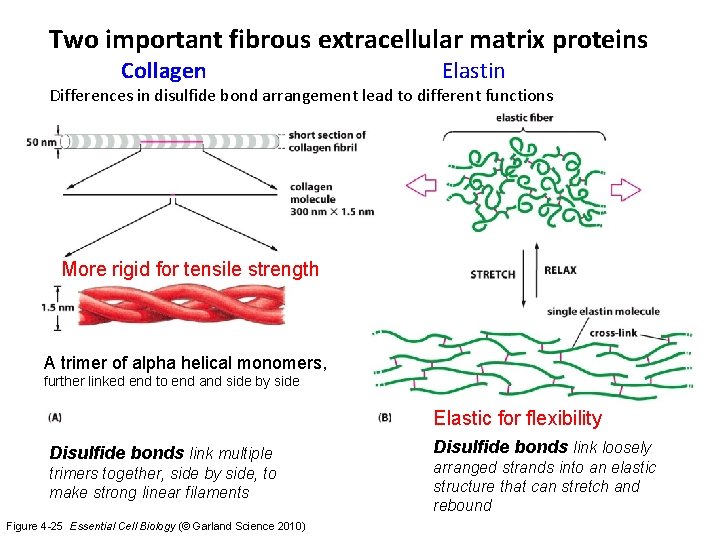
Two important fibrous extracellular matrix proteins Collagen Elastin Differences in disulfide bond arrangement lead to different functions More rigid for tensile strength A trimer of alpha helical monomers, further linked end to end and side by side Elastic for flexibility Disulfide bonds link multiple trimers together, side by side, to make strong linear filaments Figure 4 -25 Essential Cell Biology (© Garland Science 2010) Disulfide bonds link loosely arranged strands into an elastic structure that can stretch and rebound
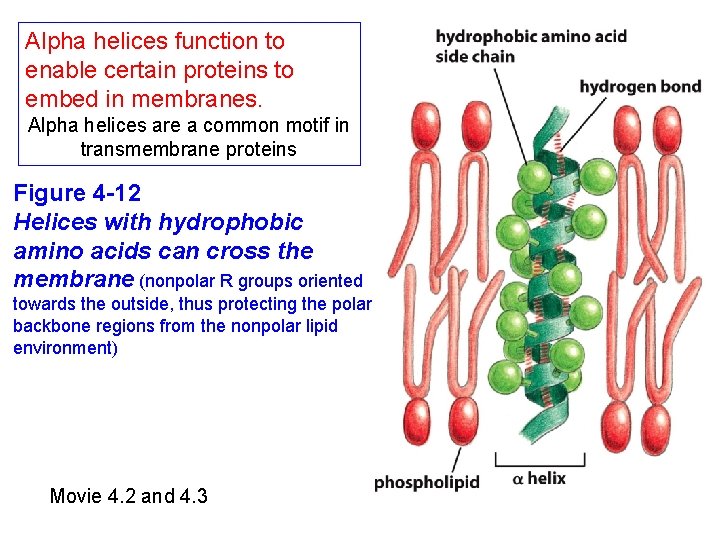
Alpha helices function to enable certain proteins to embed in membranes. Alpha helices are a common motif in transmembrane proteins Figure 4 -12 Helices with hydrophobic amino acids can cross the membrane (nonpolar R groups oriented towards the outside, thus protecting the polar backbone regions from the nonpolar lipid environment) Movie 4. 2 and 4. 3
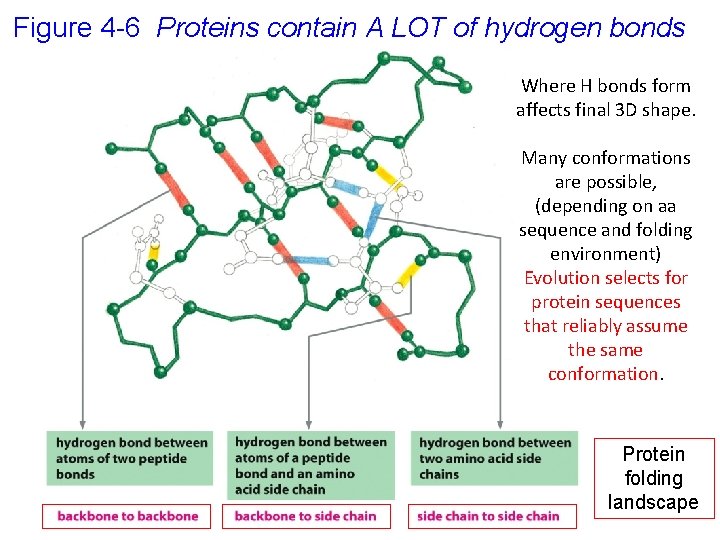
Figure 4 -6 Proteins contain A LOT of hydrogen bonds Where H bonds form affects final 3 D shape. Many conformations are possible, (depending on aa sequence and folding environment) Evolution selects for protein sequences that reliably assume the same conformation. Protein folding landscape
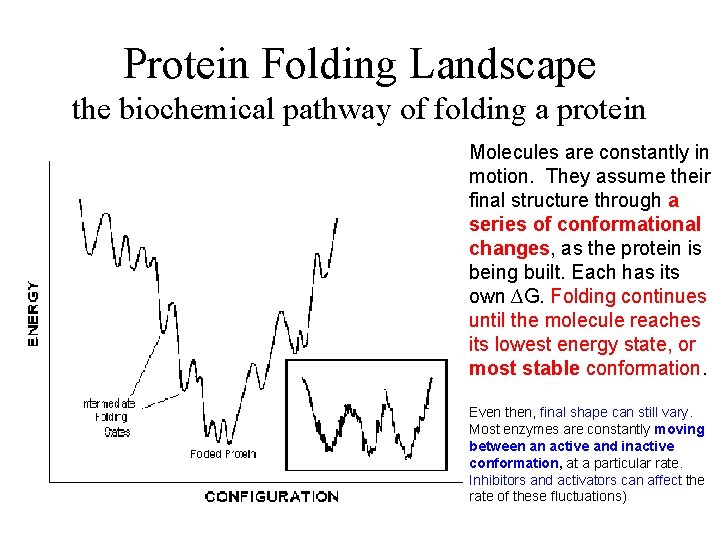
Protein Folding Landscape the biochemical pathway of folding a protein Molecules are constantly in motion. They assume their final structure through a series of conformational changes, as the protein is being built. Each has its own ∆G. Folding continues until the molecule reaches its lowest energy state, or most stable conformation. Even then, final shape can still vary. Most enzymes are constantly moving between an active and inactive conformation, at a particular rate. Inhibitors and activators can affect the rate of these fluctuations)
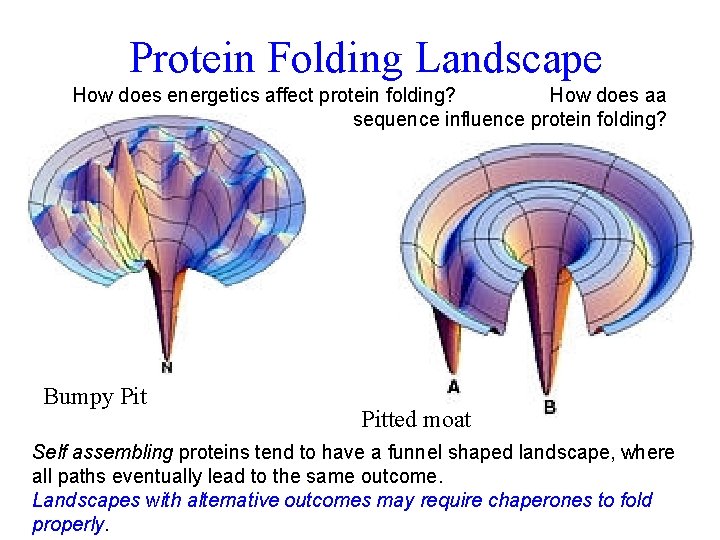
Protein Folding Landscape How does energetics affect protein folding? How does aa sequence influence protein folding? Bumpy Pitted moat Self assembling proteins tend to have a funnel shaped landscape, where all paths eventually lead to the same outcome. Landscapes with alternative outcomes may require chaperones to fold properly.
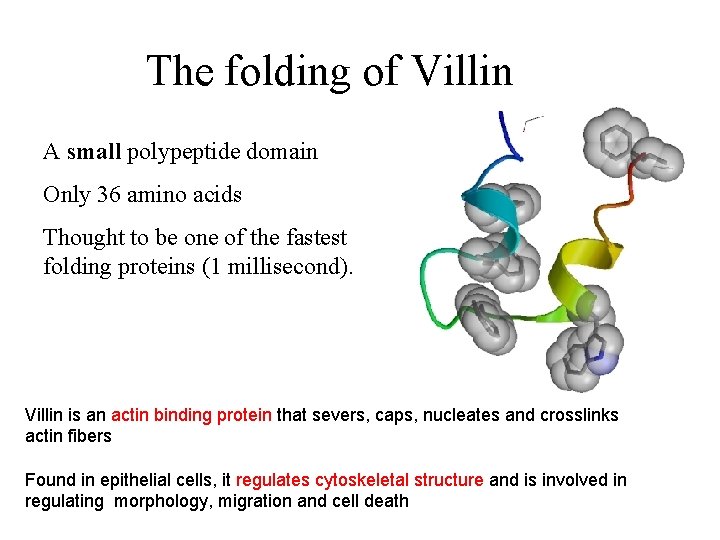
The folding of Villin A small polypeptide domain Only 36 amino acids Thought to be one of the fastest folding proteins (1 millisecond). Villin is an actin binding protein that severs, caps, nucleates and crosslinks actin fibers Found in epithelial cells, it regulates cytoskeletal structure and is involved in regulating morphology, migration and cell death
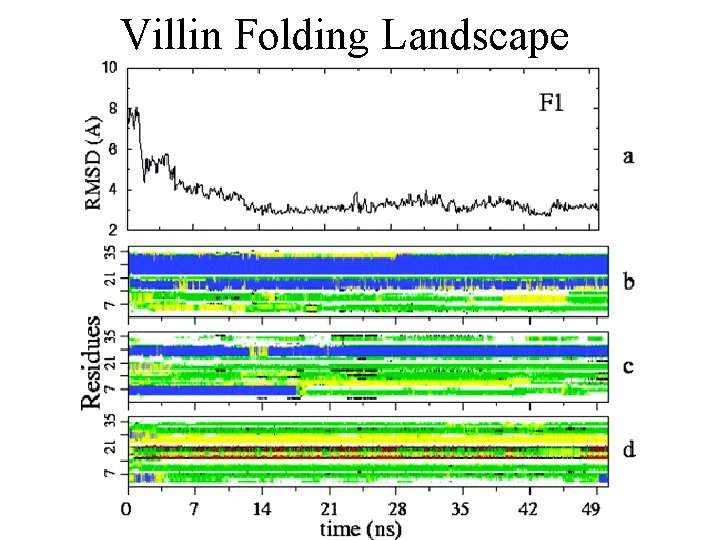
Villin Folding Landscape
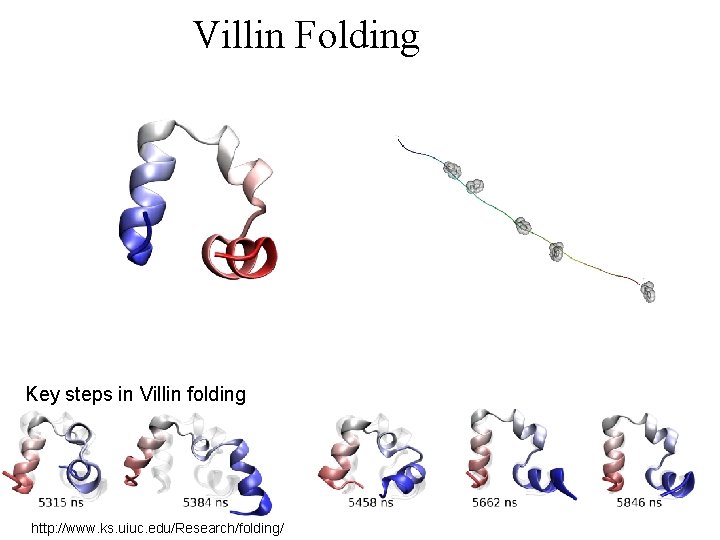
Villin Folding Key steps in Villin folding http: //www. ks. uiuc. edu/Research/folding/
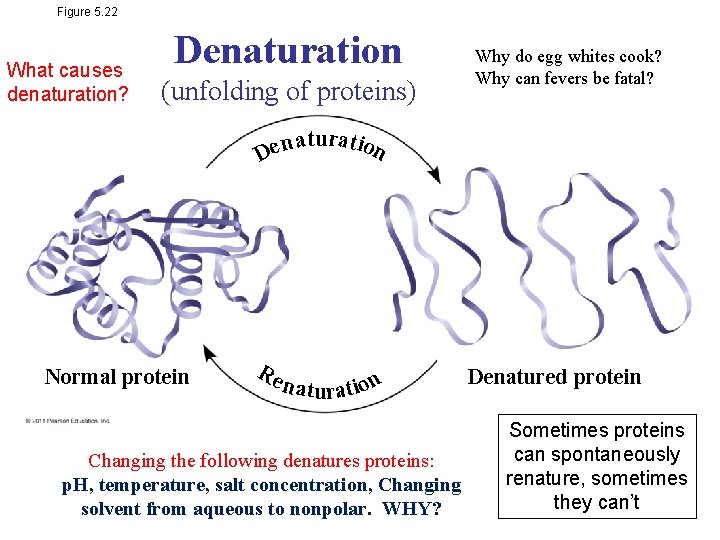
Figure 5. 22 What causes denaturation? Denaturation (unfolding of proteins) Why do egg whites cook? Why can fevers be fatal? n a tu ra t ion e D Normal protein Re n a tura t ion Changing the following denatures proteins: p. H, temperature, salt concentration, Changing solvent from aqueous to nonpolar. WHY? Denatured protein Sometimes proteins can spontaneously renature, sometimes they can’t
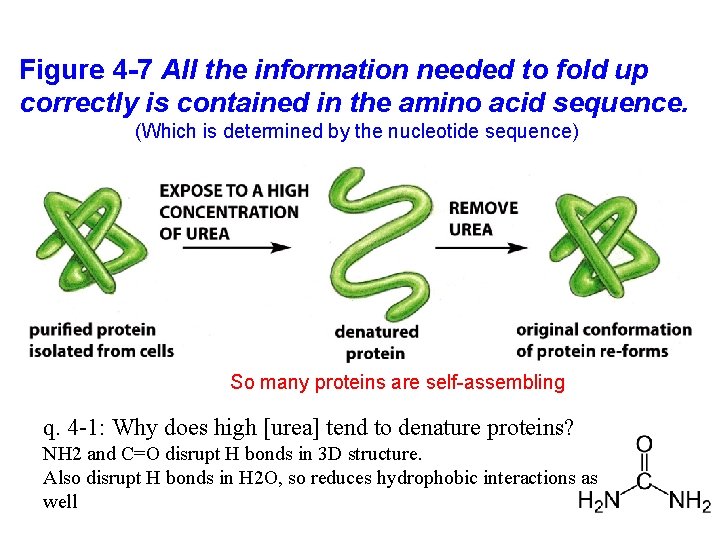
Figure 4 -7 All the information needed to fold up correctly is contained in the amino acid sequence. (Which is determined by the nucleotide sequence) So many proteins are self-assembling q. 4 -1: Why does high [urea] tend to denature proteins? NH 2 and C=O disrupt H bonds in 3 D structure. Also disrupt H bonds in H 2 O, so reduces hydrophobic interactions as well
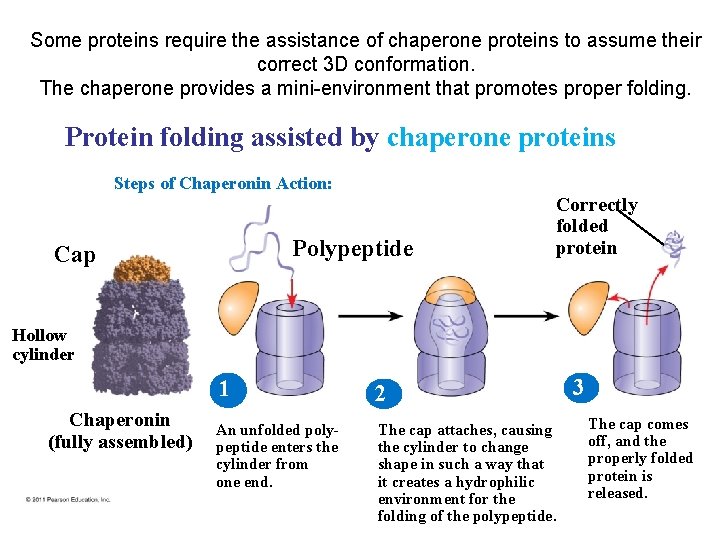
Some proteins require the assistance of chaperone proteins to assume their correct 3 D conformation. The chaperone provides a mini-environment that promotes proper folding. Protein folding assisted by chaperone proteins Steps of Chaperonin Action: Polypeptide Cap Correctly folded protein Hollow cylinder Chaperonin (fully assembled) 1 2 An unfolded polypeptide enters the cylinder from one end. The cap attaches, causing the cylinder to change shape in such a way that it creates a hydrophilic environment for the folding of the polypeptide. 3 The cap comes off, and the properly folded protein is released.

Chaperone proteins • Once thought all newly made proteins folded spontaneously • Now understand Chaperone proteins help many proteins to fold correctly • It is hard to predict a protein’s structure from its primary structure – Most proteins probably go through several stages on their way to a stable structure • Chaperones are protein molecules that assist the proper folding of other proteins – Diseases such as Alzheimer’s, Parkinson’s, and mad cow disease are associated with misfolded proteins – Cystic fibrosis is a hereditary disorder • In some individuals, protein appears to have correct amino acid sequence but fails to fold properly, so does not get delivered to membrane
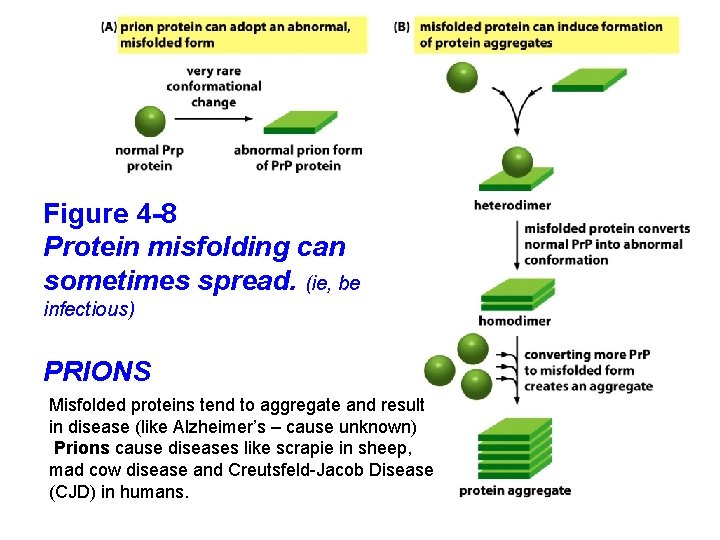
Figure 4 -8 Protein misfolding can sometimes spread. (ie, be infectious) PRIONS Misfolded proteins tend to aggregate and result in disease (like Alzheimer’s – cause unknown) Prions cause diseases like scrapie in sheep, mad cow disease and Creutsfeld-Jacob Disease (CJD) in humans.
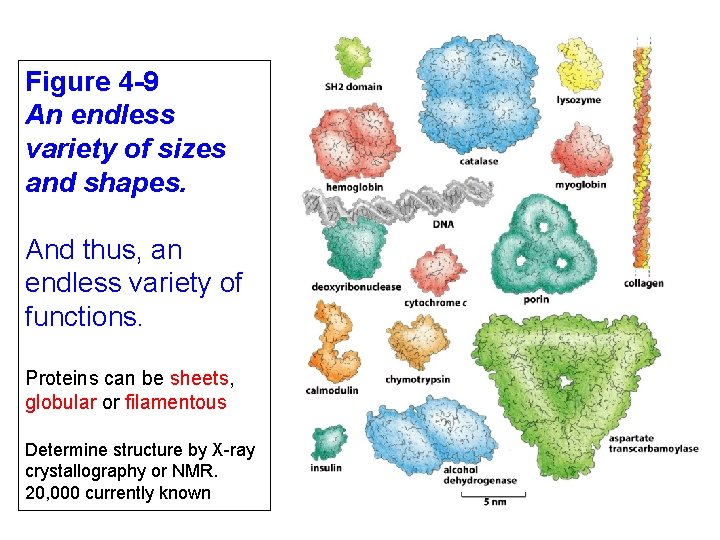
Figure 4 -9 An endless variety of sizes and shapes. And thus, an endless variety of functions. Proteins can be sheets, globular or filamentous Determine structure by X-ray crystallography or NMR. 20, 000 currently known

Four Levels of Protein Structure ie, how do proteins assume their final functional conformation? Primary Structure = the linear sequence of amino acids. (determined by order of DNA nucleotides in the gene) Secondary Structure = due to hydrogen bonding between backbone atoms (not due to R group interactions). Tertiary Structure = determines overall 3 dimensional shape of the polypeptide. (due to noncovalent, and covalent, interactions involving R groups) Quaternary Structure = actual structure formed when multiple polypeptides bind together. (not all proteins have 4 o structure)

The primary structure of a protein The sequence of amino acids Proteins are directional molecules What determines which of the 20129 possibilities of aa order this protein will assume?
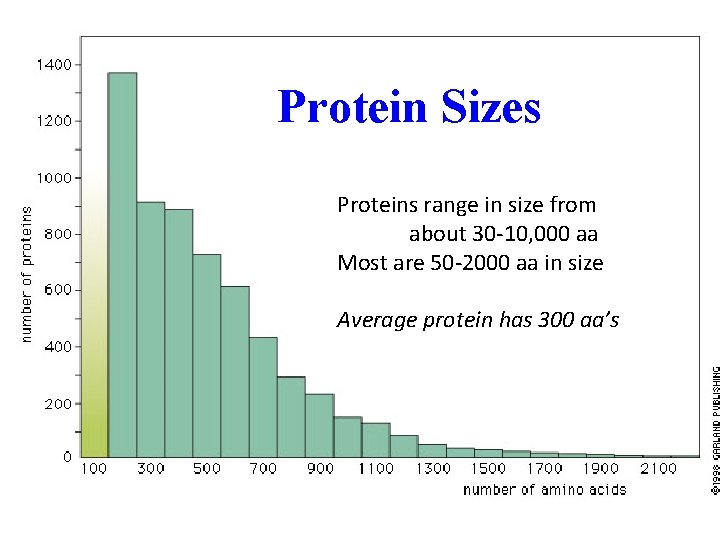
Protein Sizes Proteins range in size from about 30 -10, 000 aa Most are 50 -2000 aa in size Average protein has 300 aa’s
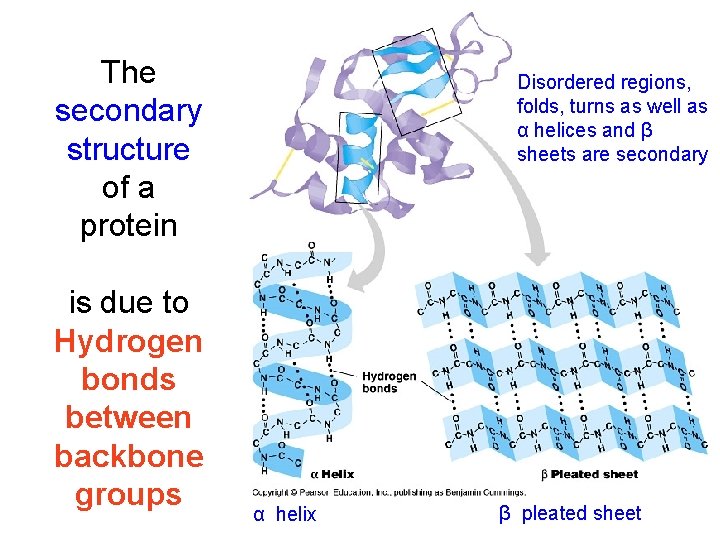
The secondary structure of a protein is due to Hydrogen bonds between backbone groups Disordered regions, folds, turns as well as α helices and β sheets are secondary α helix β pleated sheet
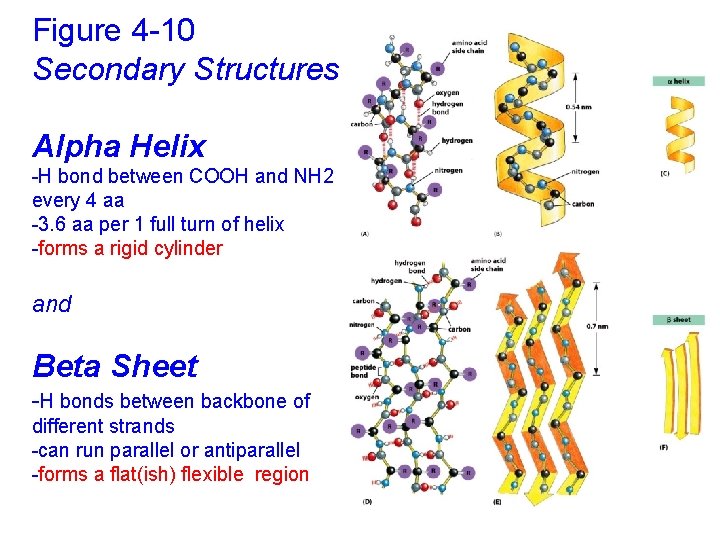
Figure 4 -10 Secondary Structures Alpha Helix -H bond between COOH and NH 2 every 4 aa -3. 6 aa per 1 full turn of helix -forms a rigid cylinder and Beta Sheet -H bonds between backbone of different strands -can run parallel or antiparallel -forms a flat(ish) flexible region
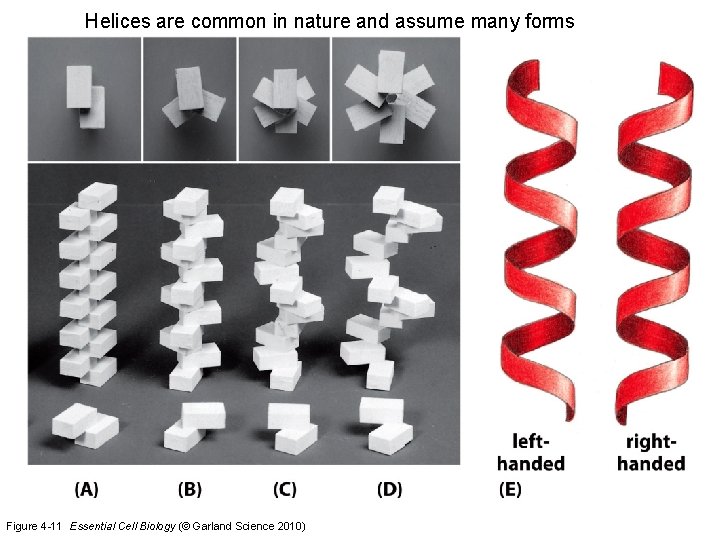
Helices are common in nature and assume many forms Figure 4 -11 Essential Cell Biology (© Garland Science 2010)
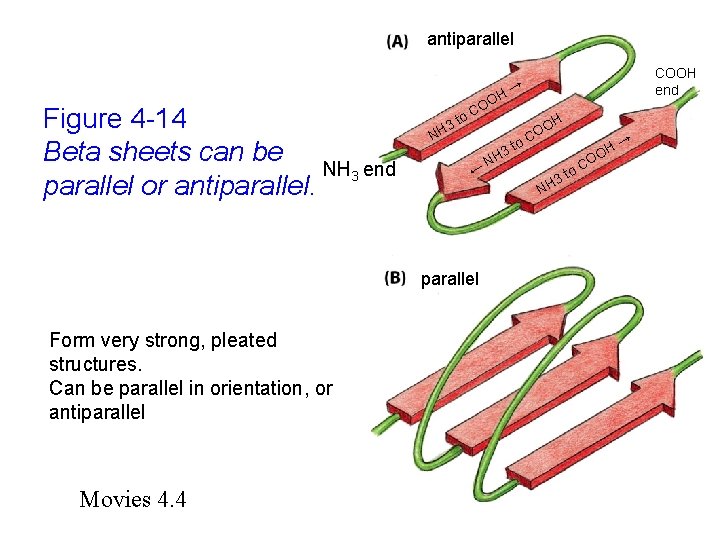
antiparallel Figure 4 -14 Beta sheets can be NH 3 end parallel or antiparallel. H OO C to 3 H N ← parallel Form very strong, pleated structures. Can be parallel in orientation, or antiparallel Movies 4. 4 COOH end → t H 3 o O CO H N NH o 3 t C H OO →
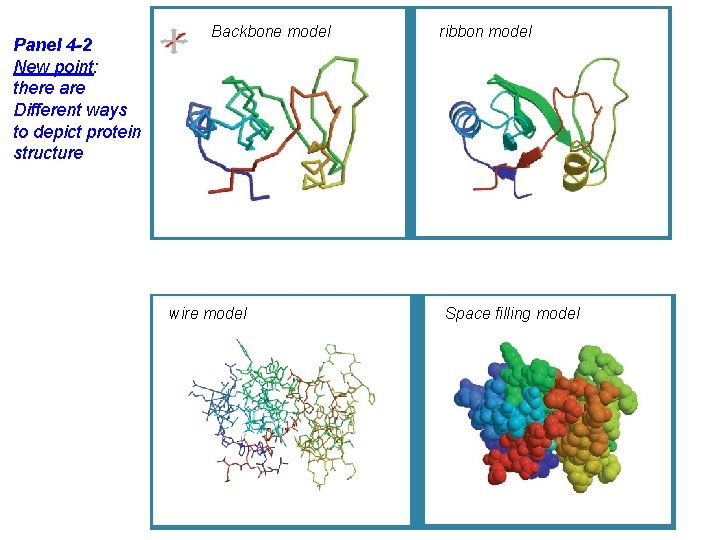
Panel 4 -2 New point: there are Different ways to depict protein structure Backbone model wire model ribbon model Space filling model

Movie 4. 2 alpha helix Movie 4. 4 Beta sheet

Reminder: β sheets and α helices are both examples of 2 o protein structure. interactions between aa backbone groups 2 o structures occur only due to Hydrogen bond formation (between carboxy and amino groups of the backbone) 3 o protein structure is due to interactions of R groups with other R groups OR of R groups with backbone groups 3 o interactions can occur via multiple types of noncovalent interactions (such as hydrogen bonds, vanderwaals forces, ionic interactions) OR can occur by covalent disulfide bonds between SH containing R groups
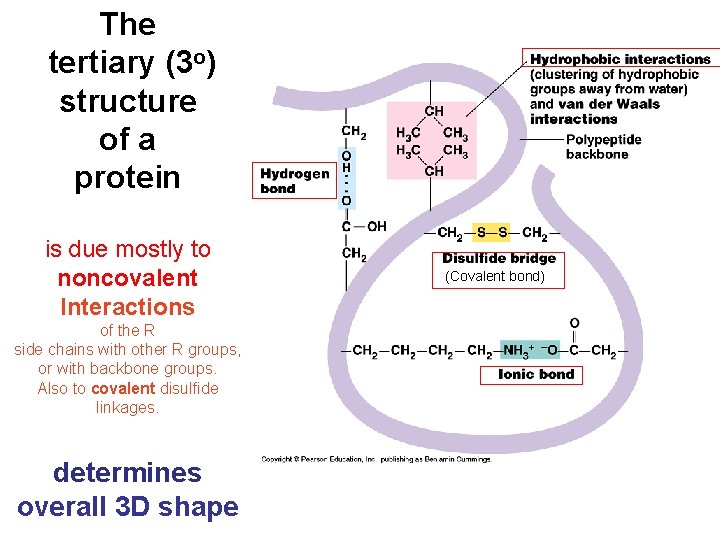
The tertiary (3 o) structure of a protein is due mostly to noncovalent Interactions of the R side chains with other R groups, or with backbone groups. Also to covalent disulfide linkages. determines overall 3 D shape (Covalent bond)
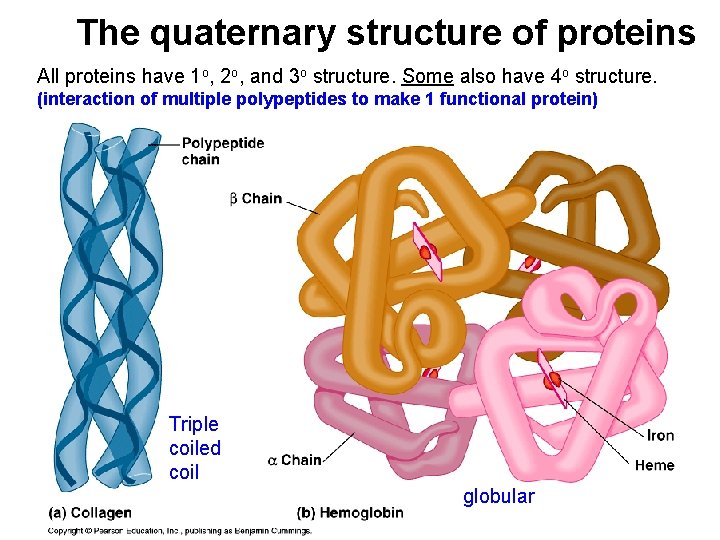
The quaternary structure of proteins All proteins have 1 o, 2 o, and 3 o structure. Some also have 4 o structure. (interaction of multiple polypeptides to make 1 functional protein) Triple coiled coil globular
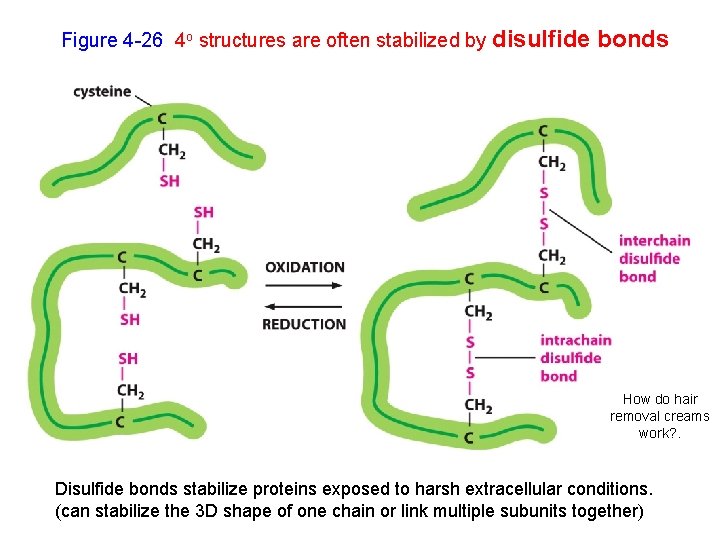
Figure 4 -26 4 o structures are often stabilized by disulfide bonds How do hair removal creams work? . Disulfide bonds stabilize proteins exposed to harsh extracellular conditions. (can stabilize the 3 D shape of one chain or link multiple subunits together)
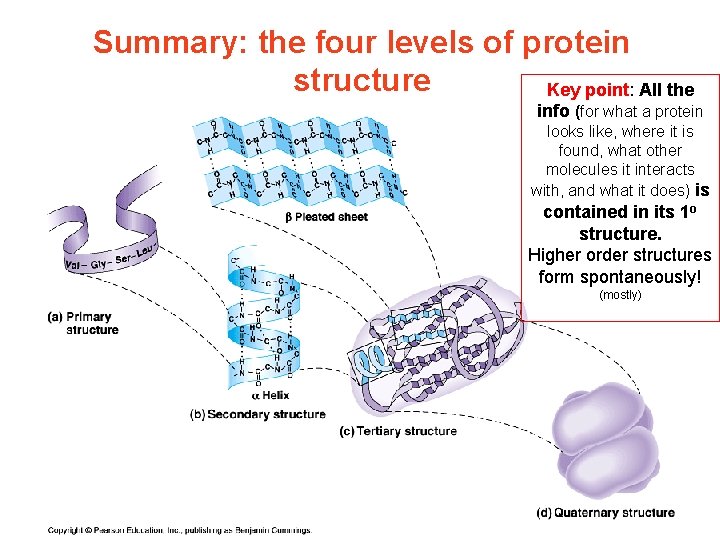
Summary: the four levels of protein structure Key point: All the info (for what a protein looks like, where it is found, what other molecules it interacts with, and what it does) is contained in its 1 o structure. Higher order structures form spontaneously! (mostly)
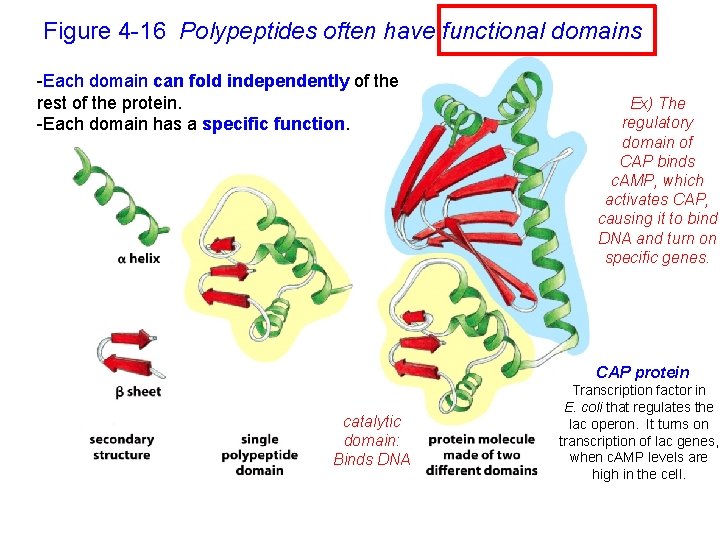
Figure 4 -16 Polypeptides often have functional domains -Each domain can fold independently of the rest of the protein. -Each domain has a specific function. Ex) The regulatory domain of CAP binds c. AMP, which activates CAP, causing it to bind DNA and turn on specific genes. CAP protein catalytic domain: Binds DNA Transcription factor in E. coli that regulates the lac operon. It turns on transcription of lac genes, when c. AMP levels are high in the cell.
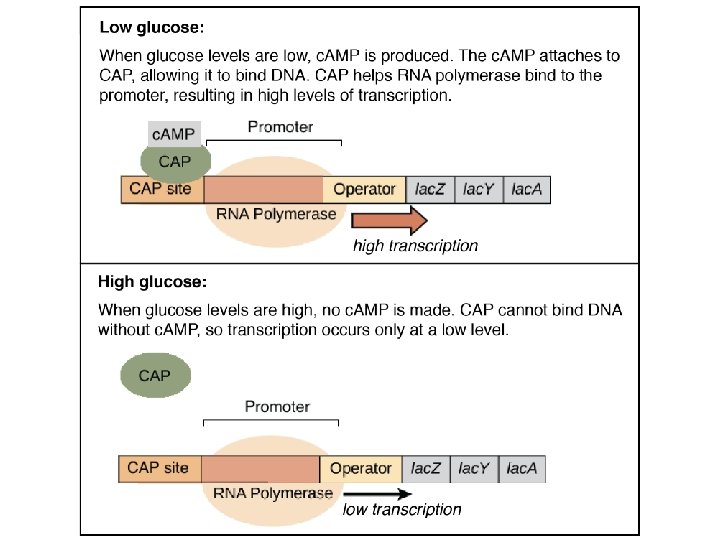
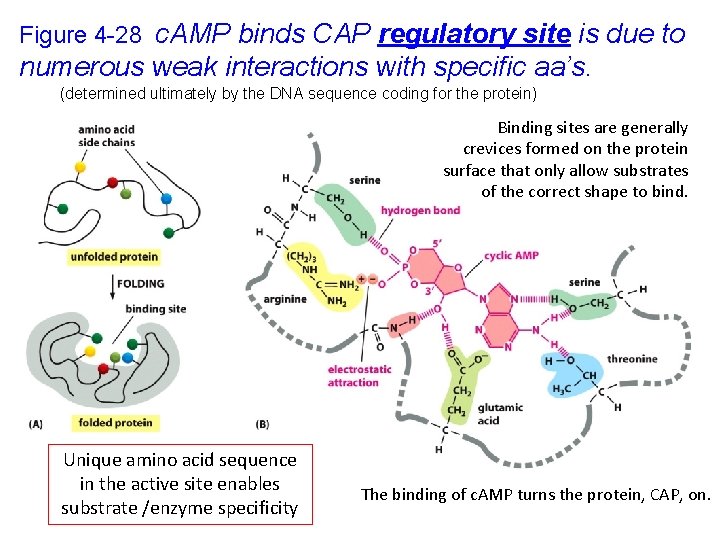
Figure 4 -28 c. AMP binds CAP regulatory site is due to numerous weak interactions with specific aa’s. (determined ultimately by the DNA sequence coding for the protein) Binding sites are generally crevices formed on the protein surface that only allow substrates of the correct shape to bind. Unique amino acid sequence in the active site enables substrate /enzyme specificity The binding of c. AMP turns the protein, CAP, on.

How proteins are regulated 3 Important domains Binding domain: recognizes correct substrates (this is the active site). This site regulates the affinity of the enzyme for its substrate Regulatory domain: binds regulatory molecules (like c. AMP, product, inhibitors, activators, etc to influence activity of the protein) - can be an allosteric site or the active site (the binding domain) -competitive inhibitors bind the active site - noncompetitive inhibitors bind a different site (the allosteric site) -binding here regulates how active an enzyme is (is it currently “on” or “off”) Catalytic domain: (enzymes have this) this is the site that changes the chemical nature of the substrate (ie, catalyzes the reaction) by inducing a shape change to the activated intermediate which results catalyzing a chemical reaction. -can be a separate site, or can be the binding domain (active site) itself
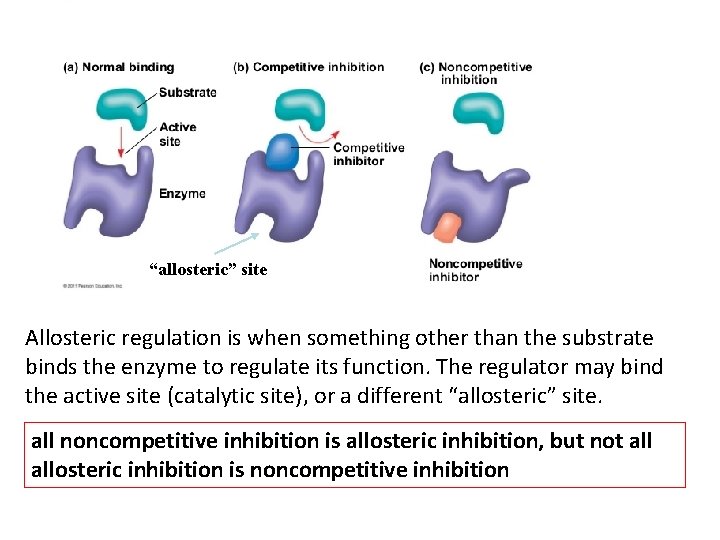
“allosteric” site Allosteric regulation is when something other than the substrate binds the enzyme to regulate its function. The regulator may bind the active site (catalytic site), or a different “allosteric” site. all noncompetitive inhibition is allosteric inhibition, but not allosteric inhibition is noncompetitive inhibition
- Slides: 69