Bioinformatics The pairwise alignment problem Srinivas Jakkidi CS

Bioinformatics: The pair-wise alignment problem Srinivas Jakkidi CS 487

Overview n n n Pair-wise alignment revisited Dynamic programming algorithm Parallel extension

Pair-wise alignment n Inexact matching: comparing two sequences while allowing for some mismatch. n n Extent of mismatch depends on type of sequence (protein vs. nucleotide) Try to minimize the number of substitutions, inserts and deletes to convert one sequence to the other

Pair-wise alignment (cont. ) n n n Insertion, deletion are considered same function – indel Each mutation has an associated penalty Try to minimize penalty (distance)

Dynamic programming algorithm n n n Dynamic programming: build solution using previous solutions for smaller subsequences Stores values corresponding to partial results in a similarity matrix We are trying to align two sequences X and Y of lengths m and n respectively.

Dynamic programming algorithm Similarity matrix SM is of size mxn SMi, j = max(SMi, j-1+ gp, SMi-1, j-1+ ss, SMi-1, j+ gp, 0) gp is the gap penalty and ss is the substitution score n
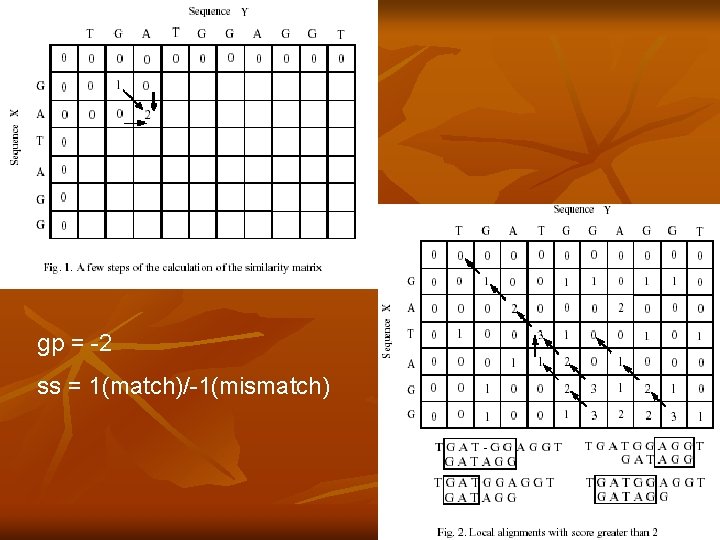
gp = -2 ss = 1(match)/-1(mismatch)
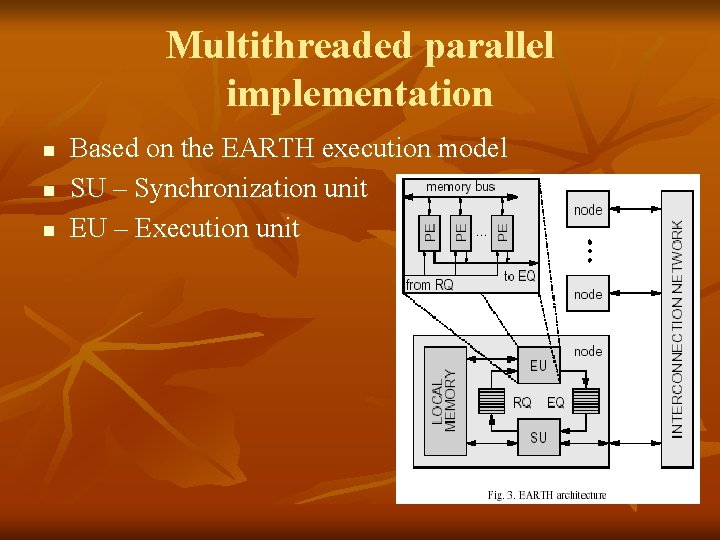
Multithreaded parallel implementation n Based on the EARTH execution model SU – Synchronization unit EU – Execution unit

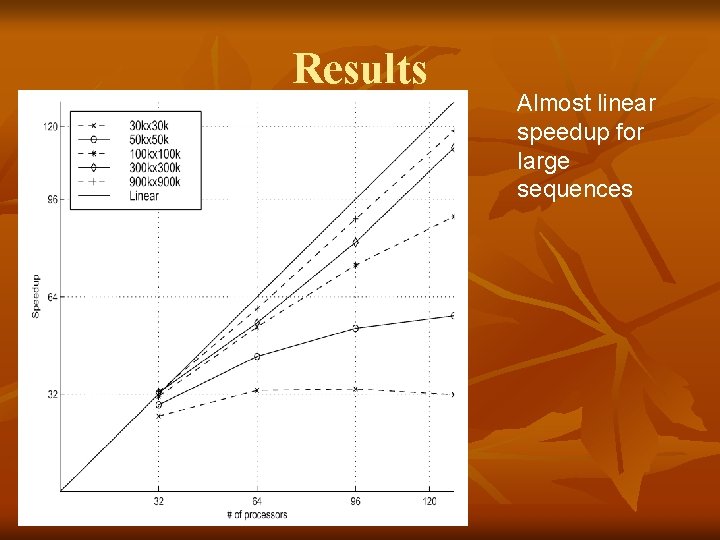
Results Almost linear speedup for large sequences
- Slides: 10