Bioinformatics Support in Trinity College Karsten Hokamp Darren

Bioinformatics Support in Trinity College Karsten Hokamp Darren Fitzpatrick Fiona Roche TBSI, 30/06/2015

Bioinformatics Support in Trinity College Position established in 2005 … to provide computational support for SFI-Biotech funded projects in Trinity College
Bioinformatics Support Pharmacology Microbiology Biochemistry Immunology Psychology Medicine Genetics Zoology Botany Already at MIT, Stanford, Harvard, Yale, Oxford, Cambridge, UCL, …
cost data Next-generation Sequencing

Major research areas: • Sequence analysis • Gene expression • Gene regulation • Computational evolutionary biology • Network and systems biology • Genome annotation • Mutations in cancer • Population genomics • Literature mining • Structural bioinformatics • High-throughput image analysis proficiency Bioinformatics

Activities - NGS RNA-seq, Ch. IP-seq, assembly, SNP detection @HWI-ST 1363: 132: C 201 RACXX: 3: 1101: 18624: 2742 1: N: 0: AGTTCC GTTTAACTTGAGTGCAAGAGGGGAGAGTGGAATTCCATGTGTAGCGGTGAAATGCGTAGAT + <@<ADDDB? FH<<EGD? ? : FFHE: C): 11811? : DHIII<? ? F>GED 7@GI@H@AB 55: 5? EBED

Activities - NGS RNA-seq, Ch. IP-seq, assembly, SNP detection @HWI-ST 1363: 132: C 201 RACXX: 3: 1101: 18624: 2742 1: N: 0: AGTTCC GTTTAACTTGAGTGCAAGAGGGGAGAGTGGAATTCCATGTGTAGCGGTGAAATGCGTAGAT + <@<ADDDB? FH<<EGD? ? : FFHE: C): 11811? : DHIII<? ? F>GED 7@GI@H@AB 55: 5? EBED

Example – Genome assembly, SNP detection

Example: RNA-seq, 20 conditions Web-browser for raw data
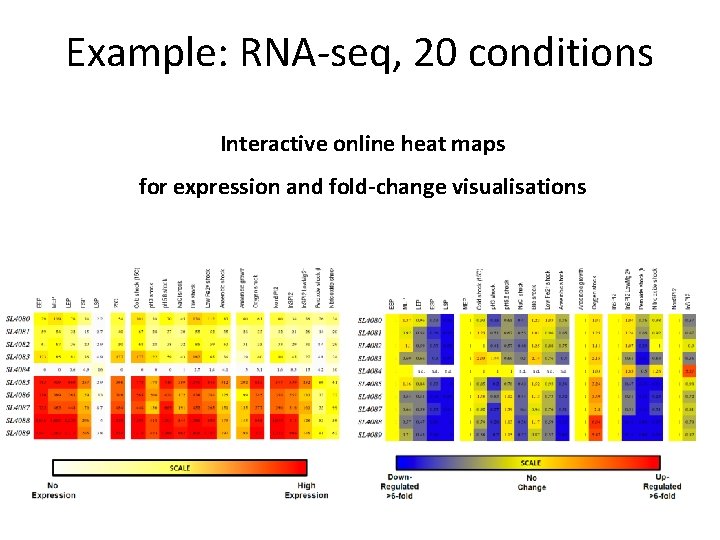
Example: RNA-seq, 20 conditions Interactive online heat maps for expression and fold-change visualisations
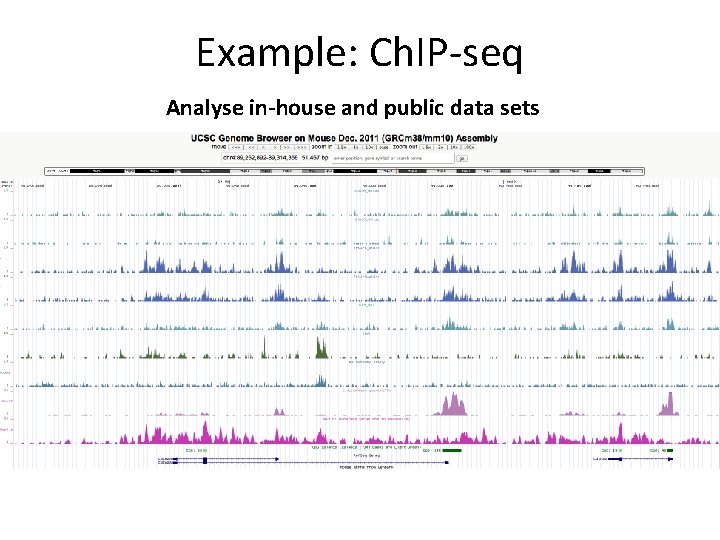
Example: Ch. IP-seq Analyse in-house and public data sets
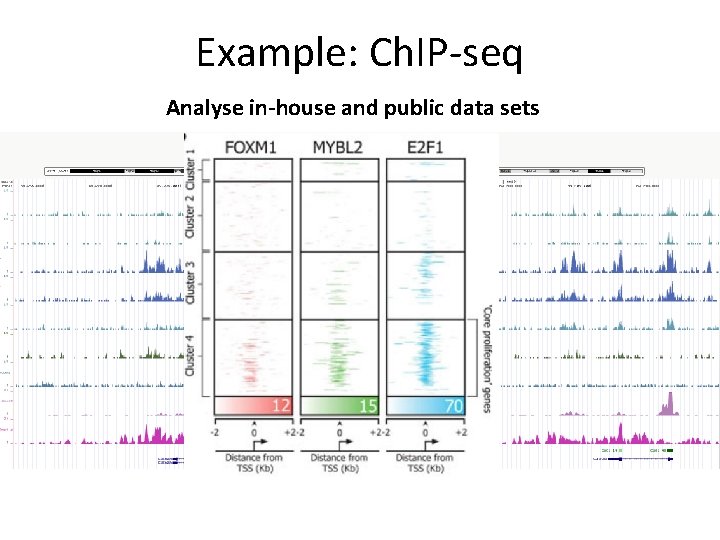
Example: Ch. IP-seq Analyse in-house and public data sets
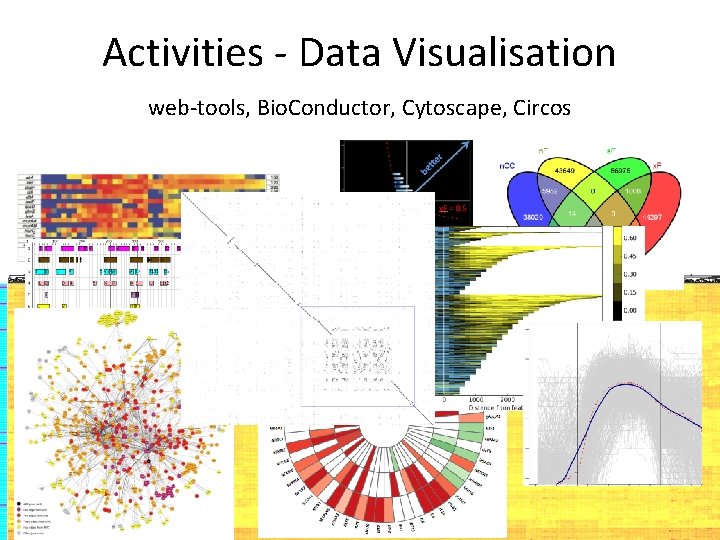
Activities - Data Visualisation web-tools, Bio. Conductor, Cytoscape, Circos

Activities - Web Servers Array. Pipe: microarray data analysis
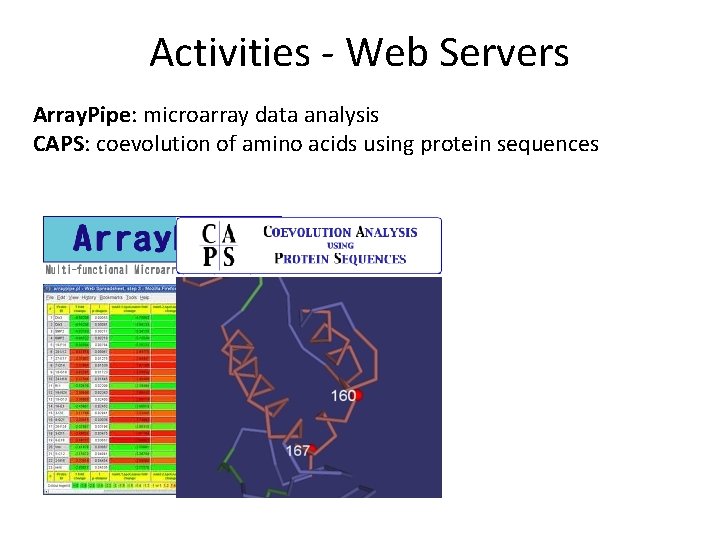
Activities - Web Servers Array. Pipe: microarray data analysis CAPS: coevolution of amino acids using protein sequences

Activities - Web Servers Array. Pipe: microarray data analysis CAPS: coevolution of amino acids using protein sequences PFPE: Phylogenetic Fooprinting Ensemble

Activities - Web Servers Array. Pipe: microarray data analysis CAPS: coevolution of amino acids using protein sequences PFPE: Phylogenetic Foo. Printing Ensemble Pub. Crawler: Literature alerting system

Activities - Web Servers Array. Pipe: microarray data analysis CAPS: coevolution of amino acids using protein sequences PFPE: Phylogenetic Foo. Printing Ensemble Pub. Crawler: Literature alerting system Sal. Com: Salmonella Typhimurium Gene Expression Compedium
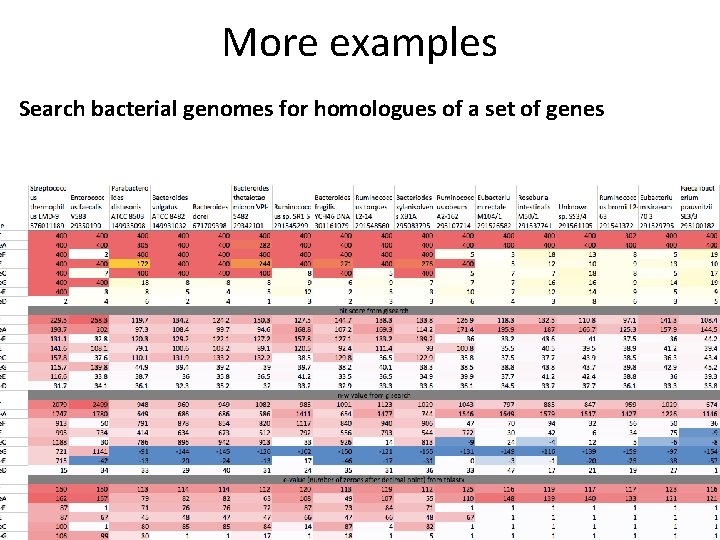
More examples Search bacterial genomes for homologues of a set of genes
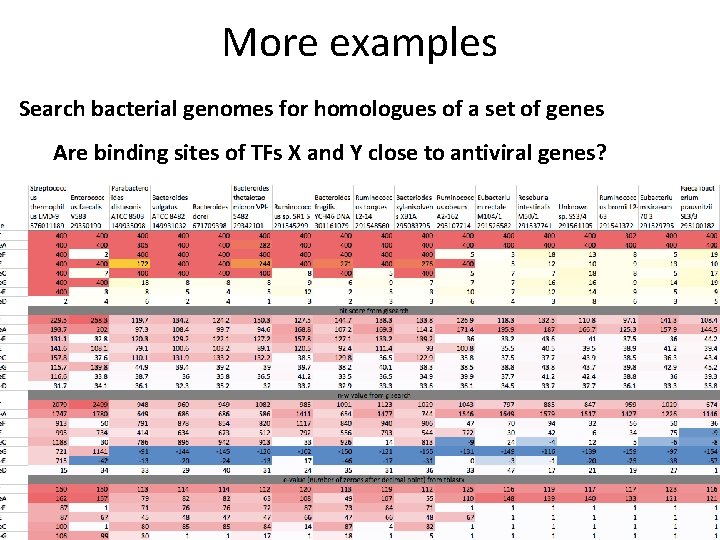
More examples Search bacterial genomes for homologues of a set of genes Are binding sites of TFs X and Y close to antiviral genes?
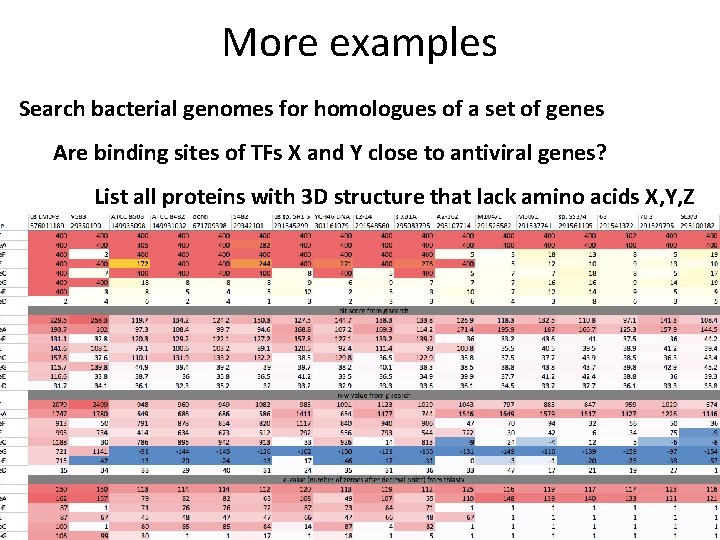
More examples Search bacterial genomes for homologues of a set of genes Are binding sites of TFs X and Y close to antiviral genes? List all proteins with 3 D structure that lack amino acids X, Y, Z

More examples Search bacterial genomes for homologues of a set of genes Are binding sites of TFs X and Y close to antiviral genes? List all proteins with 3 D structure that lack amino acids X, Y, Z Find specific motifs in a set of proteins

Introducing: Dr Fiona Roche

Background in Infectious Diseases • 1997 – 2000: Ph. D Microbiology, TCD – Staphylococcal adherence to host tissue (Prof Tim Foster) • 2000 – 2002: TCD, Postdoc – Functional genomics of Staphylococcal genome • 2002 – 2006: SFU Canada, Postdoc – Development of a bioinformatics platform for pathogenomics project (Prof. Fiona Brinkman) • 2007 – 2015: HPSC, Data Manager – Development of national surveillance systems to monitor healthcare associated infections

Background in Infectious Diseases • 1997 – 2000: Ph. D Microbiology, TCD – Staphylococcal adherence to host tissue (Prof Tim Foster) • 2000 – 2002: TCD, Postdoc – Functional genomics of Staphylococcal genome • 2002 – 2006: SFU Canada, Postdoc – Development of a bioinformatics platform for pathogenomics project (Prof. Fiona Brinkman) • 2007 – 2015: HPSC, Data Manager – Development of national surveillance systems to monitor healthcare associated infections

Background in Infectious Diseases • 1997 – 2000: Ph. D Microbiology, TCD – Staphylococcal adherence to host tissue (Prof Tim Foster) • 2000 – 2002: TCD, Postdoc – Functional genomics of Staphylococcal genome • 2002 – 2006: SFU Canada, Postdoc – Development of a bioinformatics platform for pathogenomics project (Prof. Fiona Brinkman) • 2007 – 2015: HPSC, Surveillance – Development of national surveillance systems to monitor healthcare associated infections

Bioinformatics for Biologists A web-based genome analysis platform designed for experimental biologists http: //galaxyproject. org/

Analysis Categories • • Statistics Visualisation Sequence Analysis NGS data analysis Proteomics Metagenomics Computational Chemistry

Workflows Example chip-seq workflow Data input fastq Quality control Fast. QC Mapping to genome Bowtie Peak calling MACS

Workflows Example chip-seq workflow Data input fastq Quality control Fast. QC Mapping to genome Bowtie Peak calling MACS

Galaxy Workshops @TCD • Local instance at TCD • Introductory session • Galaxy Workshop

Darren J. Fitzpatrick, Ph. D Education B. A. (Mod) Genetics Trinity College Dublin M. Res Computational Biology University of York “In silico approach to the prediction of Antibody Thermostability” Ph. D Statistical Genetics University College Dublin “The effect of combinations of genetic variants”

Research Experiences GWAS: plink, e. QTLs, epistasis (BMC Genomics, 2015), gene expression, synergy, genotype imputation, population stratification, recombination Population Genetics: genetic ancestry, ancestry prediction, visualisation of population data (Ancestry. Mapper – PLo. S One, 2012) Data Integration: integrating novel and public data – SNPs, CNVs, epigenetic modifications, DNA-protein interactions from ENCODE, NIH Roadmap, Chromosome Capture Experiments, etc. Pharmacology: combinatorial modeling of the effects of multiple drugs on platelet activation (Plo. S Computational Biology, 2015) Structural Biology: antibodies, tessellation, prediction of biophysical properties, machine learning

Interests: • Statistical & Population Genetics, Complex Traits, Epigenetics, Next-Gen Technologies, Molecular Evolution The Skills: • Managing ‘Big Data’: data organising, cleaning, formatting and general ‘wrangling’ – (Python, my. SQL, Shell Scripting) • Analysis (BIG & small data): hypothesis tests, statistical modeling, supervised/unsupervised learning, data visualisation – (R, Bioconductor, ggplot 2, Cytoscape) Current Projects: • Ph. D Leftovers: GPCR gene expression in Alzheimer’s, Recombination in Autism • Ch. IP-chip analysis of yeast epigenetic modifications • Developing tool to visualise comparisons amongst many publicly available Ch. IP-Seq data sets

Teaching/Workshops • Teaching experience: R & statistics to 1 st year Ph. D students, Introduction to Bioinformatics to undergraduates Planned Workshops • Python programming and R statistical computing workshops for Ph. D students, Post-Docs and PIs in August/September – How to work computationally with your data (Bye bye Excel!). – How to explore, analyse and visualise your data. Office: Westland Row, Smurfit Institute of Genetics Contact: fitzpadj@tcd. ie

Activities - Training Proposed introductory Workshops: • Programming with Python • Statistics with R • Bioinformatics with Galaxy

TBSI Office

Thanks! Questions / suggestions?

Fiona Roche fmroche@tcd. ie Karsten Hokamp kahokamp@tcd. ie bioinf. gen. tcd. ie Darren Fitzpatrick fitzpadj@tcd. ie
- Slides: 39