Bioinformatics Scope Applications Definition The application of computer

Bioinformatics Scope & Applications

Definition • The application of computer technology to the management of biological information. Specifically, it is the science of • developing computer databases • algorithms • softwares to facilitate and expedite biological research

Contd…. • The mathematical, statistical and computing methods that aim to solve biological problems using DNA and amino acid sequences and related information.

Broader aspects • There are other fields-for example medical imaging / image analysis which might be considered part of bioinformatics. There is also a whole other discipline of biologically -inspired computation; genetic algorithms, AI, neural networks

How to take bioinformatics • Biological background but little computation? – Learn to use the tools. – Use the supplementary texts – Try to learn some of the CS related issues • Quantitative background but little biology? – Appreciate the diversity of application problems – Make sure you understand the biology. . Consider buying a supplementary biology text
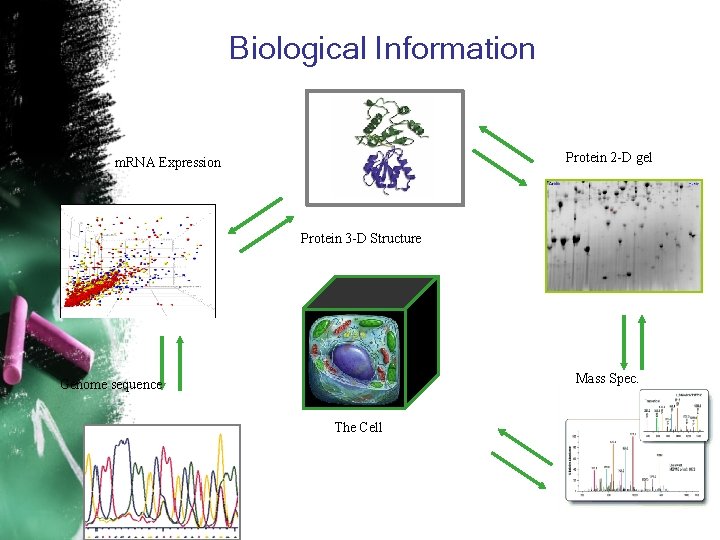
Biological Information Protein 2 -D gel m. RNA Expression Protein 3 -D Structure Mass Spec. Genome sequence The Cell

Definitions of Fields Related to Bioinformatics • Computational Biology ~ Computational biologists interest themselves more with evolutionary, population and theoretical biology.

Genomics: ~Genomics is any attempt to analyze or compare the entire genetic complement of a species Proteomics: (The Protein complement of the genome ) Qualitative and quantitative studies of gene expression at the level of the functional proteins themselves

Pharmacogenomics: • Pharmacogenomics is the application of genomic approaches and technologies to the identification of drug targets

Cheminformatics: The combination of chemical synthesis, biological screening, and data-mining approaches used to guide drug discovery and development Medical Informatics: The study, invention, and implementation of structures and algorithms to improve communication, understanding and management of medical information

Nature of Biological data • • Heterogeneous Unorganized Voluminous Dynamic

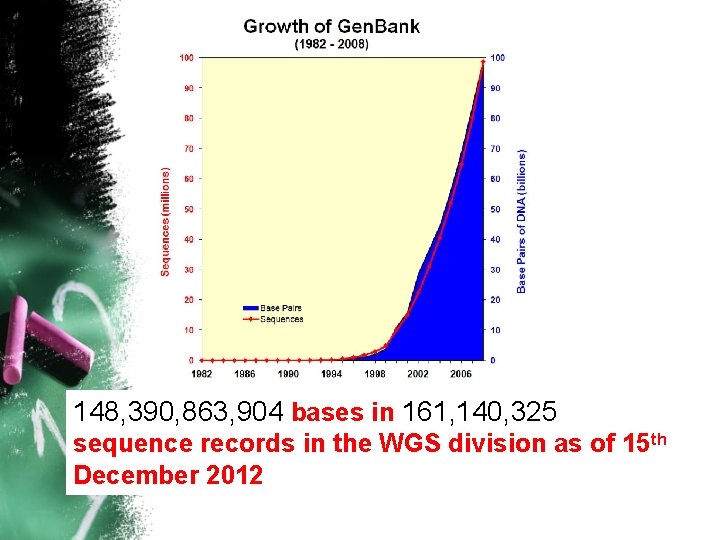
148, 390, 863, 904 bases in 161, 140, 325 sequence records in the WGS division as of 15 th December 2012

What a computer can do ? • Speed -> Accuracy -> Diligence • Organize the heterogeneous data • Means to store voluminous data (Biological databases) • Data analysis tools

Biological databases and organization • Storage of gene and protein sequences • Storage of Protein (and other) structures • Ontology • Others (custom or specialised databases)

How can biological data be organized ? • Public databases available on the internet • Data analysis tools • Web interface

What is bioinformatics? • Sequence analysis – Geneticists/ molecular biologists analyze genome sequence information to understand disease processes • Molecular modeling – Crystallographers/ biochemists design drugs using computer-aided tools • Phylogeny/evolution – Geneticists obtain information about the evolution of organisms by looking for similarities in gene sequences • Ecology and population studies – Bioinformatics is used to handle large amounts of data obtained in population studies • Medical informatics – Personalised medicine

Promises of genomics and bioinformatics • Medicine – Knowledge of protein structure facilitates drug design – Understanding of genomic variation allows the tailoring of medical treatment to the individual’s genetic make-up – Genome analysis allows the targeting of genetic diseases

Contd. • The effect of a disease or of a therapeutic on RNA and protein levels can be elucidated • The same techniques can be applied to biotechnology, crop and livestock improvement, etc. . .
What kind of information ? http: //www. ncbi. nlm. nih. gov
Sequences Genebank http: //www. ncbi. nlm. nih. gov/Genbank. Ove rview. html • DNA sequences. • Primary data generators submit to Genebank. • Annotation issues. • Heart of most genomics projects. http: //www. ncbi. nlm. nih. gov
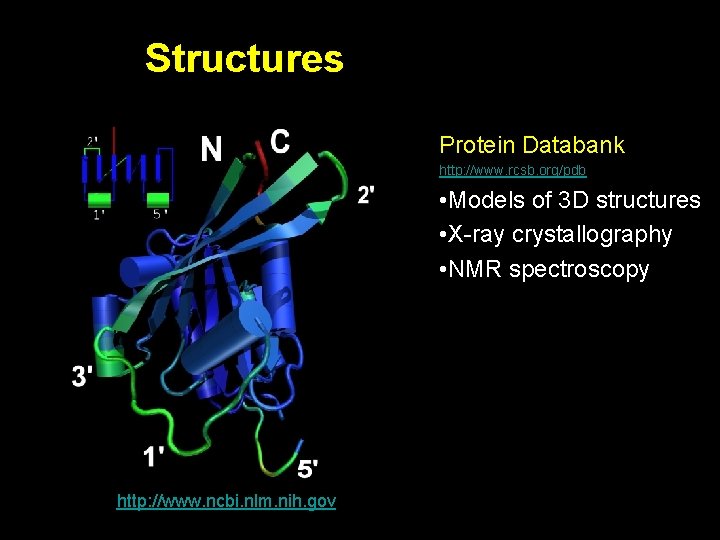
Structures Protein Databank http: //www. rcsb. org/pdb / • Models of 3 D structures • X-ray crystallography • NMR spectroscopy http: //www. ncbi. nlm. nih. gov
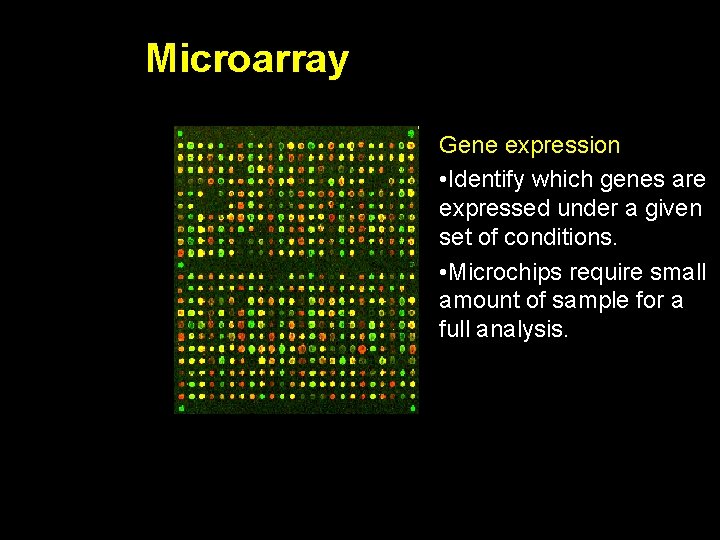
Microarray Gene expression • Identify which genes are expressed under a given set of conditions. • Microchips require small amount of sample for a full analysis.

What can we do with sequences? Multiple sequence alignments Principle Character in sequences can be substituted randomly. Alignment position homologous position together. Unlikely that an ultimate alignment tool will ever be made. http: //www. ncbi. nlm. nih. gov
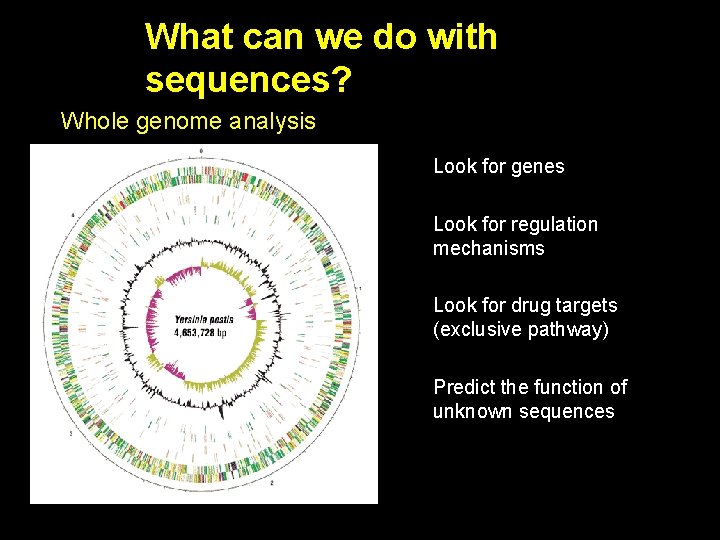
What can we do with sequences? Whole genome analysis Look for genes Look for regulation mechanisms Look for drug targets (exclusive pathway) Predict the function of unknown sequences
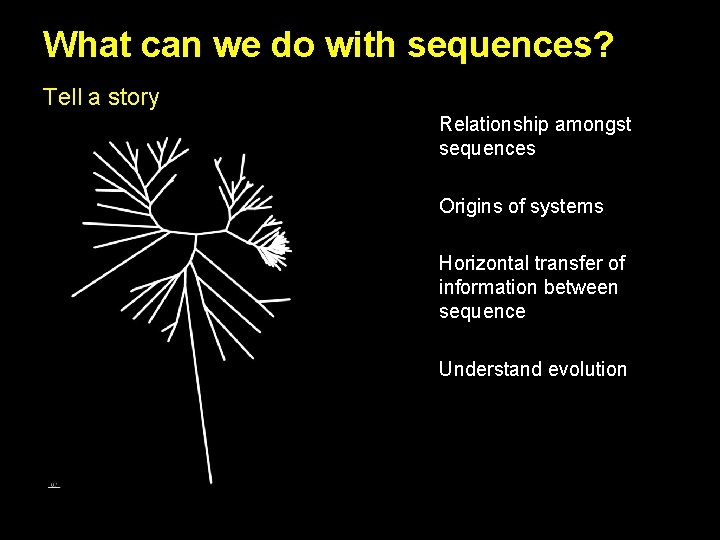
What can we do with sequences? Tell a story Relationship amongst sequences Origins of systems Horizontal transfer of information between sequence Understand evolution
Life – Origins

What can we do with structures? What is its function? What is the mechanism? Does it relate to other known structures Can we design a drug to enhance/suppress its function? Predict the structure of related proteins. http: //www. ks. uiuc. edu/Research/vmd/
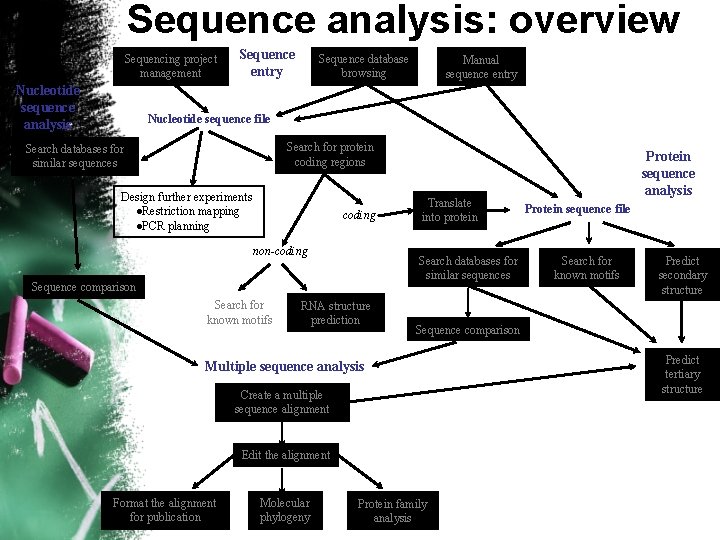
Sequence analysis: overview Sequencing project management Nucleotide sequence analysis Sequence entry Sequence database browsing Manual sequence entry Nucleotide sequence file Search for protein coding regions Search databases for similar sequences Design further experiments l. Restriction mapping l. PCR planning coding non-coding Search databases for similar sequences Sequence comparison Search for known motifs Translate into protein RNA structure prediction Create a multiple sequence alignment Edit the alignment Molecular phylogeny Protein sequence file Search for known motifs Predict secondary structure Sequence comparison Multiple sequence analysis Format the alignment for publication Protein sequence analysis Protein family analysis Predict tertiary structure

1 atgcgttata ttcgcctgtg tattatctcc ctgttagcca ccctgccgct ggcggtacac 61 gccagcccgc agccgcttga gcaaattaaa ctaagcgaaa gccagctgtc gggccgcgta 121 ggcatgatag aaatggatct ggccagcggc cgcacgctga ccgcctggcg cgccgatgaa 181 cgctttccca tgatgagcac ctttaaagta gtgctctgcg gcgcagtgct ggcgcgggtg 241 gatgccggtg acgaacagct ggagcgaaag atccactatc gccagcagga tctggtggac 301 tactcgccgg tcagcgaaaa acatcttgcc gacggcatga cggtcggcga actctgcgcc 361 gccgccatta ccatgagcga taacagcgcc gccaatctgc tgctggccac cgtcggcggc 421 cccgcaggat tgactgcctt tttgcgccag atcggcgaca acgtcacccg ccttgaccgc 481 tgggaaacgg aactgaatga ggcgcttccc ggcgacgccc gcgacaccac taccccggcc 541 agcatggccg cgaccctgcg caagctgctg accagc gtctgagcgc ccgttcgcaa 601 cggcagctgc tgcagtggat ggtggacgat cgggtcgccg gaccgttgat ccgctccgtg 661 ctgccggcgg gctggtttat cgccgataag accggagctg gcgaacgggg tgcgcgcggg 721 attgtcgccc tgcttggccc gaataacaaa gcagagcgca ttgtggtgat ttatctgcgg 781 gatacgccgg cgagcatggc cgagcgaaat cagcaaatcg ccgggatcgg cgcggcgctg 841 atcgagcact ggcaacgcta a

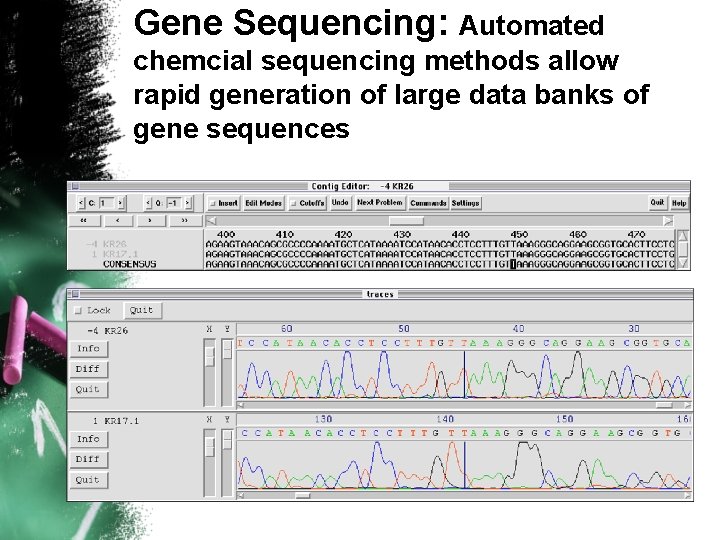
Gene Sequencing: Automated chemcial sequencing methods allow rapid generation of large data banks of gene sequences

Database similarity searching: The BLAST program has been written to allow rapid comparison of a new gene sequence with the 100 s of 1000 s of gene sequences in data bases Sequences producing significant alignments: (bits) Value gnl|PID|e 252316 (Z 74911) ORF YOR 003 w [Saccharomyces cerevisiae] 112 gi|603258 (U 18795) Prb 1 p: vacuolar protease B [Saccharomyces ce. . . 106 gnl|PID|e 264388 (X 59720) YCR 045 c, len: 491 [Saccharomyces cerevi. . . 69 gnl|PID|e 239708 (Z 71514) ORF YNL 238 w [Saccharomyces cerevisiae] 30 gnl|PID|e 239572 (Z 71603) ORF YNL 327 w [Saccharomyces cerevisiae] 29 gnl|PID|e 239737 (Z 71554) ORF YNL 278 w [Saccharomyces cerevisiae] 29 7 e-26 5 e-24 7 e-13 0. 66 1. 1 1. 5 gnl|PID|e 252316 (Z 74911) ORF YOR 003 w [Saccharomyces cerevisiae] Length = 478 Score = 112 bits (278), Expect = 7 e-26 Identities = 85/259 (32%), Positives = 117/259 (44%), Gaps = 32/259 (12%) Query: 2 QSVPWGISRVQAPAAHNRG-----LTGSGVKVAVLDTGIST-HPDLNIRGG-ASFV 50 + PWG+ RV G G GV VLDTGI T H D R + + Sbjct: 174 EEAPWGLHRVSHREKPKYGQDLEYLYEDAAGKGVTSYVLDTGIDTEHEDFEGRAEWGAVI 233 Query: 51 PGEPSTQDGNGHGTHVAGTIAALNNSIGVLGVAPSAELYXXXXXXXXXQGLE 110 P D NGHGTH AG I + + GVA + ++ +G+E Sbjct: 234 PANDEASDLNGHGTHCAGIIGSKH-----FGVAKNTKIVAVKVLRSNGEGTVSDVIKGIE 288
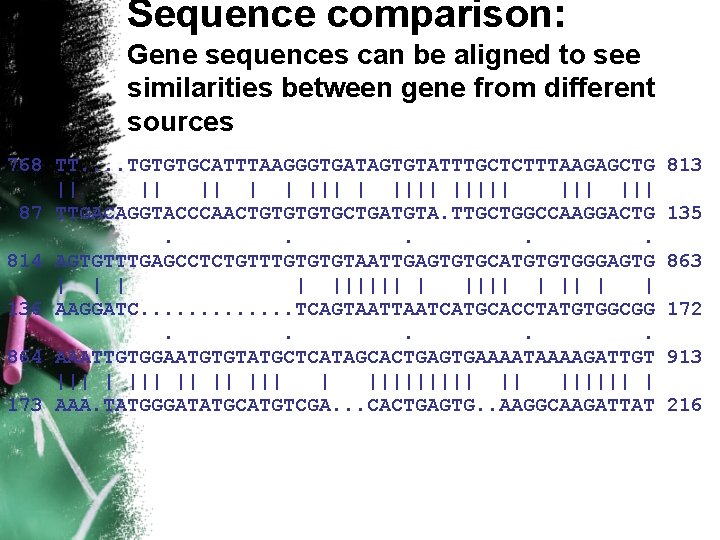
Sequence comparison: Gene sequences can be aligned to see similarities between gene from different sources 768 TT. . TGTGTGCATTTAAGGGTGATAGTGTATTTGCTCTTTAAGAGCTG || || || | | |||| ||| ||| 87 TTGACAGGTACCCAACTGTGCTGATGTA. TTGCTGGCCAAGGACTG. . . 814 AGTGTTTGAGCCTCTGTTTGTGTGTAATTGAGTGTGCATGTGTGGGAGTG | | |||||| | | 136 AAGGATC. . . TCAGTAATCATGCACCTATGTGGCGG. . . 864 AAATTGTGGAATGTGTATGCTCATAGCACTGAGTGAAAATAAAAGATTGT ||| || ||||| || |||||| | 173 AAA. TATGGGATATGCATGTCGA. . . CACTGAGTG. . AAGGCAAGATTAT 813 135 863 172 913 216

Restriction mapping: Genes can be analysed to detect gene sequences that can be cleaved with restriction enzymes 50 Ace. III Alu. I Alw. I Apo. I Ban. II Bfa. I Bfi. I Bsa. XI Bsg. I Bsi. HKAI Bsp 1286 I Bsr. FI Cje. I Cvi. JI Cvi. RI Dde. I Dpn. I Eco. RI Hinf. I Mae. III Mnl. I Mse. I Msp. I Nde. I Sau 3 AI Sst. I Tfi. I Tsp 45 I Tsp 509 I Tsp. RI 100 150 200 250 1 2 1 2 1 1 1 2 4 1 2 2 1 1 2 1 2 1 3 1 CAGCTCnnnnnnn’nnn. . . AG’CT GGATCnnnn’n_ r’AATT_y G_r. GCy’C C’TA_G ACTGGG ACnnnnn. CTCC GTGCAGnnnnnn. . . G_w. GCw’C G_d. GCh’C ACTG_Gn’ r’CCGG_y CCAnnnnnn. GTnnnnnn. . . r. G’Cy TG’CA C’Tn. A_G GA’TC G’AATT_C G’An. T_C ’GTn. AC_ CCTCnnnnnn_n’ T’TA_A C’CG_G CA’TA_TG ’GATC_ G_AGCT’C G’Aw. T_C ’GTs. AC_ ’AATT_ CAGTGnn’

PCR Primer Design: Oligonucleotides for use in the polymerization chain reaction can be designed using computer based programs OPTIMAL primer length MINIMUM primer length MAXIMUM primer length OPTIMAL primer melting temperature MINIMUM acceptable melting temp MAXIMUM acceptable melting temp MINIMUM acceptable primer GC% MAXIMUM acceptable primer GC% Salt concentration (m. M) DNA concentration (n. M) MAX no. unknown bases (Ns) allowed MAX acceptable self-complementarity MAXIMUM 3' end self-complementarity GC clamp how many 3' bases --> --> --> --> 20 18 22 60. 000 57. 000 63. 000 20. 000 80. 000 50. 000 0 12 8 0
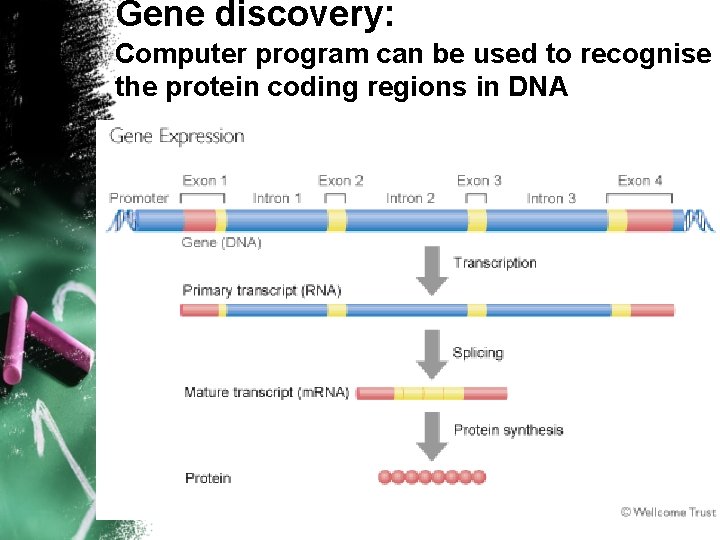
Gene discovery: Computer program can be used to recognise the protein coding regions in DNA
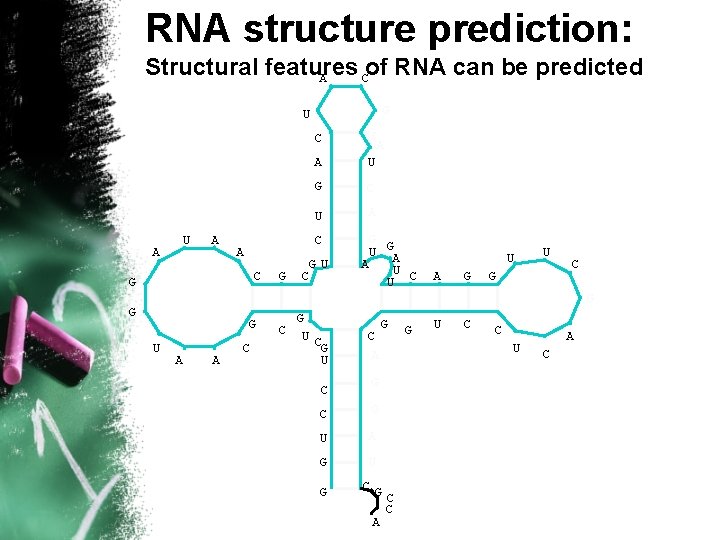
RNA structure prediction: Structural features of RNA can be predicted A C G U A G A U G C U A C A C G G G U A A C GU C G U GA A U U C C A G G U A G U G CG A A G C C U G C G U A C
Multiple sequence alignment: Sequences of proteins from different organisms can be aligned to see similarities and differences Alignment formatted using Mac. Boxshade
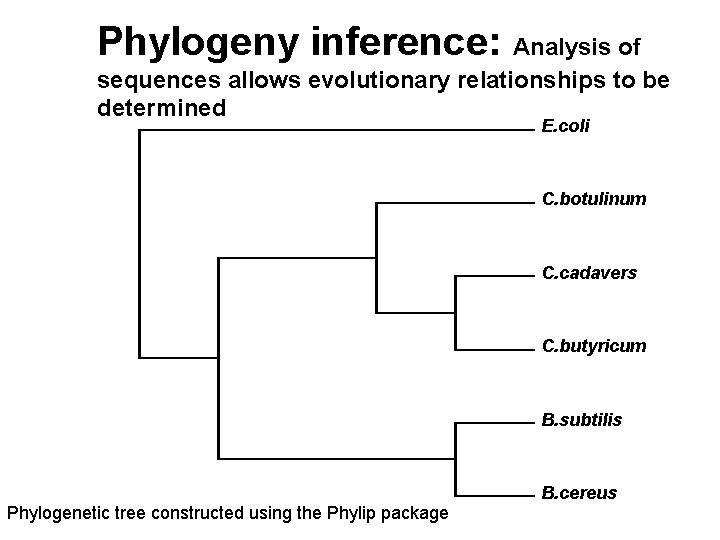
Phylogeny inference: Analysis of sequences allows evolutionary relationships to be determined E. coli C. botulinum C. cadavers C. butyricum B. subtilis Phylogenetic tree constructed using the Phylip package B. cereus

Large scale bioinformatics: genome projects • Function assignment Using database searches, • Mapping pattern searches, protein family analysis and structure Identifying the location of prediction to assign a clones and markers on the function to each predicted chromosome by genetic gene linkage analysis and physical mapping Data mining • Sequencing Searching for relationships Assembling clone sequence and correlations in the information reads into large (eventually complete) genome • Genome comparison sequences Comparing different • Gene discovery complete genomes to infer Identifying coding regions in evolutionary history and genomic DNA by database genome rearrangements searching and other methods

Finding genes in genome sequence is not easy • About 1% of human DNA encodes functional genes. • Genes are interspersed among long stretches of non-coding DNA. • Repeats, pseudo-genes, and introns confound matters

DNA chip microarrays • Put a large number (~100 K) of c. DNA sequenc or synthetic DNA oligomers onto a glass slide (or other substrate) in known locations on a gr • Label an RNA sample and hybridize • Measure amounts of RNA bound to each squa in the grid • Make comparisons – Cancerous vs. normal tissue – Treated vs. untreated – Time course • Many applications in both basic and clinical research

Spot your own Chip Robot spotter Ordinary glass microscope slide
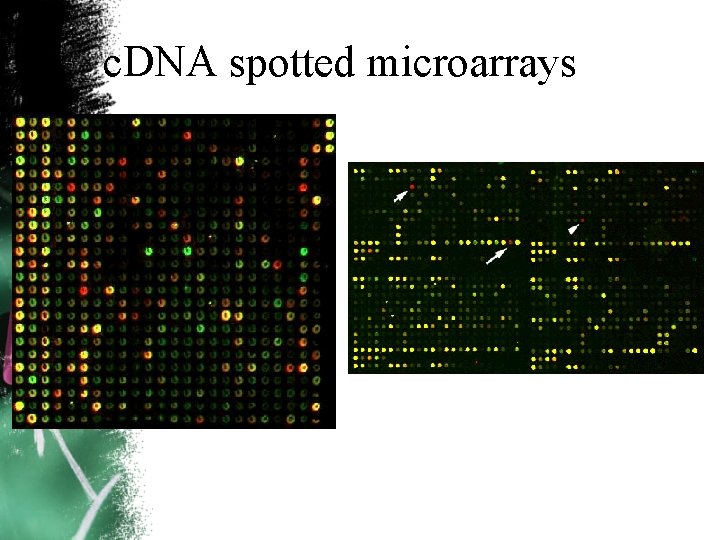
c. DNA spotted microarrays


Goal of Microarray experiments • Microarrays are a very good way of identifying a bunch of genes involved in a disease process – Differences between cancer and normal tissue – Tuberculosis infected vs resistant lung cells • Mapping out a pathway – Co-regulated genes • Finding function for unknown genes – Involved these processes

Direct Medical Applications • Diagnosis – Type of cancer – Aggressive or benign? • Monitor treatment outcome – Is a treatment having the desired effect on the target tissue?
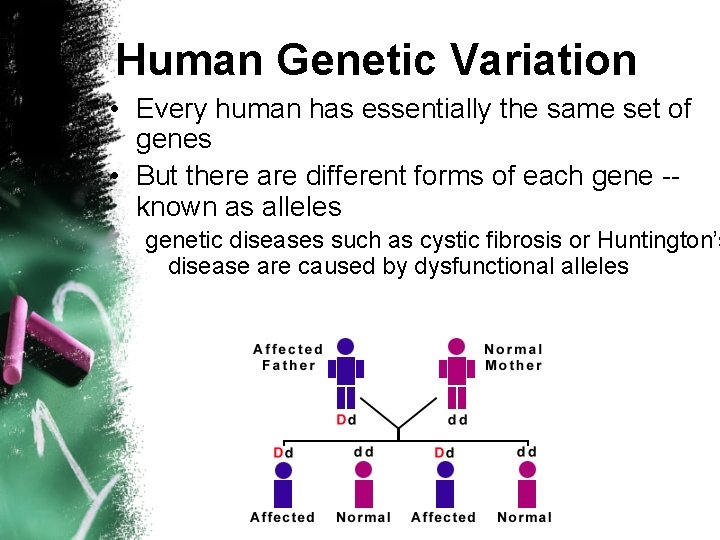
Human Genetic Variation • Every human has essentially the same set of genes • But there are different forms of each gene -known as alleles genetic diseases such as cystic fibrosis or Huntington’s disease are caused by dysfunctional alleles

• On chromosome four the sequence CAG is repeated six or more times. • The other strand of DNA in this gene has repeats of GTC. • This sequence happens to lie within a gene, called huntingtin. • longer the repeat of CAG, the greater is the probability mispairing happening. • individual born with extra CAGs in the huntingtin gene is likely to develop Huntington's chorea. The more extra CAGs there are in the gene, the earlier in life the disease will show up

Clinical Manifestations of Genetic Variation (All disease has a genetic component) • Susceptibility vs. resistance • Variations in disease severity or symptoms • Reaction to drugs (pharmacogenetics) All of these traits can be traced back to particular genes (or sets of genes)

Pharmacogenomics • People react differently to drugs – Side effects – Variable effectiveness • There are genes that control these reactions • SNP markers can be used to identify these genes (profiles)

Use the Profiles • Genetic profiles of new patients can then be used to prescribe drugs more effectively & avoid adverse reactions. – Sell a drug with a gene test • Can also speed clinical trials by testing on those who are likely to respond well.

Toxicogenomics ¨ There a number of common pathways for drug toxicity (or environmental tox. ) ¨ It is possible to compile genomic signatures (gene expression data) for these pathways. ¨ Candidate drug molecules can be screened in cell culture or in animals for induction of these toxicity pathways.
Planning for a Genomics Revolution • Bioinformatics support must be integral in the planning process for the development of new genomics research facilities. • Genome Project sequencing centers have more staff and more $$$ spent on data analysis than on the sequencing itself. • Microarray facilities will be even more skewed toward data analysis • It is an information-intensive business!

Implications for Biomedicine • Physicians will use genetic information to diagnose and treat disease. • Virtually all medical conditions have a genetic component. • Faster drug development research • Individualized drugs • Gene therapy • All Biologists will use gene sequence information in their daily work

Training "computer savvy" scientists u Know the right tool for the job u Get the job done with tools available u u Network connection is the lifeline of the scientist Jobs change, computers change, projects change, scientists need to be adaptable

Long Term Implications A "periodic table for biology" will lead to an explosion of research and discoveries - we will finally have the tools to start making systematic analyses of biological processes (quantitative biology). u Understanding the genome will lead to the ability to change it - to modify the characteristics of organisms and people in a wide variety of ways u

Genomics Education • Genomics scientists need basic training in both Molecular Biology and Computing • Specific training in the use of automated laboratory equipment, the analysis of large datasets, and bioinformatics algorithms • Particularly important for the training of medical doctors - at least a familiarity with the technology

Genomics in Medical Education The explosion of information about the new genetics will create a huge problem in health education. Most physicians in practice have had not a single hour of education in genetics and are going to be severely challenged to pick up this new technology and run with it.
- Slides: 61