BIOINFORMATICS Paulin Hogeweg1979 Biobiology InformatiqueData processing Study of

BIOINFORMATICS

• Paulin Hogeweg(1979) – • Bio-biology, Informatique-Data processing • Study of information content and flow in a biological system with the help of IT. • Multidisciplinary branch • Historical background: • 1961 -identified m. RNA- Jacob, Meselson, Francois Jacob • 1977 -DNA Sequencing-Sanger, Gilbert • 1983 -Release of GEN Bank • 1983 -Sequence database sequence algorithm

• • • 1985 Similarity searching—FASTP&FASTN 1988 NCBI By NIH/NLM 1990 -BLAST-Pairwise sequence similarity search 1991 -Expressed sequence tag sequencing 1994 EMBL 1995 First bacterial genome analysed 1996 Yeast genome analysed 2000 -Pseudomonas genome , A. thaliana genome 2001 -Human Genome 2005 -Rice Genome sequenced.

BIOLOGICAL DATABASES • Database is a System to store, search and retrieve any type of data • Biological database-Information of DNA, RNA, Proteins, 3 D images, Organism specific informations. • PRIMARY, SECONDARY &DERIVATIVE DATABASES

• PRIMARY DATABASES: Information of Nucleic acid, Protein, and their structure. Original submission by experimentalists and controlled by original submitter. Eg: Gen Bank, SWISS PROT, PDB SECONDARY DATABASES: With additional information from analysis of primary databases. Tr. EMBL, PROSITE DERIVATIVE DATABASES/COMPOSITE DATABASES: From primary databases. controlled by third party. Eg. NCBI, Ref. Seq, Uni. Gene.

EMBL(EUROPEAN MOLECULAR BIOLOGY LABORATORY) • Nucleotide sequence database • Collect and present nucleotide sequence and annotation with comprehensive global coverage. • Integrate nucleotide sequences and given to EBI • Through SRS(Sequence retriesval system) data can be viewed over 200 libraries. • Genome webserver includes 2494 completed genomes.

GEN BANK • Collection of DNA sequences • 106, 533, 756 bases in 108, 431, 92 records • Collaboration with DDBJ, EMBL and NCBI

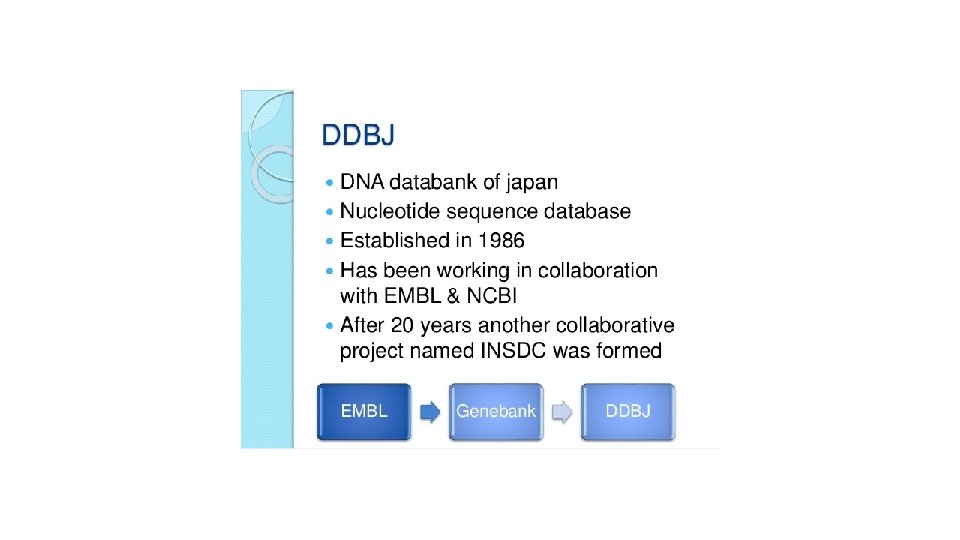
DDBJ(DNA DATABASE OF JAPAN) • Established in 1986 at NIG(National Institute of Genetics) • Collaboration with EMBL and Gen Bank • Improve the quality of INSD(International Nucleotide Database) • Collect information from Scientists of Japan and other countries. • Sequence submission tool is SAKURA

PROTEIN SEQUENCE DATABASE PDB( PROTEIN DATA BANK) Information portal to macromolecules-proteins, nucleic acids Estabished in Brookhaven National Laboratory Maintained by RCSB(Research Collaboratory for Structural Bioinformatics) Study the function through the study of 3 D structure of macromolecules of different living organisms. • Obtained by x-ray crystallography or NMR spectroscopy • 65075 structures in PDB • URL is http: //www. pdb. org • • •



Uni. Prot

PIR(PROTEIN INFORMATION RESOURCE) • Established in 1984 by NBRF(National Biomedical Research Foundation) • Located at Georgetown University Medical Centre. • Assist researchers in the identification and interpretation of Protein sequence information. • Support proteomic, genomic and systems biology research. • URL is http: //pir. Georgetown. edu/.








ORGANISMAL DATABASE • HUMAN GENOME DATABASES: • Repository of human genes, clones, STSs, polymorphisms and maps. by GDB Map viewer Search methods Find an object with a known GDB Accession number Find an object with a known sequence database accession number Find objects having specific name Find objects that contain one or more keywords anywhere in their text.

BIODIVERSITY DATABASE-Species 2000

SEQUENCE ANALYSIS/SEQUENCE ALIGNMENT • Compare two or more sequences of nucleic acids or protein to find out the similarity. • Homologous • Orthologous • Paralogous • Xenologous: sequences in different species arose by horizontal or lateral gene transfer

TYPES OF SEQUENCE ALIGNMENT • PAIRWISE ALIGNMENT • MULTIPLE SEQUENCE ALIGNMENT • According to the length of sequences • GLOBAL SEQUENCE ALIGNMENT • LOCAL SEQUENCE ALIGNMENT

TOOLS FOR SEQUENCE ALIGNMENT • BLAST: Basic Local Alignment Search Tool • Developed by Altshchul et al • For pairwise alignment • Uses in DNA and protein sequence databases • Available with NCBI website • Types: • BLAST P-Protein against protein • BLAST N-Nucleotide against nucleotide

• BLAST X-NUCLEOTIDE QUERY TRANSLATED TO PROTEIN AGAINST PROTEIN DATABASE • TBALSTN: PROTEIN AGAINST NUCLEOTIDE DATABASE AFTERTRANSLATING NUCLEOTIDE DATABASE SEQUENCE IN TO PROTEIN • TBLASTX: NUCLEOTIDE DATABASE TRANSLTED TO PROTEIN AGAINST NUCLEOTIDE TRANSLATED PROTEIN QUERY.

Clustal w • Multiple sequence alignment program for DNA and PROTEINS. • Described by Higgins and Sharp • It calculates best match for specific sequences and finds similarities, identities and differences • Alignment method: • PAIRWISE ALIGNMENT of different sequnces • PREPARATION OF PHYLOGENETIC TREE • Used to align any group of proteins or nucleic acids









- Slides: 39