Bioinformatics Overview School of BI TCD May 2010

Bioinformatics Overview School of B&I TCD May 2010

Who, me? • • Andrew Lloyd atlloyd@tcd. ie 087 -225 -9850, 053 -9255717, 01 -896 -2450 Director INCBI 1993 -2000 Population genetics, evolution Whole genome analysis Immunology, chickens, FIRM

Definition/scope • Storage, retrieval and analysis of biological (sequence) information. • Insert better definition here • Case can be made for microarray analysis • NOT – ecoinformatics (ecology) – Image analysis – Bar-coding hospital sheets

Philosophy “Nothing worth learning can be taught” Oscar Wilde

Getting bioinformation • Type it in: A, T, C, C, G, T, C, A (1991) • Access databases – Literature (Pubmed) – Medical (OMIM) – DNA sequence (EMBL/Gen. Bank) – Protein sequence (Uni. Prot, Swiss. Prot, PIR) – 3 -D structure (PDB)

Annotation • In any DB, half is data and half context. – Gene ontology (language) – Parsing sequence (ORF, RBS, Intron, -helix) – Recognising similar sequences (evolution!) – Complementary info : DB cross-referencing • (DNA -> Protein -> 3 D structure -> motifs)

Secondary databases • • Protein motifs, domains, families RNA structures (16 S ribosomal RNA…) Taxonomy/classification Metabolic pathways (KEGG) Enzymes (Brenda, TCD, Ireland) SNPs: mutations and variants Disease DBs (OMIM) Immuno, epitope DBs

Complete genomes • Ensembl (complex, basically vertebrate) – Uniform look-and-feel; cross-refs • UCSC Golden. Path browser • Plants • Bacterial genomes – Including mitochondrial, chloroplast – Eubacteria vs Archaea vs Eukaryotes

Annotated/known genes • What does my gene do? • Blast (fasta) against the DB • SRS/Entrez to access databases – Neighboring (similar things in same DB) • DB cross-references – full picture of attributes – What biochemical pathway?
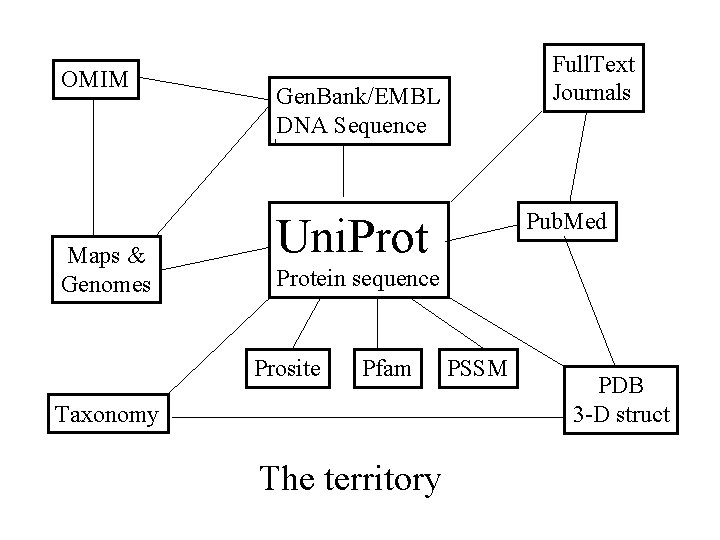
OMIM Maps & Genomes Full. Text Journals Gen. Bank/EMBL DNA Sequence Pub. Med Uni. Protein sequence Prosite Pfam Taxonomy The territory PSSM PDB 3 -D struct

Databases • BIG • EMBL/Gen. Bank 200 Gbp, 100 m entries, 2500 complete genomes, 200 K species • Encycl. Britannica 180 m letters. 40 m words • EMBL 1 km of Britannica Volumes • Doubling every 14 -18 mo • Human genome is X bp?

Intrinsic vs Context Internal • DNA, protein sequence – DNA: Purine/Pyrimidine – AAs: small, hydrophobic, aromatic, polar – Variants: SNPs, Indels, Alt Splicing • 2 ndry structure – DNA: stem/loops – Protein: helix, sheet, turn, loop

Intrinsic vs Context External, context for your molecule • In other species (homologs, phylog trees) • In which cellular location (GO) • Molecular complex (dimers) • Which pathway (KEGG) • Where in genome (neighbors, synteny)

New Unknown Gene • • Blast homology searching Genomic location/neighboring genes Where is it expressed? How regulated (control sequences) Intron/exon structure Domain structure Restriction sites etc. Primer design

DNA/gene structure • Four bases A T C G U – 2 pyrimidine, 2 purine – LOTS of them: how many? • • Open reading frame 5’ signals, 3’ signals Introns/exons Neighbours (operons)

Two sequences • Alignment – Local – Global • Dotplot • Threading

One seq vs many • • • Homology search vs database Special case of 2 -seq alignment Blast vs fasta Limit by species/taxon Substitution matrices Low complexity masking

Multiple sequence alignment • MSA • Progressive alignment • Clustal. W or (better) T-Coffee
Phylogenetic trees • Computationally intensive • Distance matrix methods – Neighbor-joining (NJ) – UPGMA • Minimum evolution • Maximum parsimony • Maximum likelihood – Bayesian methods

Genefinding • Special case of DNA analysis • How to annotate a genome • Bacterial – Find open reading frames (ORFs) – With start/stop codons – With promoter, RBS, CAAT, TATA • Eukaryotic – As above PLUS – Introns/exons – Alternative splicing
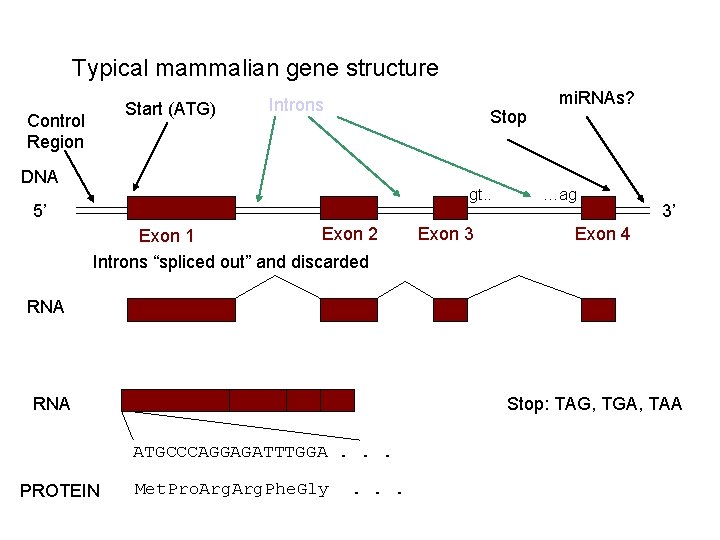
Typical mammalian gene structure Start (ATG) Control Region Introns Stop DNA gt. . 5’ Exon 2 Exon 1 Introns “spliced out” and discarded Exon 3 mi. RNAs? …ag 3’ Exon 4 RNA Stop: TAG, TGA, TAA ATGCCCAGGAGATTTGGA. . . PROTEIN Met. Pro. Arg. Phe. Gly . . .

Protein substructure • DNA makes protein and protein (enzymes) make everything else. • 20 Amino acids • Amino acid properties • Motifs • Domains • Biological units
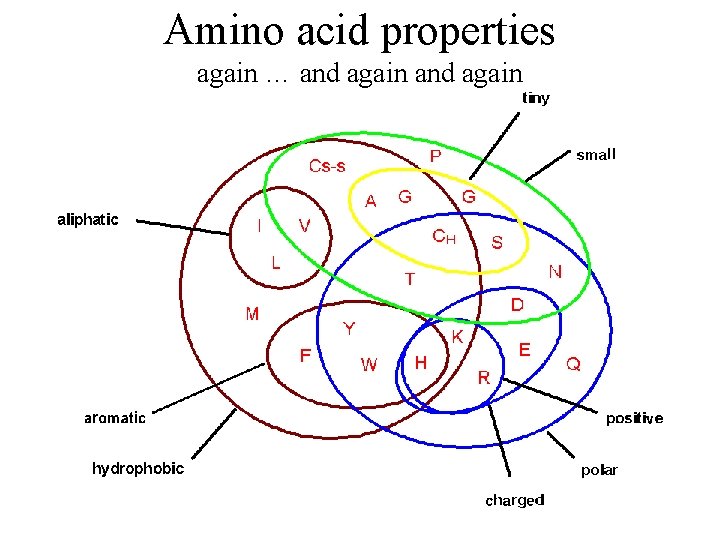
Amino acid properties again … and again
Protein 3 -D structure • Relationship between sequence & structure • Secondary structure – Alpha helix – Beta sheet – Coil – Turn • Threading sequence to homologous structure

Gene Expression • • EST SAGE Micro. Array Clustering of same expressed genes

Genomics • Complete DNA seq for a species • Gene order • Gene clusters/operons – Missing operons • Gene duplication • Whole genome duplication (WGD)

SNPs • Key issue in genetics is that two organisms are both the same and different: – Humans vs chimps vs mouse – Parent vs offspring vs co-national vs human • Single nucleotide polymorphisms • Variation between individuals • Pharmacogenetics – Personal tailored medicine

Summary/take home • Course designed to give you access to databases, software tools • …and ways of thinking about data
- Slides: 28