Bioinformatics Metabolic pathway analysis Jacques van Helden jvanhelducmb

Bioinformatics Metabolic pathway analysis Jacques van Helden jvanheld@ucmb. ulb. ac. be

Graph-based analysis of biochemical networks Examples of metabolic pathways Jacques van Helden jvanheld@ucmb. ulb. ac. be
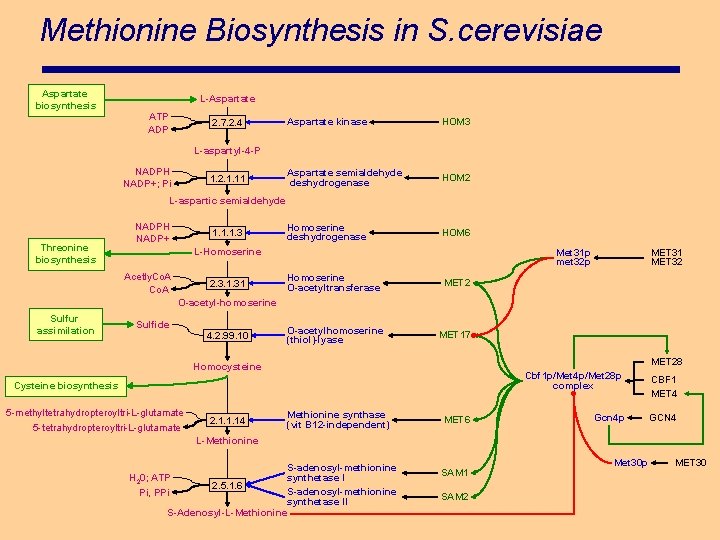
Methionine Biosynthesis in S. cerevisiae Aspartate biosynthesis L-Aspartate ATP ADP 2. 7. 2. 4 Aspartate kinase HOM 3 Aspartate semialdehyde deshydrogenase HOM 2 Homoserine deshydrogenase HOM 6 L-aspartyl-4 -P NADPH NADP+; Pi 1. 2. 1. 11 L-aspartic semialdehyde Threonine biosynthesis NADPH NADP+ 1. 1. 1. 3 L-Homoserine Acetly. Co. A 2. 3. 1. 31 Met 31 p met 32 p Homoserine O-acetyltransferase MET 2 O-acetylhomoserine (thiol)-lyase MET 17 MET 31 MET 32 O-acetyl-homoserine Sulfur assimilation Sulfide 4. 2. 99. 10 MET 28 Homocysteine Cysteine biosynthesis 5 -methyltetrahydropteroyltri-L-glutamate 5 -tetrahydropteroyltri-L-glutamate 2. 1. 1. 14 Methionine synthase (vit B 12 -independent) MET 6 Cbf 1 p/Met 4 p/Met 28 p complex CBF 1 MET 4 Gcn 4 p GCN 4 L-Methionine S-adenosyl-methionine synthetase I H 20; ATP 2. 5. 1. 6 S-adenosyl-methionine Pi, PPi synthetase II S-Adenosyl-L-Methionine SAM 1 SAM 2 Met 30 p MET 30
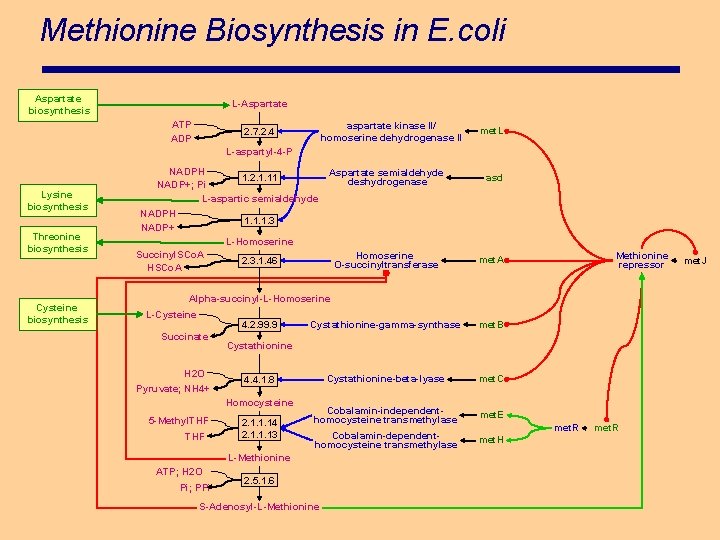
Methionine Biosynthesis in E. coli Aspartate biosynthesis L-Aspartate ATP ADP aspartate kinase II/ homoserine dehydrogenase II 2. 7. 2. 4 met. L L-aspartyl-4 -P Lysine biosynthesis Threonine biosynthesis Cysteine biosynthesis NADPH 1. 2. 1. 11 NADP+; Pi L-aspartic semialdehyde NADPH NADP+ Aspartate semialdehyde deshydrogenase asd 1. 1. 1. 3 L-Homoserine Succinyl. SCo. A Homoserine O-succinyltransferase met. A Cystathionine-gamma-synthase met. B Cystathionine-beta-lyase met. C Cobalamin-independenthomocysteine transmethylase met. E Cobalamin-dependenthomocysteine transmethylase met. H 2. 3. 1. 46 Methionine repressor Alpha-succinyl-L-Homoserine L-Cysteine 4. 2. 99. 9 Succinate H 2 O Pyruvate; NH 4+ Cystathionine 4. 4. 1. 8 Homocysteine 5 -Methyl. THF 2. 1. 1. 14 2. 1. 1. 13 L-Methionine ATP; H 2 O Pi; PPi 2. 5. 1. 6 S-Adenosyl-L-Methionine met. R met. J
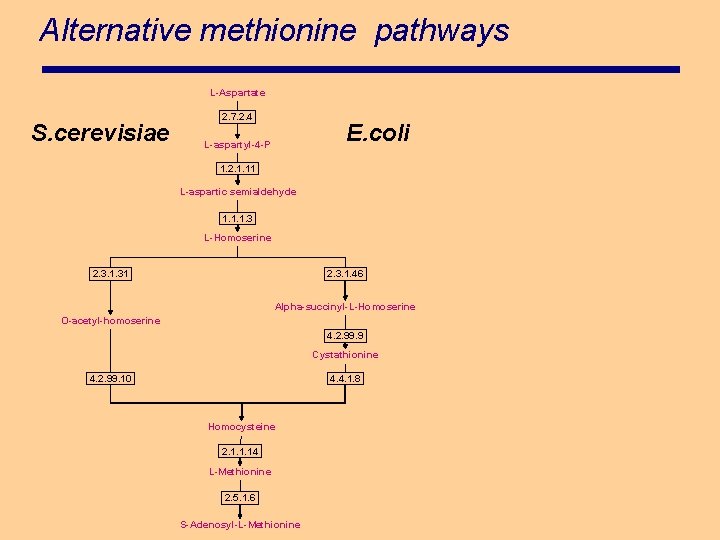
Alternative methionine pathways L-Aspartate S. cerevisiae 2. 7. 2. 4 E. coli L-aspartyl-4 -P 1. 2. 1. 11 L-aspartic semialdehyde 1. 1. 1. 3 L-Homoserine 2. 3. 1. 31 2. 3. 1. 46 Alpha-succinyl-L-Homoserine O-acetyl-homoserine 4. 2. 99. 9 Cystathionine 4. 2. 99. 10 4. 4. 1. 8 Homocysteine 2. 1. 1. 14 L-Methionine 2. 5. 1. 6 S-Adenosyl-L-Methionine
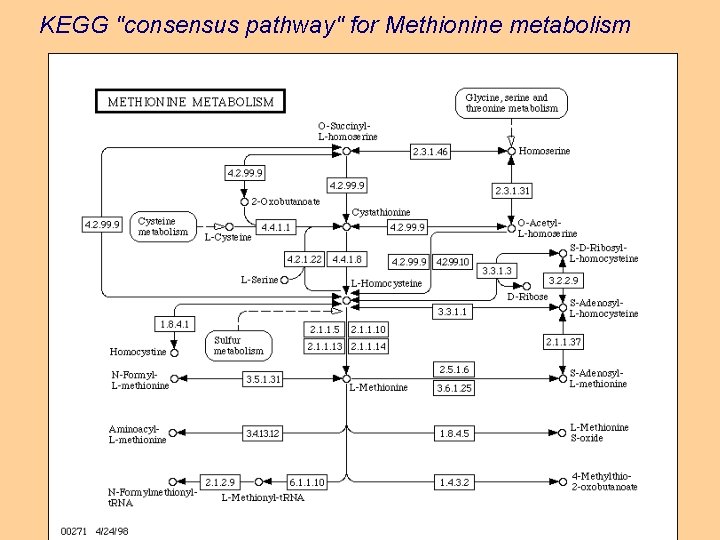
KEGG "consensus pathway" for Methionine metabolism
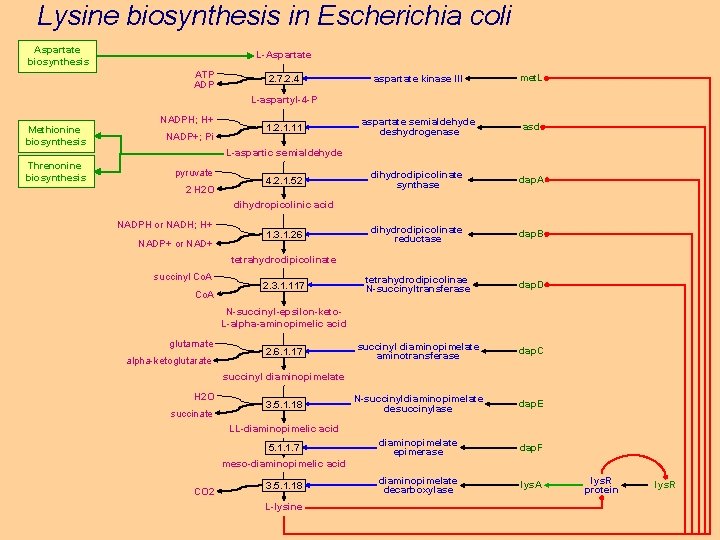
Lysine biosynthesis in Escherichia coli Aspartate biosynthesis L-Aspartate ATP ADP 2. 7. 2. 4 aspartate kinase III met. L aspartate semialdehyde deshydrogenase asd dihydrodipicolinate synthase dap. A dihydrodipicolinate reductase dap. B tetrahydrodipicolinae N-succinyltransferase dap. D succinyl diaminopimelate aminotransferase dap. C N-succinyldiaminopimelate desuccinylase dap. E diaminopimelate epimerase dap. F diaminopimelate decarboxylase lys. A L-aspartyl-4 -P Methionine biosynthesis Threnonine biosynthesis NADPH; H+ NADP+; Pi 1. 2. 1. 11 L-aspartic semialdehyde pyruvate 2 H 2 O 4. 2. 1. 52 dihydropicolinic acid NADPH or NADH; H+ NADP+ or NAD+ 1. 3. 1. 26 tetrahydrodipicolinate succinyl Co. A 2. 3. 1. 117 N-succinyl-epsilon-keto. L-alpha-aminopimelic acid glutamate alpha-ketoglutarate 2. 6. 1. 17 succinyl diaminopimelate H 2 O succinate 3. 5. 1. 18 LL-diaminopimelic acid 5. 1. 1. 7 meso-diaminopimelic acid CO 2 3. 5. 1. 18 L-lysine lys. R protein lys. R
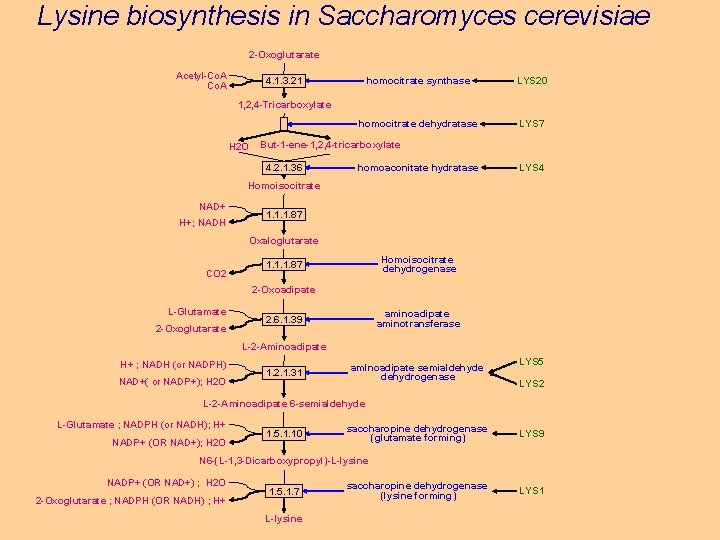
Lysine biosynthesis in Saccharomyces cerevisiae 2 -Oxoglutarate Acetyl-Co. A homocitrate synthase LYS 20 homocitrate dehydratase LYS 7 4. 1. 3. 21 1, 2, 4 -Tricarboxylate H 2 O But-1 -ene-1, 2, 4 -tricarboxylate 4. 2. 1. 36 homoaconitate hydratase LYS 4 Homoisocitrate NAD+ H+; NADH 1. 1. 1. 87 Oxaloglutarate CO 2 Homoisocitrate dehydrogenase 1. 1. 1. 87 2 -Oxoadipate L-Glutamate 2 -Oxoglutarate aminoadipate aminotransferase 2. 6. 1. 39 L-2 -Aminoadipate H+ ; NADH (or NADPH) NAD+( or NADP+); H 2 O 1. 2. 1. 31 amlnoadipate semialdehyde dehydrogenase LYS 5 LYS 2 L-2 -Aminoadipate 6 -semialdehyde L-Glutamate ; NADPH (or NADH); H+ NADP+ (OR NAD+); H 2 O 1. 5. 1. 10 saccharopine dehydrogenase (glutamate forming) LYS 9 N 6 -(L-1, 3 -Dicarboxypropyl)-L-lysine NADP+ (OR NAD+) ; H 2 O 2 -Oxoglutarate ; NADPH (OR NADH) ; H+ 1. 5. 1. 7 L-lysine saccharopine dehydrogenase (lysine forming) LYS 1
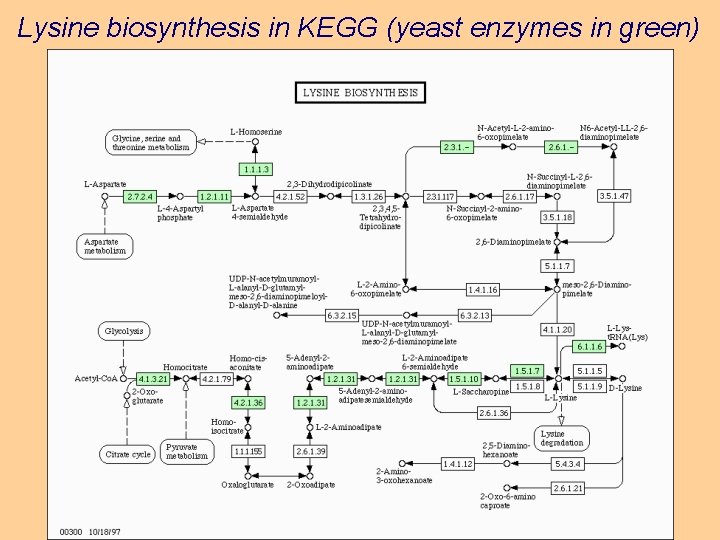
Lysine biosynthesis in KEGG (yeast enzymes in green)
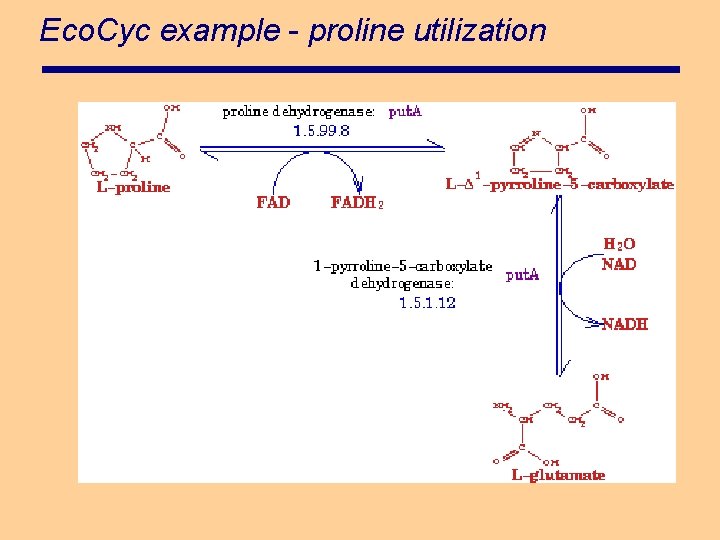
Eco. Cyc example - proline utilization
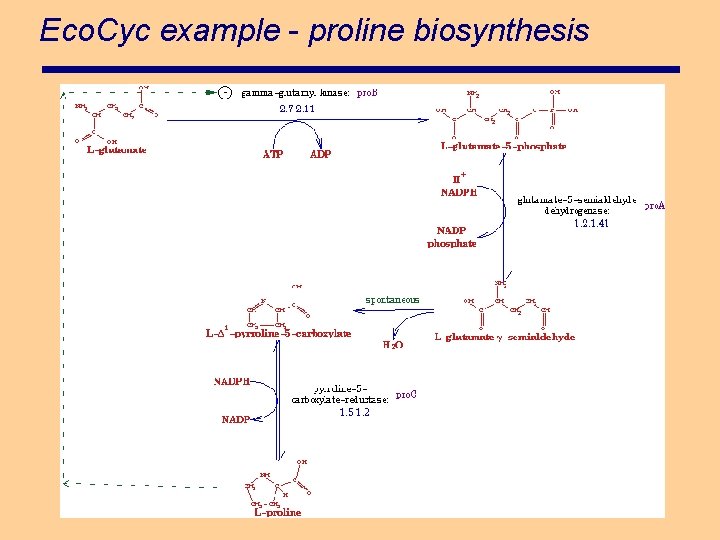
Eco. Cyc example - proline biosynthesis
Ecocyc - metabolic overview
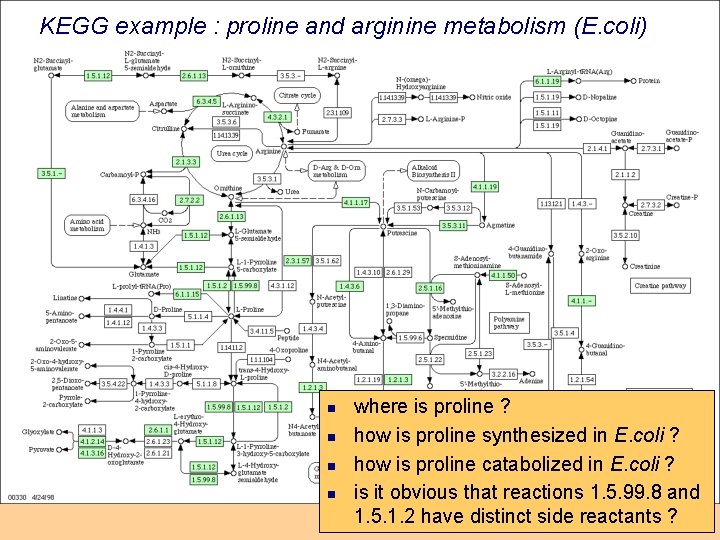
KEGG example : proline and arginine metabolism (E. coli) n n where is proline ? how is proline synthesized in E. coli ? how is proline catabolized in E. coli ? is it obvious that reactions 1. 5. 99. 8 and 1. 5. 1. 2 have distinct side reactants ?

Graph-based analysis of biochemical networks Pathway reconstruction by reaction clustering Jacques van Helden jvanheld@ucmb. ulb. ac. be
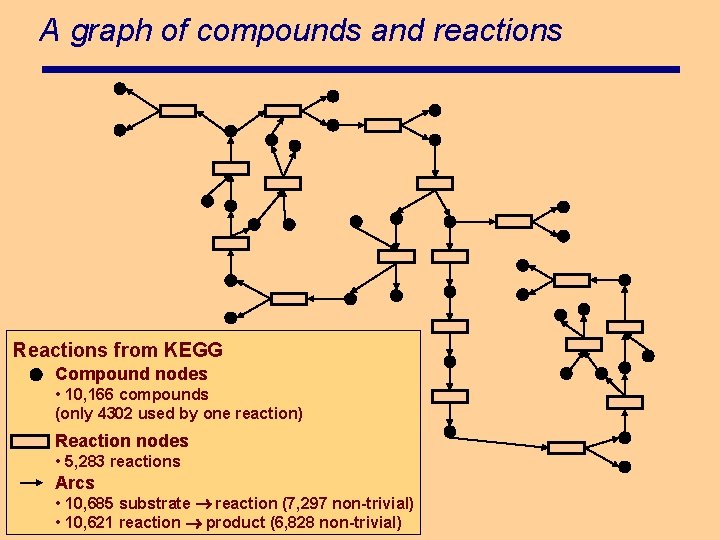
A graph of compounds and reactions Reactions from KEGG Compound nodes • 10, 166 compounds (only 4302 used by one reaction) Reaction nodes • 5, 283 reactions Arcs • 10, 685 substrate reaction (7, 297 non-trivial) • 10, 621 reaction product (6, 828 non-trivial)
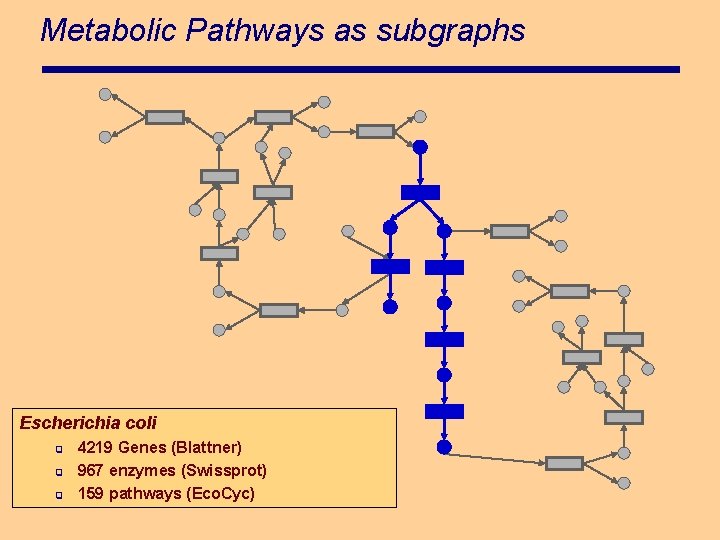
Metabolic Pathways as subgraphs Escherichia coli q q q 4219 Genes (Blattner) 967 enzymes (Swissprot) 159 pathways (Eco. Cyc)

Reconstructing a pathway from a subset of reactions n Input: q n a set of reactions (the seed reactions) Output: q a metabolic pathway including • • q the seed reactions, together with their substrates and products optionally, some additional reactions, interaalated to improve the pathway connectivity the pathway can either be connected, or contain several unconnected components
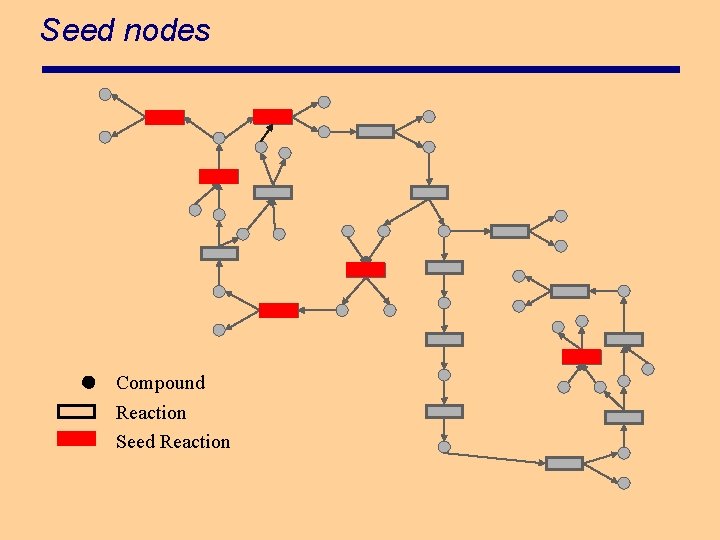
Seed nodes Compound Reaction Seed Reaction
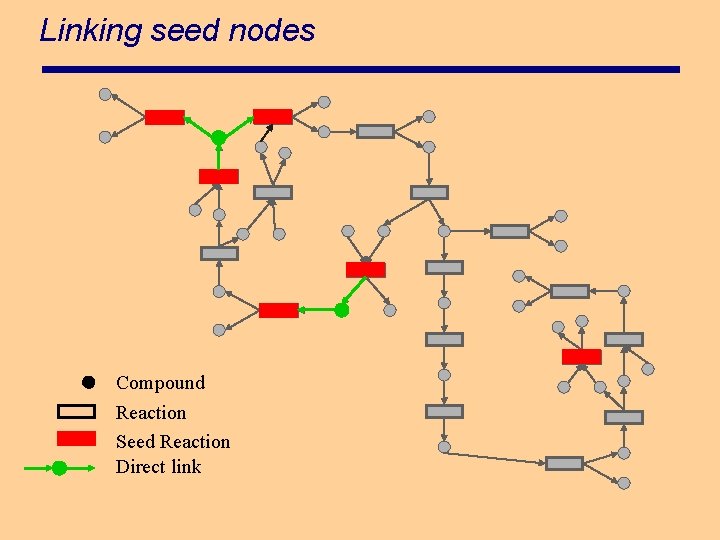
Linking seed nodes Compound Reaction Seed Reaction Direct link
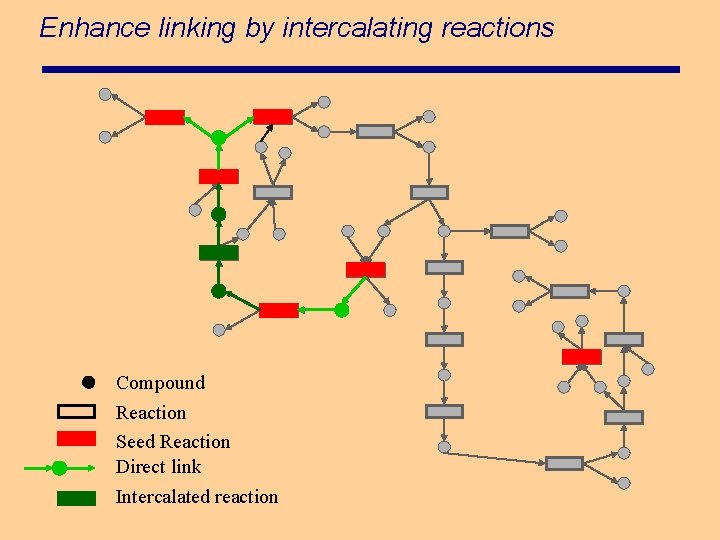
Enhance linking by intercalating reactions Compound Reaction Seed Reaction Direct link Intercalated reaction
Subgraph extraction

Validation of the method n Take a set of experimentally characterized pathways, and for each one q q Select a subset of enzymes Use the reactions catalysed by these enzymes as seed nodes Extract the subgraph Compare with known pathway
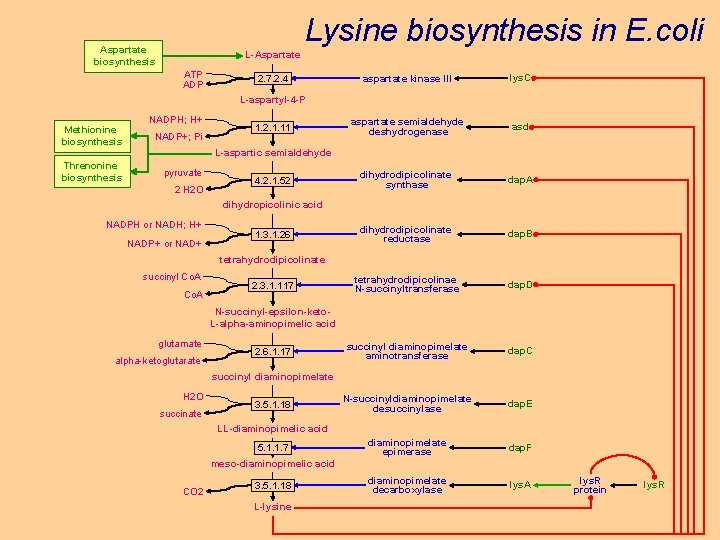
Lysine biosynthesis in E. coli Aspartate biosynthesis L-Aspartate ATP ADP 2. 7. 2. 4 aspartate kinase III lys. C aspartate semialdehyde deshydrogenase asd dihydrodipicolinate synthase dap. A dihydrodipicolinate reductase dap. B tetrahydrodipicolinae N-succinyltransferase dap. D succinyl diaminopimelate aminotransferase dap. C N-succinyldiaminopimelate desuccinylase dap. E diaminopimelate epimerase dap. F diaminopimelate decarboxylase lys. A L-aspartyl-4 -P Methionine biosynthesis Threnonine biosynthesis NADPH; H+ NADP+; Pi 1. 2. 1. 11 L-aspartic semialdehyde pyruvate 2 H 2 O 4. 2. 1. 52 dihydropicolinic acid NADPH or NADH; H+ NADP+ or NAD+ 1. 3. 1. 26 tetrahydrodipicolinate succinyl Co. A 2. 3. 1. 117 N-succinyl-epsilon-keto. L-alpha-aminopimelic acid glutamate alpha-ketoglutarate 2. 6. 1. 17 succinyl diaminopimelate H 2 O succinate 3. 5. 1. 18 LL-diaminopimelic acid 5. 1. 1. 7 meso-diaminopimelic acid CO 2 3. 5. 1. 18 L-lysine lys. R protein lys. R
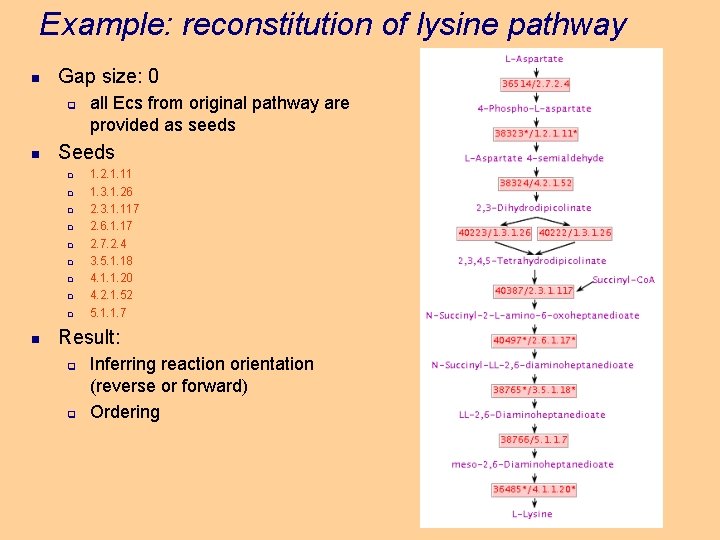
Example: reconstitution of lysine pathway n Gap size: 0 q n Seeds q q q q q n all Ecs from original pathway are provided as seeds 1. 2. 1. 11 1. 3. 1. 26 2. 3. 1. 117 2. 6. 1. 17 2. 7. 2. 4 3. 5. 1. 18 4. 1. 1. 20 4. 2. 1. 52 5. 1. 1. 7 Result: q q Inferring reaction orientation (reverse or forward) Ordering
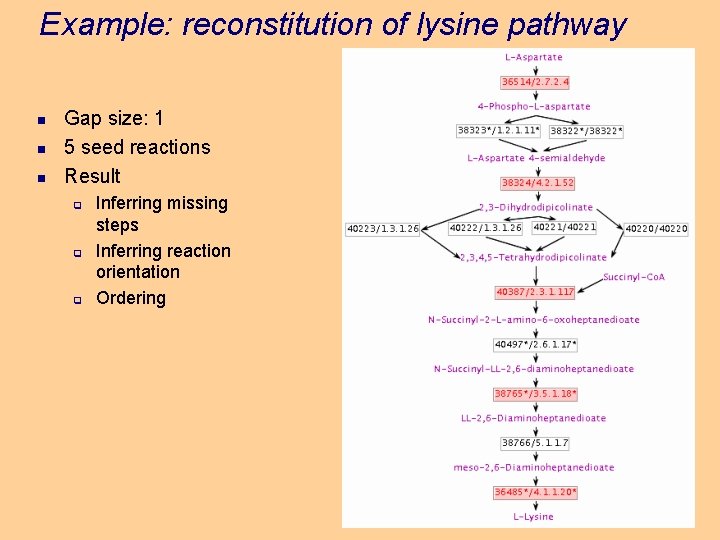
Example: reconstitution of lysine pathway n n n Gap size: 1 5 seed reactions Result q q q Inferring missing steps Inferring reaction orientation Ordering
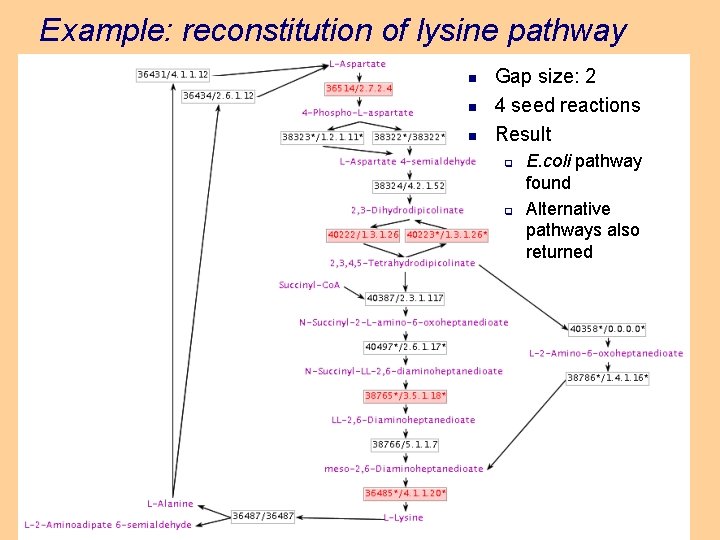
Example: reconstitution of lysine pathway n n n Gap size: 2 4 seed reactions Result q q E. coli pathway found Alternative pathways also returned
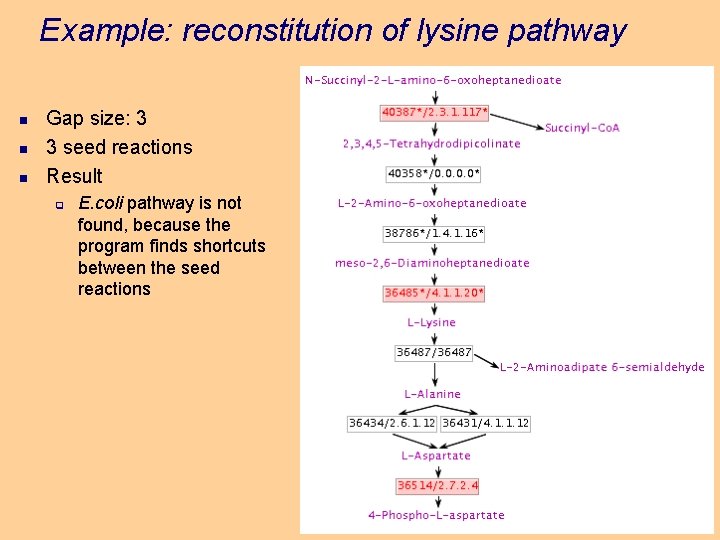
Example: reconstitution of lysine pathway n n n Gap size: 3 3 seed reactions Result q E. coli pathway is not found, because the program finds shortcuts between the seed reactions

Applications of pathway reconstruction n n We have the complete genome for dozens of bacteria, for which there is almost no experimental characterization of metabolism For these genomes, enzymes have been predicted by sequence similarity In some cases, one expects to find the same pathways as in model organisms, but in other cases, variants or completely distinct pathways For each known pathway from model organisms q q Select the subset of reactions for which an enzyme exists in the target organism If a reasonable number of reactions are present • Using these as seeds, reconstruct a pathway • Preferentially (but not exclusively) intercalate reactions for which an enzyme has been detected in the target organism

Graph-based analysis of biochemical networks From gene expression data to pathways Jacques van Helden jvanheld@ucmb. ulb. ac. be

Reaction clustering and gene expression data n n Many biochemical pathways are co-regulated at the transcriptional level. Starting from the observation that a group of genes is coregulated, try to find if they could be involved in a common pathway.
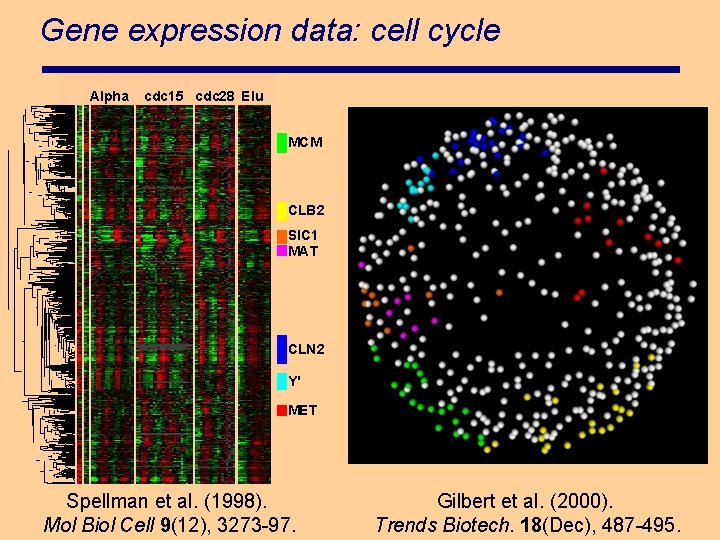
Gene expression data: cell cycle Alpha cdc 15 cdc 28 Elu MCM CLB 2 SIC 1 MAT CLN 2 Y' MET Spellman et al. (1998). Mol Biol Cell 9(12), 3273 -97. Gilbert et al. (2000). Trends Biotech. 18(Dec), 487 -495.

Study case : cluster of co-regulated genes
KEGG - gene search in pathway maps
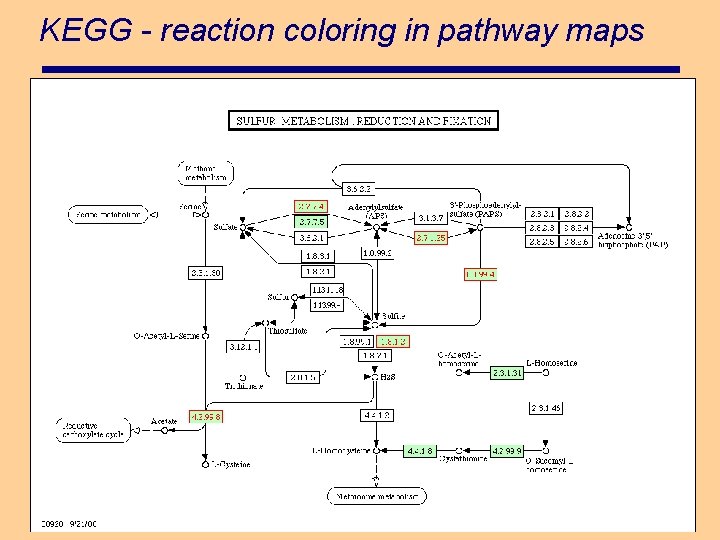
KEGG - reaction coloring in pathway maps
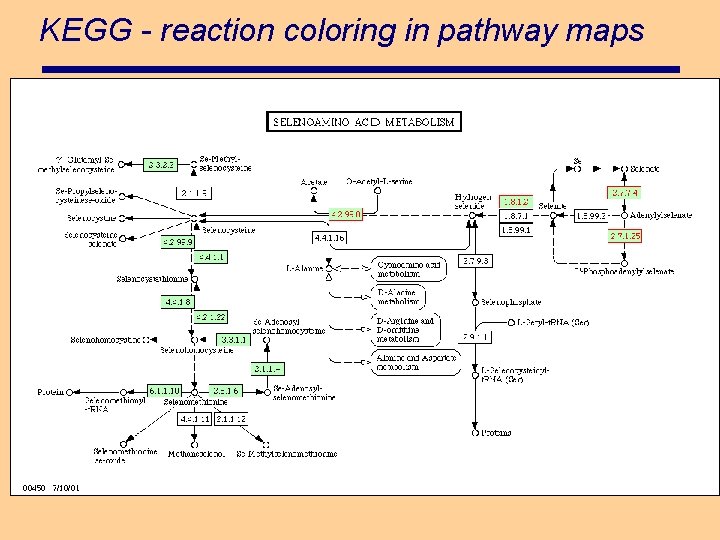
KEGG - reaction coloring in pathway maps
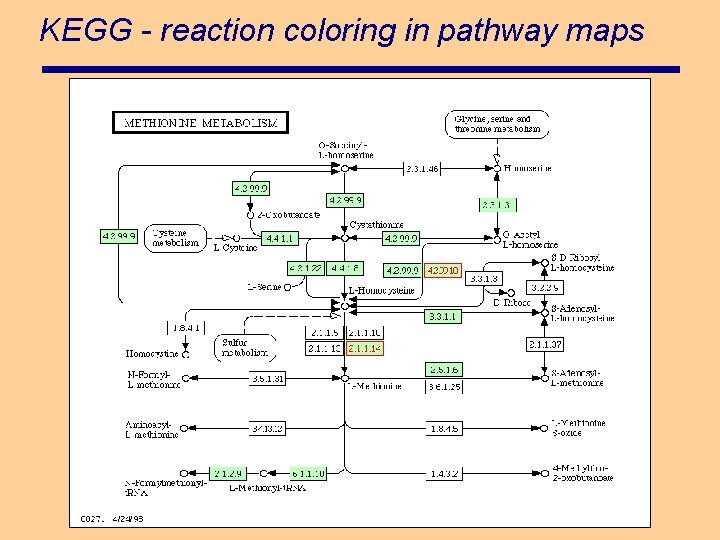
KEGG - reaction coloring in pathway maps
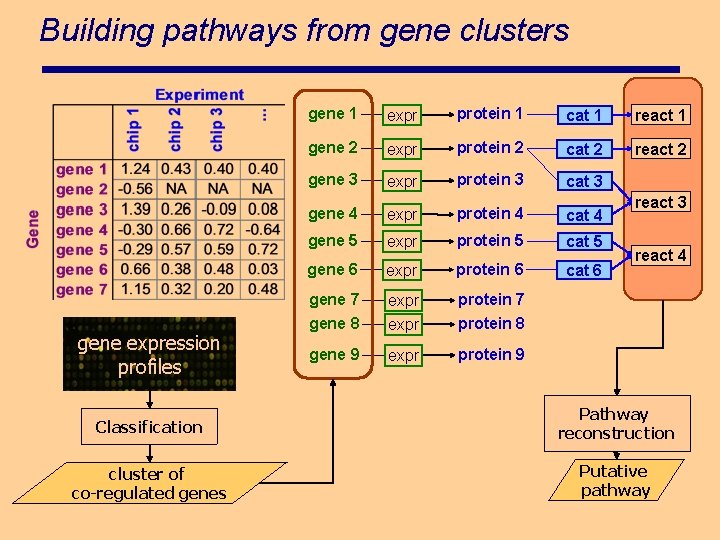
Building pathways from gene clusters gene expression profiles gene 1 expr protein 1 cat 1 react 1 gene 2 expr protein 2 cat 2 react 2 gene 3 expr protein 3 cat 3 gene 4 expr protein 4 cat 4 gene 5 expr protein 5 cat 5 gene 6 expr protein 6 cat 6 gene 7 gene 8 expr protein 7 protein 8 gene 9 expr protein 9 react 3 react 4 Classification Pathway reconstruction cluster of co-regulated genes Putative pathway
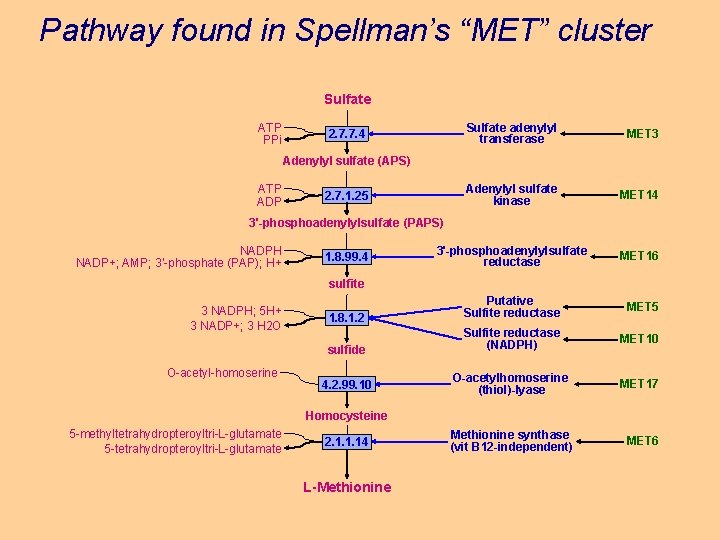
Pathway found in Spellman’s “MET” cluster Sulfate ATP PPi Sulfate adenylyl transferase MET 3 Adenylyl sulfate kinase MET 14 3'-phosphoadenylylsulfate reductase MET 16 1. 8. 1. 2 Putative Sulfite reductase MET 5 sulfide Sulfite reductase (NADPH) MET 10 4. 2. 99. 10 O-acetylhomoserine (thiol)-lyase MET 17 Methionine synthase (vit B 12 -independent) MET 6 2. 7. 7. 4 Adenylyl sulfate (APS) ATP ADP 2. 7. 1. 25 3'-phosphoadenylylsulfate (PAPS) NADPH NADP+; AMP; 3'-phosphate (PAP); H+ 1. 8. 99. 4 sulfite 3 NADPH; 5 H+ 3 NADP+; 3 H 2 O O-acetyl-homoserine Homocysteine 5 -methyltetrahydropteroyltri-L-glutamate 5 -tetrahydropteroyltri-L-glutamate 2. 1. 1. 14 L-Methionine
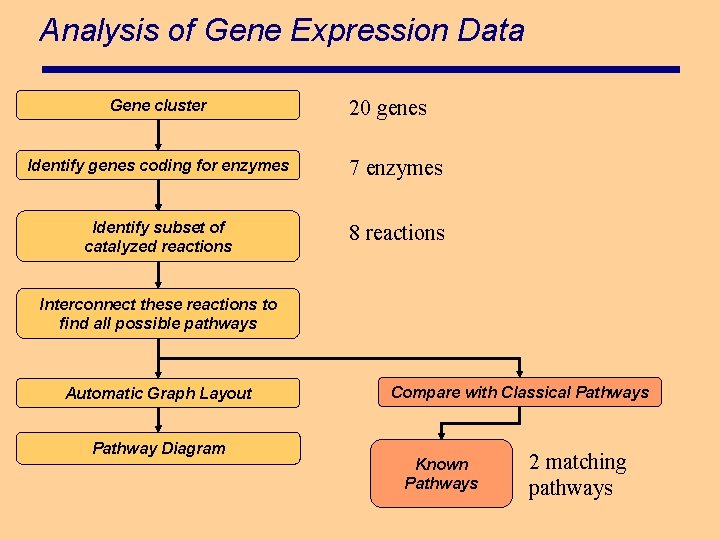
Analysis of Gene Expression Data Gene cluster 20 genes Identify genes coding for enzymes 7 enzymes Identify subset of catalyzed reactions 8 reactions Interconnect these reactions to find all possible pathways Automatic Graph Layout Pathway Diagram Compare with Classical Pathways Known Pathways 2 matching pathways
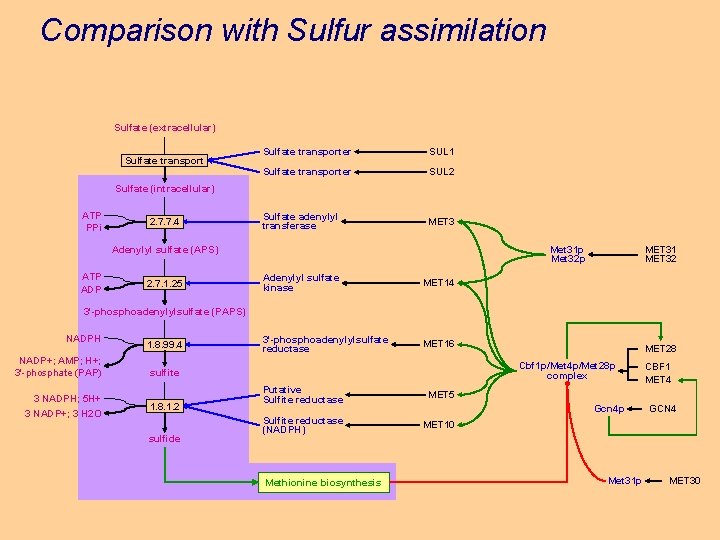
Comparison with Sulfur assimilation Sulfate (extracellular) Sulfate transporter SUL 1 Sulfate transporter SUL 2 Sulfate adenylyl transferase MET 3 Sulfate (intracellular) ATP PPi 2. 7. 7. 4 Met 31 p Met 32 p Adenylyl sulfate (APS) ATP ADP 2. 7. 1. 25 Adenylyl sulfate kinase MET 14 3'-phosphoadenylylsulfate reductase MET 16 MET 31 MET 32 3'-phosphoadenylylsulfate (PAPS) NADPH NADP+; AMP; H+; 3'-phosphate (PAP) 3 NADPH; 5 H+ 3 NADP+; 3 H 2 O 1. 8. 99. 4 Cbf 1 p/Met 4 p/Met 28 p complex sulfite 1. 8. 1. 2 sulfide MET 28 Putative Sulfite reductase MET 5 Sulfite reductase (NADPH) MET 10 Methionine biosynthesis Gcn 4 p Met 31 p CBF 1 MET 4 GCN 4 MET 30
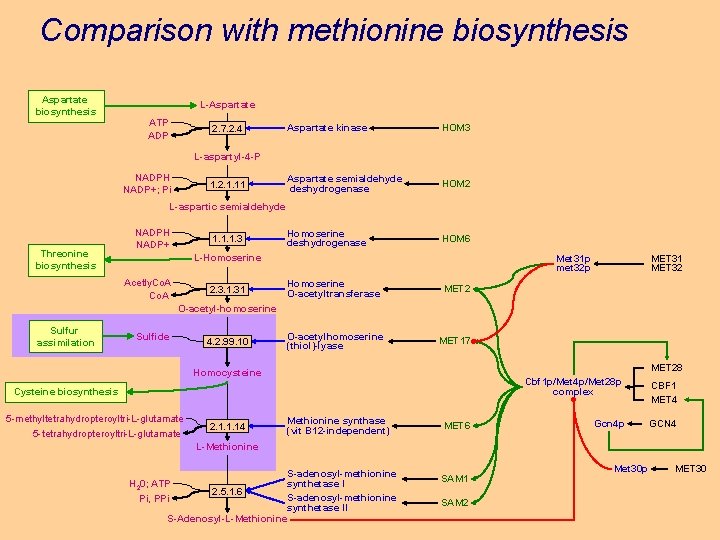
Comparison with methionine biosynthesis Aspartate biosynthesis L-Aspartate ATP ADP 2. 7. 2. 4 Aspartate kinase HOM 3 Aspartate semialdehyde deshydrogenase HOM 2 Homoserine deshydrogenase HOM 6 L-aspartyl-4 -P NADPH NADP+; Pi 1. 2. 1. 11 L-aspartic semialdehyde Threonine biosynthesis NADPH NADP+ 1. 1. 1. 3 L-Homoserine Acetly. Co. A 2. 3. 1. 31 Met 31 p met 32 p Homoserine O-acetyltransferase MET 2 O-acetylhomoserine (thiol)-lyase MET 17 MET 31 MET 32 O-acetyl-homoserine Sulfur assimilation Sulfide 4. 2. 99. 10 MET 28 Homocysteine Cysteine biosynthesis 5 -methyltetrahydropteroyltri-L-glutamate 5 -tetrahydropteroyltri-L-glutamate 2. 1. 1. 14 Methionine synthase (vit B 12 -independent) MET 6 Cbf 1 p/Met 4 p/Met 28 p complex CBF 1 MET 4 Gcn 4 p GCN 4 L-Methionine S-adenosyl-methionine synthetase I H 20; ATP 2. 5. 1. 6 S-adenosyl-methionine Pi, PPi synthetase II S-Adenosyl-L-Methionine SAM 1 SAM 2 Met 30 p MET 30

Summary n n n Starting from an unordered cluster of genes, one gets an ordered set of reactions, connected to form a pathway Should permit discovery of novel pathways, that are not stored in any pathway database yet Interpretation of intercalated reactions q q enzyme is not regulated DNA chip defect for that gene was not on the DNA chip enzyme remains to be identified in that organism

Analysis of data from Gasch et al. n n n Gasch et al (2000). Molecular Biology of the Cell, 11: 4241 -4257 6152 yeast genes Various conditions of stress (heat shock, osmotic shock, peroxide, amino acid starvation, nitrogen depletion Steady-state growth on alternative carbon sources Overexpression studies

Selected experiments MSN 2 overexpression MSN 4 overexpression YAP 1 overexpression ethanol galactose glucose mannose raffinose sucrose ethanol vs reference fructose vs reference galactose vs reference glucose vs reference mannose vs reference raffinose vs reference sucrose vs reference
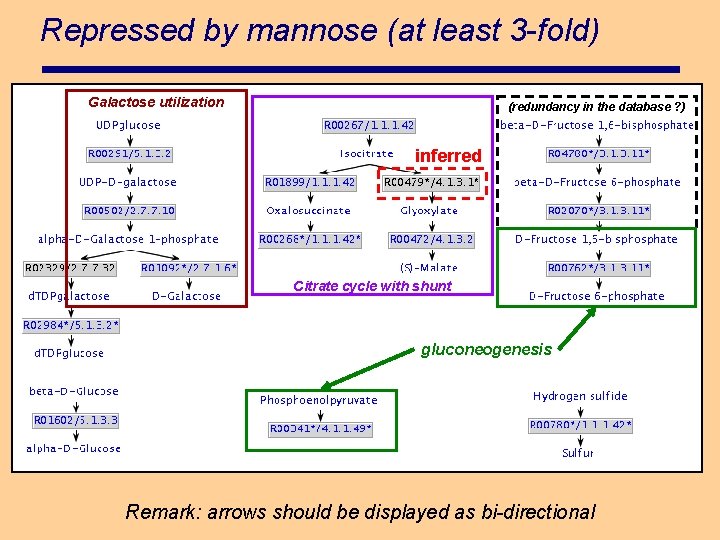
Repressed by mannose (at least 3 -fold) Galactose utilization (redundancy in the database ? ) inferred Citrate cycle with shunt gluconeogenesis Remark: arrows should be displayed as bi-directional
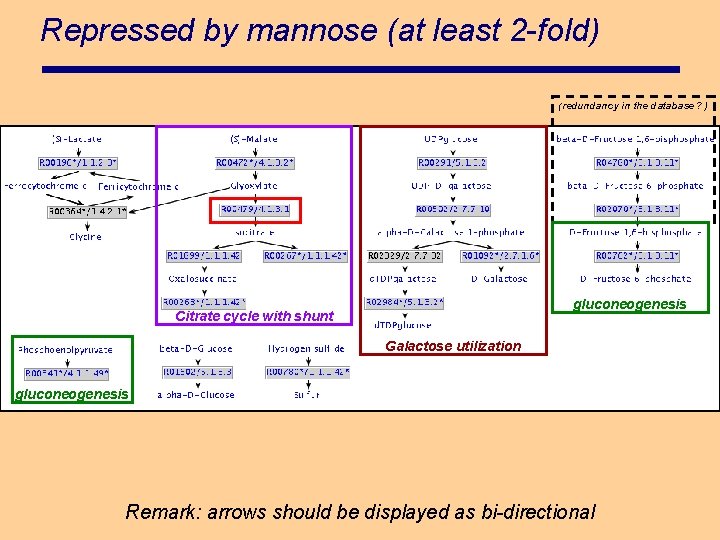
Repressed by mannose (at least 2 -fold) (redundancy in the database ? ) gluconeogenesis Citrate cycle with shunt Galactose utilization gluconeogenesis Remark: arrows should be displayed as bi-directional
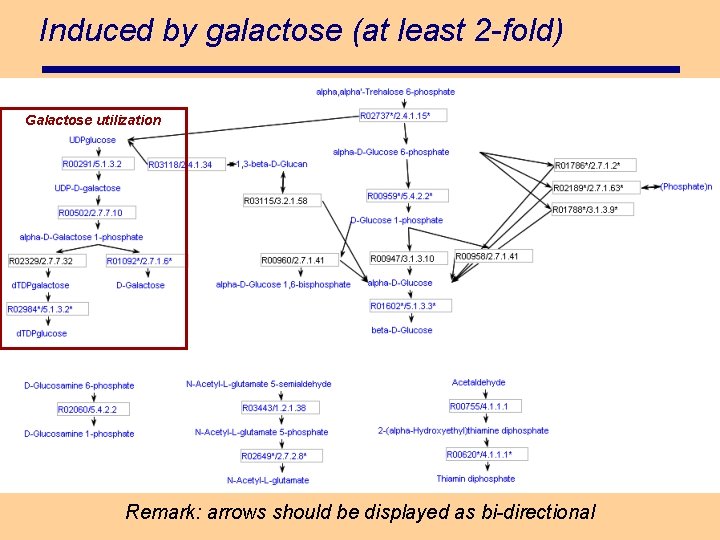
Induced by galactose (at least 2 -fold) Galactose utilization Remark: arrows should be displayed as bi-directional
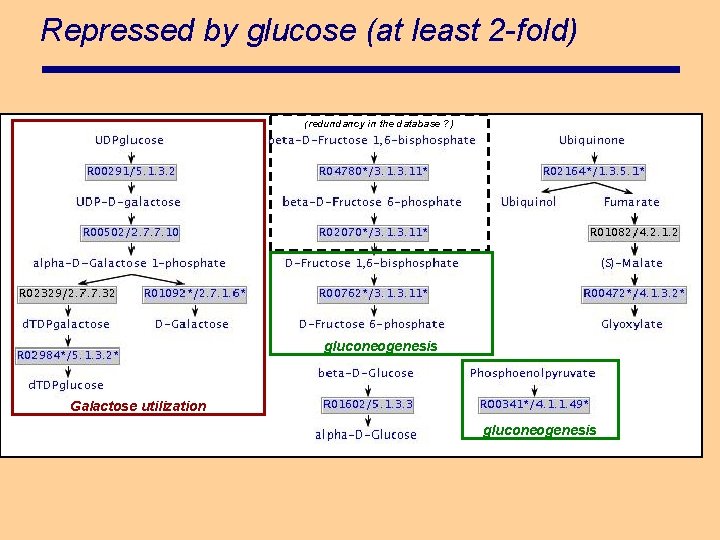
Repressed by glucose (at least 2 -fold) (redundancy in the database ? ) gluconeogenesis Galactose utilization gluconeogenesis
- Slides: 48