Bioinformatics for Targeted Metabolomics Met and Unmet Needs

Bioinformatics for Targeted Metabolomics: Met and Unmet Needs Klaus M. Weinberger Biocrates Life Sciences AG, Innsbruck, Austria 3 rd Annual Forum for SMEs Information Workshop on European Bioinformatics Resources Vienna, September 3 – 4, 2009

Agenda • Why (targeted) metabolomics? BIOCRATES • Proof-of-concept in routine clinical diagnostics • Technology platform • Workflow integration & data analysis • Issues • Acknowledgements Socrates 470 -399 BC Hippocrates 460 -377 BC Intelligence Wisdom Medicine Health “Creating Knowledge for Health”

Metabolomics is. . . the systematic identification and quantitation of all/ biologically relevant small molecules* in a given compartment, cell, tissue or body fluid. It represents the functional end-point of physiological and pathophysiological processes depicting both genetic predisposition and environmental influences like nutrition, exercise or medication. * no biopolymers (nucleic acids, polypeptides)

Why (targeted) metabolomics?
Six systems biologists examining an elephant
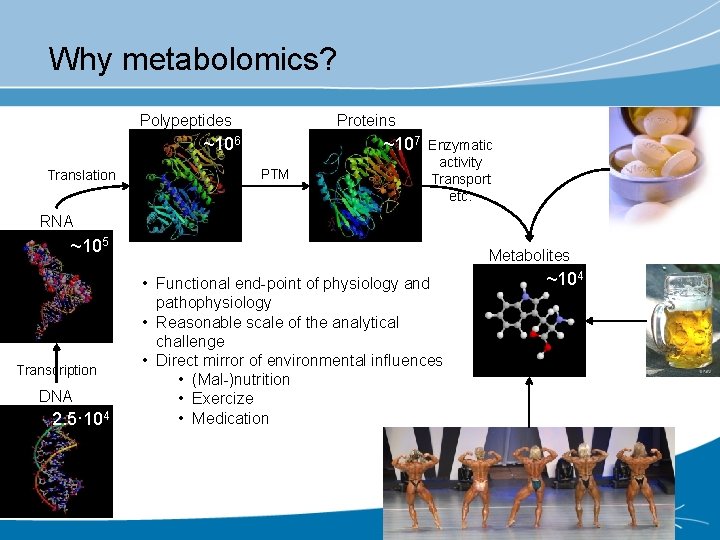
Why metabolomics? Polypeptides Proteins ~106 Translation ~107 PTM Enzymatic activity Transport etc. RNA ~105 Transcription DNA 2. 5· 104 Metabolites • Functional end-point of physiology and pathophysiology • Reasonable scale of the analytical challenge • Direct mirror of environmental influences • (Mal-)nutrition • Exercize • Medication ~104
Metabolomics approaches Sample cohorts Metabolic profiling (e. g. full scan LC-MS) Differential pattern information
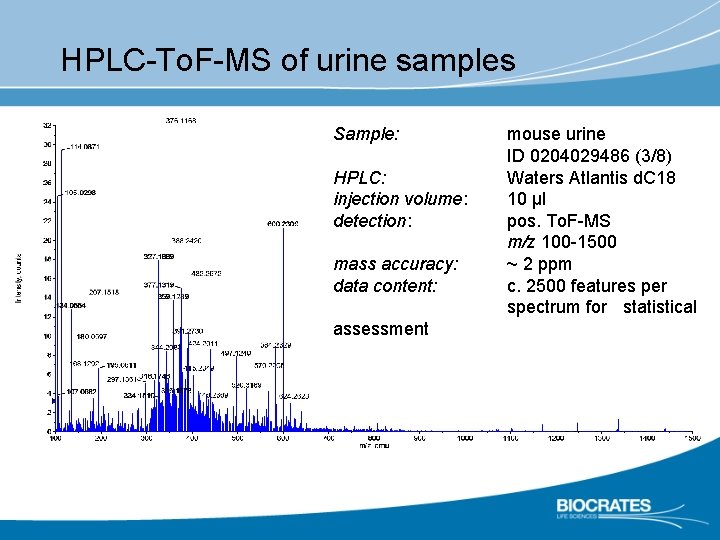
HPLC-To. F-MS of urine samples Sample: HPLC: injection volume: detection: mass accuracy: data content: assessment mouse urine ID 0204029486 (3/8) Waters Atlantis d. C 18 10 µl pos. To. F-MS m/z 100 -1500 ~ 2 ppm c. 2500 features per spectrum for statistical
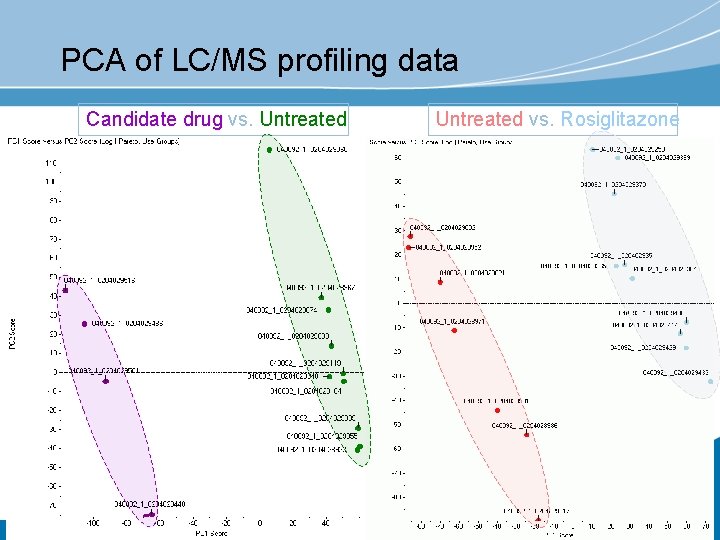
PCA of LC/MS profiling data Candidate drug vs. Untreated vs. Rosiglitazone
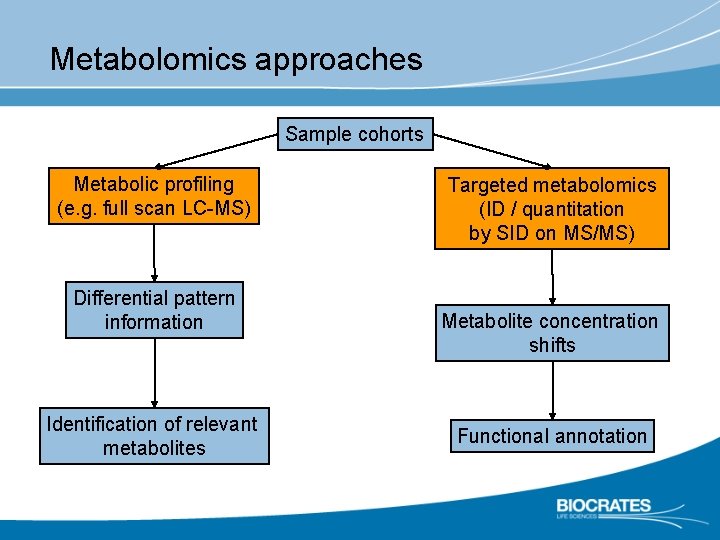
Metabolomics approaches Sample cohorts Metabolic profiling (e. g. full scan LC-MS) Differential pattern information Identification of relevant metabolites Targeted metabolomics (ID / quantitation by SID on MS/MS) Metabolite concentration shifts Functional annotation
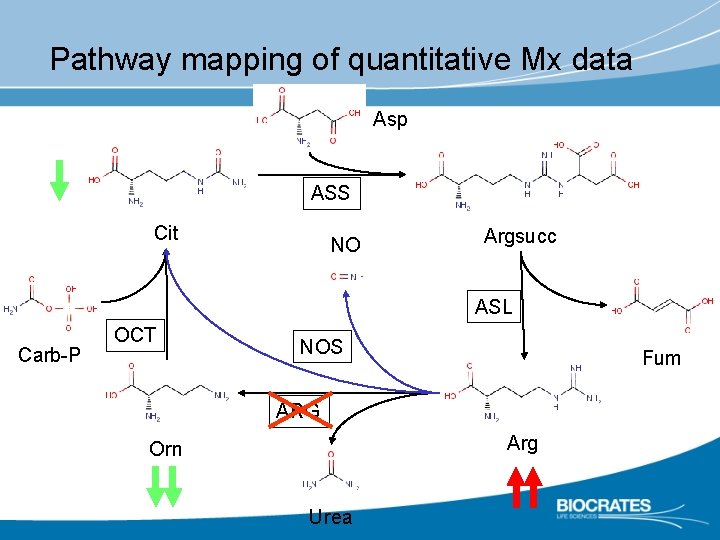
Pathway mapping of quantitative Mx data Asp ASS Cit NO Argsucc ASL Carb-P OCT NOS Fum ARG Arg Orn Urea

Areas of application Ø Basic research - Functional genomics in biochemistry, physiology, cell biology, microbiology, ecology, … Ø Agricultural & nutrition industry - Plant intermediary metabolism - Health effects of functional food products Ø Biotechnology - Optimization and monitoring of fermentation processes Ø Pharmaceutical R&D - Pathobiochemistry / characterization of disease models - Safety / toxicology - Efficacy / pharmacodynamics and mode-of-action Ø Clinical diagnostics & theranostics - Early diagnosis and accurate staging - Specific monitoring of therapeutic effects

History and proof-of-concept in clinical diagnostics

Sir Archibald Edward Garrod • • • 1857, London – 1936, Cambridge Educated in Marlborough, Oxford, and London Postgraduate studies at the AKH in Vienna in 1884/85 Publications on chemical pathology (e. g. of alkaptonuria, cystinuria, pentosuria) One gene – one enzyme hypothesis Concept of inborn errors of metabolism (Croonian lectures to the Royal College of Physicians, 1908)

Proof-of-concept in neonatology • Newborn screening for inborn metabolic disorders • • replaced expensive monoparametric assays simultaneous detection of 40 - 60 metabolites (amino acids, acylcarnitines) simultaneous diagnosis of 20 - 30 monogenic diseases (AA metabolism, FATMO) with immediate treatment options total incidence > 1: 2000 unprecedented sensitivity, specificity, ppv co-pioneered in the mid-90 s by BIOCRATES founder Bert Roscher > 1, 300, 000 newborns screened in Munich similar labs worldwide

Lessons from newborn screening 1) Quantitative tandem mass spectrometry (stable isotope dilution) is able to meet the most stringent quality criteria (precision, accuracy) for routine diagnostics 2) The concept of multiparametric biomarkers improving assay sensitivity and, particularly, specificity is valid for many monogenic (and multifactorial) diseases 3) MS-based diagnostics can save costs despite a wider analytical panel and improved diagnostic quality Also true for therapeutic drug monitoring of immunosuppressants, antidepressants, antiretrovirals. . .
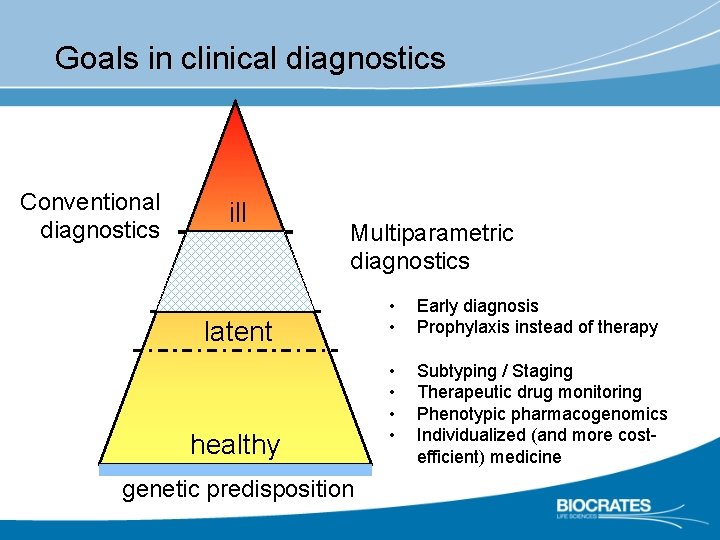
Goals in clinical diagnostics Conventional diagnostics ill Multiparametric diagnostics latent healthy genetic predisposition • • Early diagnosis Prophylaxis instead of therapy • • Subtyping / Staging Therapeutic drug monitoring Phenotypic pharmacogenomics Individualized (and more costefficient) medicine

Technology, workflow integration & data analysis
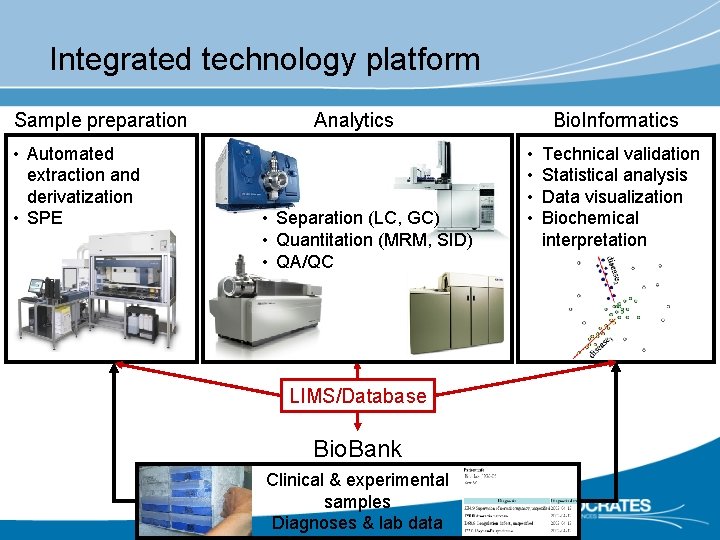
Integrated technology platform Sample preparation • Automated extraction and derivatization • SPE Analytics • Separation (LC, GC) • Quantitation (MRM, SID) • QA/QC LIMS/Database Bio. Bank Clinical & experimental samples Diagnoses & lab data Bio. Informatics • • Technical validation Statistical analysis Data visualization Biochemical interpretation
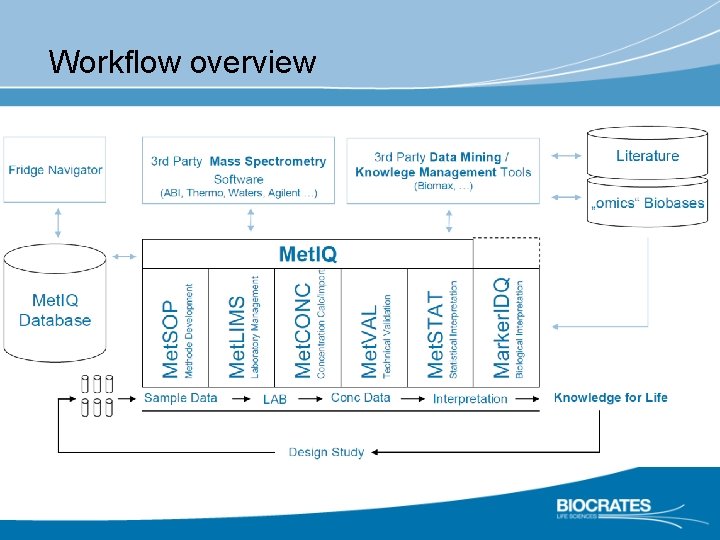
Workflow overview
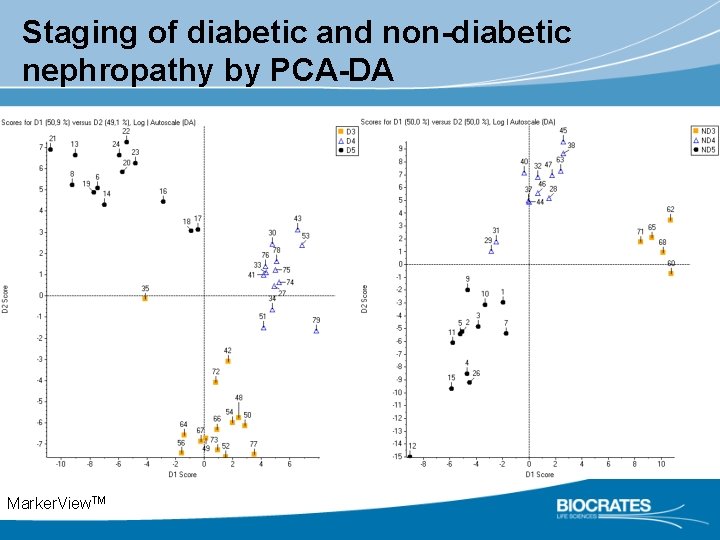
Staging of diabetic and non-diabetic nephropathy by PCA-DA Marker. View. TM
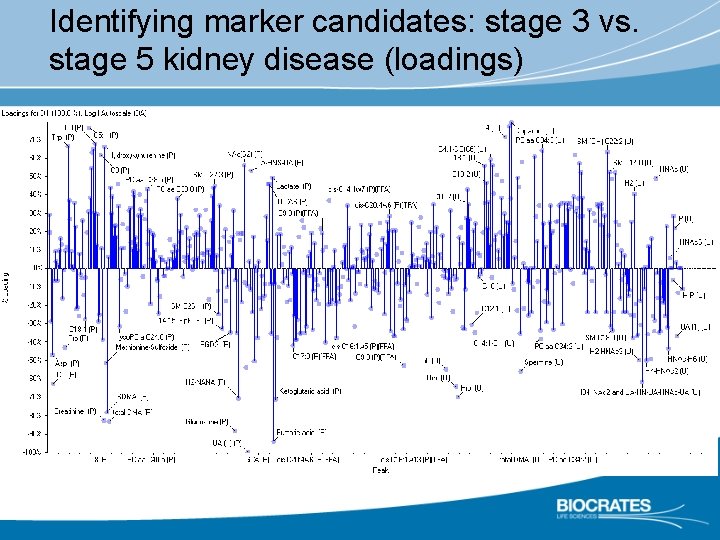
Identifying marker candidates: stage 3 vs. stage 5 kidney disease (loadings)
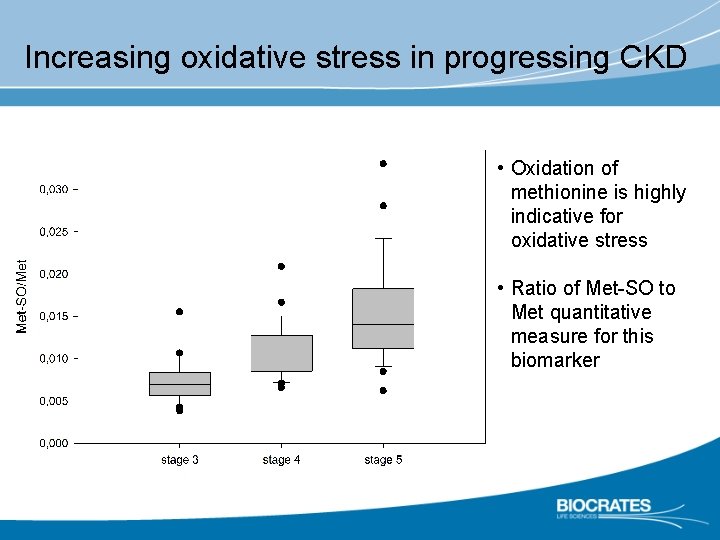
Increasing oxidative stress in progressing CKD • Oxidation of methionine is highly indicative for oxidative stress • Ratio of Met-SO to Met quantitative measure for this biomarker
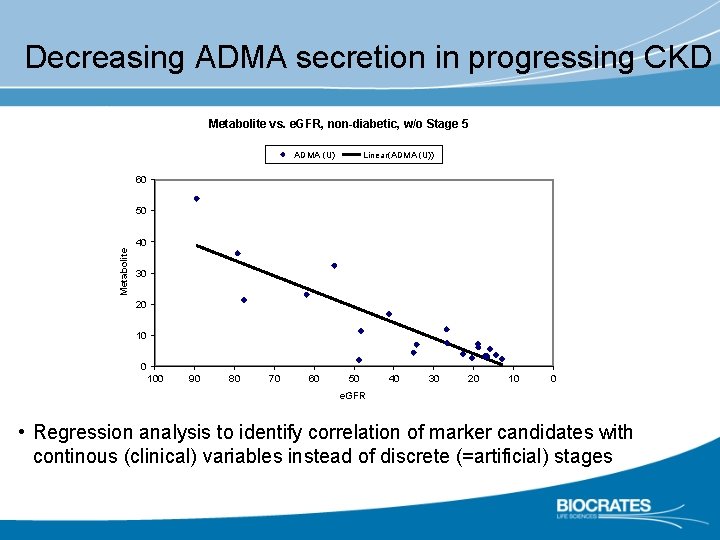
Decreasing ADMA secretion in progressing CKD Metabolite vs. e. GFR, non-diabetic, w/o Stage 5 ADMA (U) Linear(ADMA (U)) 60 Metabolite 50 40 30 20 100 90 80 70 60 50 40 30 20 10 0 e. GFR • Regression analysis to identify correlation of marker candidates with continous (clinical) variables instead of discrete (=artificial) stages R = 0. 7523 2
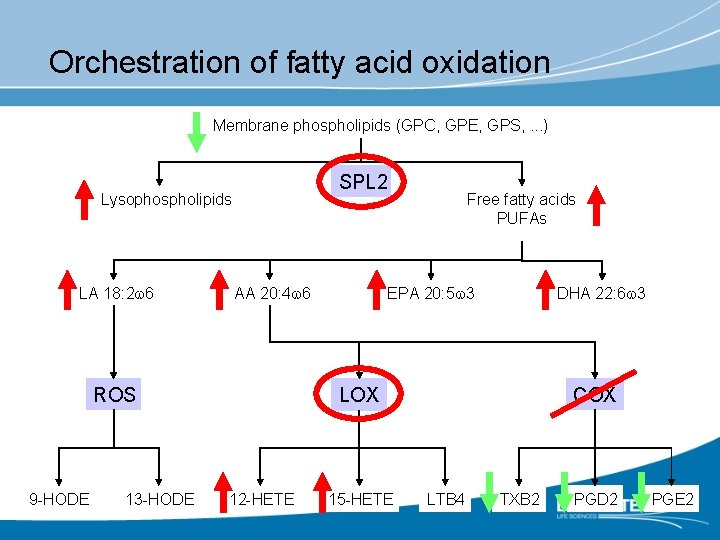
Orchestration of fatty acid oxidation Membrane phospholipids (GPC, GPE, GPS, . . . ) SPL 2 Lysophospholipids LA 18: 2 w 6 AA 20: 4 w 6 13 -HODE EPA 20: 5 w 3 DHA 22: 6 w 3 LOX ROS 9 -HODE Free fatty acids PUFAs 12 -HETE 15 -HETE COX LTB 4 TXB 2 PGD 2 PGE 2
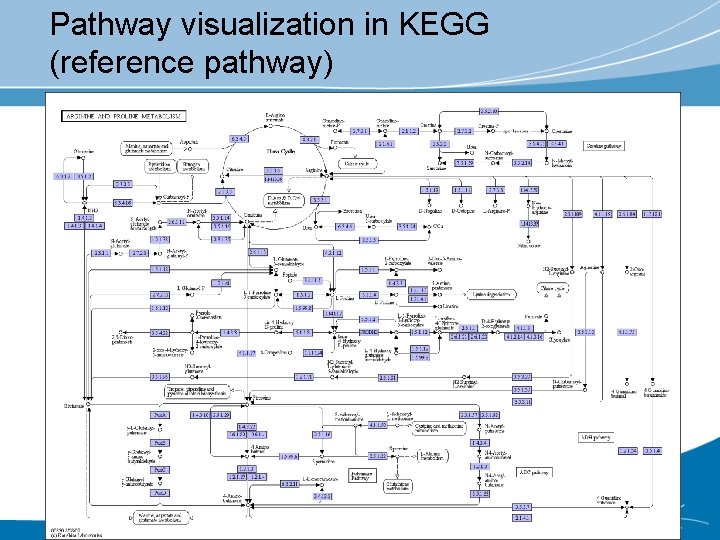
Pathway visualization in KEGG (reference pathway)
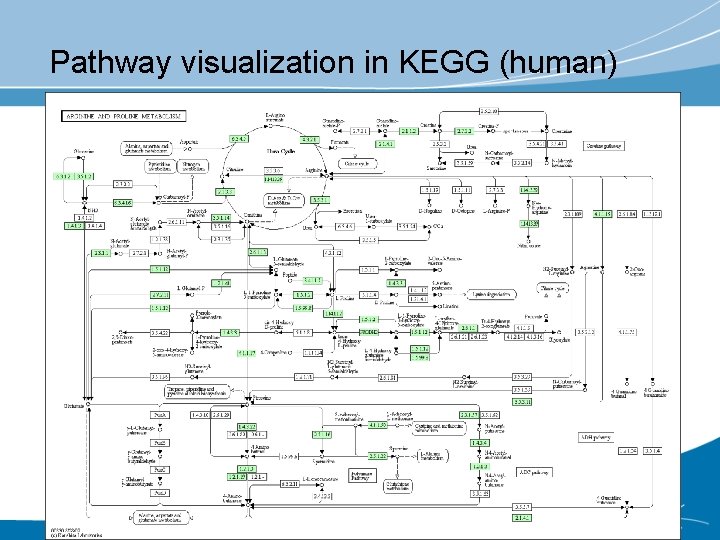
Pathway visualization in KEGG (human)
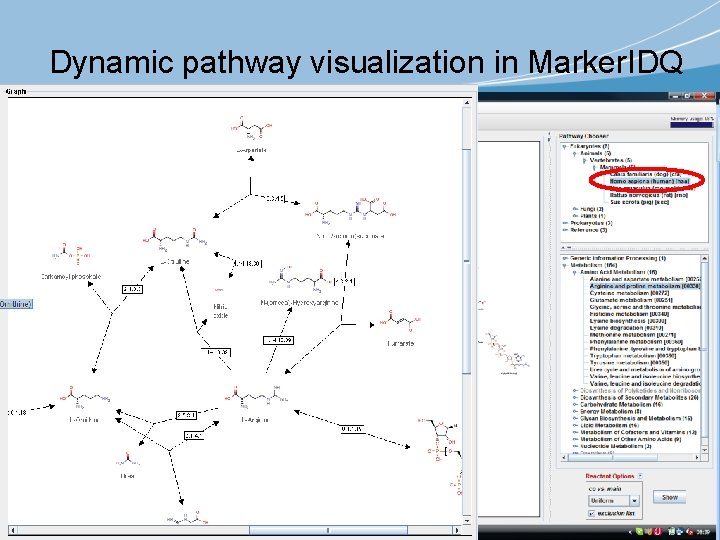
Dynamic pathway visualization in Marker. IDQ
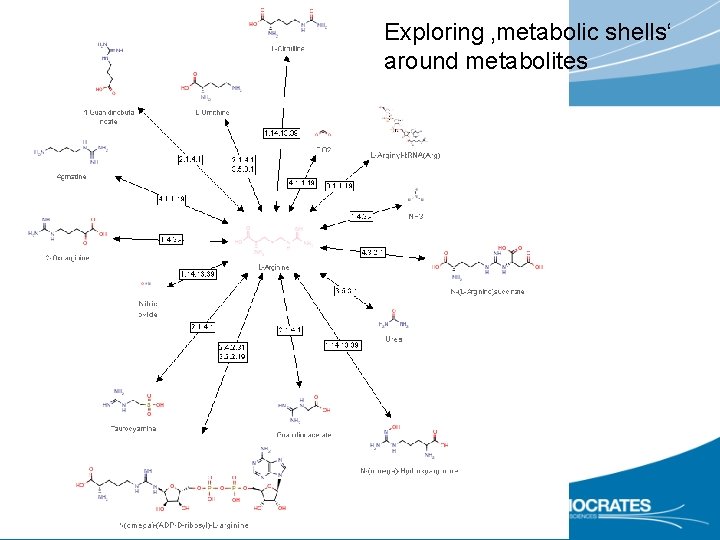
Exploring ‚metabolic shells‘ around metabolites
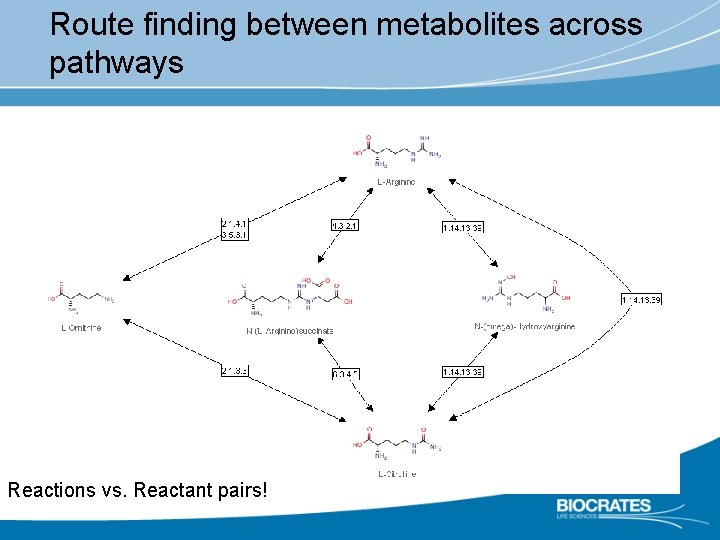
Route finding between metabolites across pathways Reactions vs. Reactant pairs!

Issues I: Databases Ø Parallel / competing initiatives with incompatible / proprietary data formats Ø KEGG Ø Meta. Cyc, Human. Cyc, etc. Ø Reactome Ø HMDB Ø OMIM Ø Lipidomics consortia Ø. . . Ø Compartmentalization not well depicted Ø Incompleteness / generic entries (phospholipids, acylcarnitines, etc. ) Ø Lack of curation Ø Lack of publication

Issues II: Standardization and normalization Ø Standardization Ø Instrument vendors oppose common data formats Ø What meta-data to record? Ø No valid guidelines for quantitation of endogenous metabolites (FDA guidance was developed for xenobiotics) Ø Nomenclature vs. analytical reality (sum signals, isomers, etc. ) Ø Normalization Ø Absolute quantitation overcomes the need for analytical normalization Ø Role of sample types (plasma, CSF, urine, tissue homogenates, cell extracts, . . . ) Ø How can biological normalization work? Are there ‚housekeeping metabolites‘?

Issues III: Biostatistics Ø Overfitting & correction Ø Suitable clustering algorithms for multivariate data sets? Ø Metabolites are no equivalent independent variables Ø Analytical validity/variability are usually not considered Ø Often, groups of metabolites are synthesized or degraded by the same enzyme(s) Ø Consecutive reactions within a pathway/network depend on each other (flux analysis!) Ø How to incorporate this in biostatistics? Weighting? Derived parameters, ratios, etc. ? Ø How to exploit this in (automated) plausibility checks?

Summary I • Metabolomics depicts the functional end-point of genetics and environment • Targeted metabolomics data are analytically reproducible and allow immediate biochemical interpretation • Proof-of-concept has been achieved in routine diagnostics of inborn errors of metabolism • Many metabolic biomarkers are valid across species and enable translational research • Comprehensive targeted metabolomics bridges the gap to open profiling approaches
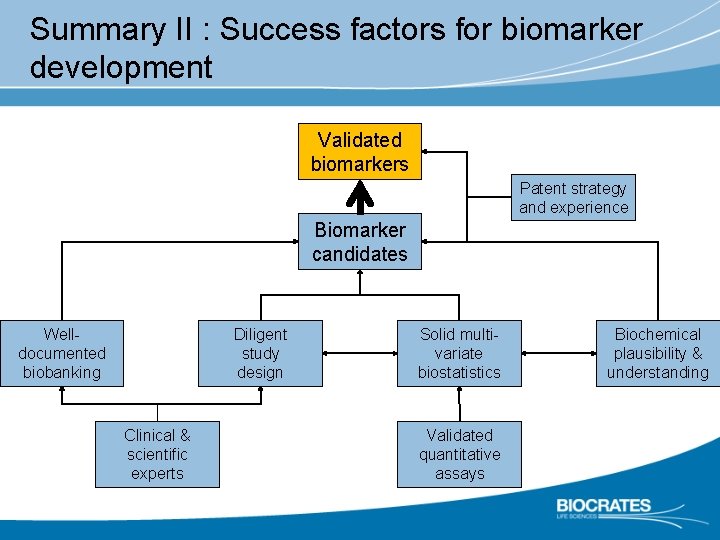
Summary II : Success factors for biomarker development Validated biomarkers Patent strategy and experience Biomarker candidates Welldocumented biobanking Diligent study design Clinical & scientific experts Solid multivariate biostatistics Validated quantitative assays Biochemical plausibility & understanding

Selected partners

Analytics Stefanie Gstrein Sascha Dammeier Hai Pham Tuan Cornelia Röhring Therese Koal Ali Alchalabi Verena Forcher Ines Unterwurzacher Stefan Urban Doreen Kirchberg Ralf Bogumil Patrizia Hofer Lisa Körner Peter Enoh Acknowledgements Brad Morie Doris Gigele Elgar Schnegg Admin, IT & Biz. Dev Anton Grones Ingrid Sandner Georg Debus Wolfgang Samsinger Patricia Aschacher Bioinformatics Daniel Andres Olivier Lefèvre Paolo Zaccaria Florian Bichteler Marc Breit Manuel Gogl Bernd Haas Mattias Bair Robert Eller Hamza Ovacin Gerd Lorünser Yi Zao Statistics & Biochemistry Ingrid Osprian Marion Beier Vera Neubauer Oliver Lutz Matthias Keller Denise Sonntag Hans-Peter Deigner Ulrika Lundin
- Slides: 38