Bioinformatics Biological Databases Revised 011006 Introduction Repository databases

Bioinformatics Biological Databases Revised 01/10/06

Introduction • Repository databases – Redundant – Cutting edge information • Curated databases – Manual & automatic curation – Organization of information important – An attempt to be nonredundant – Comprehensive in some cases
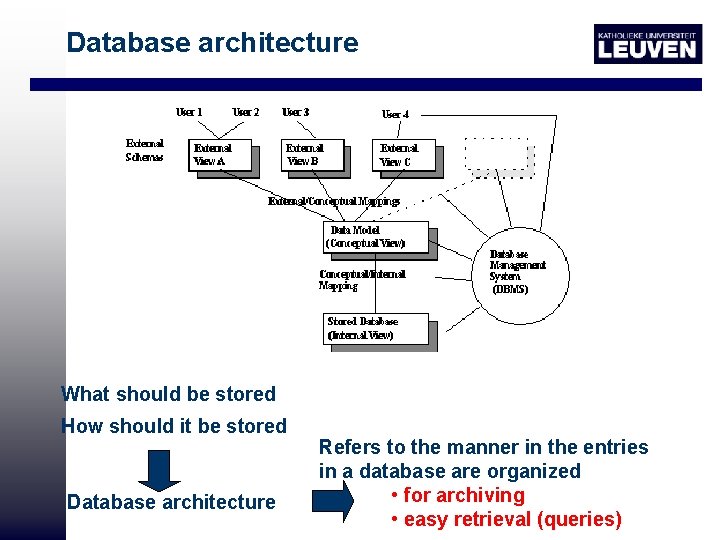
Database architecture What should be stored How should it be stored Database architecture Refers to the manner in the entries in a database are organized • for archiving • easy retrieval (queries)

Relational database Data are stores in tables Relationships between records can be many to one or many to many. In the latter case an index is required. All records in a table have identical features A record is identified by its table and record identifier Navarro et al. , 2003
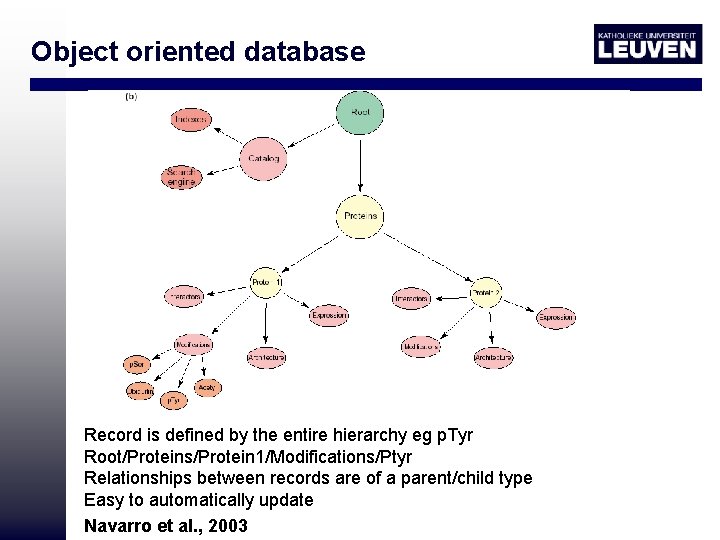
Object oriented database Record is defined by the entire hierarchy eg p. Tyr Root/Proteins/Protein 1/Modifications/Ptyr Relationships between records are of a parent/child type Easy to automatically update Navarro et al. , 2003
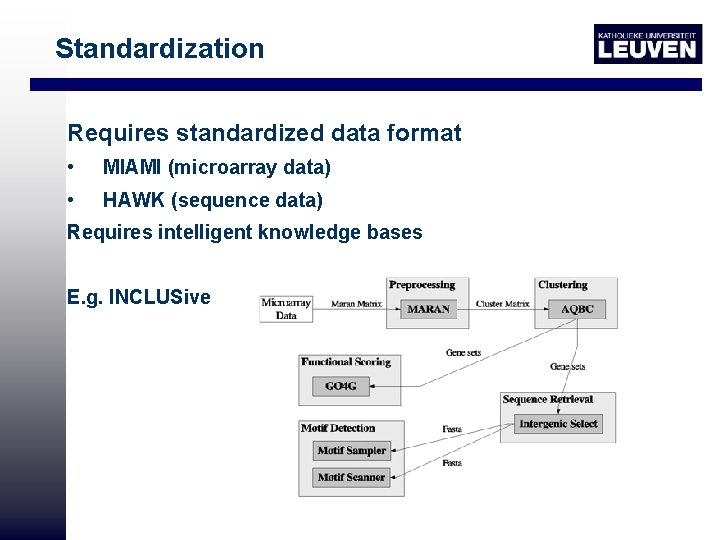
Standardization Requires standardized data format • MIAMI (microarray data) • HAWK (sequence data) Requires intelligent knowledge bases E. g. INCLUSive

Sequence databases

Sequence Formats • A sequence file needs to be recognized by a computer program, • special formats have been invented – Fast. A – Gen. Bank

Sequence formats Gen. Bank

Sequence Repositories Sequence repositories: redundant databases National Center for Biotechnology Information (NCBI) – Gen. Bank • For each sequence multiple entries in Gen. Bank • EST, c. DNA, genomic sequence – EST database (Ncbi) – Swiss. Prot (Curated)

Sequence Repositories at Ncbi • http: //www. ncbi. nih. gov/Database/index. html • Gen. Bank uses a relational model • New sequences can be submitted by a submission page. • Gen. Bank also accepts submission of sequences with a high error rate and provides curated databases (99% accuracy) • 200000 users a day, 4 million queries a day

Sequence Repositories at Ncbi ENTREZ, a resource prepared by NCBI is used to retrieve a DNA or protein sequence or Medline from the databases at NCBI.

Sequence Repositories at Ncbi: Gen. Bank Redundant number of entries => need for a comprehensive database
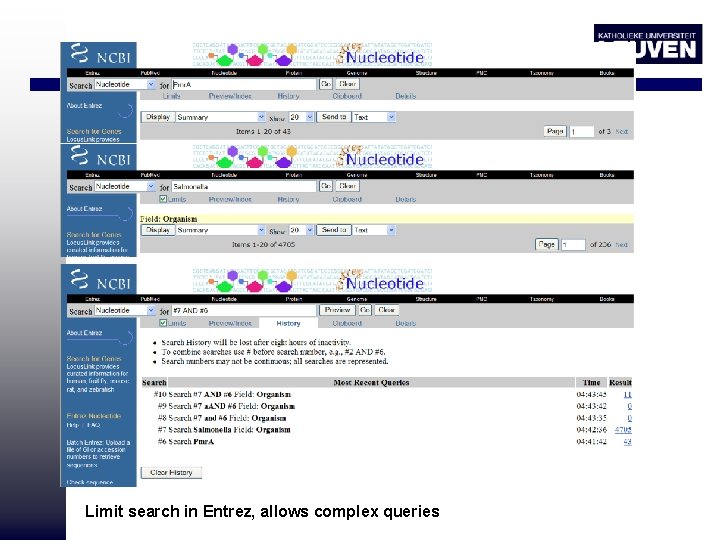
Limit search in Entrez, allows complex queries

Gen. Bank format: DNA sequence
Gen. Bank format: protein sequence
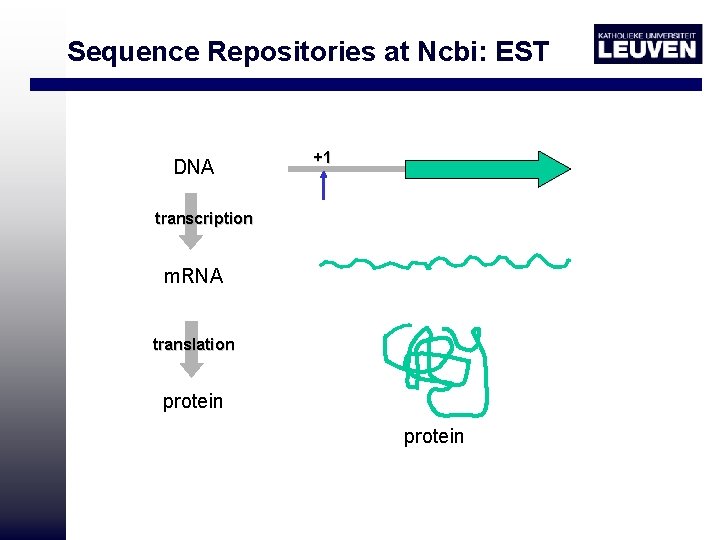
Sequence Repositories at Ncbi: EST DNA +1 transcription m. RNA translation protein

EST represent first pass sequences with an error rate as high as 1 in 100, including incorrectly identified bases and insertions http: //www. ncbi. nlm. nih. gov/db. EST/

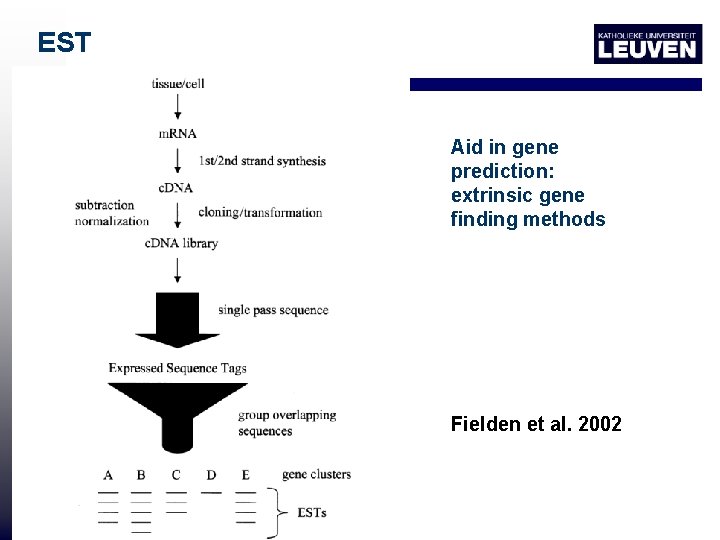
EST Aid in gene prediction: extrinsic gene finding methods Fielden et al. 2002

Comprehensive databases Curated databases • Unigene (Ncbi): automatic partitioning of Gen. Bank into a nonredundant set of gene-oriented clusters • Ref. Seq (Ncbi): • ENSEMBL/VEGA (Ebi): Integrate the information as such that for a locus in the genome a complete description is given that is no longer redundant Provide a comprehensive non redundant set of sequences including genomic DNA, transcript and protein products for major research organisms
Comprehensive DB: Uni. Gene
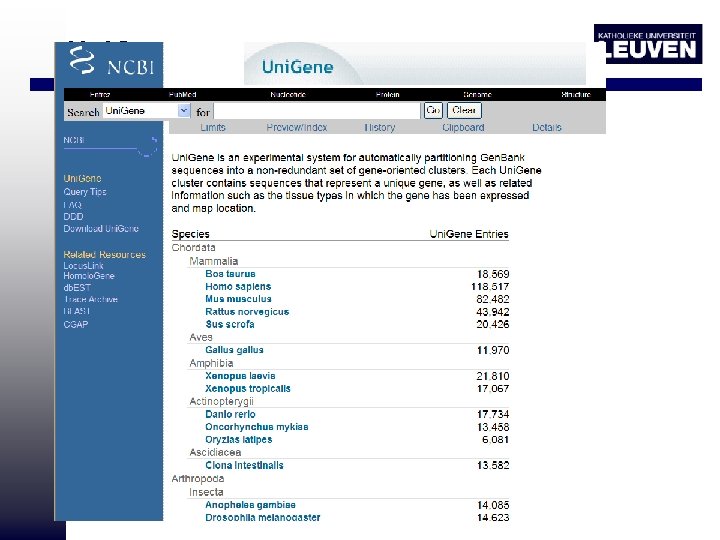
Uni. Gene

Comprehensive DB: Uni. Gene • Uni. Gene is an experimental system for automatically partitioning Gen. Bank sequences into a non-redundant set of gene-oriented clusters • Each Uni. Gene cluster contains sequences that represent a unique gene as well as related information such as the tissue types in which the gene has been expressed and map location. • These clusters represent the same gene based on the alignment of EST sequences with each other and with the genome sequences of the organism. • no attempt has been made to produce contigs – splicing variants for a gene are put into the same set. – Moreover, EST-containing sets often contain 5' and 3' reads from the same c. DNA clone, but these sequences do not always overlap.
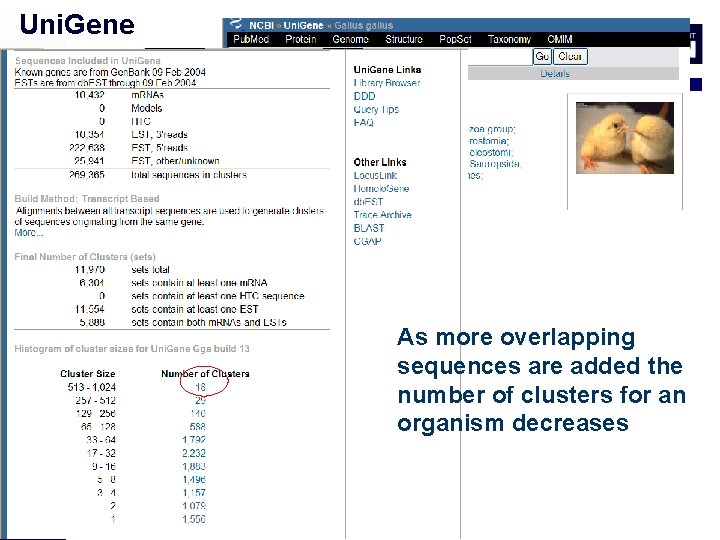
Uni. Gene As more overlapping sequences are added the number of clusters for an organism decreases

Comprehensive DB: Uni. Gene

Comprehensive DB: REf. Seq • For a particular gene many independent redundant records might exist in Gen. Bank • All this information is integrated as such that for a particular locus in the genome a complete description is given that is no longer redundant: the locuslink • Redundant Gen. Bank entries e. g. representing distinct indications on the transcript of a gene (incomplete c. DNA sequences, ESTs) are unified to a single refseq that represents the complete transcript • A Refseq sequence – protein (starting with NP_) – a genomic sequence (starting with NG_) – All Ref. Seq sequences that belong to the same locus on the genome receive the same locus link – Additional links to other interesting databases containing additional functional annotation or information are made (e. g to Gene Ontology,

Ref. Seq
Gene: Ref. Seq

Comprehensive DB: Ensembl
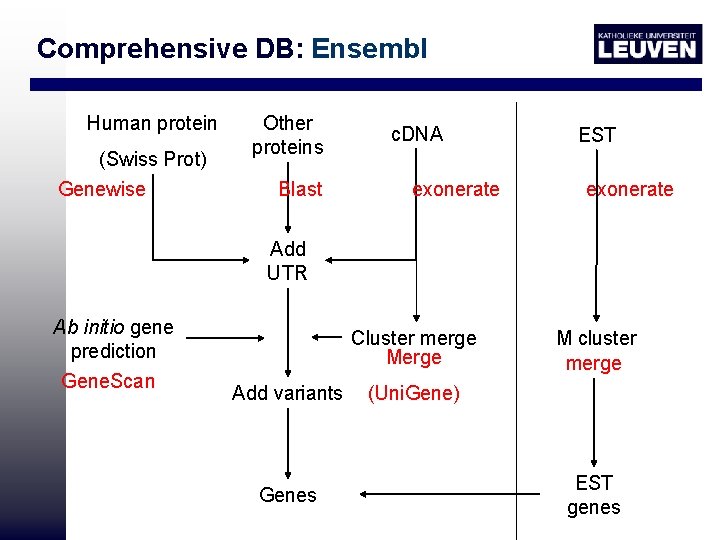
Comprehensive DB: Ensembl Human protein (Swiss Prot) Genewise Other proteins Blast c. DNA exonerate EST exonerate Add UTR Ab initio gene prediction Gene. Scan Cluster merge Merge Add variants Genes M cluster merge (Uni. Gene) EST genes
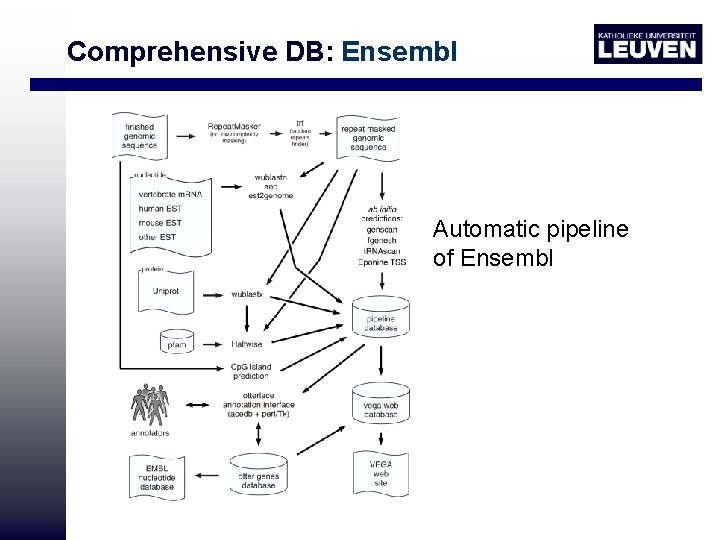
Comprehensive DB: Ensembl Automatic pipeline of Ensembl

Ensembl • Ab initio gene scan: doesn’t use protein/c. DNA/EST evidence • More genomes available: gene predictions will improve • ENSEMBL: 70 -75% genes annotated • EST genes used to help predicting UTR and splice variants • Problem automatic annotation: pseudogenes Processed (with poly A tail) pseudogene Unprocessed (rearrangement, duplication)

Ensembl • ENSEMBL: automatic analysis flow • VEGA (vertebrate genome annotation database) database: manual curation • ref. Seq: best curated database for c. DNAs (no integration with ESTs (<-> VEGA) AUTOMATIC • Weeks • Use draft sequence • No pseudogenes MANUAL • Months • Need finished sequence • Pseudogenes • Consult public databases/ literature
Vega

Other databases

Expression Databases • Microarray database: – SMD (Stanford) – Miami express (Ebi) – GEO (Ncbi) • SAGE data base • EST based expression database • Proteome database
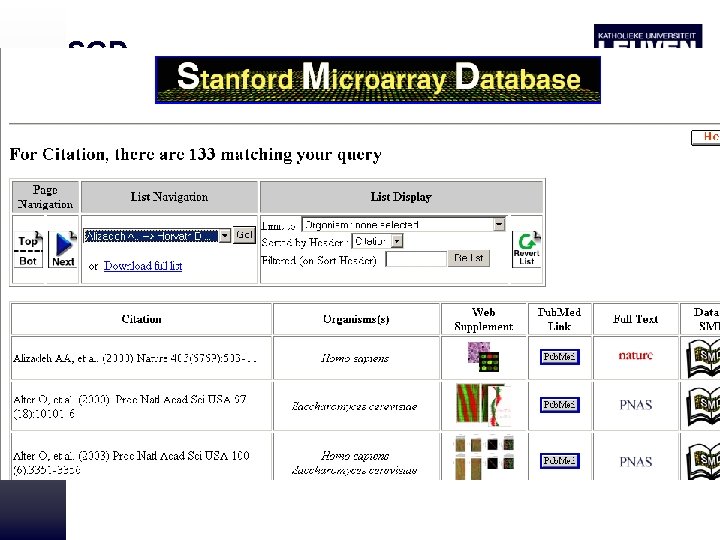
SGD
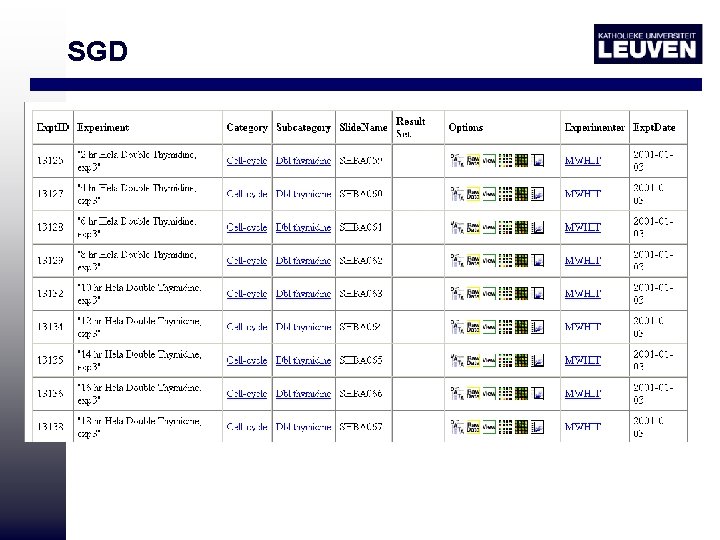
SGD

SGD
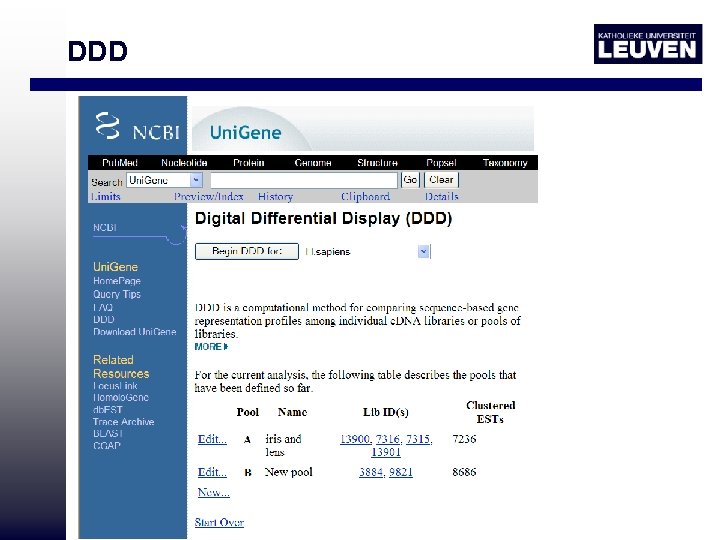
DDD • http: //www. ncbi. nlm. nih. gov/Uni. Gene/ddd. cgi? ORG=Hs
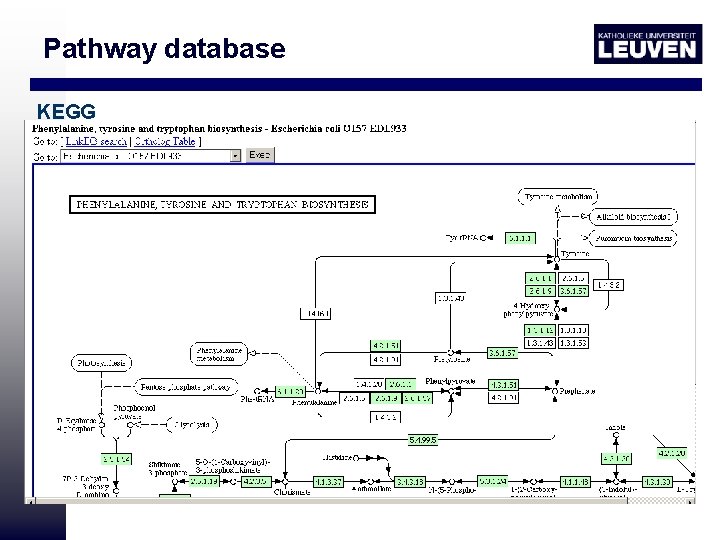
Pathway database KEGG

Ontologies Controlled vocabularies Tree structured Describe gene products and associated processes Species independent • Gene Ontology • Ecocyc

Ontologies GO: gene ontology • Organize biological information about proteins classes and functions into a hierarchical classification using controlled vocabulary http: //www. ensembl. org/Homo_sapiens/goview? query=GO%3 A 0003700
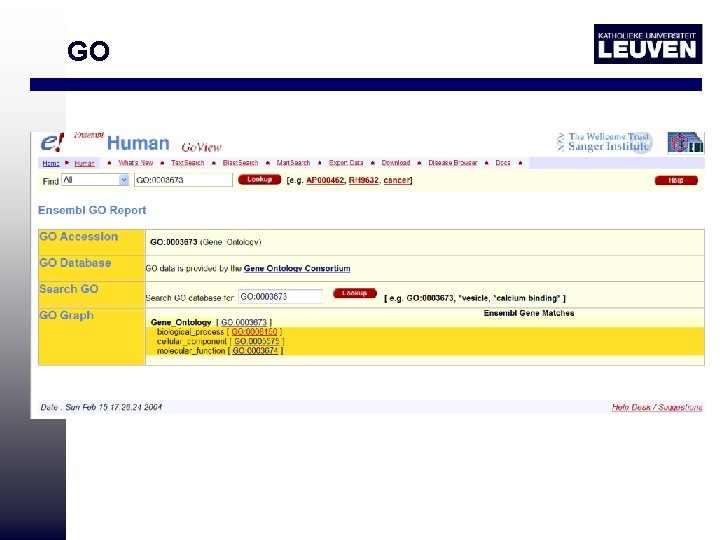
GO
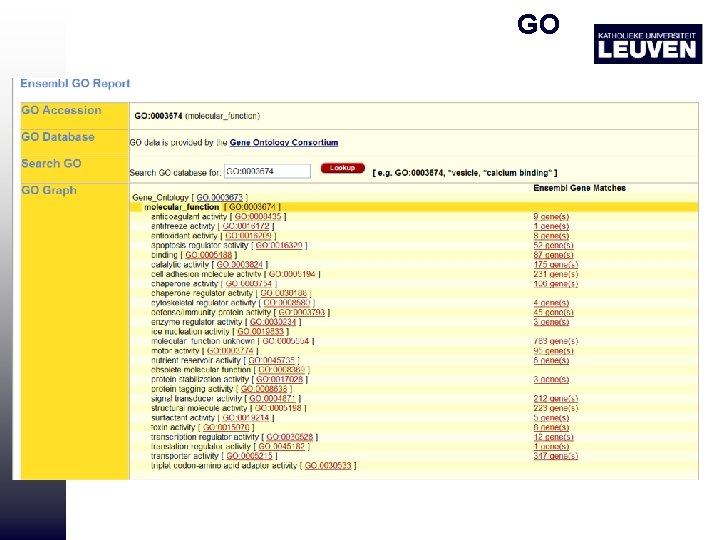
GO
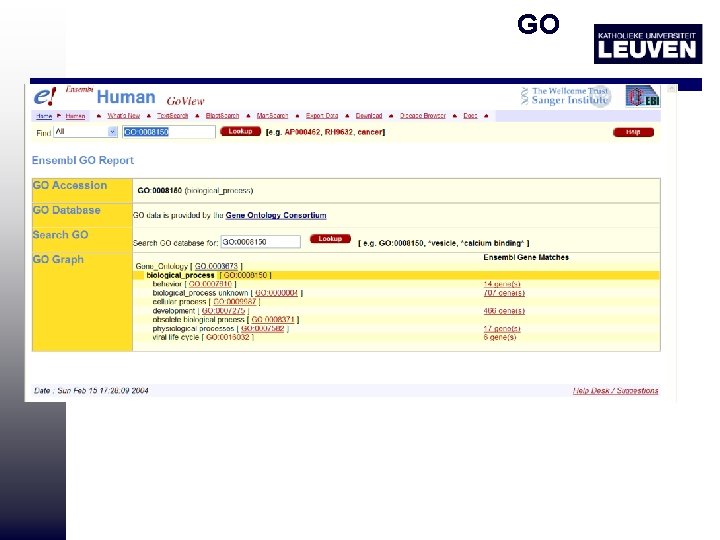
GO
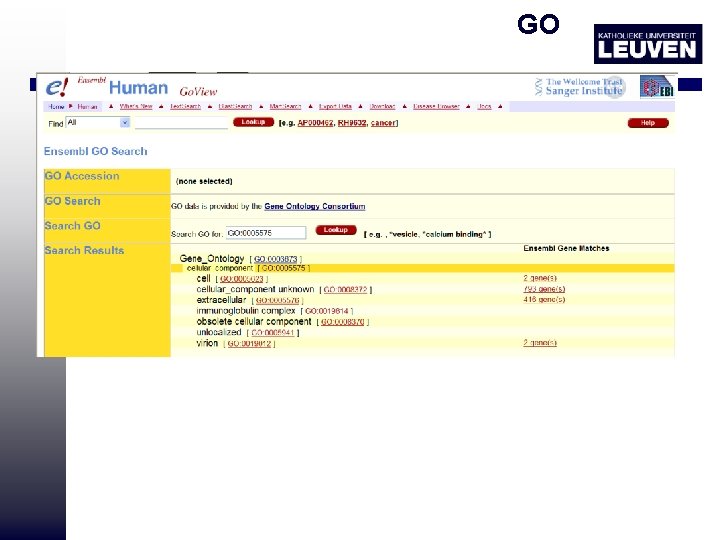
GO

GO

Eco. CYC

Databases with regulatory motifs DNA motifs • Transfac • Regulon. DB Protein Motifs • PFAM • Prosite


• http: //www. ncbi. nlm. nih. gov/Tools/



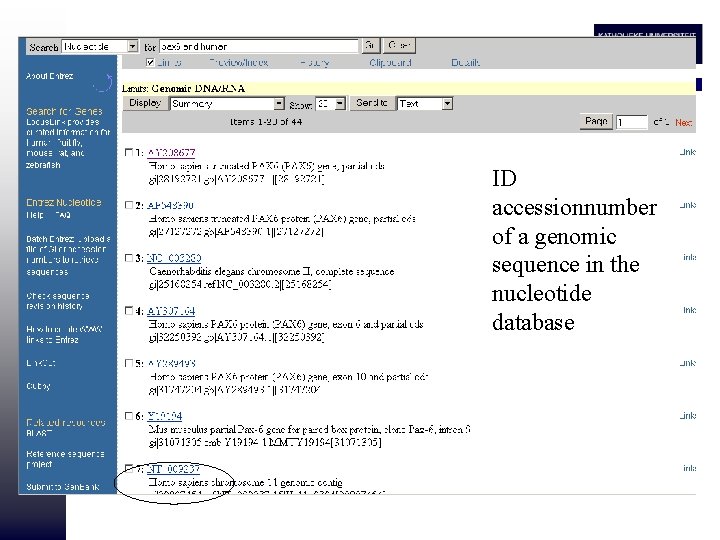
ID accessionnumber of a genomic sequence in the nucleotide database


• Many databases with sequences that give information on the same locus • Need for comprehensive databases • ENSEMBL • Locus. Link/Ref. Seq (ncbi)
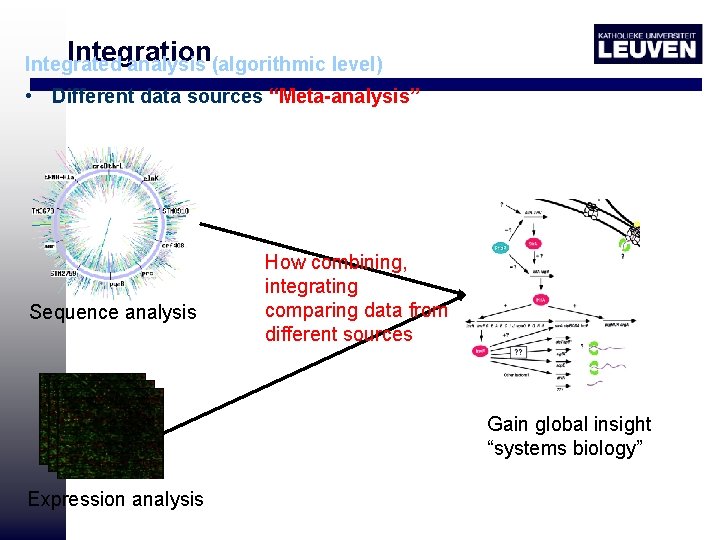
Integration Integrated analysis (algorithmic level) • Different data sources “Meta-analysis” Sequence analysis How combining, integrating comparing data from different sources Gain global insight “systems biology” Expression analysis
- Slides: 60