Bioinformatics and Grids Professor Carole Goble University of

Bioinformatics and Grids Professor Carole Goble, University of Manchester, UK carole@cs. man. ac. uk Director of my. Grid e-Science project Co-director ESNW e-Science Regional Centre

Roadmap n n Post Genome biology Challenges for bioinformatics n n n Why biology isn’t physics Information-centric Grids An example: my. Grid Other projects Take home

Take home n n n Complexity & Diversity - Size isn’t everything. Computation is important but information and knowledge services dominate. Integration, curation, annotation, fusion Automating support for integration and fusion means moving from… … human interaction to machine interaction. … machine readable to machineunderstandable. Metadata using ontologies for finding, managing & controlling services & content
Functional Genomics n n n An integrated view of how organisms work and interact in growth, development and pathogenesis From single gene to whole genome From single biochemical reactions to whole physiological and developmental systems What do genes do? How do they interact?
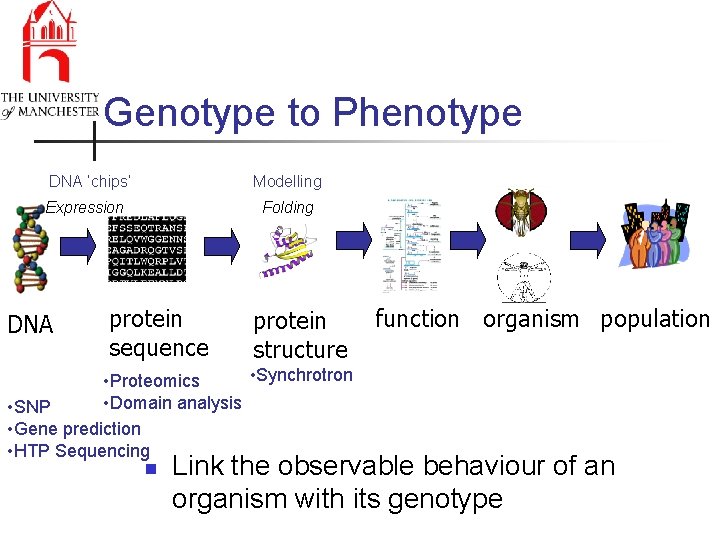
Genotype to Phenotype DNA ‘chips’ Modelling Expression DNA Folding protein sequence protein structure • Synchrotron • Proteomics • Domain analysis • SNP • Gene prediction • HTP Sequencing n function organism population Link the observable behaviour of an organism with its genotype
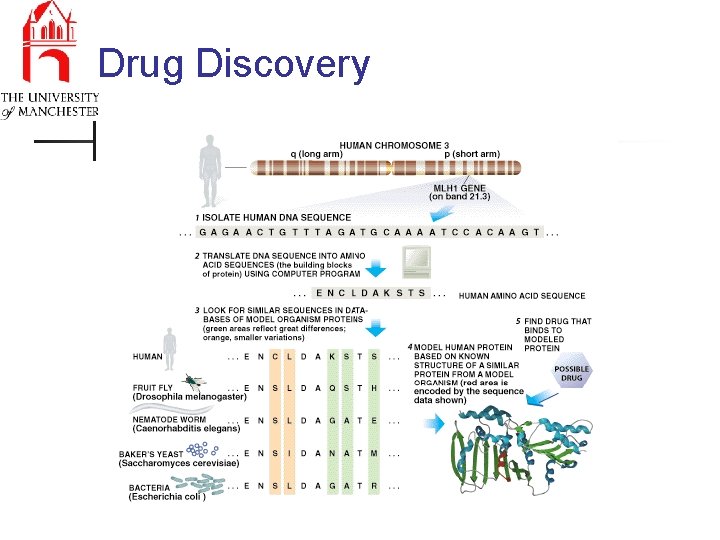
Drug Discovery
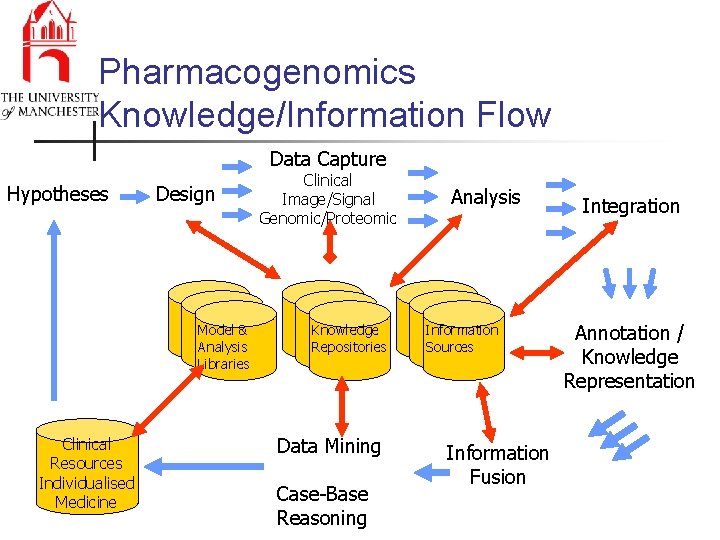
Pharmacogenomics Knowledge/Information Flow Data Capture Hypotheses Design Model & Analysis Libraries Clinical Resources Individualised Medicine Clinical Image/Signal Genomic/Proteomic Knowledge Repositories Data Mining Case-Base Reasoning Analysis Information Sources Information Fusion Integration Annotation / Knowledge Representation

Use Cases (I 3 C) n n n Show me all the genes in the glucose metabolism pathway and get their Gen. Bank accession numbers Find all the citations for the HOX gene family for human and mouse Find all the kinase genes from Wormbase and retrieve the DNA sequence

Use Cases Show me Nucleotide binding proteins in mouse Answer: n P 12345 in Swiss-Prot is an ATPase n Terri Attwood is an expert on this n Jackson labs have a database but you need to register n A paper has just been published in Proteins by the Stanford lab on this.
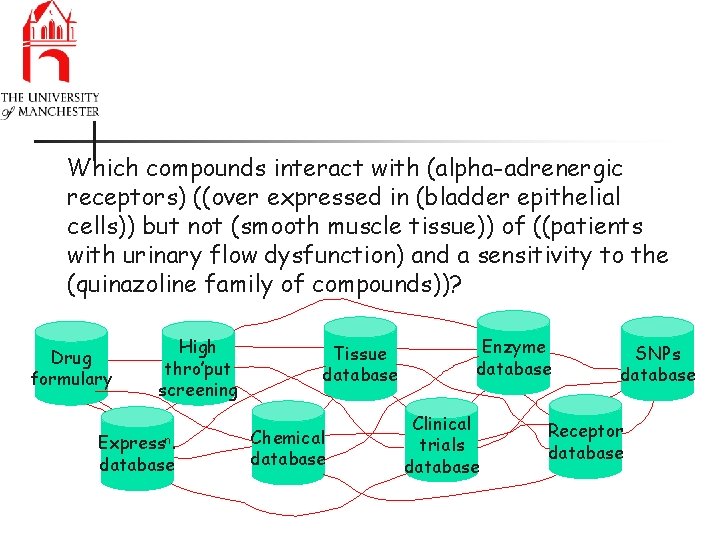
Which compounds interact with (alpha-adrenergic receptors) ((over expressed in (bladder epithelial cells)) but not (smooth muscle tissue)) of ((patients with urinary flow dysfunction) and a sensitivity to the (quinazoline family of compounds))? Drug formulary High thro’put screening Expressn. database Tissue database Chemical database Enzyme database Clinical trials database SNPs database Receptor database
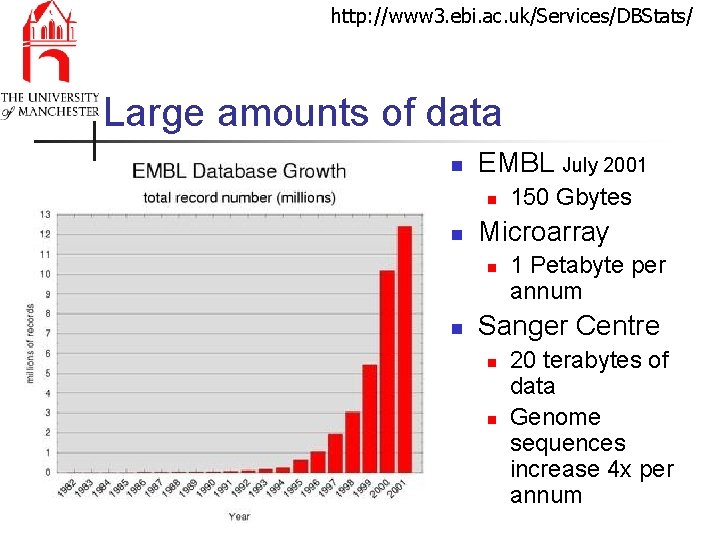
http: //www 3. ebi. ac. uk/Services/DBStats/ Large amounts of data n EMBL July 2001 n n Microarray n n 150 Gbytes 1 Petabyte per annum Sanger Centre n n 20 terabytes of data Genome sequences increase 4 x per annum
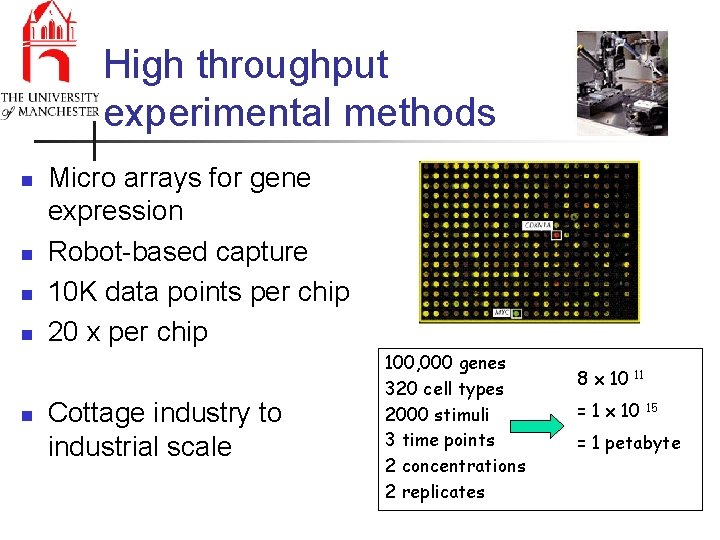
High throughput experimental methods n n n Micro arrays for gene expression Robot-based capture 10 K data points per chip 20 x per chip Cottage industry to industrial scale 100, 000 genes 320 cell types 2000 stimuli 3 time points 2 concentrations 2 replicates 8 x 10 11 = 1 x 10 15 = 1 petabyte

Heterogeneity n n Data types & forms Community Autonomy Over 500 different databases n n Different formats, structure, schemas, coverage… Web interfaces, flat file distribution, …
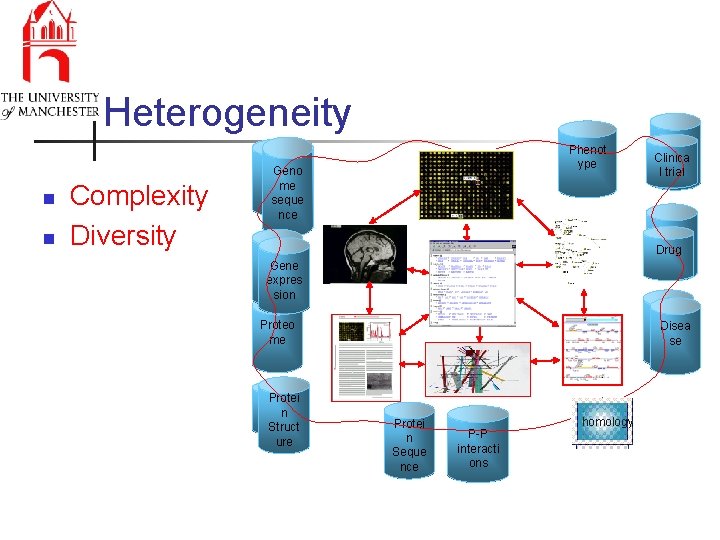
Heterogeneity n n Complexity Diversity Phenot ype Geno seque me nce seque nce Disea se Drug Gene expres sion Diseas Disea e se Proteo me Protei nn Struct ure Disea se Clinica l trial Protei n Seque nce P-P interacti ons homology
Heterogeneity n n Complexity Diversity Phenot ype Geno seque me nce seque nce Disea se Clinica l trial Disea se Drug Gene expres sion Genomic, proteomic, transcriptomic, metabalomic, protein-protein Proteo interactions, regulatory biome networks, alignments, disease, patterns & motifs, protein structure, Protei nn Protei protein classifications, Struct specialist n ure Seque proteins (enzymes, receptors), … nce Diseas Disea e se P-P interacti ons homology

Heterogeneous Data n n n Multimedia Images & Video Text annotations & literature Descriptive as well as numeric Knowledge-based Text Extraction

SWISSPROT: TET 9_ENTFA ID TET 9_ENTFA STANDARD; PRT; 639 AA. AC P 21598; DT 01 -MAY-1991 (REL. 18, CREATED) DT 01 -MAY-1991 (REL. 18, LAST SEQUENCE UPDATE) DT 01 -OCT-1993 (REL. 27, LAST ANNOTATION UPDATE) DE TETRACYCLINE RESISTANCE PROTEIN TETM (TRANSPOSON TN 916). GN TETM(916). OS ENTEROCOCCUS FAECALIS (STREPTOCOCCUS FAECALIS). RA BURDETT V. ; RL NUCLEIC ACIDS RES. 18: 6137 -6137(1990). CC -!- FUNCTION: ABOLISH THE INHIBITORY EFFECT OF TETRACYCLIN ON PROTEIN CC SYNTHESIS BY A NON-COVALENT MODIFICATION OF THE RIBOSOMES. CC -!- SIMILARITY: VERY HIGH TO OTHER TETM/TETO PROTEINS. CC -!- SIMILARITY: TO GTP-BINDING ELONGATION FACTORS. DR EMBL; X 56353; G 47062; -. DR PIR; S 13142. DR PROSITE; PS 00301; EFACTOR_GTP; 1. KW PROTEIN BIOSYNTHESIS; ANTIBIOTIC RESISTANCE; GTP-BINDING; KW TRANSPOSABLE ELEMENT. FT NP_BIND 10 17 GTP (BY SIMILARITY). FT NP_BIND 74 78 GTP (BY SIMILARITY). SQ SEQUENCE 639 AA; 72464 MW; 523 F 1359 CRC 32; >TET 9_ENTFA MKIINIGVLAHVDAGKTTLTESLLYNSGAITELGSVDKGTTRTDNTLLERQRGITIQTGI TSFQWENTKVNIIDTPGHMDFLAEVYRSLSVLDGAILLISAKDGVQAQTRILFHALRKMG IPTIFFINKIDQNGIDLSTVYQDIKEKLSAEIVIKQKVELYPNVCVTNFTESEQWDTVIE GNDDLLEKYMSGKSLEALELEQEESIRFQNCSLFPLYHGSAKSNIGIDNLIEVITNKFYS STHRGPSELCGNVFKIEYTKKRQRLAYIRLYSGVLHLRDSVRVSEKEKIKVTEMYTSING ELCKIDRAYSGEIVILQNEFLKLNSVLGDTKLLPQRKKIENPHPLLQTTVEPSKPEQREM LLDALLEISDSDPLLRYYVDSTTHEIILSFLGKVQMEVISALLQEKYHVEIEITEPTVIY MERPLKNAEYTIHIEVPPNPFWASIGLSVSPLPLGSGMQYESSVSLGYLNQSFQNAVMEG IRYGCEQGLYGWNVTDCKICFKYGLYYSPVSTPADFRMLAPIVLEQVLKKAGTELLEPYL SFKIYAPQEYLSRAYNDAPKYCANIVDTQLKNNEVILSGEIPARCIQEYRSDLTFFTNGR SVCLTELKGYHVTTGEPVCQPRRPNSRIDKVRYMFNKIT Swiss-Prot

Heterogeneity n Lymphocyte associated receptor of death n n n n n LARD WSL-LR WSL-S 1 WSL-S 2 proteins WSL-1 protein precursor Apoptosis-mediating receptor DR 3 Apoptosis-mediating receptor TRAMP Death Domain receptor 3 WSL protein apoptosis inducing receptor AIR APO-3

Functional genomics Tissue Structural Genomics Disease Population Genetics Genome Clinical Data Clinical trial sequence n n n Data resources have been built introspectively for human researchers Information is machine readable not machine understandable CONTROLLED VOCABULARIES & ONTOLOGIES
Shared data-> shared meaning Service provider Service provider
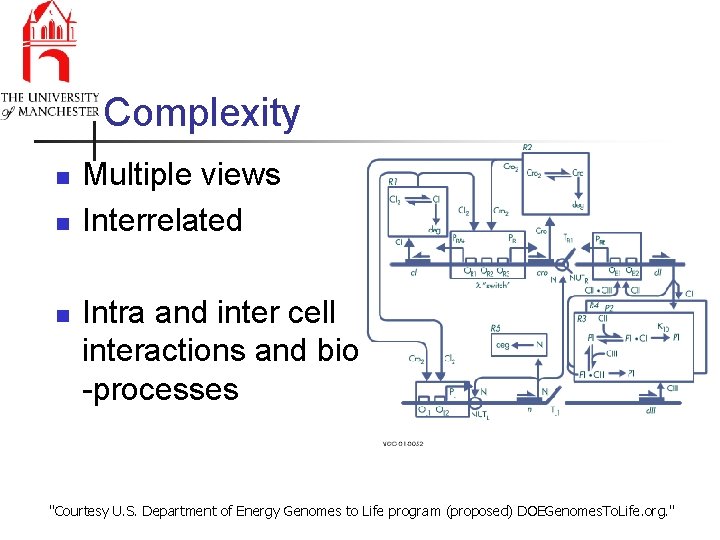
Complexity n n n Multiple views Interrelated Intra and inter cell interactions and bio -processes "Courtesy U. S. Department of Energy Genomes to Life program (proposed) DOEGenomes. To. Life. org. "

Instability & Quality n n Exploring the unknown n At least 5 definitions of a gene n The sequence is a model n Other models are “work in progress” Names unstable Data unstable Models unstable n “the problem in the field is not a lack of good integrating software, Smith says. The packages usually end up leading back to public databases. "The problem is: the databases are God-awful, " he told Bio. Med. Net. … If the data is still fundamentally flawed, then better algorithms add little. ” Temple Smith, director of the Molecular Engineering Research Center at Boston University,
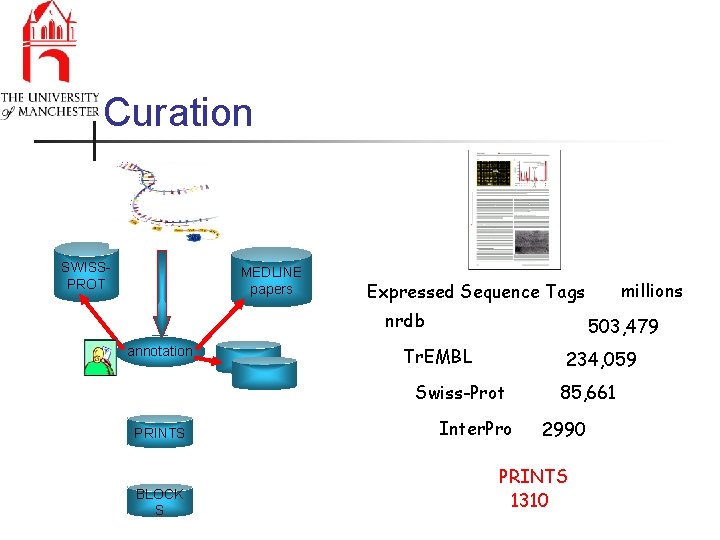
Curation SWISSPROT MEDLINE papers nrdb annotation 503, 479 Tr. EMBL 234, 059 Swiss-Prot PRINTS BLOCK S millions Expressed Sequence Tags Inter. Pro 85, 661 2990 PRINTS 1310
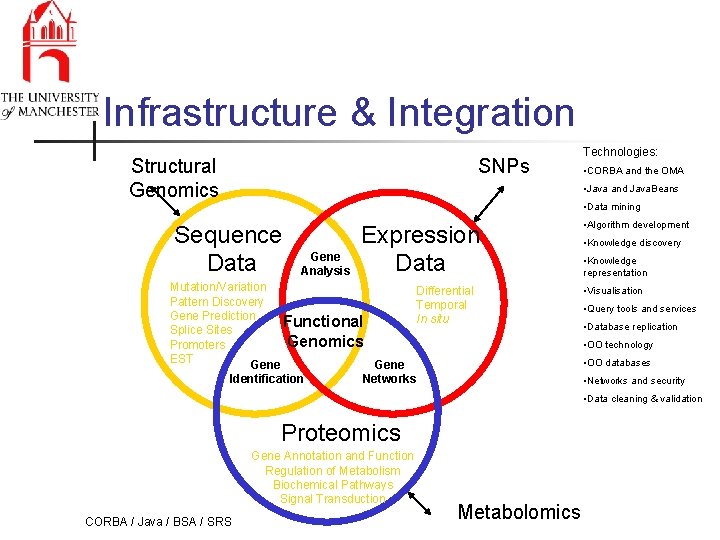
Infrastructure & Integration Structural Genomics SNPs Technologies: • CORBA and the OMA • Java and Java. Beans • Data mining Sequence Data Gene Analysis Expression Data Mutation/Variation Differential Pattern Discovery Temporal Gene Prediction In situ Functional Splice Sites Genomics Promoters EST Gene Identification Networks • Algorithm development • Knowledge discovery • Knowledge representation • Visualisation • Query tools and services • Database replication • OO technology • OO databases • Networks and security • Data cleaning & validation Proteomics Gene Annotation and Function Regulation of Metabolism Biochemical Pathways Signal Transduction CORBA / Java / BSA / SRS Metabolomics

Bioinformatics Analysis n Different algorithms n n Different implementations n n BLAST, FASTA, p. SW WU-BLAST, NCBI-BLAST Different service providers n NCBI, EBI, DDBJ

In silico experimentation
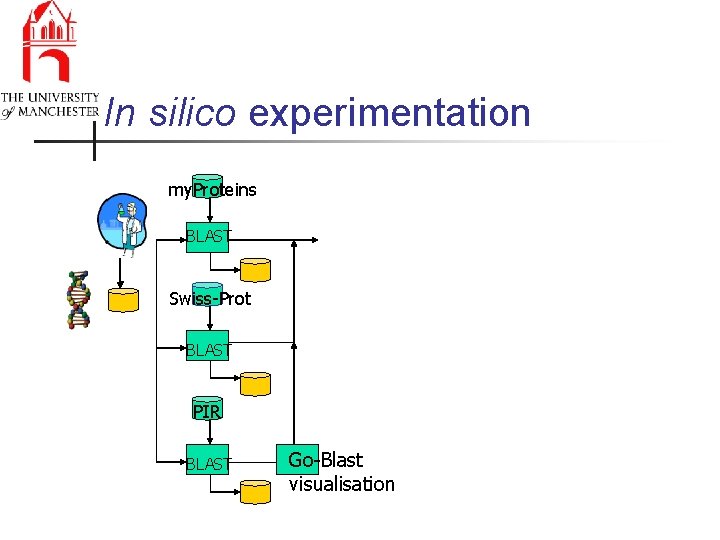
In silico experimentation my. Proteins BLAST Swiss-Prot BLAST PIR BLAST Go-Blast visualisation
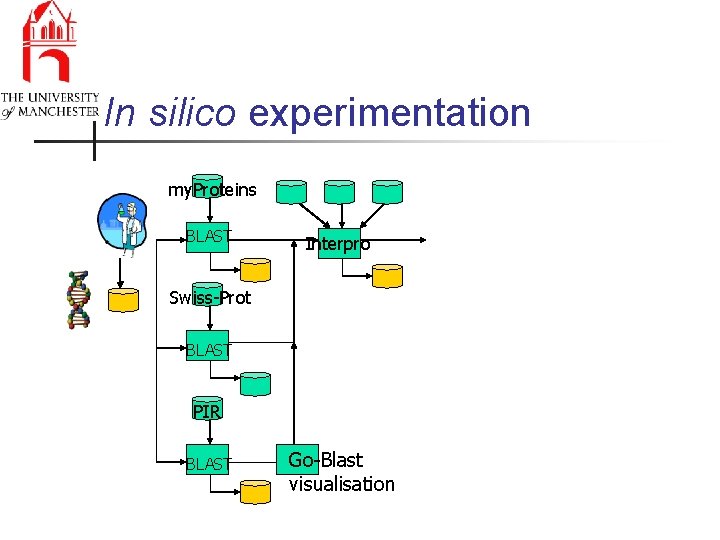
In silico experimentation my. Proteins BLAST Interpro Swiss-Prot BLAST PIR BLAST Go-Blast visualisation
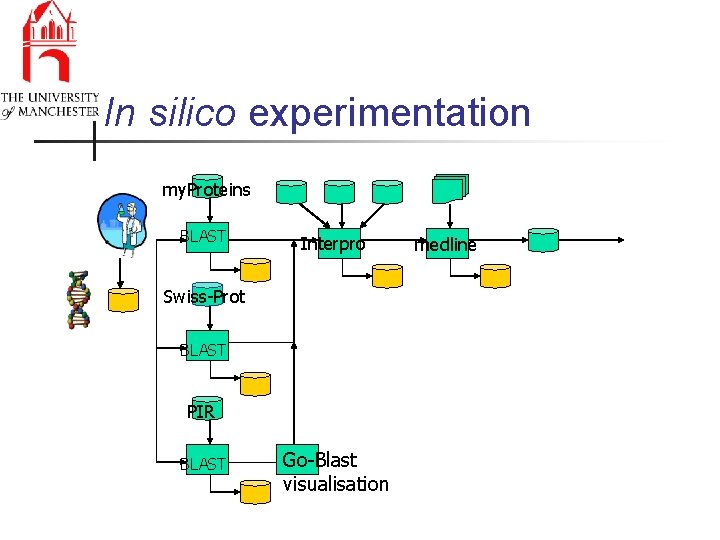
In silico experimentation my. Proteins BLAST Interpro Swiss-Prot BLAST PIR BLAST Go-Blast visualisation medline
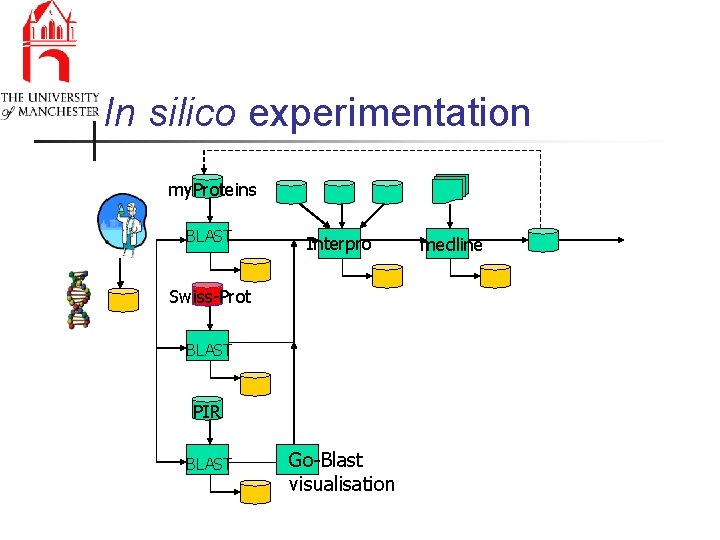
In silico experimentation my. Proteins BLAST Interpro Swiss-Prot BLAST PIR BLAST Go-Blast visualisation medline
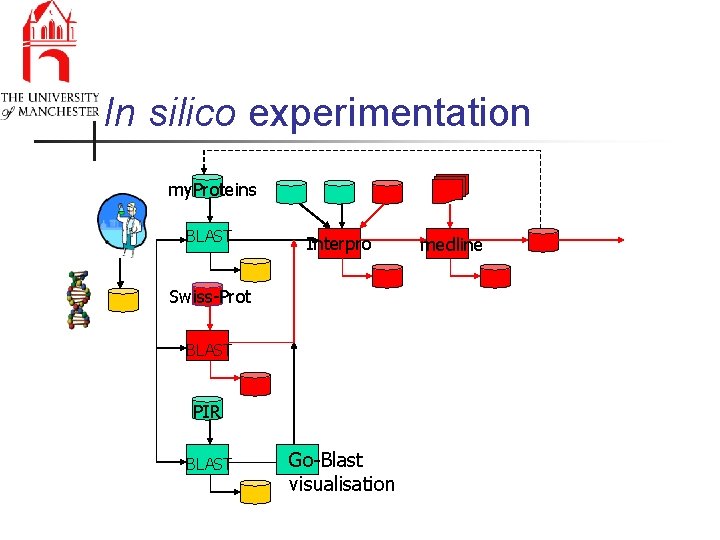
In silico experimentation my. Proteins BLAST Interpro Swiss-Prot BLAST PIR BLAST Go-Blast visualisation medline

In silico experimentation n n Discovery, interoperation, fusion, sharing of data, knowledge and workflows Explicit management of workflows n n Improving quality of experiments & data n n provenance & propagating change Scientific discovery is personal & global n n information & processes & best practice personalisation & collaborative working Security, ownership -> valuable assets

my. Grid n Personalised extensible environments for data-intensive in silico experiments in biology n http: //www. mygrid. org. uk n n n

my. Grid n n UK e-Science Grid programme pilot (EPSRC) Generic middleware Bioinformatics & Genomics setting 1 st October 2001 -- 31 st March 2005 n n (36 months funded in 42 execution period) 16 full-time researchers/developers

my. Grid Partners m
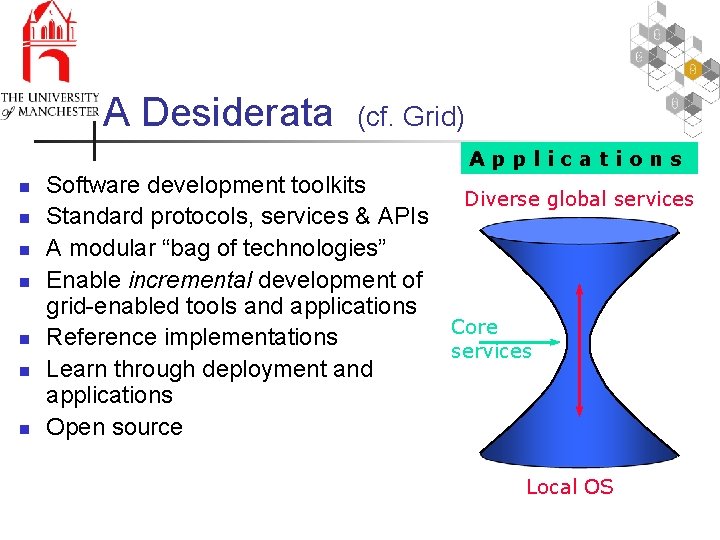
A Desiderata (cf. Grid) Applications n n n n Software development toolkits Diverse global services Standard protocols, services & APIs A modular “bag of technologies” Enable incremental development of grid-enabled tools and applications Core Reference implementations services Learn through deployment and applications Open source Local OS
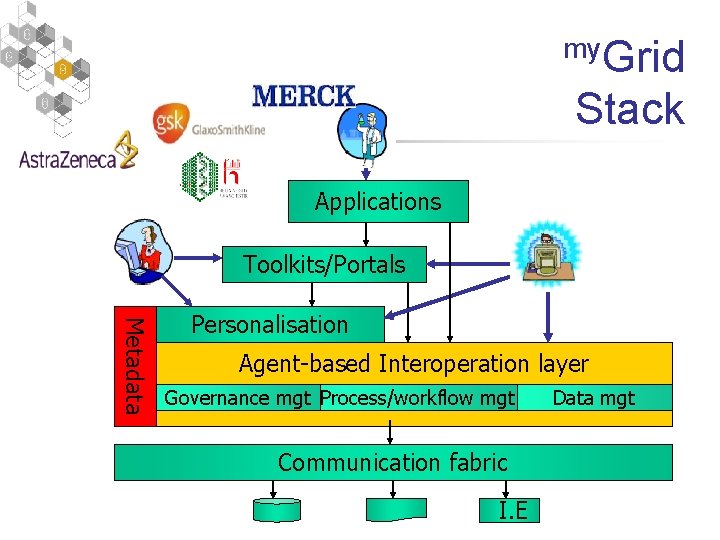
my. Grid Stack Approach Applications Toolkits/Portals Metadata Personalisation Agent-based Interoperation layer Governance mgt Process/workflow mgt Communication fabric I. E Data mgt

my. Grid 1. e-Scientists n n n 2. Outcomes Environment built on toolkits for service access, personalisation & community Gene function expression analysis using S. cerevisiae Annotation workbench for the PRINTS pattern database Developers n n Protocols and service descriptions my. Grid-in-a-Box developers kit Re-purposing DAS, App. Lab and Open. BSA … Integrating ISYS & Glaxo. Smith. Kline platforms

my. Grid 1. 2. 3. 4. 5. 6. 7. tech outcomes Services, service descriptions (ontologies), message protocols & APIs Database access from the Grid Process enactment on the Grid Personalisation services Provenance services Metadata services ~ DAML+OIL, RDF(S) Laying the foundations for Agent Services
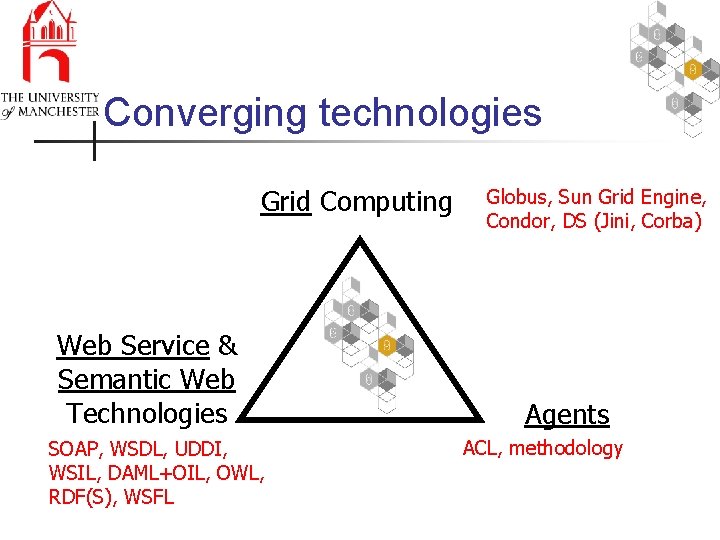
Converging technologies Grid Computing Web Service & Semantic Web Technologies SOAP, WSDL, UDDI, WSIL, DAML+OIL, OWL, RDF(S), WSFL Globus, Sun Grid Engine, Condor, DS (Jini, Corba) Agents ACL, methodology
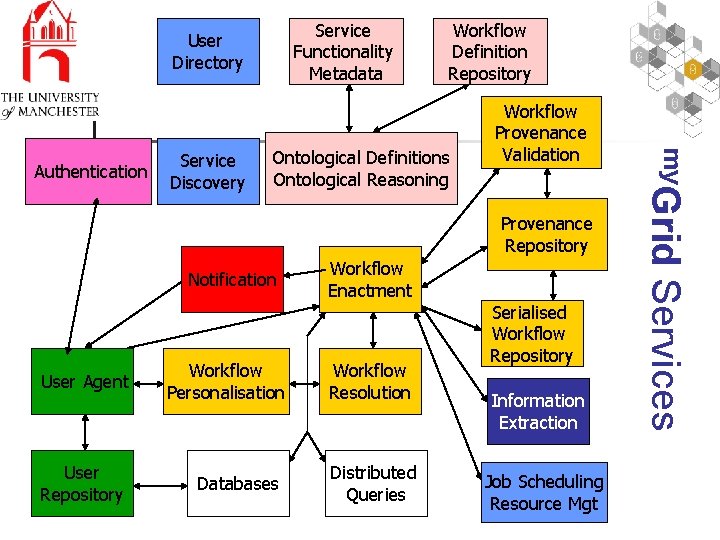
Service Functionality Metadata User Directory Service Discovery Ontological Definitions Ontological Reasoning Workflow Provenance Validation Provenance Repository User Agent User Repository Workflow Personalisation Databases Workflow Enactment Workflow Resolution Distributed Queries Serialised Workflow Repository Information Extraction Job Scheduling Resource Mgt Services Notification my. Grid Authentication Workflow Definition Repository
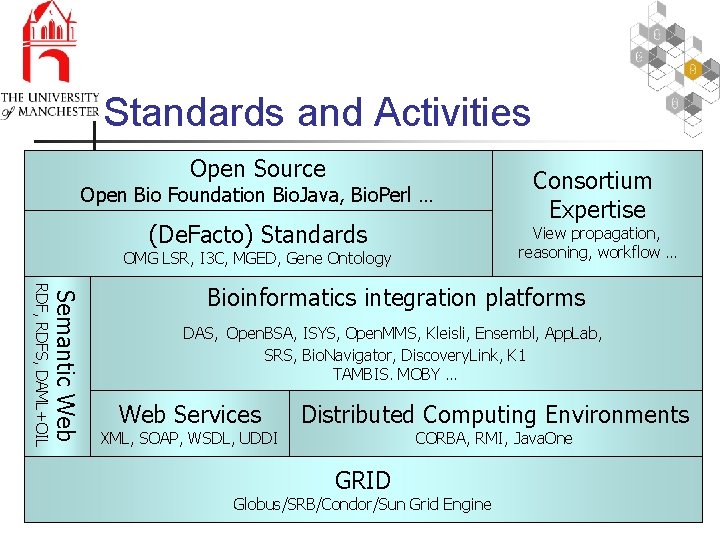
Standards and Activities Open Source Open Bio Foundation Bio. Java, Bio. Perl … (De. Facto) Standards Consortium Expertise View propagation, reasoning, workflow … OMG LSR, I 3 C, MGED, Gene Ontology Semantic Web RDF, RDFS, DAML+OIL Bioinformatics integration platforms DAS, Open. BSA, ISYS, Open. MMS, Kleisli, Ensembl, App. Lab, SRS, Bio. Navigator, Discovery. Link, K 1 TAMBIS. MOBY … Web Services XML, SOAP, WSDL, UDDI Distributed Computing Environments CORBA, RMI, Java. One GRID Globus/SRB/Condor/Sun Grid Engine

Other Bio. Grids n n n n n Bio. Opera North Carolina Bio. Grid Novartis Grid Scientific Annotation Middleware project Entropia AIDS modelling Grid …. Discovery. Net Proteomics analysis Protein structure prediction Biodiversity CLEF Clinical records …

my. Grid n n n Summary my. Grid aims to develop infrastructure middleware for an e-Biologist’s workbench The setting is bioinformatics but the results are intended to be generally applicable to e-Science A mix of standard, vanguard and bleed edge technologies, advanced development and (some) research Academic & commercial partnership my. Grid project is timely & reflects a community desire to “collaborate, or die”

Take home reprise n n n Complexity & Diversity - Size isn’t everything. Computation is important but information and knowledge services dominate. Integration, curation, annotation, fusion Automating support for integration and fusion means moving from… … human interaction to machine interaction. … machine readable to machineunderstandable. Metadata using ontologies for finding, managing & controlling services & content

Acknowledgements n n n Colleagues on my. Grid Robert Stevens Norman Paton Alan Robinson at EMBL-EBI I 3 C Interoperable Informatics Infrastructure Consortium http: //www. i 3 c. org

URLs n EBI n n LSR n n http: //www. omg. org/homepages/lsr/ Open-Bio n n http: //www. ebi. ac. uk/ http: //www. open-bio. org/ I 3 C n http: //www. i 3 c. org/

"Molecular biologists appear to have eyes for data that are bigger than their stomachs. As genomes near completion, as DNA arrays on chips begin to reveal patterns of gene sequences and expressions, as researchers embark on characterising all known proteins, the anticipated flood of data vastly exceeds in scale anything biologists have been used to. " (Editorial Nature, June 10, 1999)

n n Presented over the Access. Grid to the CSC Finnish IT Centre for Science Grid Seminar Otaniemi, Espoo, Finland 6 th March 2002 http: //www. csc. fi/suomi/tapahtumat/Grid Seminar/
- Slides: 49