Bioinformatics and Data Warehousing 1 Introduction to Bioinformatics

Bioinformatics and Data Warehousing 1) Introduction to Bioinformatics 2) FASTA File Format 3) Searching Gene Sequences (BLAST) 4) Data Management in Biomedical Informatics Michael Kane, Ph. D. Computer & Information Technology Bindley Bioscience Center Purdue University
DNA is Information Storage

“Zipped Files” Decompression “Executable Files”
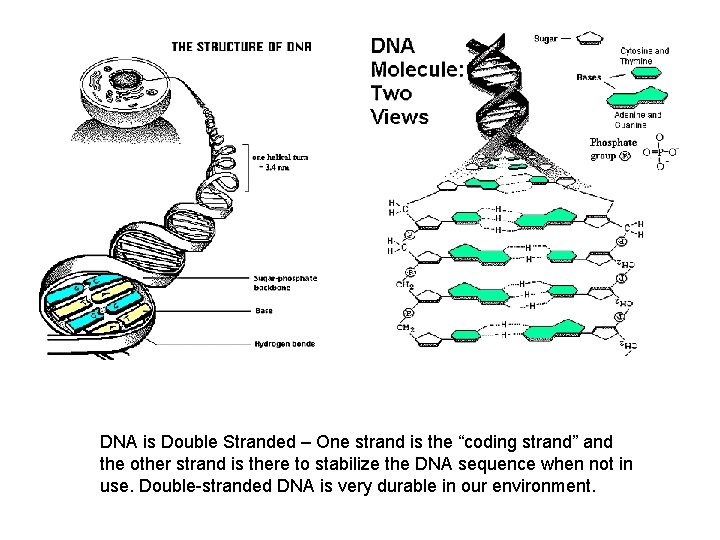
DNA is Double Stranded – One strand is the “coding strand” and the other strand is there to stabilize the DNA sequence when not in use. Double-stranded DNA is very durable in our environment.

CAGGACCATGGAACTCAGCGTCCTCCTCTTCCTTGCACTCCTCACAGGACTCTTGCTAC TCCTGGTTCAGCGCCACCCTAACACCCATGACCGCCTCCCACCAGGGCCCCGCCCTCT GCCCCTTTTGGGAAACCTTCTGCAGATGGATAGAAGAGGCCTACTCAAATCCTTTCTGA GGTTCCGAGAGAAATATGGGGACGTCTTCACGGTACACCTGGGACCGAGGCCCGTGGT CATGCTGTGTGGAGTAGAGGCCATACGGGAGGCCCTTGTGGACAAGGCTGAGGCCTTC TCTGGCCGGGGAAAAATCGCCATGGTCGACCCATTCTTCCGGGGATATGGTGTGATCTT TGCCAATGGAAACCGCTGGAAGGTGCTTCGGCGATTCTCTGTGACCACTATGAGGGAC TTCGGGATGGGAAAGCGGAGTGTGGAGGAGCGGATTCAGGAGGAGGCTCAGTGTCTG ATAGAGGAGCTTCGGAAATCCAAGGGGGCCCTCATGGACCCCACCTTCCTCTTCCAGT CCATTACCGCCAACATCATCTGCTCCATCGTCTTTGGAAAACGATTCCACTACCAAGAT CAAGAGTTCCTGAAGATGCTGAACTTGTTCTACCAGACTTTTTCACTCATCAGCTCTGTA TTCGGCCAGCTGTTTGAGCTCTTCTCTGGCTTCTTGAAATACTTTCCTGGGGCACACAG GCAAGTTTACAAAAACCTGCAGGAAATCAATGCTTACATTGGCCACAGTGTGGAGAAG CACCGTGAAACCCTGGACCCCAGCGCCCCCAAGGACCTCATCGACACCTGCTCC ACATGGAAAAAGAGAAATCCAACGCACACAGTGAATTCAGCCACCAGAACCTCAACCT CAACACGCTCTTCTTTGCTGGCACTGAGACCACCAGCACCACTCTCCGCTACG GCTTCCTGCTCAAATACCCTCATGTTGCAGAGTCTACAGGGAGATTGAA CAGGTGATTGGCCCACATCGCCCTCCAGAGCTTCATGACCGAGCCAAAATGCCATACA CAGAGGCAGTCATCTATGAGATTCAGAGATTTTCCGACCTTCTCCCCATGGGTGTGCCC CACATTGTCACCCAACACACCAGCTTCCGAGGGTACATCATCCCCAAGGACACAGAAG TATTTCTCATCCTGAGCACTGCTCTCCATGACCCACACTA

THEREDCAT_HSDKLSD_WASNOTHOTBUT_WKKNASDN KSAOJ. ASDNALKS_WASWET_ASDFLKSDOFIJEIJKNAW DFN_ANDMAD_WERN. JSNDFJN_YETSAD_MNSFDGPOIJ D_BUTTHEFOX_SDKMFIDSJIR. JER_GOTWET_JSN. DFOI AMNJNER_ANDATEHIM.
Start with a thin 2 x 4 lego block… Add a 2 x 2 lego block… Add a 2 x 3 lego block… Add a 2 x 4 lego block…
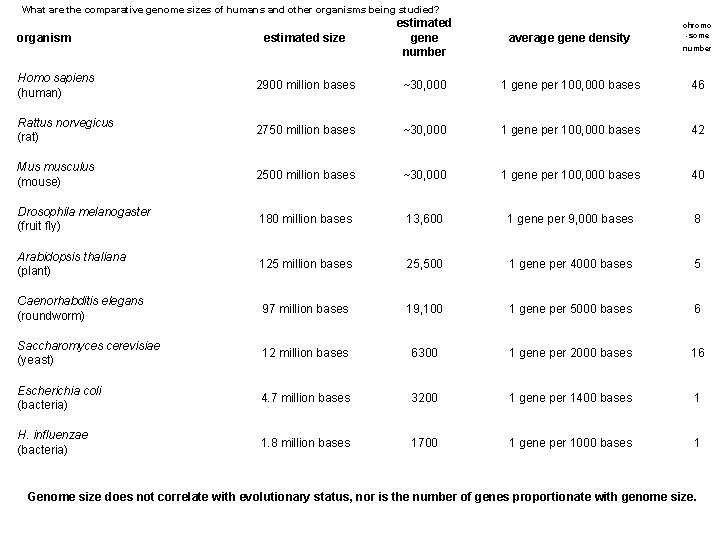
What are the comparative genome sizes of humans and other organisms being studied? estimated size estimated gene number average gene density Homo sapiens (human) 2900 million bases ~30, 000 1 gene per 100, 000 bases 46 Rattus norvegicus (rat) 2750 million bases ~30, 000 1 gene per 100, 000 bases 42 Mus musculus (mouse) 2500 million bases ~30, 000 1 gene per 100, 000 bases 40 Drosophila melanogaster (fruit fly) 180 million bases 13, 600 1 gene per 9, 000 bases 8 Arabidopsis thaliana (plant) 125 million bases 25, 500 1 gene per 4000 bases 5 Caenorhabditis elegans (roundworm) 97 million bases 19, 100 1 gene per 5000 bases 6 Saccharomyces cerevisiae (yeast) 12 million bases 6300 1 gene per 2000 bases 16 Escherichia coli (bacteria) 4. 7 million bases 3200 1 gene per 1400 bases 1 H. influenzae (bacteria) 1. 8 million bases 1700 1 gene per 1000 bases 1 organism chromo -some number Genome size does not correlate with evolutionary status, nor is the number of genes proportionate with genome size.

![FASTA File Format Tiny. Seq XML >gi|1924940|emb|CAA 67058. 1| myosin-IF [Homo sapiens] QEKLTSRKMDSRWGGRSESINVTLNVEQAAYTRDALAKGLYARLFDFLVEAINRAMQKPQEEYSIGVLDI YGFEIFQKNGFEQFCINFVNEKLQQIFIELTLKAEQEEYVQEGIRWTPIQYFNNKVVCDLIENKLSPPGI FASTA File Format Tiny. Seq XML >gi|1924940|emb|CAA 67058. 1| myosin-IF [Homo sapiens] QEKLTSRKMDSRWGGRSESINVTLNVEQAAYTRDALAKGLYARLFDFLVEAINRAMQKPQEEYSIGVLDI YGFEIFQKNGFEQFCINFVNEKLQQIFIELTLKAEQEEYVQEGIRWTPIQYFNNKVVCDLIENKLSPPGI](http://slidetodoc.com/presentation_image_h/103057150b3e164c1683b5dbecbffabe/image-10.jpg)
FASTA File Format Tiny. Seq XML >gi|1924940|emb|CAA 67058. 1| myosin-IF [Homo sapiens] QEKLTSRKMDSRWGGRSESINVTLNVEQAAYTRDALAKGLYARLFDFLVEAINRAMQKPQEEYSIGVLDI YGFEIFQKNGFEQFCINFVNEKLQQIFIELTLKAEQEEYVQEGIRWTPIQYFNNKVVCDLIENKLSPPGI MSVLDDVCATMHATGGGADQTLLQKLQAAVGTHEHFNSWSAGFVIHHYAGKVSYDVSGFCERNRDVLFSD LIELMQSSDQAFLRMLFPEKLDGDKKGRPSTAGSKIKKQANDLVATLMRCTPHYIRCIKPNETKHARDWE ENRVQHQVEYLGLKENIRVRRAGFAYRRQFAKFLQRYAILTPETWPRWRGDERQGVQHLLRAVNMEPDQY QMGSTKVFVKNPESLFLLEEVRERKFDGFARTIQKAWRRHVAVRKYEEMREEASNILLNKKERRRNSINR NFVGDYLGLEERPELRQFLGKKERVDFADSVTKYDRRFKPIKRDLILTPKCVYVIGREKMKKGPEKGPVC EILKKKLDIQALRGVSLSTRQDDFFILQEDAADSFLESVFKTEFVSLLCKRFEEATRRPLPLTFSDTLQF RVKKEGWGGGGTRSVTFSRGFGDLAVLKVGGRTLTVSVGDGLPKNSKPTGKGLAKGKPRRSSQAPTRAAP GAPQGMDRNGAPLCPQGGAPCPLEKFIWPRGHPQASPALRPHPWDASRRPRARPPSEHNTEFLNVPDQGM AGMQRKRSVGQRPVPVGRPKPQPRTHGPRCRALYQYVGQDVDELSFNVNEVIEILMEDPSGWWKGRLHGQ EGLFPGNYVEKI <? xml version="1. 0"? > <!DOCTYPE TSeq PUBLIC "-//NCBI TSeq/EN" "http: //www. ncbi. nlm. nih. gov/dtd/NCBI_TSeq. dtd"> <TSeq_seqtype value="nucleotide"/> <TSeq_gi>1924939</TSeq_gi> <TSeq_accver>X 98411. 1</TSeq_accver> <TSeq_taxid>9606</TSeq_taxid> <TSeq_orgname>Homo sapiens</TSeq_orgname> <TSeq_defline>Homo sapiens partial m. RNA for myosin-IF</TSeq_defline> <TSeq_length>2711</TSeq_length> <TSeq_sequence>CAGGAGAAGCTGACCAGCCGCAAGATGGACAGCCGCTGGGGCGCAGCGAGTCCATCAATGT…… </TSeq>

FASTA File Format DATABASE (DATA WAREHOUSE) >GENE NUMBER ONE AGCTGCTAGTAGAGTCGCTCGGATAGGACTGCTAGCTGC. . . . >GENE NUMBER TWO AGCTGCTAGTAGAGTCGCTCGGATAGGACTGCTAGCTGC. . . . >GENE NUMBER THREE AGCTGCTAGTAGAGTCGCTCGGATAGGACTGCTAGCTGC. . . . >GENE NUMBER FOUR AGCTGCTAGTAGAGTCGCTCGGATAGGACTGCTAGCTGC. . . . >GENE NUMBER FIVE AGCTGCTAGTAGAGTCGCTCGGATAGGACTGCTAGCTGC. . . . >GENE NUMBER SIX AGCTGCTAGTAGAGTCGCTCGGATAGGACTGCTAGCTGC. . . . >GENE NUMBER SEVEN AGCTGCTAGTAGAGTCGCTCGGATAGGACTGCTAGCTGC. . . >GENE NUMBER TWENTY MILLION AGCTGCTAGTAGAGTCGCTCGGATAGGACTGCTAGCTGC. . . .

DYNAMIC PROGRAMMING and SEQUENCE SEARCHES 'Dynamic programming' is an efficient programming technique for solving certain combinatorial problems. It is particularly important in bioinformatics as it is the basis of sequence alignment algorithms for comparing protein and DNA sequences. In the bioinformatics application Dynamic Programming gives a spectacular efficiency gain over a purely recursive algorithm. Don't expect much enlightenment from the etymology of the term 'dynamic programming, ' though. Dynamic programming was formalized in the early 1950 s by mathematician Richard Bellman, who was working at RAND Corporation on optimal decision processes. He wanted to concoct an impressive name that would shield his work from US Secretary of Defense Charles Wilson, a man known to be hostile to mathematics research. His work involved time series and planning—thus 'dynamic' and 'programming' (note, nothing particularly to do with computer programming). Bellman especially liked 'dynamic' because "it's impossible to use the word dynamic in a derogatory sense"; he figured dynamic programming was "something not even a Congressman could object to. ”

DYNAMIC PROGRAMMING and SEQUENCE SEARCHES The following is an example of global sequence alignment using Needleman/ Wunsch techniques. For this example, the two sequences to be globally aligned are: G A A T T C A G T T A (sequence #1) G G A T C G A (sequence #2) Initialization Step Since this example assumes there is no gap opening or gap extension penalty, the first row and first column of the matrix can be initially filled with 0.

DYNAMIC PROGRAMMING and SEQUENCE SEARCHES Matrix Fill Step
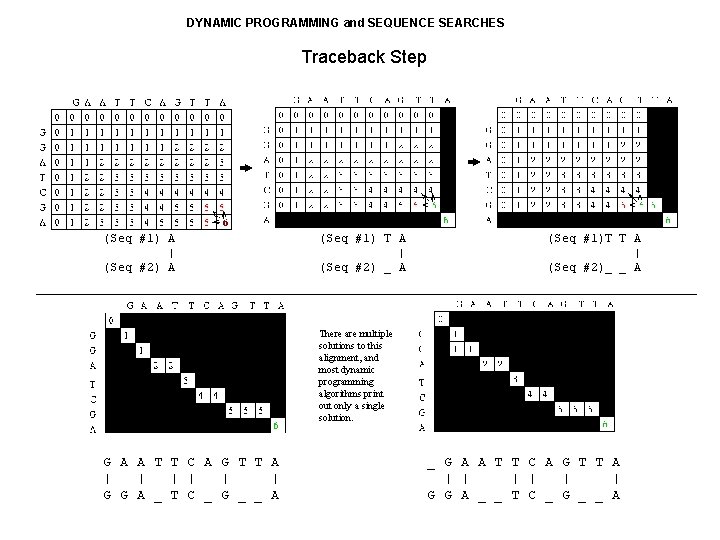
DYNAMIC PROGRAMMING and SEQUENCE SEARCHES Traceback Step (Seq #1) A | (Seq #2) A (Seq #1) T A | (Seq #2) _ A (Seq #1)T T A | (Seq #2)_ _ A There are multiple solutions to this alignment, and most dynamic programming algorithms print out only a single solution. G A A T T C A G T T A | | | G G A _ T C _ G _ _ A _ G A A T T C A G T T A | | | G G A _ _ T C _ G _ _ A

BLAST (Basic Local Alignment Search Tool) Why is BLAST so fast? By preindexing all the possible 11 -letter words into the database records (411 = 4, 194, 304). . . GTCGTAGTCG ATCGTAGTCG CTCGTAGTCG. . Steps: 1) Find all the 11 -letter words in your query sequence, plus a few variations. 2) Look these up in the 11 -letter-word index. 3) Retrieve all sequences containing those words. 4) Use a rigorous algorithm (e. g. Smith-Waterman) to extend the match in both directions

http: //www. ncbi. nlm. nih. gov/ >UNKNOWN GENE SEQUENCE AGGACCATGGAACTCAGCGTCCTCCTCTTCCTTGCACTCCTCACAGGACTCTTGCTACTC CTGGTTCAGCGCCACCCTAACACCCATGACCGCCTCCCACCAGGGCCCCGCCCTCTGCC CCTTTTGGGAAACCTTCTGCAGATGGATAGAAGAGGCCTACTCAAATCCTTTCTGAGGTT CCGAGAGAAATATGGGGACGTCTTCACGGTACACCTGGGACCGAGGCCCGTGGTCATG CTGTGTGGAGTAGAGGCCATACGGGAGGCCCTTGTGGACAAGGCTGAGGCCTTCTCTG GCCGGGGAAAAATCGCCATGGTCGACCCATTCTTCCGGGGATATGGTGTGATCTTTGCC AATGGAAACCGCTGGAAGGTGCTTCGGCGATTCTCTGTGACCACTATGAGGGACTTCGG GATGGGAAAGCGGAGTGTGGAGGAGCGGATTCAGGAGGAGGCTCAGTGTCTGATAGA GGAGCTTCGGAAATCCAAGGGGGCCCTCATGGACCCCACCTTCCTCTTCCAGTCCATTA CCGCCAACATCATCTGCTCCATCGTCTTTGGAAAACGATTCCACTACCAAGATCAAGAGT TCCTGAAGATGCTGAACTTGTTCTACCAGACTTTTTCACTCATCAGCTCTGTATTCGGCCA GCTGTTTGAGCTCTTCTCTGGCTTCTTGAAATA
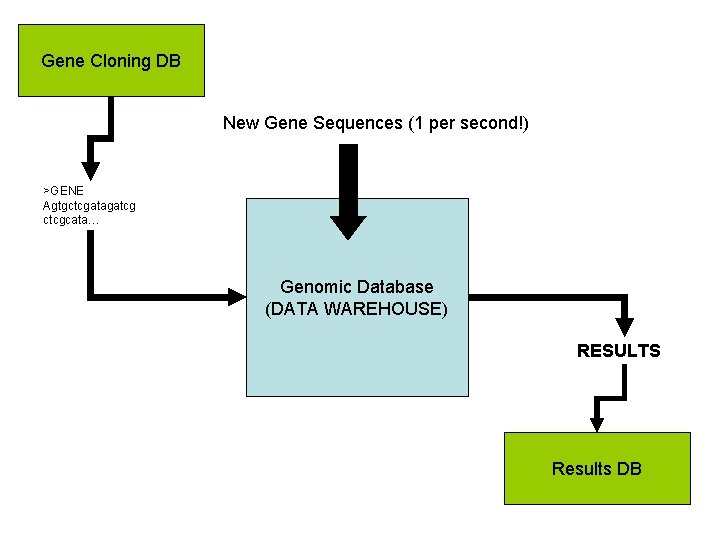
Gene Cloning DB New Gene Sequences (1 per second!) >GENE Agtgctcgatagatcg ctcgcata… Genomic Database (DATA WAREHOUSE) RESULTS Results DB
- Slides: 18