Bioinformatics and Computational Molecular Biology Geoff Barton http

Bioinformatics and Computational Molecular Biology Geoff Barton http: //www. compbio. dundee. ac. uk

Practical Tutorial • Dr David Martin practical tutorial on the use of pymol molecular graphics software. • In this lecture I will show lots of protein structures – use www. ebi. ac. uk/msd to find them, and/or scop domains database (find with google).

Similarities in Proteins • Lecture 1 – Overview of data in molecular biology – Protein modelling – Similarities of Protein Sequence, Structure, Function

Introduction to Sequence Comparison • Lecture 2: – Why compare sequences? – Methods for sequence comparison/alignment. – Multiple alignment – Database searching - FASTA/BLAST – Iterative searching - PSI-BLAST

Practical/WWW references • Organised by Drs Martin – Good preparation would be to look at: http: //www. ebi. ac. uk/Tools and http: //www. ncbi. nlm. nih. gov – Look at BLAST and FASTA on these sites as well as database access facilities.
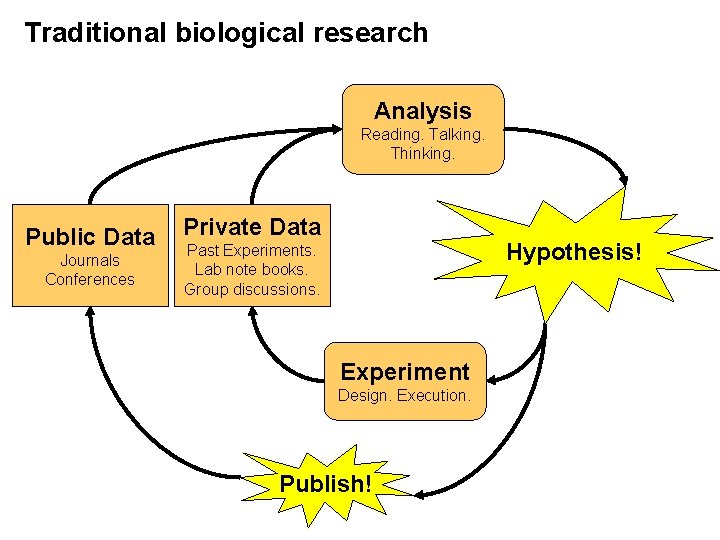
Traditional biological research Analysis Reading. Talking. Thinking. Public Data Journals Conferences Private Data Hypothesis! Past Experiments. Lab note books. Group discussions. Experiment Design. Execution. Publish!
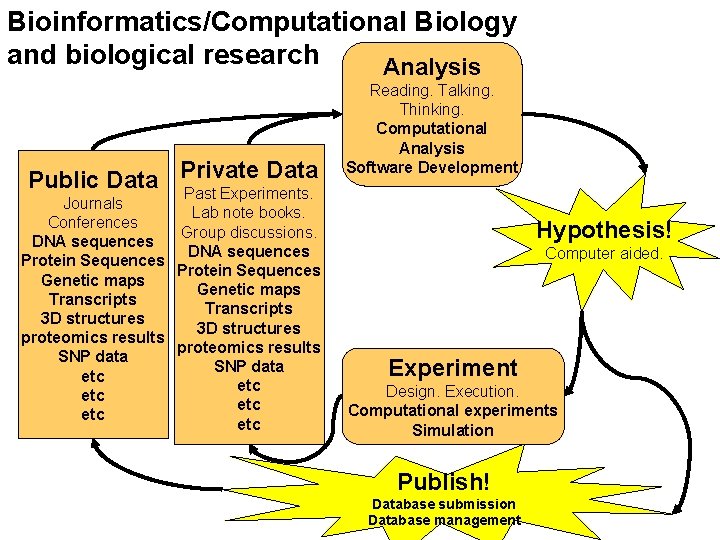
Bioinformatics/Computational Biology and biological research Analysis Data Public Data Private Past Experiments. Journals Lab note books. Conferences Group discussions. DNA sequences Protein Sequences Genetic maps Transcripts 3 D structures proteomics results SNP data etc etc etc Reading. Talking. Thinking. Computational Analysis Software Development Hypothesis! Computer aided. Experiment Design. Execution. Computational experiments Simulation Publish! Database submission Database management
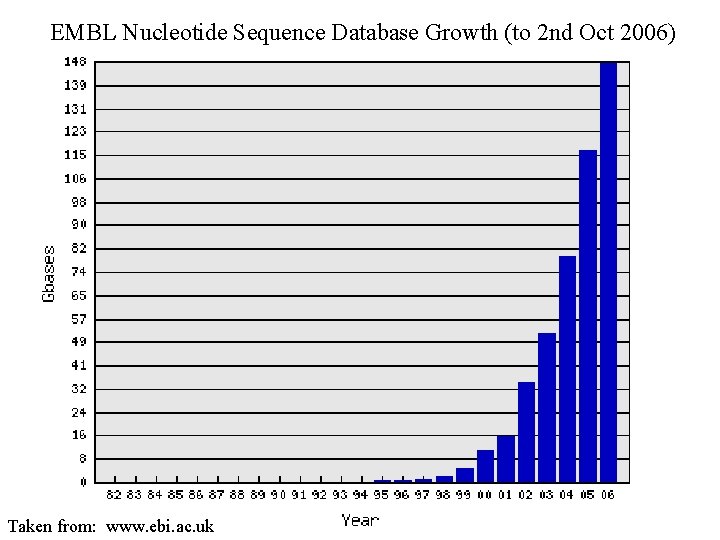
EMBL Nucleotide Sequence Database Growth (to 2 nd Oct 2006) Taken from: www. ebi. ac. uk

Protein Sequences Approx 3, 500, 000 known for all species (Oct. 2006. ) 25, 000 for Human (not counting splice variants and posttranslational modifications)

Protein 3 D Structures Approx 39, 000 known (much duplication)

Biological data in context
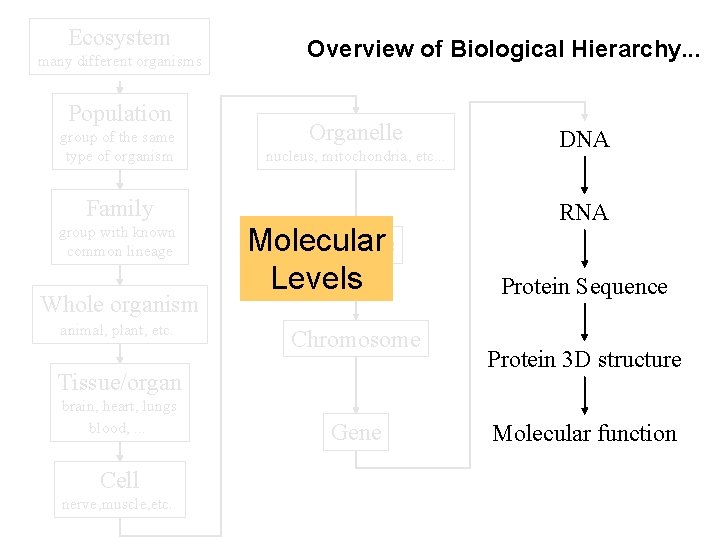
Ecosystem many different organisms Population group of the same type of organism Overview of Biological Hierarchy. . . Organelle nucleus, mitochondria, etc. . . Family group with known common lineage Whole organism animal, plant, etc. Molecular Nucleus Levels Chromosome Tissue/organ brain, heart, lungs blood, . . . Cell nerve, muscle, etc. . Gene DNA RNA Protein Sequence Protein 3 D structure Molecular function
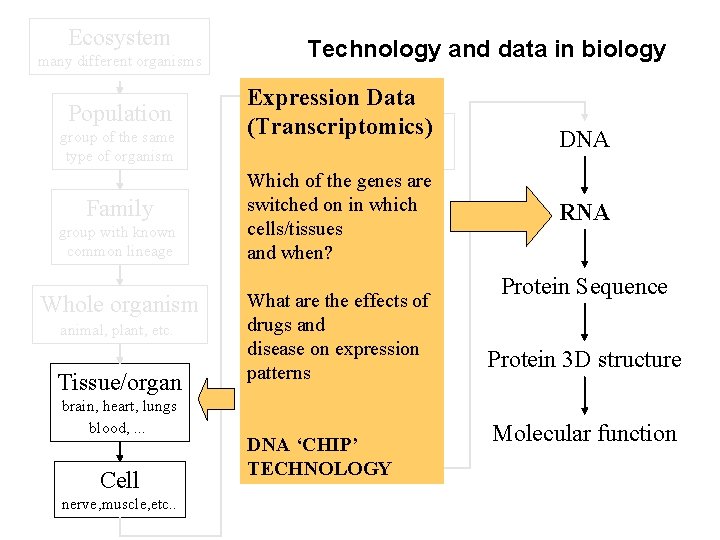
Ecosystem many different organisms Population group of the same type of organism Family group with known common lineage Whole organism animal, plant, etc. Tissue/organ brain, heart, lungs blood, . . . Cell nerve, muscle, etc. . Technology and data in biology Expression Data (Transcriptomics) Organelle nucleus, mitochondria, etc. . . Which of the genes are switched on in which cells/tissues Nucleus and when? What are the effects of drugs and Chromosome disease on expression patterns Gene DNA ‘CHIP’ TECHNOLOGY DNA RNA Protein Sequence Protein 3 D structure Molecular function
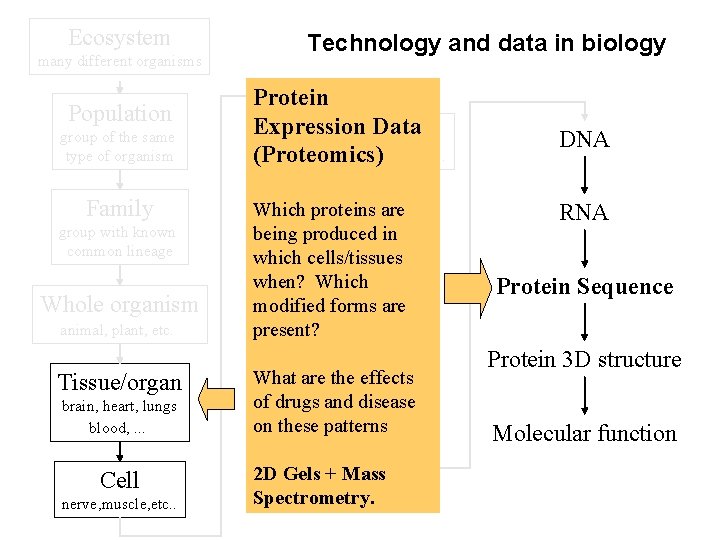
Ecosystem many different organisms Population group of the same type of organism Family group with known common lineage Whole organism animal, plant, etc. Tissue/organ brain, heart, lungs blood, . . . Cell nerve, muscle, etc. . Technology and data in biology Protein Expression Data Organelle nucleus, mitochondria, etc. . . (Proteomics) Which proteins are being produced in Nucleus which cells/tissues when? Which modified forms are present? Chromosome What are the effects of drugs and disease on these patterns Gene 2 D Gels + Mass Spectrometry. DNA RNA Protein Sequence Protein 3 D structure Molecular function
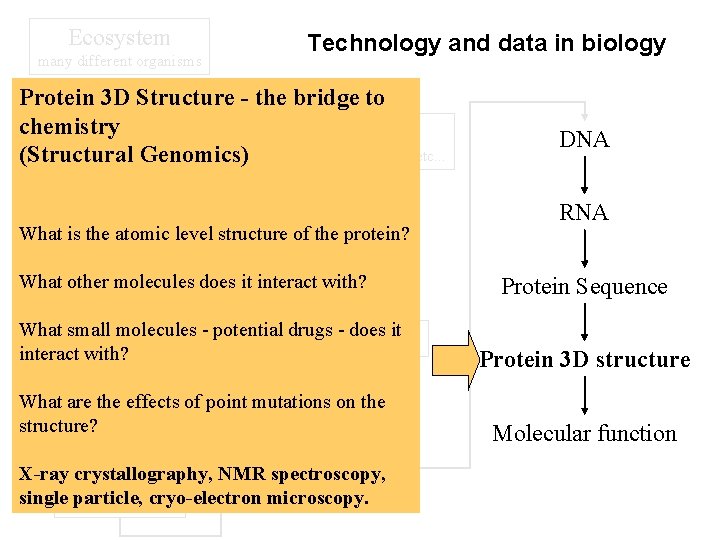
Ecosystem many different organisms Technology and data in biology Protein 3 D Structure - the bridge to Population chemistry Organelle group of the same type of organism (Structural Genomics) nucleus, mitochondria, etc. . . Family knownlevel structure of the protein? Whatgroup is thewith atomic common lineage DNA RNA Nucleus What other molecules does it interact with? Whole organism etc. Whatanimal, small plant, molecules - potential drugs - does it Chromosome interact with? Tissue/organ Whatbrain, are the effects heart, lungs of point mutations on the structure? blood, . . . Gene X-ray crystallography, NMR spectroscopy, Cell singlenerve, muscle, etc. . particle, cryo-electron microscopy. Protein Sequence Protein 3 D structure Molecular function
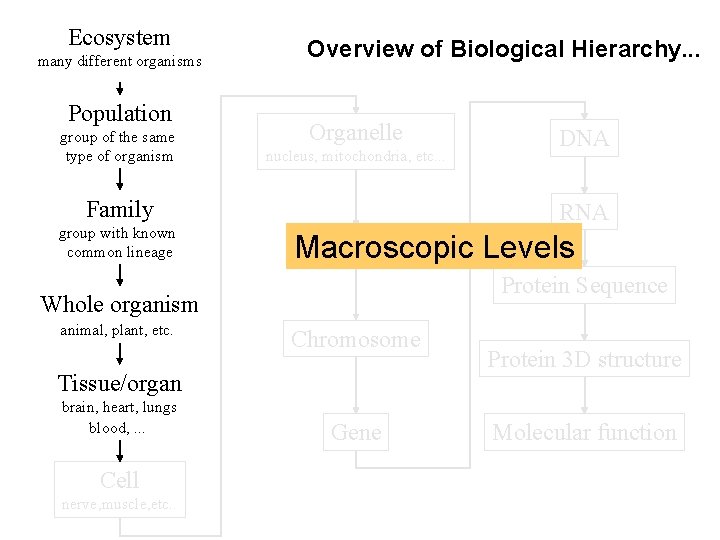
Ecosystem many different organisms Population group of the same type of organism Overview of Biological Hierarchy. . . Organelle nucleus, mitochondria, etc. . . Family group with known common lineage RNA Nucleus Macroscopic Levels Protein Sequence Whole organism animal, plant, etc. Chromosome Tissue/organ brain, heart, lungs blood, . . . Cell nerve, muscle, etc. . DNA Gene Protein 3 D structure Molecular function

Biology is now a data intensive science To do good science, you need to know how to use (and not abuse) computational tools.

Protein Structure Prediction • ‘Homology’ modelling – Relies on the fact that similarity of sequence implies similarity of 3 D structure.
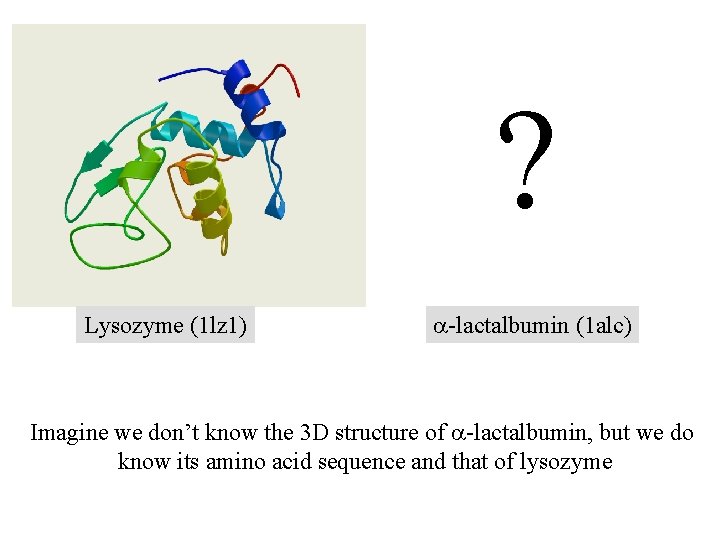
? Lysozyme (1 lz 1) a-lactalbumin (1 alc) Imagine we don’t know the 3 D structure of a-lactalbumin, but we do know its amino acid sequence and that of lysozyme
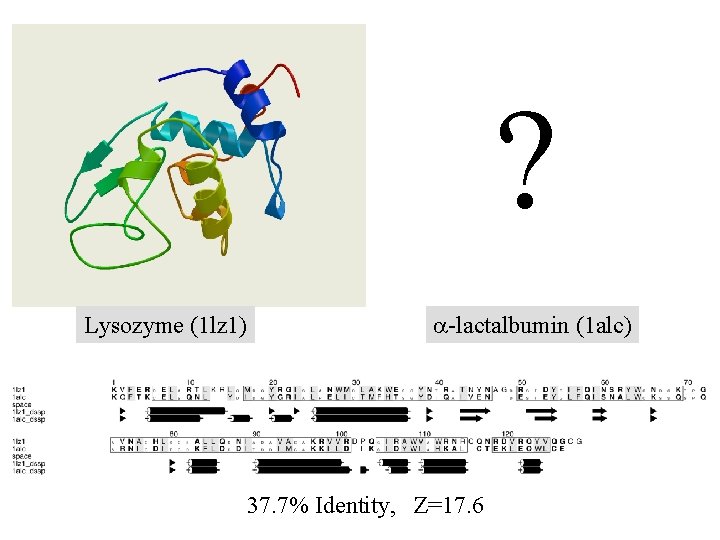
? Lysozyme (1 lz 1) a-lactalbumin (1 alc) 37. 7% Identity, Z=17. 6

Protein structure prediction (Homology Modelling) • Align sequence of protein of unknown structure to sequence of protein of known structure. • In ‘conserved core’ of protein, substitute the amino acid types into the known structure. • Deal with ‘loops’ between the core elements of structure.
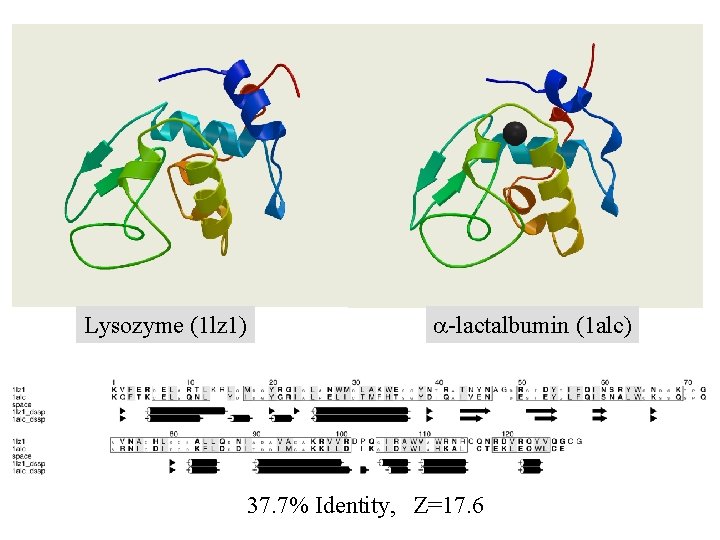
Lysozyme (1 lz 1) a-lactalbumin (1 alc) 37. 7% Identity, Z=17. 6

Protein structure prediction (Homology modelling) • Problems: – Need protein of known structure that is similar in sequence. – Building loops where there are deletions. – Verifying model. • Key is getting a good alignment in the first place – Bad alignment => bad model.

Good alignment on its own can: • Identify key residues (absolutely conserved) • Identify likely protein core (conserved hydrophobic residues) • Help predict protein secondary structure (not this lecture).

Sequence alignment is a fundamental technique in molecular biology. • May predict proteins of common function even when no 3 D structure is known. • May be used to predict 3 D structure and so help understanding of mutants. • Some examples of where this is right and wrong. . .

Prediction of structure and function by similarity to known sequences and structures Assumption is that similar sequence implies similar structure and function. But what do we mean by “similar”? Does similarity of sequence really imply similarity of function?
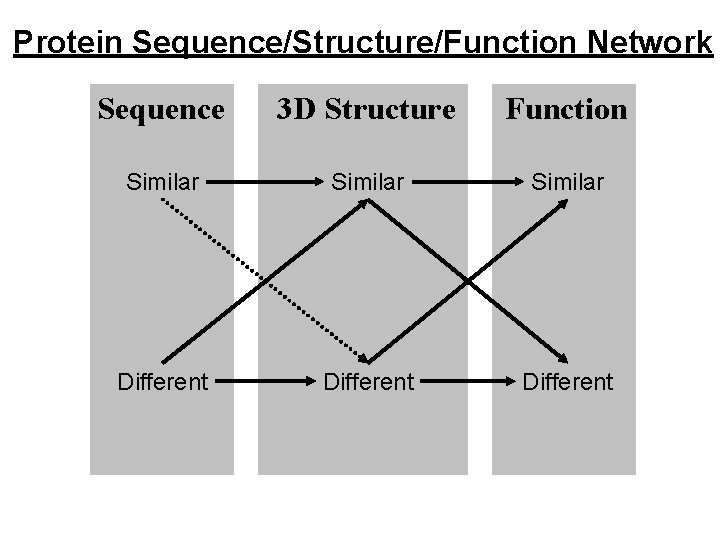
Protein Sequence/Structure/Function Network Sequence 3 D Structure Function Similar Different
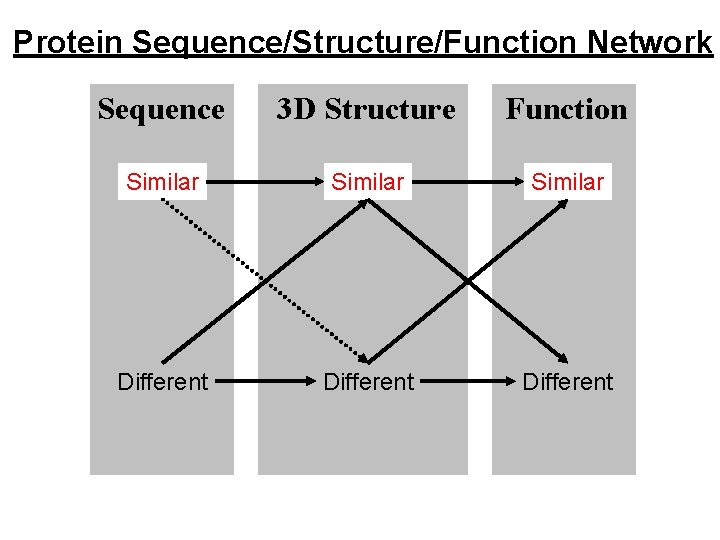
Protein Sequence/Structure/Function Network Sequence 3 D Structure Function Similar Different

Similar Sequence, Similar Structure, Similar Function. e. g. Trypsin-like Serine Proteinases Same fold, same catalytic mechanism. But DIFFERENT specificity. e. g. Immunoglobulin variable domains. Same fold, similar binding function. But DIFFERENT specificity. True of all examples. Similarities only give clues to function, differences in specificity can be regarded as differences of function.
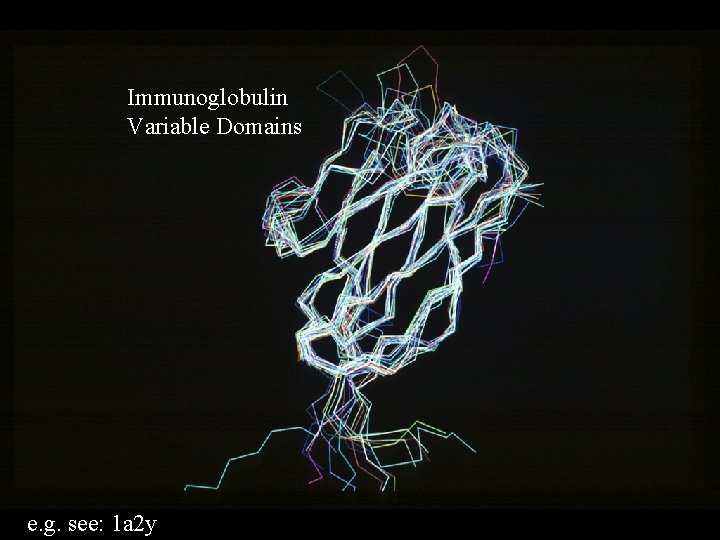
Immunoglobulin Variable Domains e. g. see: 1 a 2 y
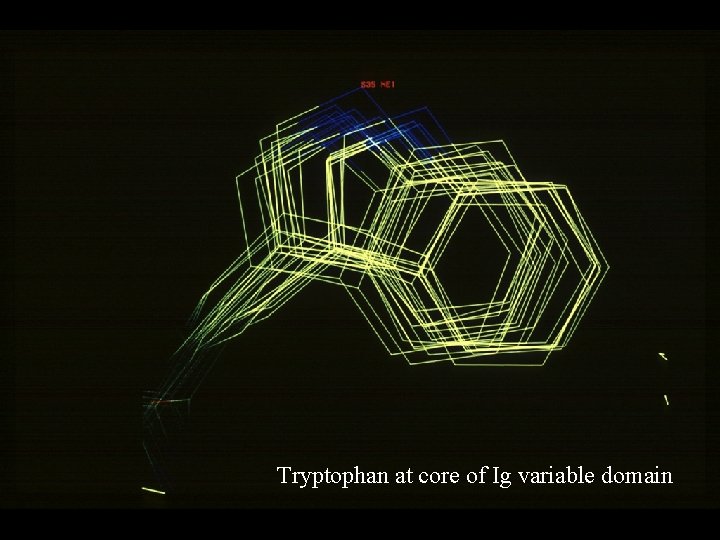
Tryptophan at core of Ig variable domain
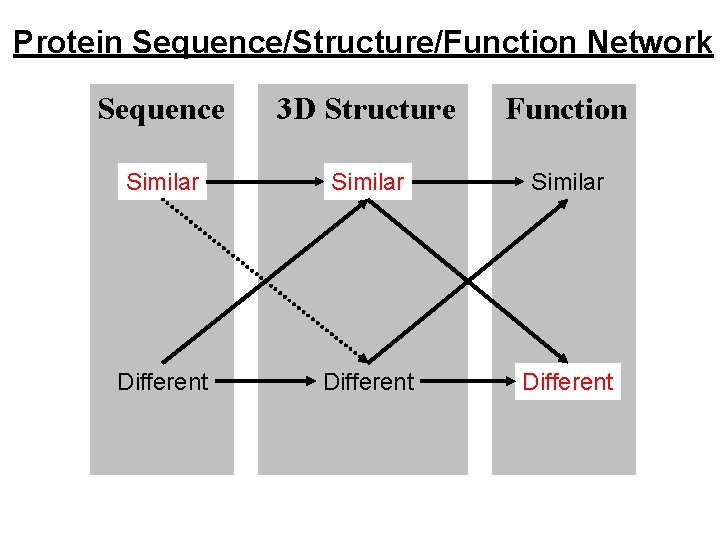
Protein Sequence/Structure/Function Network Sequence 3 D Structure Function Similar Different
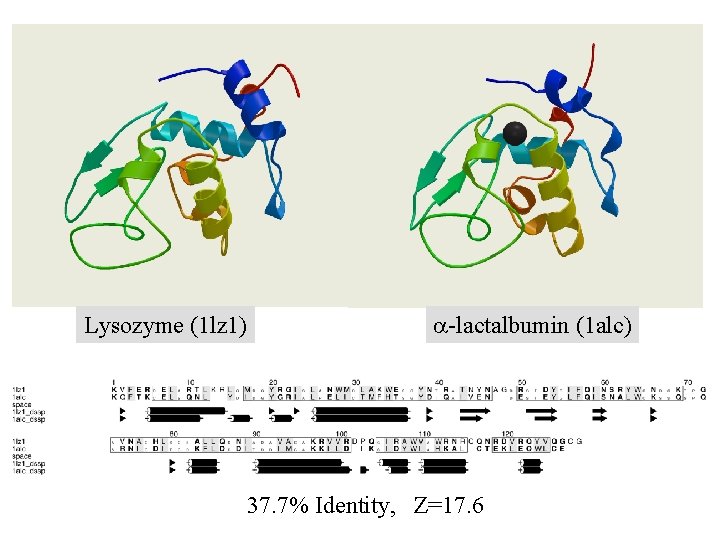
Lysozyme (1 lz 1) a-lactalbumin (1 alc) 37. 7% Identity, Z=17. 6
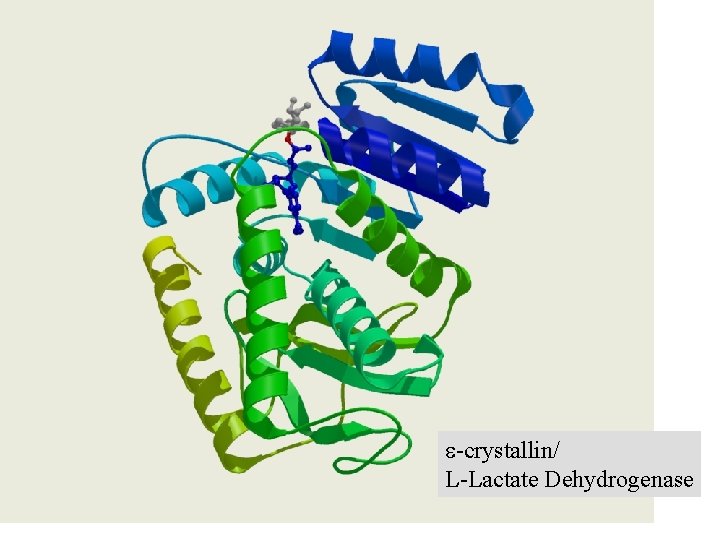
e-crystallin/ L-Lactate Dehydrogenase
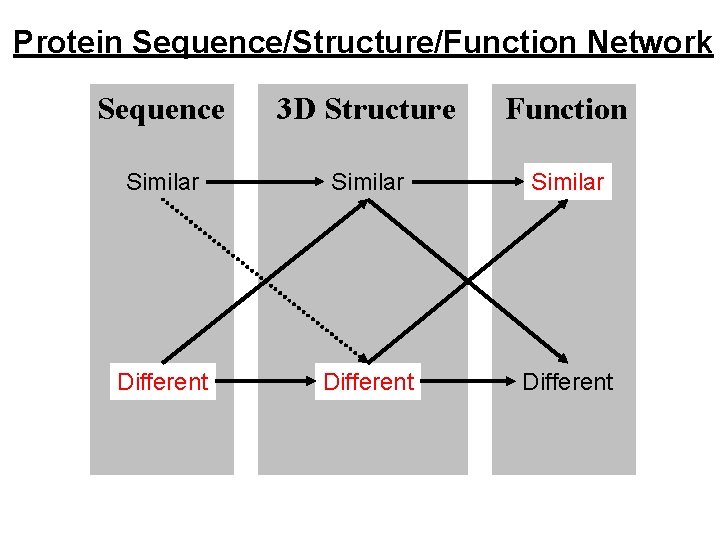
Protein Sequence/Structure/Function Network Sequence 3 D Structure Function Similar Different
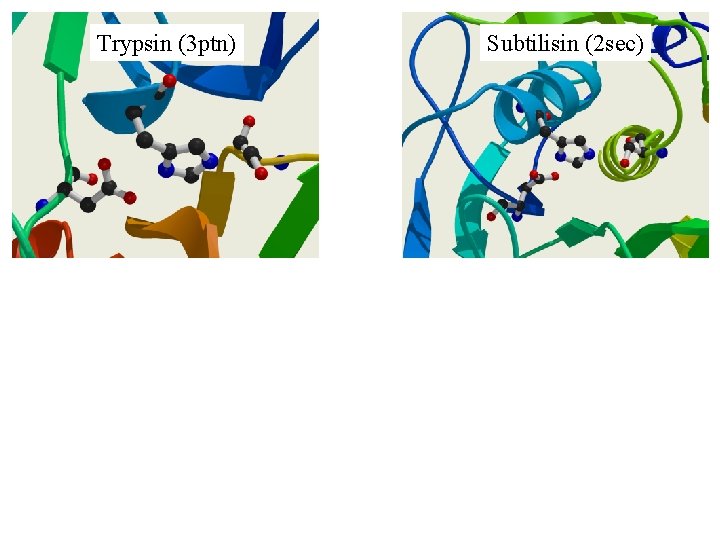
Trypsin (3 ptn) Subtilisin (2 sec)
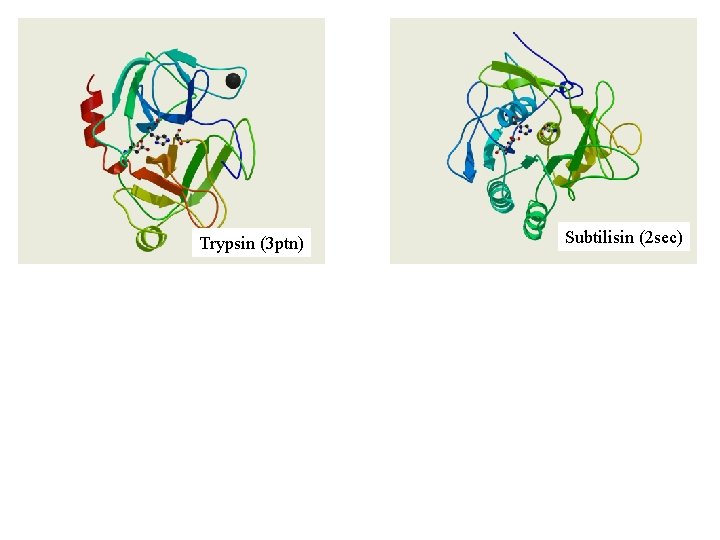
Trypsin (3 ptn) Subtilisin (2 sec)
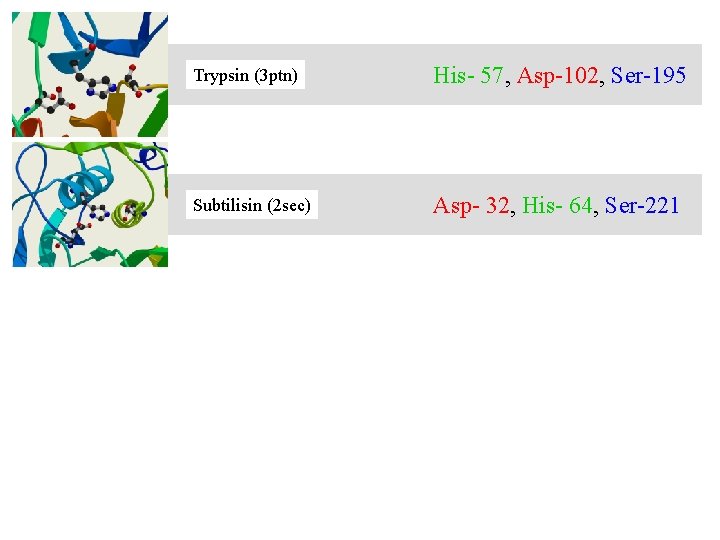
Trypsin (3 ptn) His- 57, Asp-102, Ser-195 Subtilisin (2 sec) Asp- 32, His- 64, Ser-221
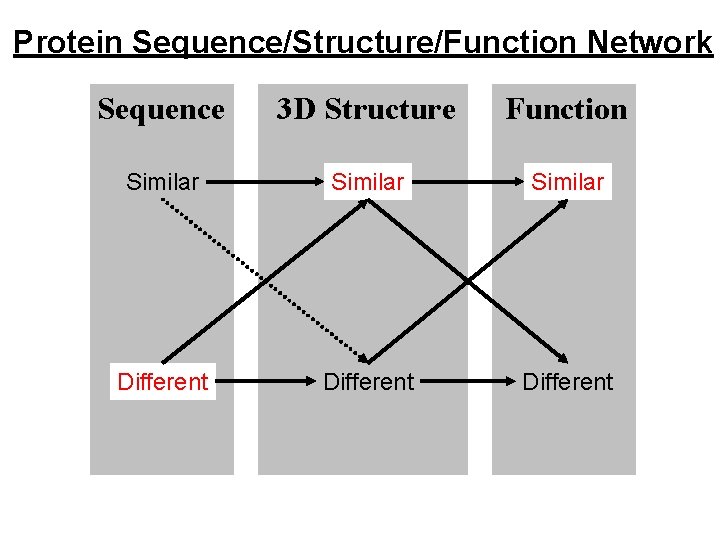
Protein Sequence/Structure/Function Network Sequence 3 D Structure Function Similar Different
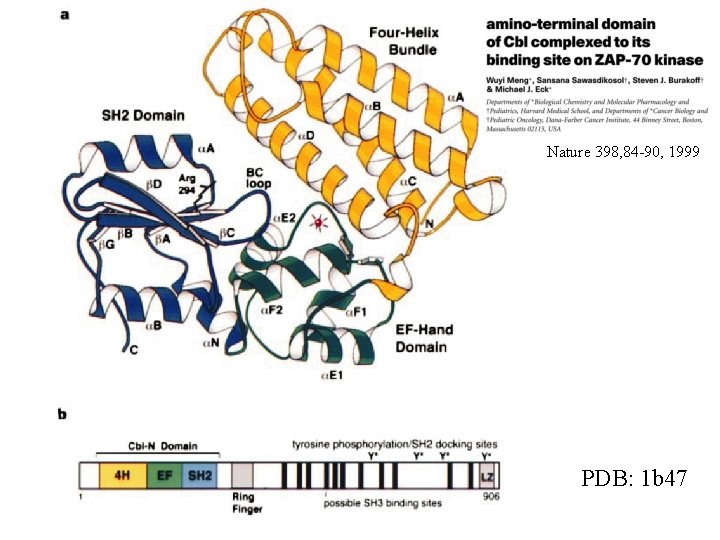
Nature 398, 84 -90, 1999 PDB: 1 b 47
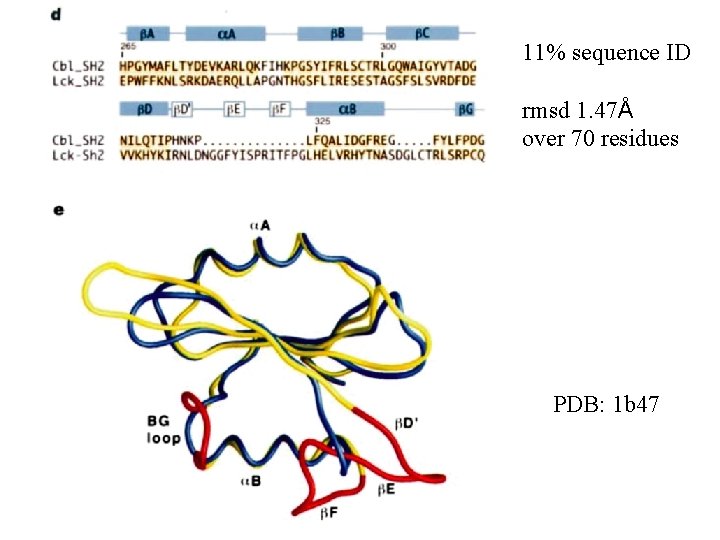
11% sequence ID rmsd 1. 47Å over 70 residues PDB: 1 b 47
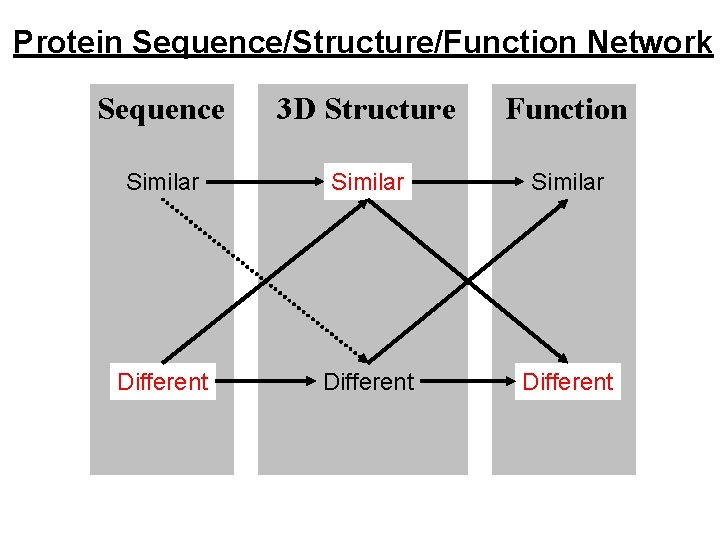
Protein Sequence/Structure/Function Network Sequence 3 D Structure Function Similar Different
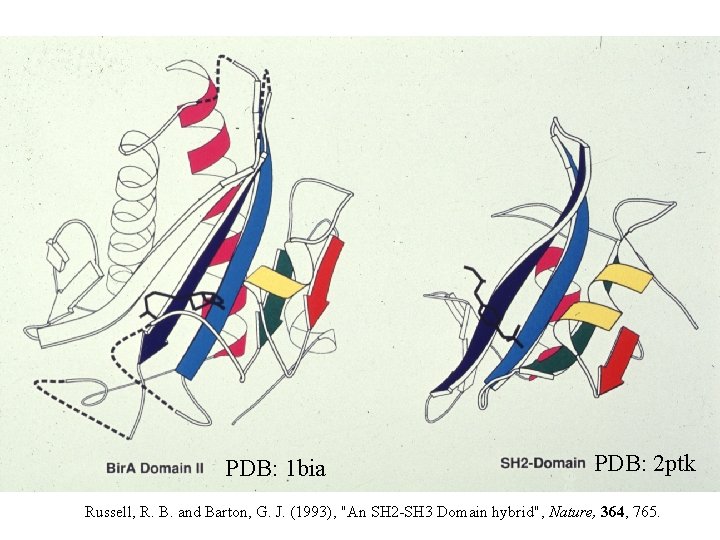
PDB: 1 bia PDB: 2 ptk Russell, R. B. and Barton, G. J. (1993), "An SH 2 -SH 3 Domain hybrid", Nature, 364, 765.
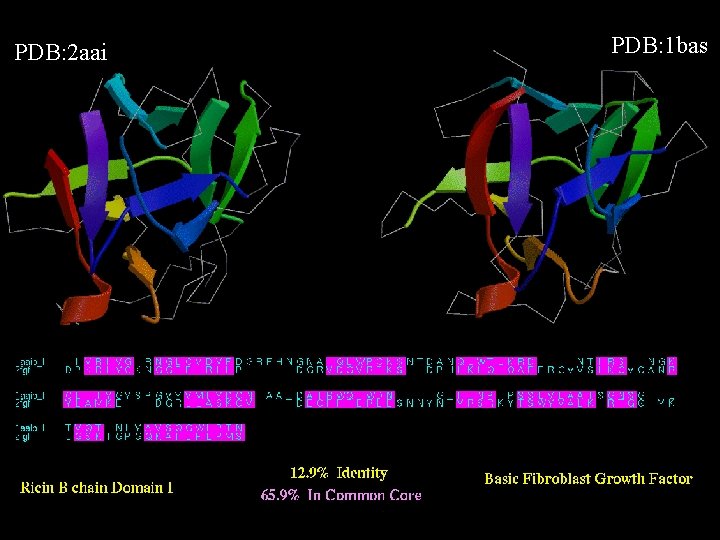
PDB: 2 aai PDB: 1 bas
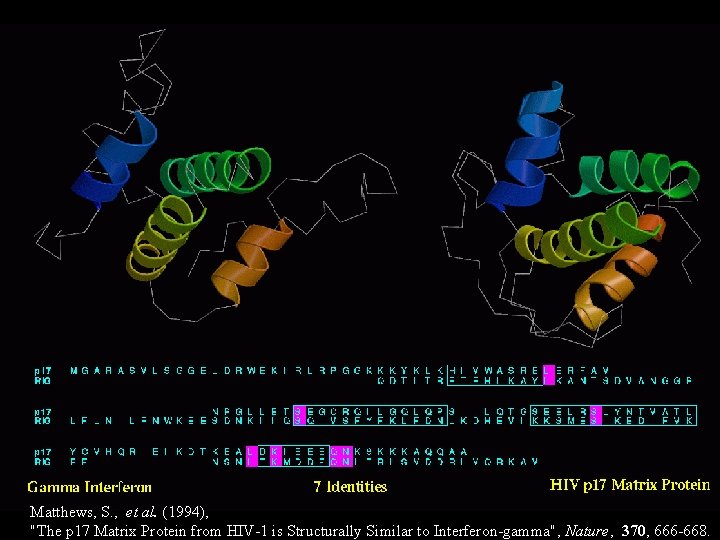
Matthews, S. , et al. (1994), "The p 17 Matrix Protein from HIV-1 is Structurally Similar to Interferon-gamma", Nature, 370, 666 -668.
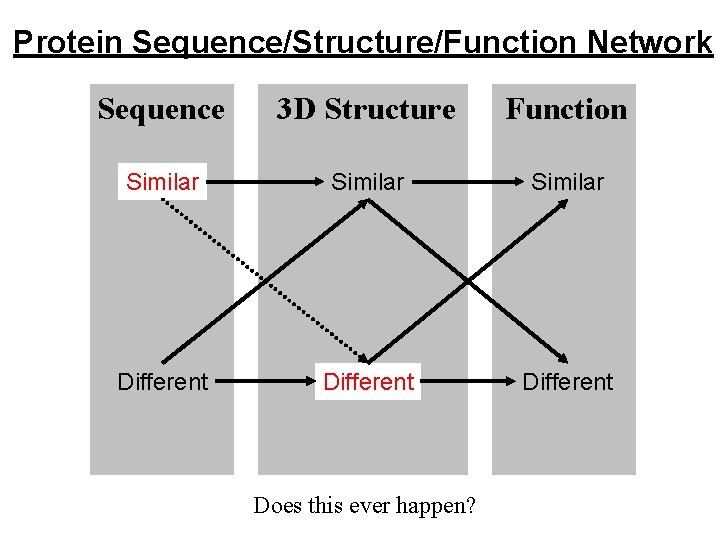
Protein Sequence/Structure/Function Network Sequence 3 D Structure Function Similar Different Does this ever happen?
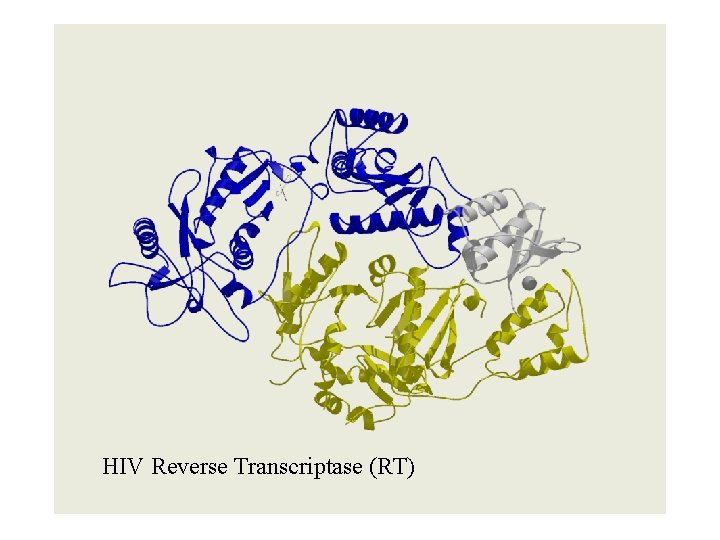
HIV Reverse Transcriptase (RT)
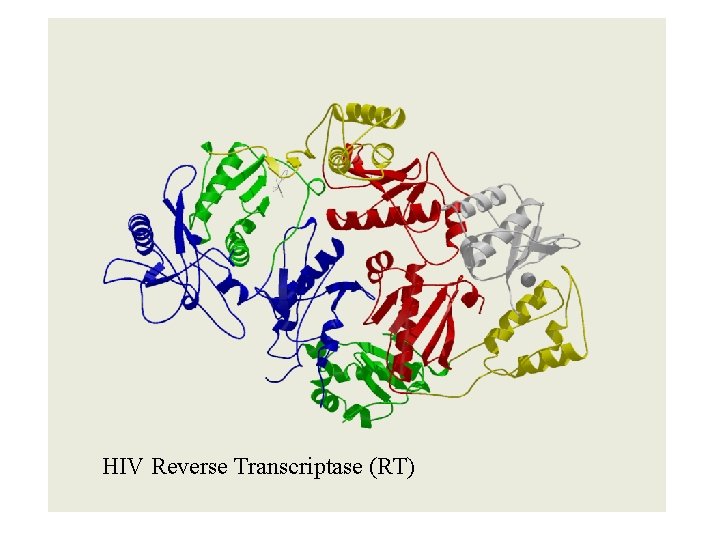
HIV Reverse Transcriptase (RT)
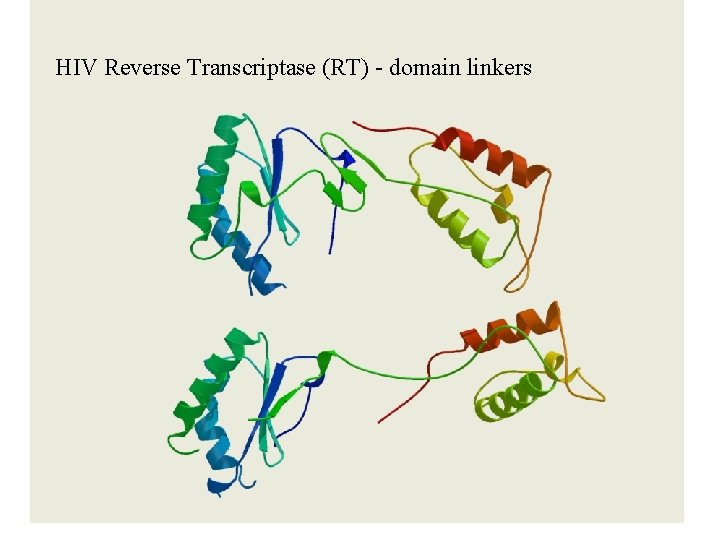
HIV Reverse Transcriptase (RT) - domain linkers

Protein Sequence and Structural Similarity
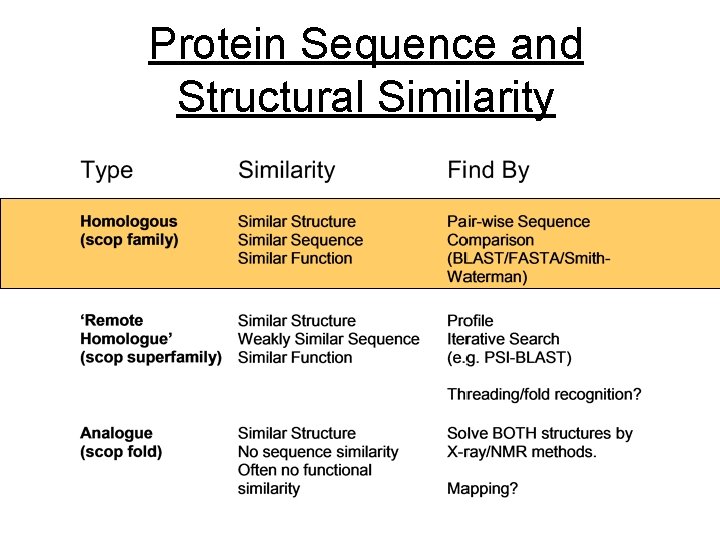
Protein Sequence and Structural Similarity

Barton, G. J. et al, (1992), "Human Platelet Derived Endothelial Cell Growth Factor is Homologous to E. coli Thymidine Phosphorylase", Prot. Sci. , 1, 688 -690.
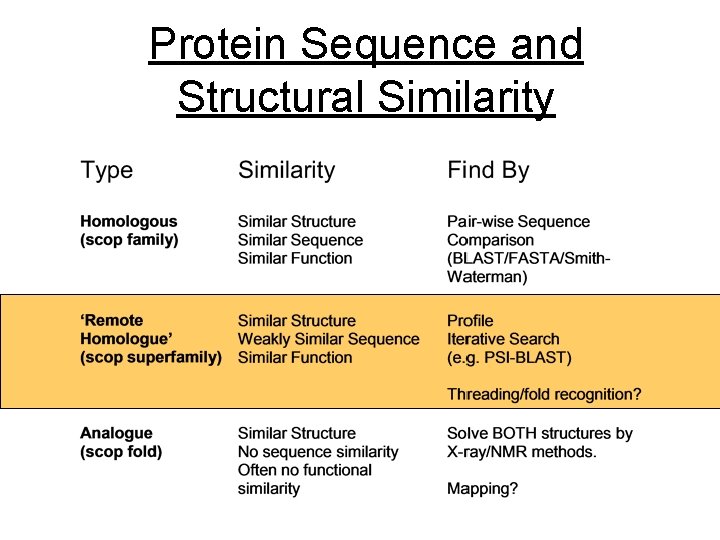
Protein Sequence and Structural Similarity
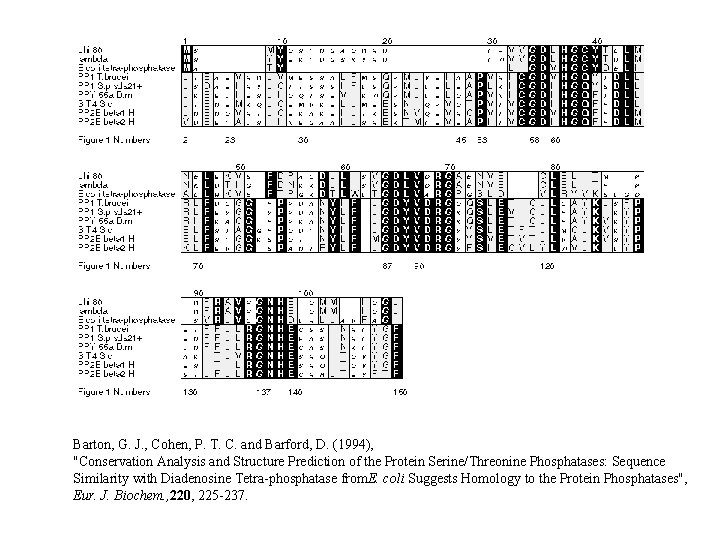
Barton, G. J. , Cohen, P. T. C. and Barford, D. (1994), "Conservation Analysis and Structure Prediction of the Protein Serine/Threonine Phosphatases: Sequence Similarity with Diadenosine Tetra-phosphatase from. E. coli Suggests Homology to the Protein Phosphatases", Eur. J. Biochem. , 220, 225 -237.
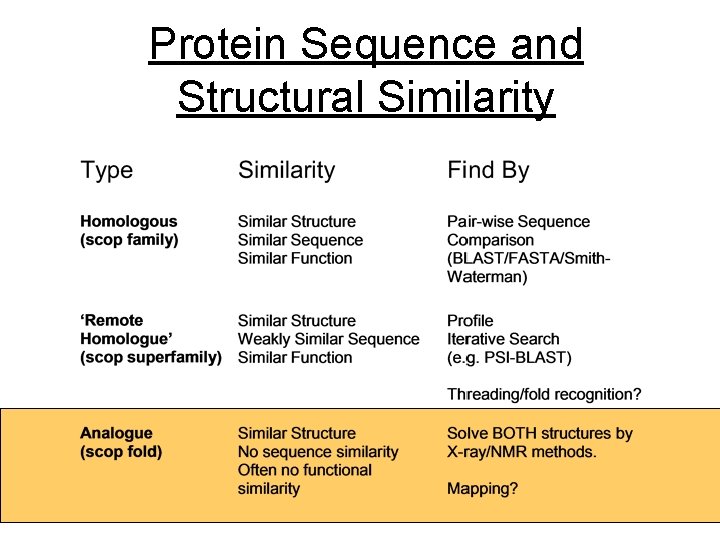
Protein Sequence and Structural Similarity
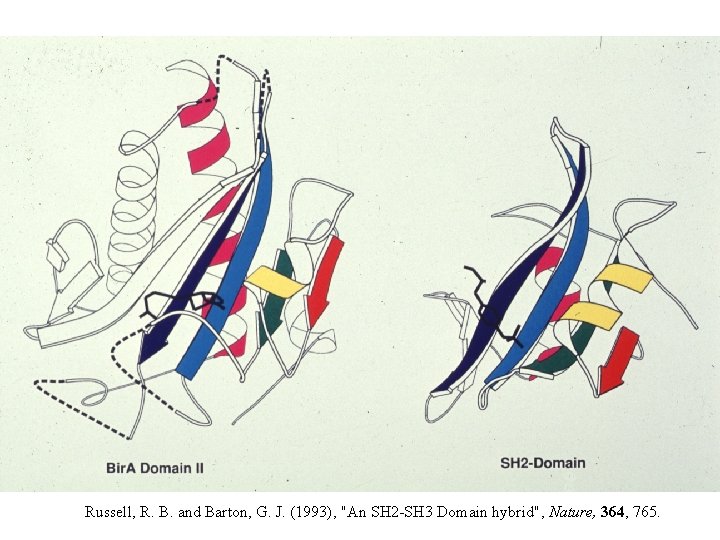
Russell, R. B. and Barton, G. J. (1993), "An SH 2 -SH 3 Domain hybrid", Nature, 364, 765.

Reading material for this lecture: This lecture itself. pdf’s for “Barton” papers: www. compbio. dundee. ac. uk/ftp/pdf/ Database statistics: http: //www. ebi. ac. uk/embl/ Structure of the amino-terminal domain of Cbl complexed to its binding site on ZAP-70 kinase Wuyi Meng, Sansana Sawasdikosol, Steven J. Burakoff, Michael J. Eck Nature 398, 84 - 90 (04 March 1999) (available on-line at www. nature. com - search for ZAP-70 kinase - republished in December on-line) Protein recognition: An SH 2 domain in disguise John Kuriyan, James E. Darnell Nature 398, 22 - 25 (04 March 1999) (news and views article for above paper) Russell, R. B. and Barton, G. J. (1993), "An SH 2 -SH 3 Domain hybrid", Nature, 364, 765. Matthews, S. , et al. (1994), "The p 17 Matrix Protein from HIV-1 is Structurally Similar to Interferon-gamma", Nature, 370, 666 -668. Barton, G. J. , Cohen, P. T. C. and Barford, D. (1994), "Conservation Analysis and Structure Prediction of the Protein Serine/Threonine Phosphatases: Sequence Similarity with Diadenosine Tetra-phosphatase from. E. coli Suggests Homology to the Protein Phosphatases", Eur. J. Biochem. , 220, 225 -237.

The end of Lecture 1 Lecture 2 will be on sequence comparison methods.
- Slides: 58