Bioinformatics an overview MingJing Hwang Institute of Biomedical

Bioinformatics: an overview Ming-Jing Hwang (黃明經) Institute of Biomedical Sciences Academia Sinica http: //gln. ibms. sinica. edu. tw/

The human genome project Year 2001

Promises “More will happen in biology in the next 10 years than in the past 50” (Craig Venter, Celera Genomics). v “We should be able to uncover the major hereditary contributions to common illnesses like diabetes and mental illness, probably in the next three to five years” (Francis Collins, head of HGP). v
Genetics & Genomics From DNA to population Source: gsk

What makes us human ? q The difference between you & chimp is ~1. 24% q The difference between you and Maggie is ~0. 1%

Hunting for disease genes Source: gsk
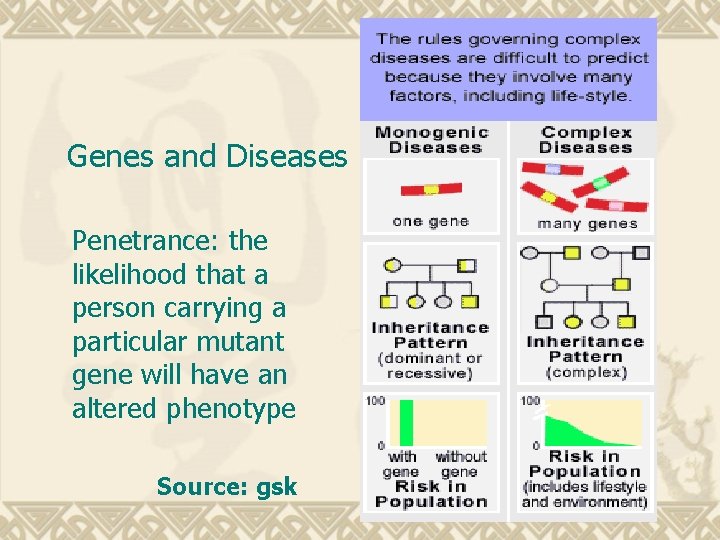
Genes and Diseases Penetrance: the likelihood that a person carrying a particular mutant gene will have an altered phenotype Source: gsk

phenotype and genotype v Many different genotypes can have same phenotype v Many genotypes do not change the phenotype v One phenotype could be due to many different genotypes -- statistical genetics

The common variant – common disease (CV-CD) hypothesis It is believed that most polygenic contributions to disease susceptibility will arise from variants that are relatively common in the susceptible population.

Genetic variations v SNP constitute 90% of human genetic variations v Other forms of variations include insertion, deletion, and differences in the copy number of tandem repeats or large genomic segments, etc.
Three phases of human genome sequencing v The genome map (draft in 2001, “finished” in 2003) v The SNP map (TSC, 2001) v The haplotype map (Hap. Map, 2005)
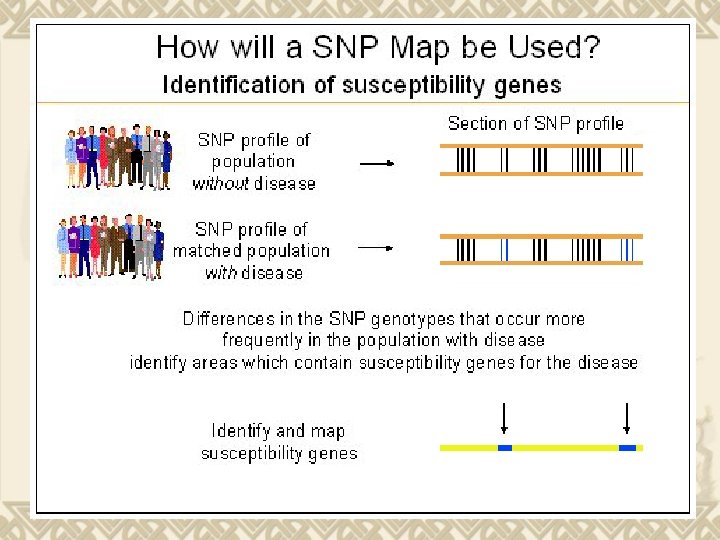
Source: gsk
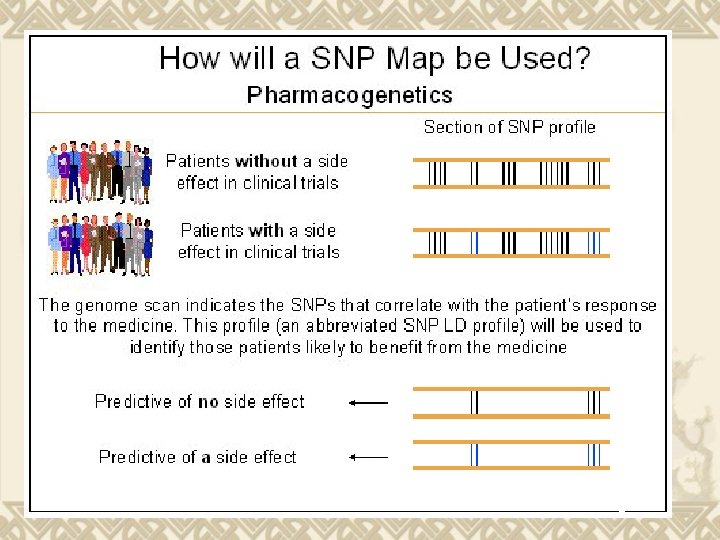
Source: gsk

pharmacogenomics 8/2 (二) 10: 00 pm on PTS (CH 13)
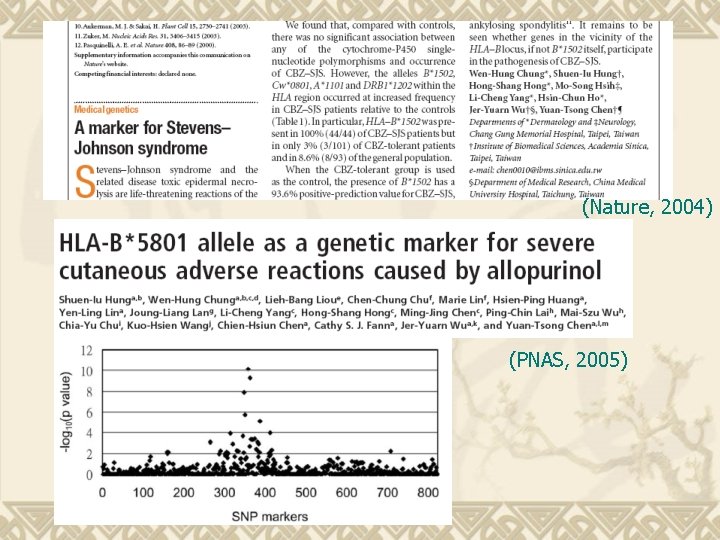
(Nature, 2004) (PNAS, 2005)

Common SNPs Kruglyak & Nickerson, 2001
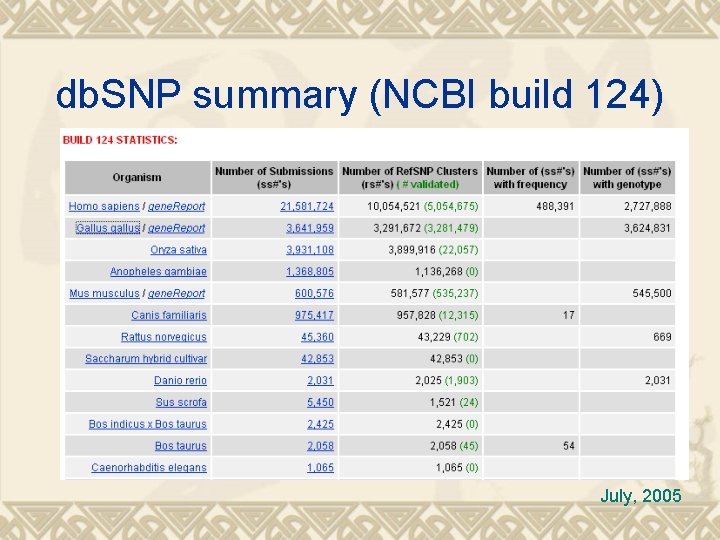
db. SNP summary (NCBI build 124) July, 2005
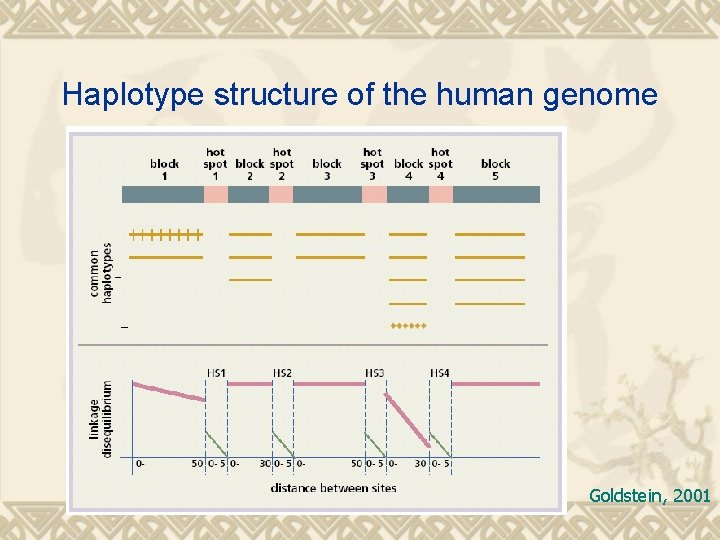
Haplotype structure of the human genome Goldstein, 2001

Rationale of Hap. Map In a given population, 55 percent of people may have one version of a haplotype, 30 percent may have another, 8 percent may have a third, and the rest may have a variety of less common haplotypes. The International Hap. Map Project is identifying these common haplotypes in four populations from different parts of the world. It also is identifying "tag" SNPs that uniquely identify these haplotypes. By testing an individual's tag SNPs (a process known as genotyping), researchers will be able to identify the collection of haplotypes in a person's DNA. The number of tag SNPs that contain most of the information about the patterns of genetic variation is estimated to be about 300, 000 to 600, 000, which is far fewer than the 10 million common SNPs.

Beyond genome

Chemical genomics


Bioinformatics has many subdisciplines v Genome Informatics (DNA sequence) v Transcriptome Informatics (expression) v Proteome informatics (ID, post-transl. mod. ) v Protein Informatics (protein struct. /funct. ) v Evolutionary Informatics v Biomedical Informatics (human disease) …

Briefings in bioinformatics (Mar 2005) v v v The many faces of sequence alignment (Altman ) Bioinformatics analysis of alternative splicing (Lee & Wang) Putting microarray in a context: Integrated analysis of diverse biological data (Troyanskaya) Bioinformatics approaches and resources for single nucleotide polymorphism functional analysis (Mooney) A survey of current work in biomedical text mining (Cohen & Hersh) Current efforts in the analysis of RNAi and RNAi target genes (Bengert & Dandekar)

Sequence alignment v The problem is still not solved. Sequence alignment methodology and tool development continue to grow, indicating that the alignment problem is still not solved. How can that be, after nearly forty years of research and literally hundreds of available tools? v Why should alignments remain an open problem? It is not a single problem but rather a collection of many quite diverse questions that all have in common the search for sequence similarity The exponential expansion of biological sequence databases – faster than Moore’s law Batzoglou, 2005

Sequence alignment challenges v Sensitivity and specificity v Speed v Evaluation v Low similarity v Rearrangements v Orthology detection v Multiple (genome) alignments
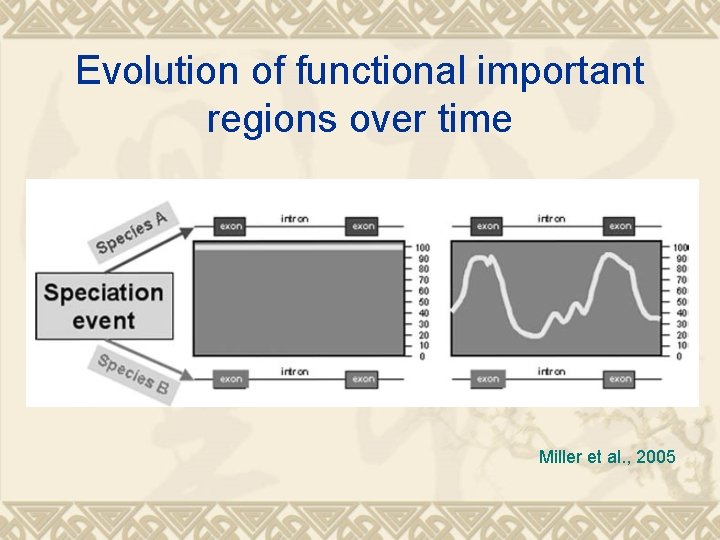
Evolution of functional important regions over time Miller et al. , 2005
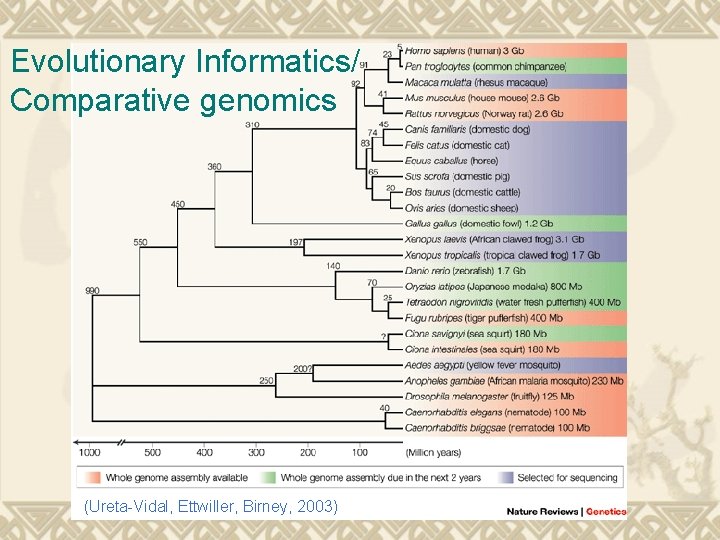
Evolutionary Informatics/ Comparative genomics (Ureta-Vidal, Ettwiller, Birney, 2003)
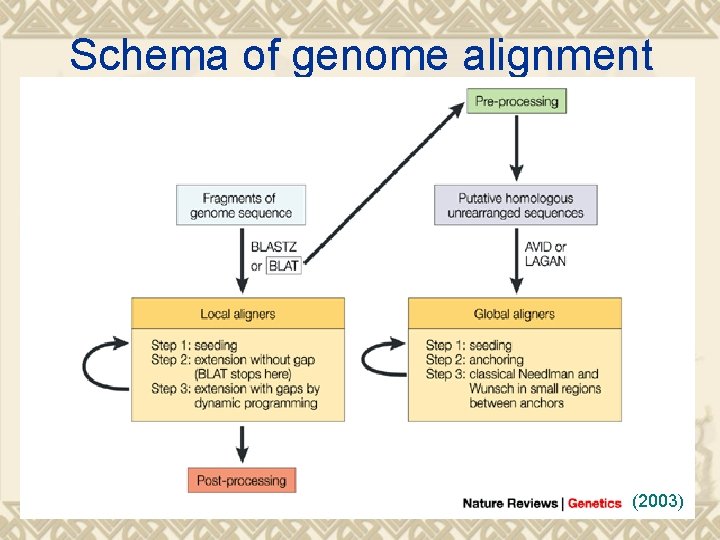
Schema of genome alignment (2003)

Genome alignment recent reviews v v v An Applications-Focused Review of Comparative Genomics Tools: Capabilities, Limitations and Future Challenges (Chain et al. Briefings in bioinformatics, 2003) The many faces of sequence alignment (Batzoglou, Briefings in bioinformatics, 2005) Comparative genomics (Miller et al. Annu. Rev. Genomics Hum. Genet. 2004)
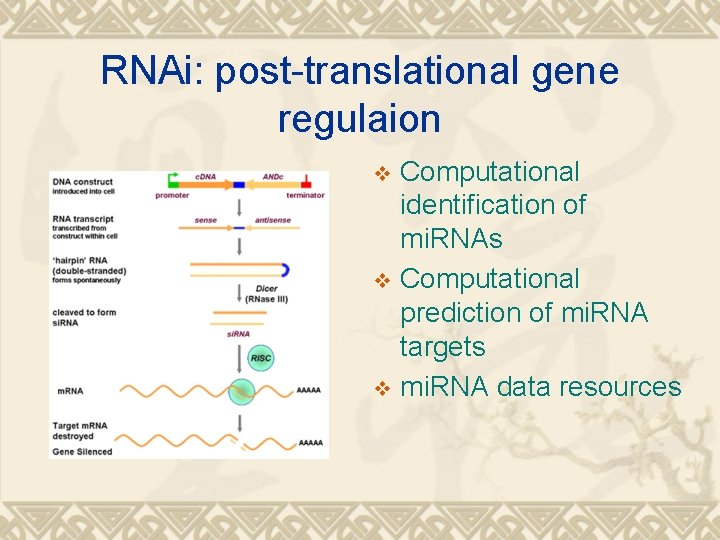
RNAi: post-translational gene regulaion Computational identification of mi. RNAs v Computational prediction of mi. RNA targets v mi. RNA data resources v
Transcriptomics: tools for understanding the body plan

Microarray & Integrated analysis Troyanskaya, 2005
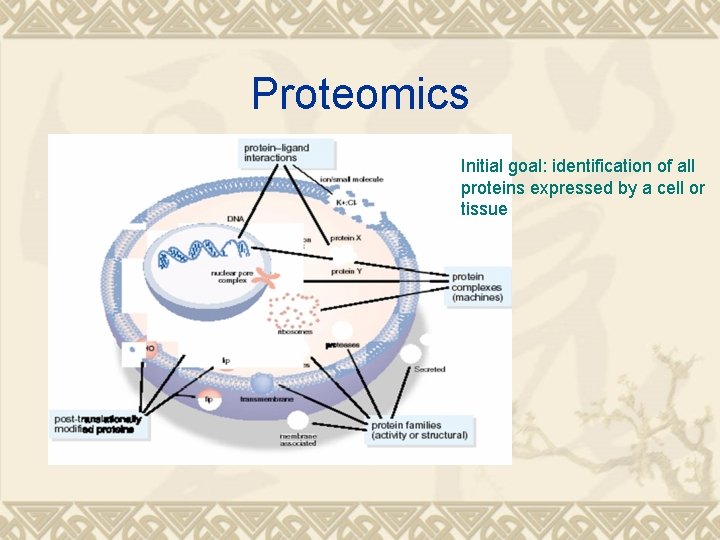
Proteomics Initial goal: identification of all proteins expressed by a cell or tissue
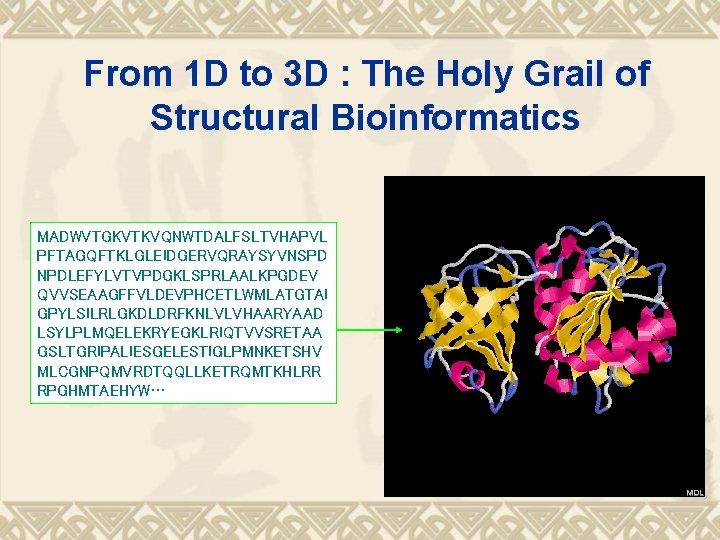
From 1 D to 3 D : The Holy Grail of Structural Bioinformatics MADWVTGKVTKVQNWTDALFSLTVHAPVL PFTAGQFTKLGLEIDGERVQRAYSYVNSPD NPDLEFYLVTVPDGKLSPRLAALKPGDEV QVVSEAAGFFVLDEVPHCETLWMLATGTAI GPYLSILRLGKDLDRFKNLVLVHAARYAAD LSYLPLMQELEKRYEGKLRIQTVVSRETAA GSLTGRIPALIESGELESTIGLPMNKETSHV MLCGNPQMVRDTQQLLKETRQMTKHLRR RPGHMTAEHYW…

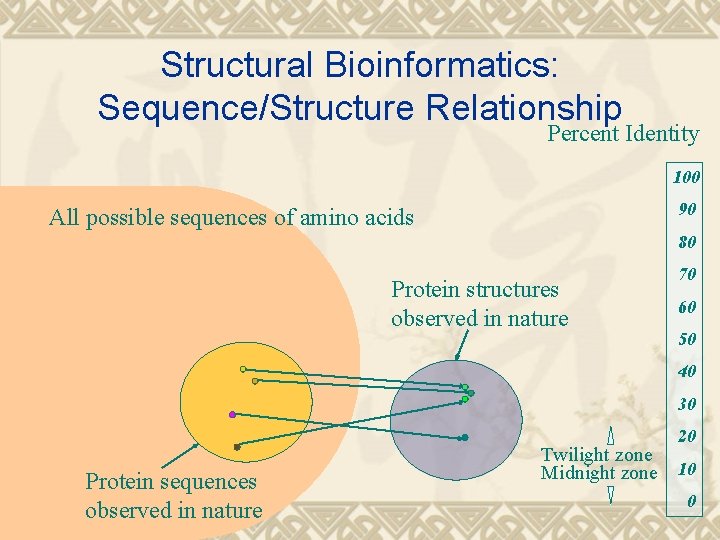
Structural Bioinformatics: Sequence/Structure Relationship Percent Identity 100 90 All possible sequences of amino acids 80 Protein structures observed in nature 70 60 50 40 30 20 Protein sequences observed in nature Twilight zone Midnight zone 10 0
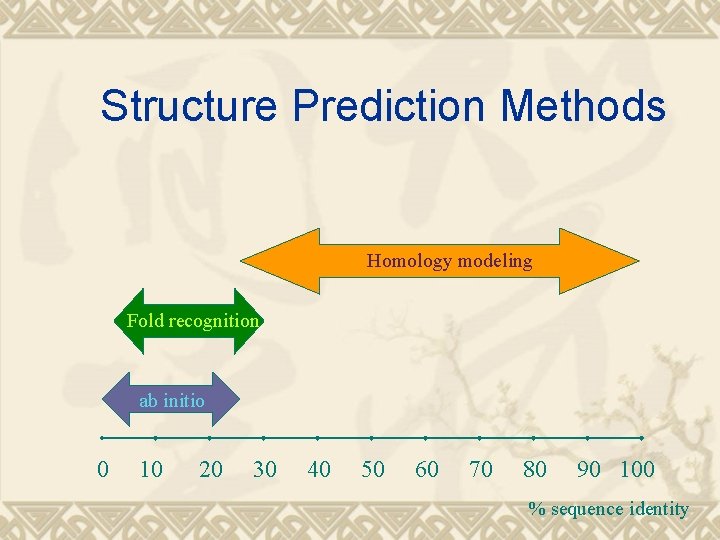
Structure Prediction Methods Homology modeling Fold recognition ab initio 0 10 20 30 40 50 60 70 80 90 100 % sequence identity
CASP Experiments
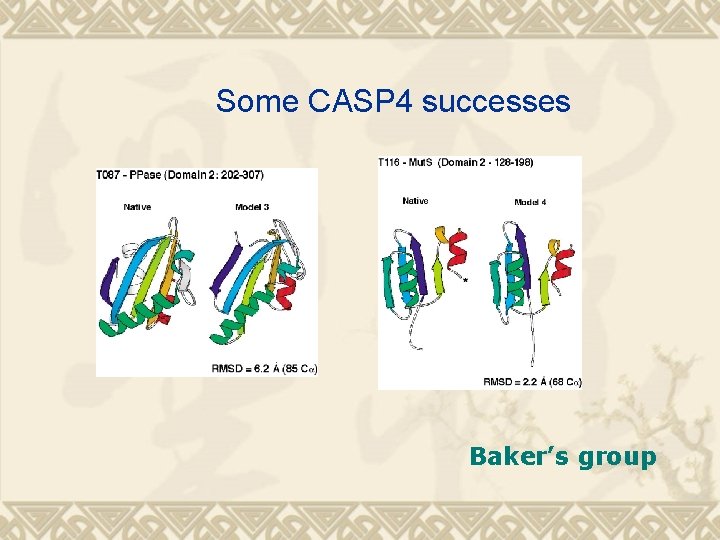
Some CASP 4 successes Baker’s group

3 D to 1 D? Science 2003
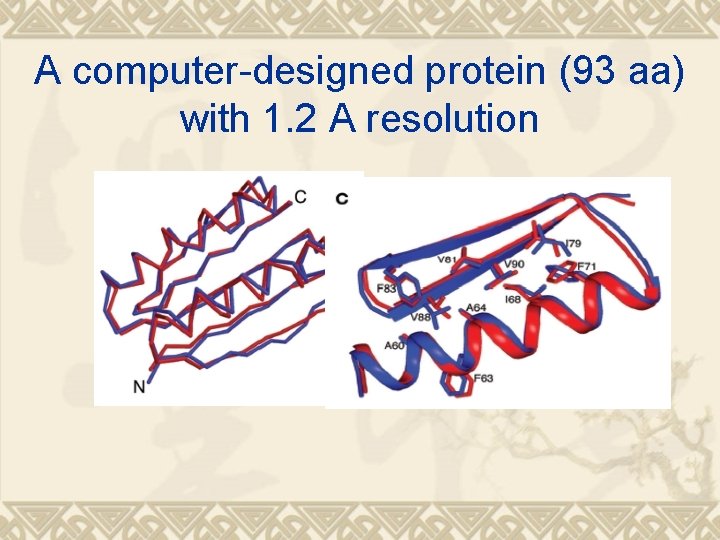
A computer-designed protein (93 aa) with 1. 2 A resolution
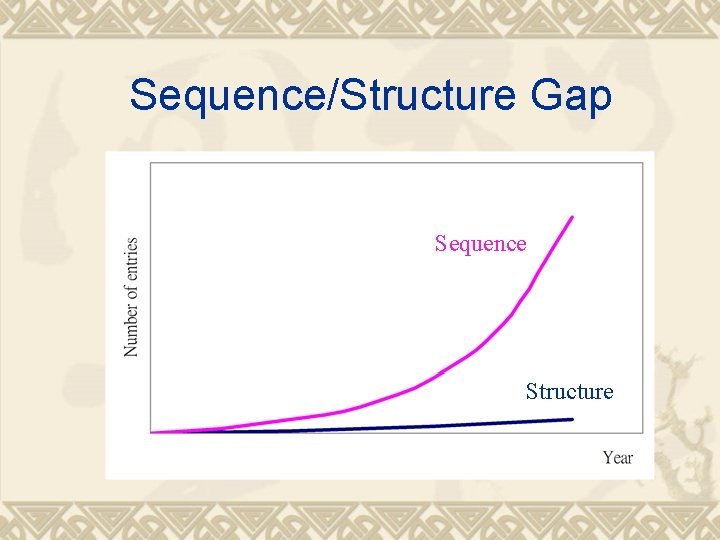
Sequence/Structure Gap Sequence Structure
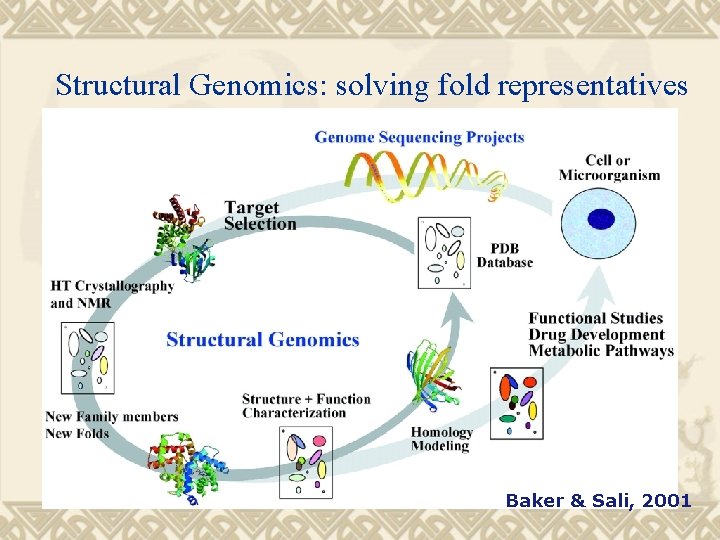
Structural Genomics: solving fold representatives Baker & Sali, 2001

Structural Genomics: overview v v v When: 1997 by Barry Honig, Wayne Henderickson and colleagues in a DOE’s Advanced Photon Source (APS) proposal Goals: 10, 000 structures (100 -200 str/center/yr) – each representing a protein family – in 5(10) years Enabling factors: genome sequences, technology advancement (synchrotron & MAD, etc. ) Cost: reducing current ~US$200, 000/str to $10, 000/str (est. $1. 5 -5 billion US) Players: academic & industry
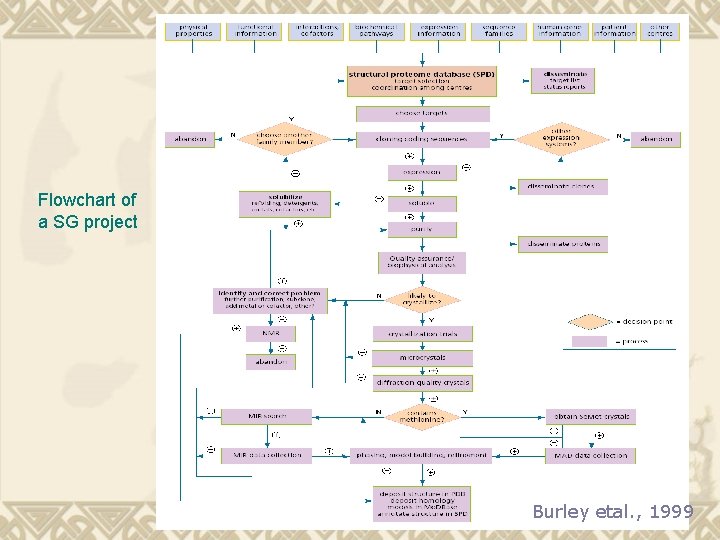
Flowchart of a SG project Burley etal. , 1999

PSI phase I (pilot) centers v v v v v Berkeley Structural Genomics Center focused on two bacterial species with extremely small genomes to study proteins essential for independent life. Principal investigator: Sung-Hou Kim, Lawrence Berkeley National Laboratory Center for Eukaryotic Structural Genomics, based in Wisconsin, focused on protein production, characterization, and structure determination from Arabidopsis thaliana, a plant that is frequently used in laboratory research and that has many genes in common with humans and animals. Principal investigator: John Markley, University of Wisconsin, Madison Joint Center for Structural Genomics, based in California, focused on novel structures from thermophilic microorganisms and on human proteins thought to be involved in cell signaling. Principal investigator: Ian Wilson, The Scripps Research Institute Midwest Center for Structural Genomics, based in Illinois, selected bacterial targets related to disease and proteins from all three kingdoms of life. The emphasis was on previously unknown folds and on proteins from disease-causing organisms. Principal investigator: Andrzej Joachimiak, Argonne National Laboratory New York Structural Genomics Research Consortium solved protein structures for disease-related proteins from eukaryotes and bacteria. Principal investigator: Stephen K. Burley, Structural Genomi. X, Inc. Northeast Structural Genomics Consortium, based in New Jersey, focused on target proteins from various model organisms, including the fruit fly, yeast, and roundworm. It used both X-ray crystallography and NMR spectroscopy. Principal investigator: Gaetano Montelione, Rutgers University The Southeast Collaboratory for Structural Genomics, based in Georgia, determined structures from the prokaryotic model organism, Pyrococcus furiosus, and the eukaryotic model organism C. elegans, as well as some human proteins. Principal investigator: Bi-Cheng Wang, University of Georgia Structural Genomics of Pathogenic Protozoa Consortium, based in Washington, solved protein structures from organisms known as protozoans, many species of which cause deadly diseases such as sleeping sickness, malaria, and Chagas' disease. Principal investigator: Wim G. J. Hol, University of Washington TB Structural Genomics Consortium, based in New Mexico, analyzed protein structures from Mycobacterium tuberculosis. Principal investigator: Thomas Terwilliger, Los Alamos National Laboratory

PSI Pilot Phase Facts at a Glance v v v Goal: To develop new approaches and tools needed to streamline and automate the steps of protein structure determination, and to incorporate those methods into high-throughput pipelines that use DNA sequence information to generate three-dimensional protein structure models Project period: September 2000 to June 2005 Funding: $270 million (funded largely by the National Institute of General Medical Sciences, with additional support from the National Institute of Allergy and Infectious Diseases) Number of Centers: 9 (6 survived to phase II) Solved protein structures: More than 1, 100 Unique structures solved (structures sharing less than 30 percent of their sequence with other known proteins): More than 700
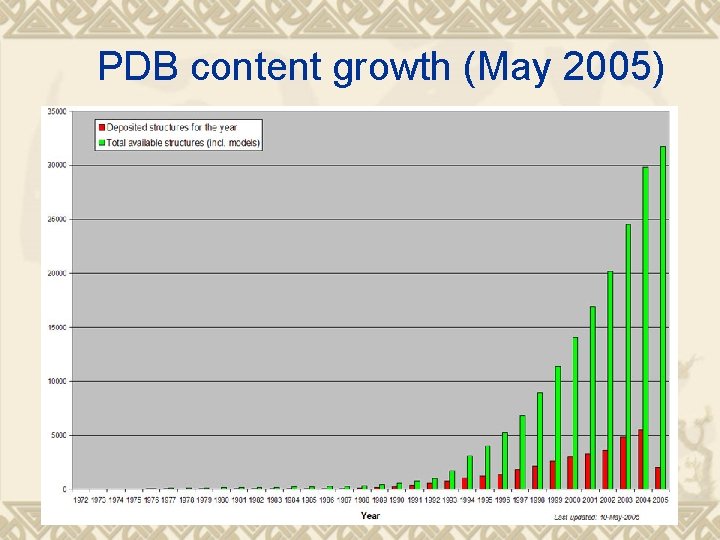
PDB content growth (May 2005)
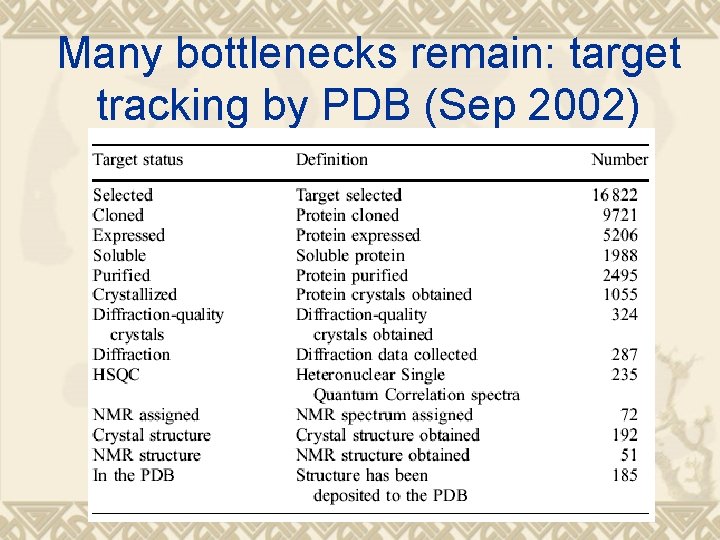
Many bottlenecks remain: target tracking by PDB (Sep 2002)

Current (phase II) PSI centers
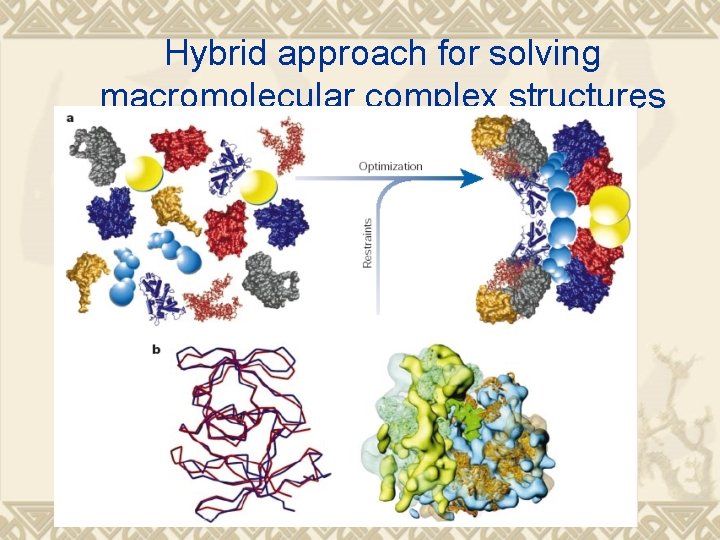
Hybrid approach for solving macromolecular complex structures
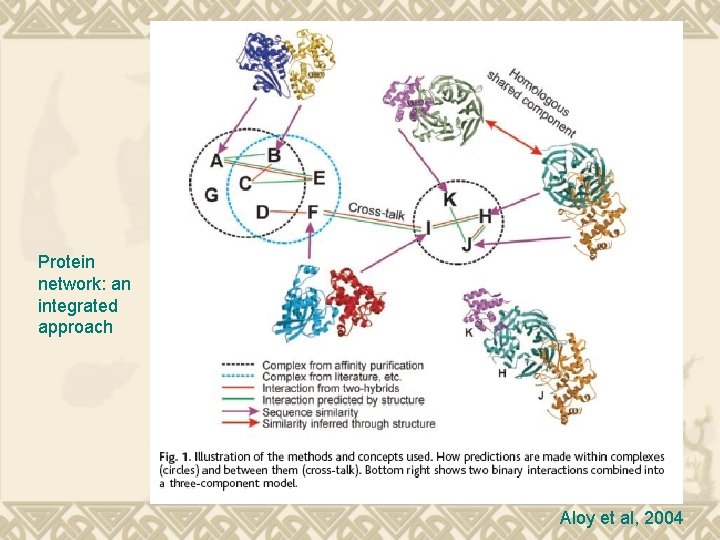
Protein network: an integrated approach Aloy et al, 2004
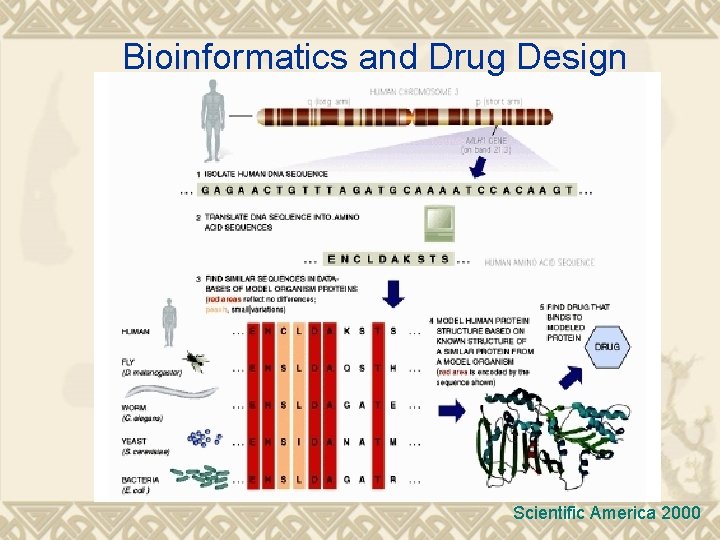
Bioinformatics and Drug Design Scientific America 2000

Yeast protein interaction network Nat Rev Genet. 2004
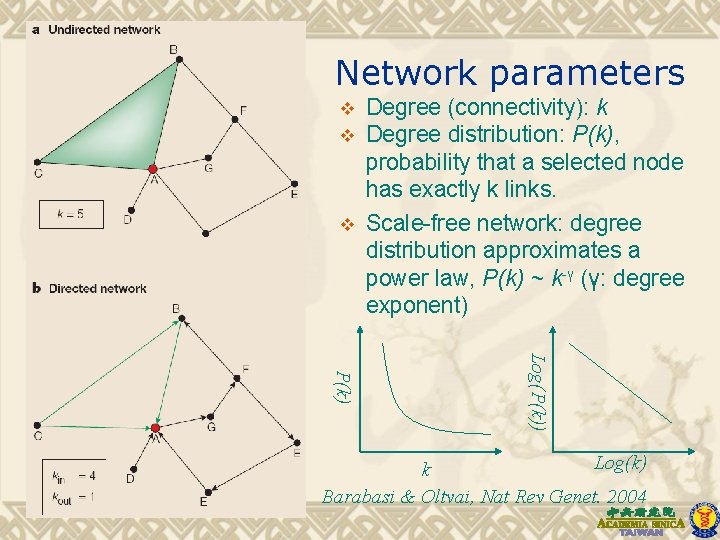
Network parameters v v v Degree (connectivity): k Degree distribution: P(k), probability that a selected node has exactly k links. Scale-free network: degree distribution approximates a power law, P(k) ~ k-γ (γ: degree exponent) Log(P(k)) P(k) Log(k) k Barabasi & Oltvai, Nat Rev Genet. 2004
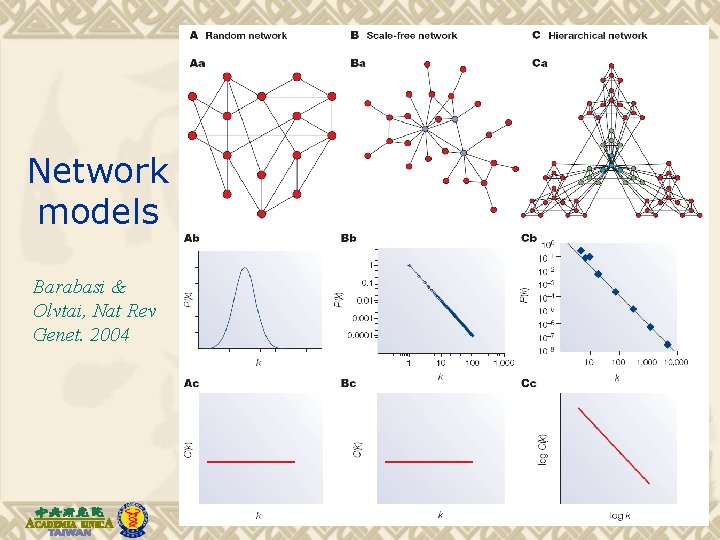
Network models Barabasi & Olvtai, Nat Rev Genet. 2004
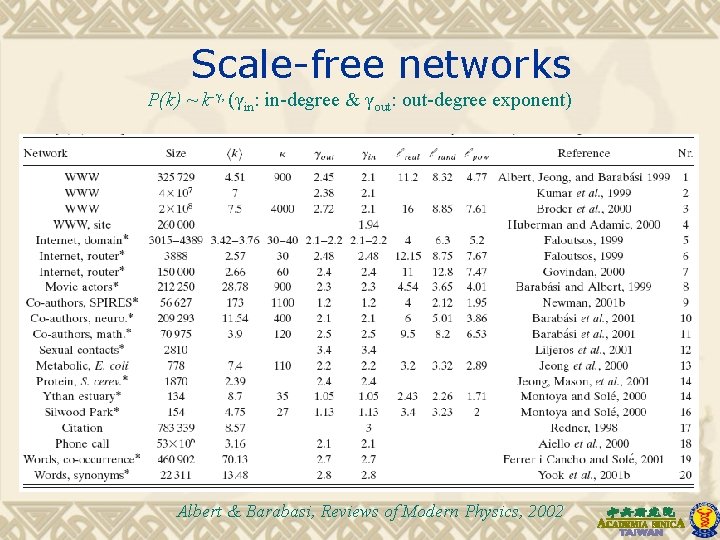
Scale-free networks P(k) ~ k-γ, (γin: in-degree & γout: out-degree exponent) Albert & Barabasi, Reviews of Modern Physics, 2002

Challenges in network biology v Network databases v Information integration v Organization characteristics and principles v Design rules v Evolution mechanisms v Validation
Neuroinformatics: neuroscience & bioinformatics The human brain project -UC Davis http: //nir. cs. ucdavis. edu/index. jsp http: //ncmir. ucsd. edu/NCDB/

Bioinformatics Journals Bioinformatics v Nucleic Acids Research v BMC Bioinformatics v Briefings in Bioinformatics v Proteins v J. Mol. Biol. v… v PNAS v PLo. S computational biology v Genome Research v… v

Scope of bioinformatics v v v v v Genome analysis Sequence analysis Phylogenetics Structural bioinformatics Gene expression Genetic and population analysis Systems biology Data and text mining Databases and ontologies

Sample articles of a recent issue v v v v v Exon–domain correlation and its corollaries Functional annotation from predicted protein interaction networks HYPROSP II-A knowledge-based hybrid method for protein secondary structure prediction based on local prediction confidence Comparative interactomics analysis of protein family interaction networks using PSIMAP (protein structural interactome map) Semi-supervised protein classification using cluster kernels A new progressive-iterative algorithm for multiple structure alignment Practical FDR-based sample size calculations in microarray experiments Mining genetic epidemiology data with Bayesian networks I: Bayesian networks and example application (plasma apo. E levels) Inferring protein–protein interactions through high-throughput interaction data from diverse organisms A latent variable model for chemogenomic profiling

NAR Database issue (Jan. 2005) Categories # 1. Nucleotide Sequence Databases 53 2. RNA Sequence Databases 34 3. Protein Sequence Databases 105 4. Structure Databases 64 5. Genomic Databases (non-human) 134 6. Metabolic Enzyme & Pathways; Signals Pathways 36 7. Human & Other Vertebrate Genomes 64 8. Human Genes & Diseases 69 9. Microarray Data & Other Gene Expression Databases 42 10. Proteomics Resources 7 11. Other Molecular Biology Databases 17 12. Organelle Databases 18 13. Plant Databases 48 14. Immunological Databases 20 Total 711 http: //nar. oupjournals. org/cgi/content/full/33/suppl_1/D 5/TBL 1

NAR Web Server Issue (July 2005) Year of publication 2004 2005 Total # 129 (137) 166 295

Computer Related (2) Bio-* Programming Tools (1) Statistics (1) DNA (57) Annotations (9) Gene Prediction (4) Mapping and Assembly (1) Phylogeny Reconstruction (4) Sequence Feature Detection (16) Sequence Polymorphisms (8) Sequence Retrieval and Submission (3) Tools For the Bench (12) Education (1) Directories and Portals (1) Expression (48) c. DNA, EST, SAGE (8) Gene Regulation (22) Microarrays (16) Splicing (2) Human Genome (13) Annotations (3) Health and Disease (3) Other Resources (2) Sequence Polymorphisms (5) Model Organisms (9) Microbes (4) Mouse and Rat (2) Plants (1) Yeast (2) Other Molecules (2) Carbohydrates (2) Protein (131) 2 -D Structure Prediction (10) 3 -D Structure Prediction, Comparison (34) 3 -D Structure Retrieval, Viewing (6) Biochemical Features (8) Domains and Motifs (25) Function (10) Interactions, Pathways, Enzymes (13) Localization and Targeting (7) Phylogeny Reconstruction (5) Proteomics (2) Sequence Features (6) Sequence Retrieval (5) RNA (15) Functional RNAs (5) Motifs (3) Sequence Retrieval (2) Structure Prediction, Visualization, and Design (5) Sequence Comparison (29) Alignment Editing and Visualization (2) Analysis of Aligned Sequences (12) Comparative Genomics (7) Multiple Sequence Alignments (2) Pairwise Sequence Alignments (2) Similarity Searching (4) Literature (5) Search Tools (3) Text Mining (2) http: //bioinformatics. ubc. ca/resources/links_directory/narweb 2005/
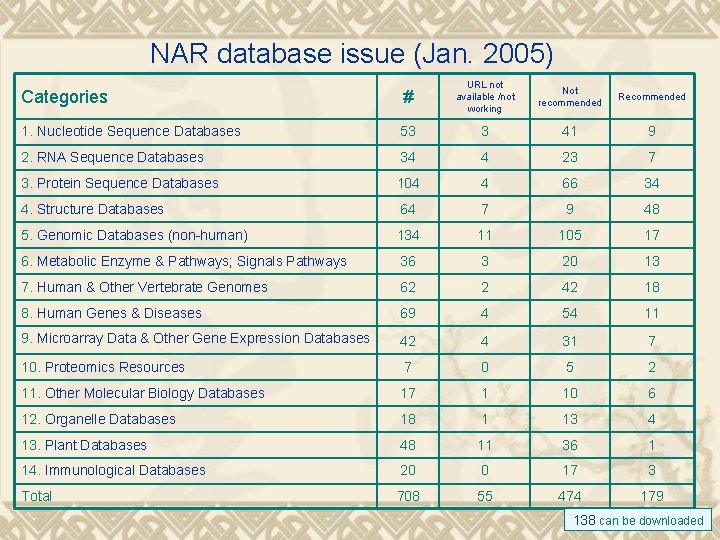
NAR database issue (Jan. 2005) Categories # URL not available /not working 1. Nucleotide Sequence Databases 53 3 41 9 2. RNA Sequence Databases 34 4 23 7 3. Protein Sequence Databases 104 4 66 34 4. Structure Databases 64 7 9 48 5. Genomic Databases (non-human) 134 11 105 17 6. Metabolic Enzyme & Pathways; Signals Pathways 36 3 20 13 7. Human & Other Vertebrate Genomes 62 2 42 18 8. Human Genes & Diseases 69 4 54 11 9. Microarray Data & Other Gene Expression Databases 42 4 31 7 10. Proteomics Resources 7 0 5 2 11. Other Molecular Biology Databases 17 1 10 6 12. Organelle Databases 18 1 13 4 13. Plant Databases 48 11 36 1 14. Immunological Databases 20 0 17 3 Total 708 55 474 179 Not recommended Recommended 138 can be downloaded

An example of our curation

Two keys in bioinformatics research v Solve a significant biological question v Develop a must-use application tool
- Slides: 70