Bioinformatics A Summary seminar with many hints for

Bioinformatics A Summary seminar (with many hints for exam questions)

1) Introduction 1. 2. 3. 4. 5. 6. The question: Transfer of information The tools: MRS, BLAST, Clustal, Databases, Swiss. Prot Amino acid knowledge: understand secondary structure Secondary structure -> protein structure Protein structure helps make alignments Alignments allow for transfer of information

Bioinformatics Necessary evil, panacea, or just a useful tool? With a month in the lab you can easily prevent having to sit an hour in front of the computer. Nothing is impossible for a biologist who doesn’t have to discover it him/her-self.
Bio + informatica
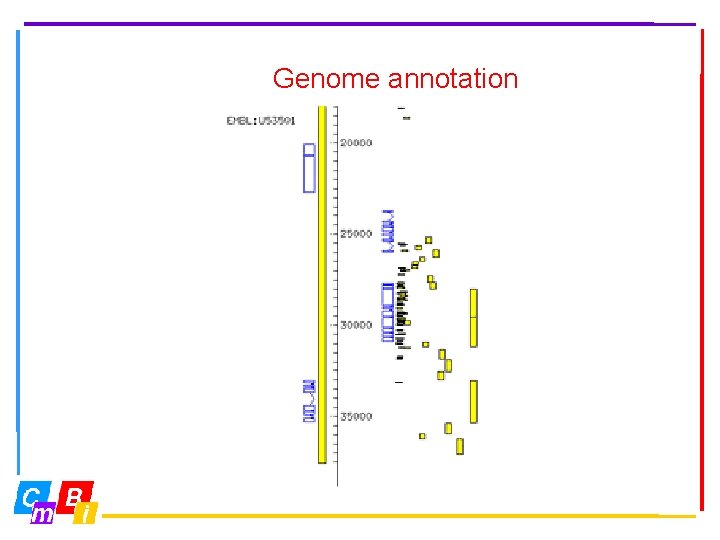
Genome annotation

Bioinformatics and medicines One day we know everything about all human (and flu) proteins and then can we start to ‘calculate’ flu-medicines.
Drug Design
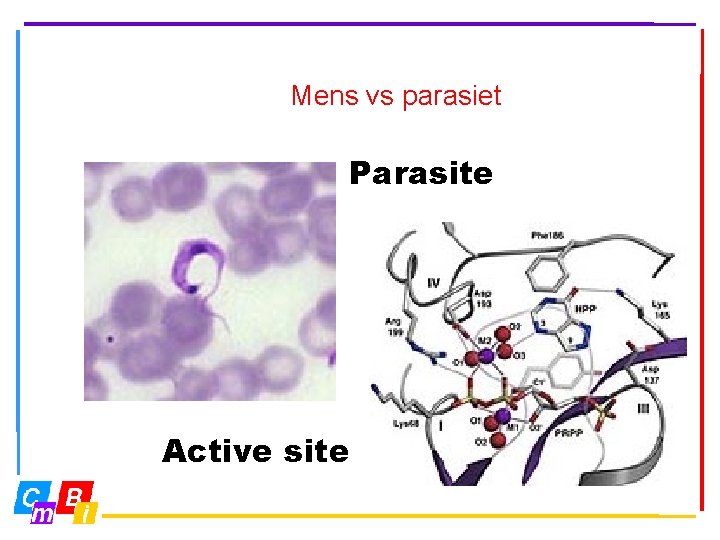
Mens vs parasiet Parasite Active site
H 1 N 1 / H 5 N 1

2) Tools MRS, kind of bio. Google BLAST to find homologs Swiss. Prot: protein sequences PDB: macromolecular structures EMBL: nucleotide sequences OMIM: genetic disorders Pro. Site: motifs (e. g. {P} [ST] {P} N )

Biological databases (1) Primary databases contain biomolecular sequences or structures (experimental data!) and associated annotation information Sequences Nucleic acid sequences Protein sequences EMBL, Genbank, DDBJ Swiss. Prot, tr. EMBL, Uni. Prot Structures Protein Structures PDB Structures of small compounds CSD Genomes Ensembl UCSC ©CMBI 2010

Databases Data must be in a certain format for software to recognize Every database can have its own format but some data elements are essential for every database: 1. Unique identifier, or accession code 2. Name of depositor 3. Literature references 4. Deposition date 5. The real data Nomenclature: • Database entry or database record • Database fields ©CMBI 2015

Swiss. Prot database • Database of protein sequences • >500. 000 sequence entries • Swiss. Prot is manually annotated and reviewed, thus of high quality, but never complete; it contains many feature descriptions and many hyperlinks to other databases; a bioinformatician always looks in Swiss. Prot first… • Obligatory deposit of in Swiss. Prot before publication • Swiss. Prot is part of Uni. Prot • The other main part of Uni. Prot is Trembl (translated EMBL). Trembl is automatically annotated and is not reviewed. ©CMBI 2015

Part III: Sequence Retrieval with MRS Google Thé best generic search and retrieval system Google searches everywhere for everything MRS Maarten’s Retrieval System (http: //mrs. cmbi. ru. nl ) MRS searches in selected data environments MRS is the Google of the biological database world Search engine (like Google) • Input/Query = word(s) • Output = entry/entries from database Other programs exist: Entrez, SRS, . . ©CMBI 2009

Transfer of information to corresponding residues BLAST finds two database hits that are annotated to have a phosphorylated serine. DRT-GHNIPLMSTRK-TYHIHIENASEERTIKLLMN DRR-GTTINLMTTKR-TYADELENASEDRTLLLNMN AEPIYYHL---LTKRETYHIHIENASEEKIIKIVVN “this serine is phorylated in a known protein from the database, so in my protein the corresponding serine is likely to be phosphorylated too”.
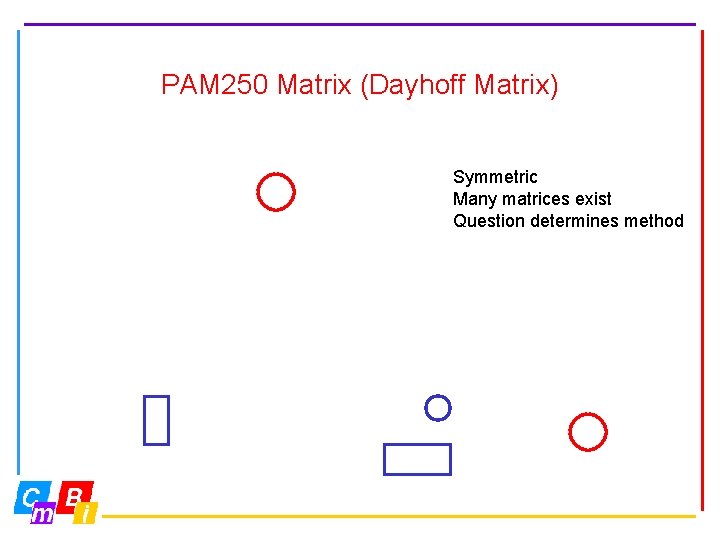
PAM 250 Matrix (Dayhoff Matrix) Symmetric Many matrices exist Question determines method

Amino Acid substitutions, some thoughts Not all 20 x 20 possible mutations occur equally often • Residues mutate more easily to similar ones (e. g. Leucine and Isoleucine) • Residues at surface mutate more easily • Aromatics mutate preferably into aromatics • Core tends to be hydrophobic; • Cysteines are dangerous at the surface • Cysteines in sulfur bridges (S-S) seldom mutate • Some amino acids have similar codons (for example TTT & TTC for Phe, TTA & TTG for Leu) • Etc etc

BLAST Output Click here to go to the corresponding swissprot entry Click here to study alignment in detail; Look here first!! A high score indicates a likely relationship A low E-value indicates that a match is unlikely to have arisen by chance

Low complexity motifs visible

3) Amino acids Hydrophobicity Entropy of water Amino acids have characteristics that determine their behaviour, and what they are being used for (Gly, Cys, His, Ser, Asp, etc).
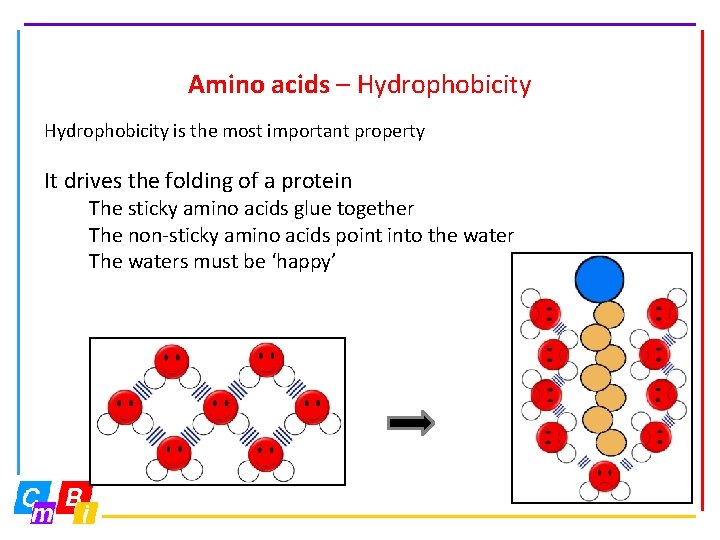
Amino acids – Hydrophobicity is the most important property It drives the folding of a protein The sticky amino acids glue together The non-sticky amino acids point into the water The waters must be ‘happy’
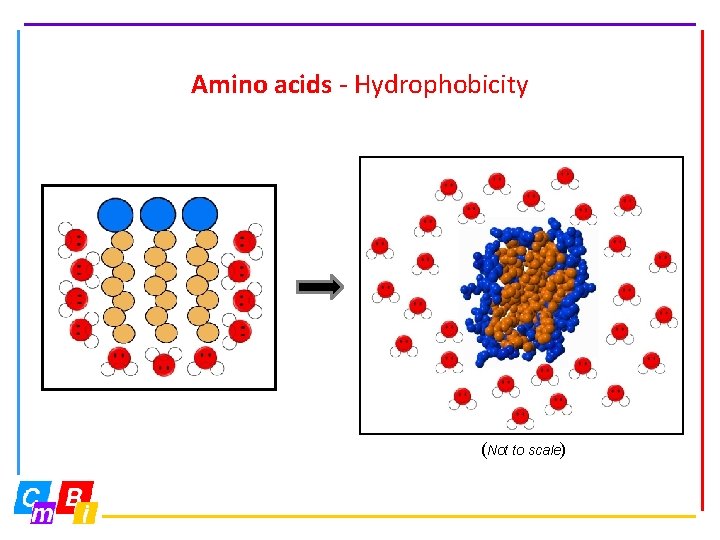
Amino acids - Hydrophobicity (Not to scale)
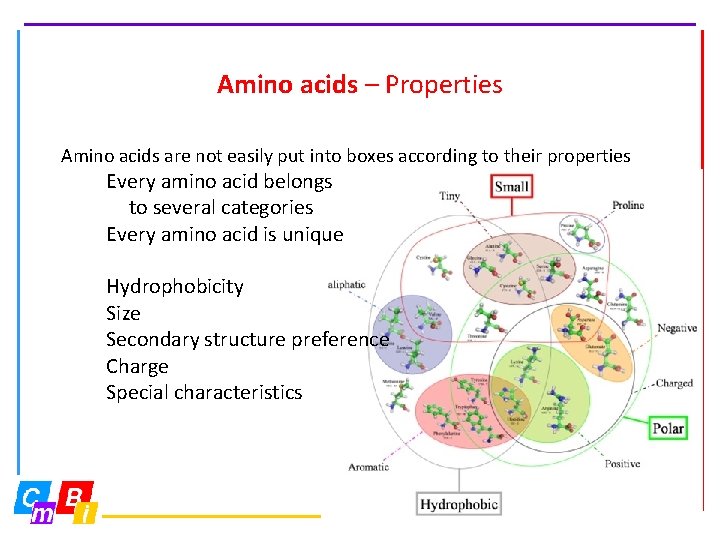
Amino acids – Properties Amino acids are not easily put into boxes according to their properties Every amino acid belongs to several categories Every amino acid is unique Hydrophobicity Size Secondary structure preference Charge Special characteristics

4) (Secondary) structure Structure data often is not available. Sequences don’t exist; structures exist. Residues at corresponding positions in structures have corresponding functions. Sequence alignment is the poor man’s solution to structure alignment. Knowledge of the structure (even if only predicted) can help improve the alignment.
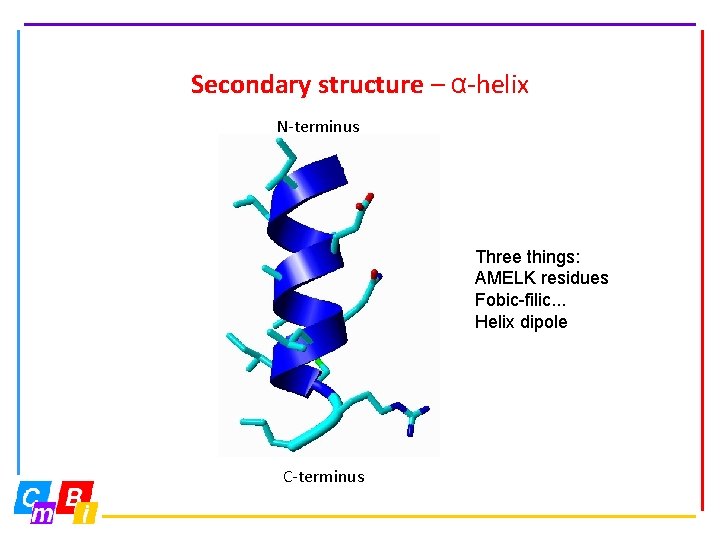
Secondary structure – α-helix N-terminus Three things: AMELK residues Fobic-filic. . . Helix dipole C-terminus
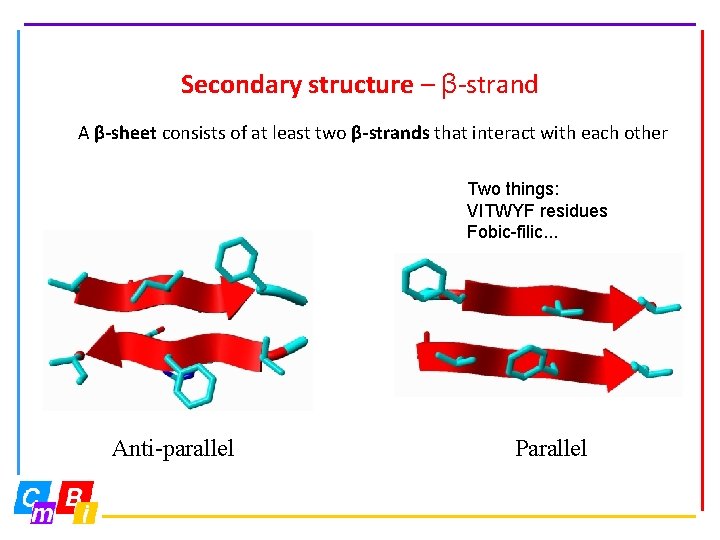
Secondary structure – β-strand A β-sheet consists of at least two β-strands that interact with each other Two things: VITWYF residues Fobic-filic. . . Anti-parallel Parallel
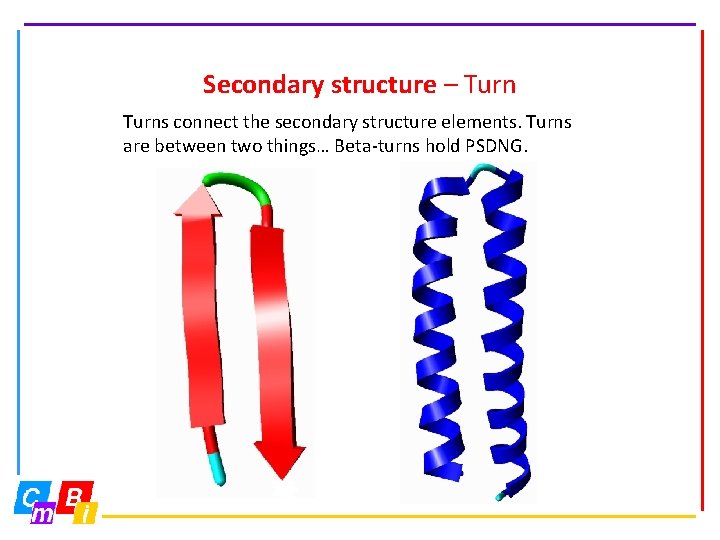
Secondary structure – Turns connect the secondary structure elements. Turns are between two things… Beta-turns hold PSDNG.
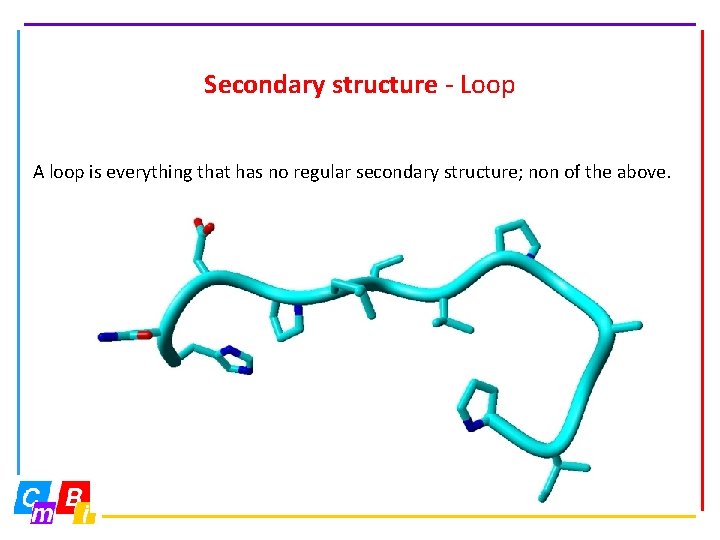
Secondary structure - Loop A loop is everything that has no regular secondary structure; non of the above.
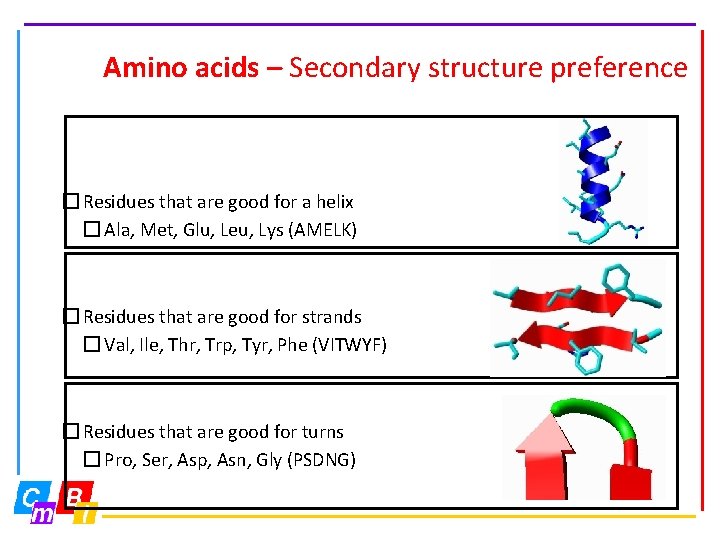
Amino acids – Secondary structure preference � Residues that are good for a helix � Ala, Met, Glu, Leu, Lys (AMELK) � Residues that are good for strands � Val, Ile, Thr, Trp, Tyr, Phe (VITWYF) � Residues that are good for turns � Pro, Ser, Asp, Asn, Gly (PSDNG)

3) Align CWEALALLAELALAAMKGSTPNGS met CWEALALLLEALMRGTTPNGG CWEALALLAELALAAMKGSTPNGS ? ? hhhhhhhh------CWEALALLLEALMR---GTTPNGG ? ? hhhhhh-----CW obviously on top of CW. Predict and align the two helices. Gap at end of helix. ©CMBI 20
5) Structure and alignment

Aquaporin

Aquaporin 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 A A A A A G P Y N Q F G G G A N S V A L G Y H H H T T > S+ <>S+ ><5 S+ 3<5 SX 5 3 < 3 S+ < + > T 3 S+ < - 0 0 0 0 0 0 0 0 0 48 32 4 65 147 72 29 0 5 22 1 29 4 37 180 82 95 597 599 599 600 600 599 601 601 601 602 601 70 91 4 77 79 72 13 0 0 9 0 83 25 47 75 3 2 AAAGDSHDTSASTGGNGASTTAAAGSSAKTNSSTSSGSAGSgggr. KKKGKRKKNSGGSKADDSSGKD YYYFDYQDYYYPPPPPLYYYIYFYPYYYQYPFYYFYRQFPQQQQVQQQPRPQQQYLLLPFAAEFLQA FFYYYYYYYYYYFYYYYYYYYYYYYYYYYYYYYY NDVATEDNNNVQQQQQQNTNQNVHQIVVQNQNTNNVEQVMEGGGEQQQHQQQQQTAEDEVQQEMAQQ RRRRKVATRRRTTRRRVRRRRKRRTRTRRRRRVVRALRRRSAAAAASAAARRTRRRAAMDRAA YYYYHYFHYYYLLLLLNYYYQYYYLYLYYYYYLHYLHLLLLLLLNLNLLLYYTYLYNNDHHLN GGGGGGGGGGGGDGGGGGGGGKNGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG AAAAAATAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN TFEETEVTVQSTTVTTVTEMEEEMVSSSGMTMEVTFTREFMAAAVTTTLTLTTTGEVEVEMMSAETM LLLVVLVVLLLVVVVVVLLLVLLLVLVVLVVVVVVVVLVVLVLVVVVVVV AHSSASAAAAAAAQAAAAASASSSNSAAAASQNQSNSAAAAAAASAASSAAAAGAA ADDAVPLVDEDPPPHHHADDHEADPDDDDDHHDDDPHNSHAHHHSPPPHPHHPPHAPADSSSHPAPS GGGGGGCGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG YYYYYYYYYYYYYYYYYYYFYYYYYYYYYYFFFYYYYF MSAs contain conserved residues, correlated mutations, and variable residues. SFTDALKNMKPYESSFTRIVN SFTASLKNLKPYCSSFTRVIG SFTDALKLIVPYESSFTDVIH SWTAVLKLMVPYLSSFTDILR SYTDALKNVKPYESSFTRVVN ©CMBI 20

The amino acids in their natural habitat Topics: • Hydrogen bonds • Secondary Structure • Alpha helix • Beta strands & beta sheets • Turns • Loop • Tertiary & Quarternary Structure • Protein Domains

6) Transfer of information GPNANGPALLEILSLIAEAAQALAGGNQDDEA Can be phosphorylated at exactly one spot by kinase X. GGLEAAKLASSAASAAELLAGDNKKKW too.

Transfer of information GPNANGPALLEILSLIAEAAQALAGGNQDDEA Can be phosphorylated at exactly one spot by kinase X. GGLEAAKLASSAASAAELLAGDNKKKW too. GPNANGPALLEILSLIAEAAQALAGGNQDDEA GGLEAAKLASSAASAAELLAGDNKKKW
- Slides: 36