Bioinformatics 2010 2011 Lecture 1 Introduction Dr Aladdin

Bioinformatics 2010 -2011 Lecture 1 Introduction Dr. Aladdin Hamwieh Khalid Al-shamaa Abdulqader Jighly Aleppo University Faculty of technical engineering Department of Biotechnology

Main Lines • Definition • Bioinformatics areas • Bioinformatics data – Data types – Applications for these data • Next generation sequencing • Bioinformatics algorithms • Joint international programming initiatives

Definition • Bioinformatics is the field of science in which biology, computer science, and information technology merge into a single discipline. • Bioinformatics is the science of managing and analyzing biological data using advanced computing techniques • Bioinformatics applies principles of information science to make the vast, diverse, and complex life sciences data more understandable and useful.

Definition • There are two extremes bioinformatics work in – Tool users (biologists): know how to press the buttons and the biology but have no clue what happens inside the program – Tool shapers (informaticians): know the algorithms and how the tool works but have no clue about the biology

Bioinformatics areas • Molecular sequence analysis 1. 2. 3. 4. 5. Sequence alignment Sequence database searching Motif discovery Gene and promoter finding Reconstruction of evolutionary relationships 6. Genome assembly and comparison

Bioinformatics areas • Molecular structural analysis 1. 2. 3. 4. 5. Protein structure analysis Nucleic acid structure analysis Comparison Classification prediction

Bioinformatics areas • Molecular functional analysis 1. gene expression profiling 2. Protein–protein interaction prediction 3. protein sub-cellular localization prediction 4. Metabolic pathway reconstruction 5. simulation
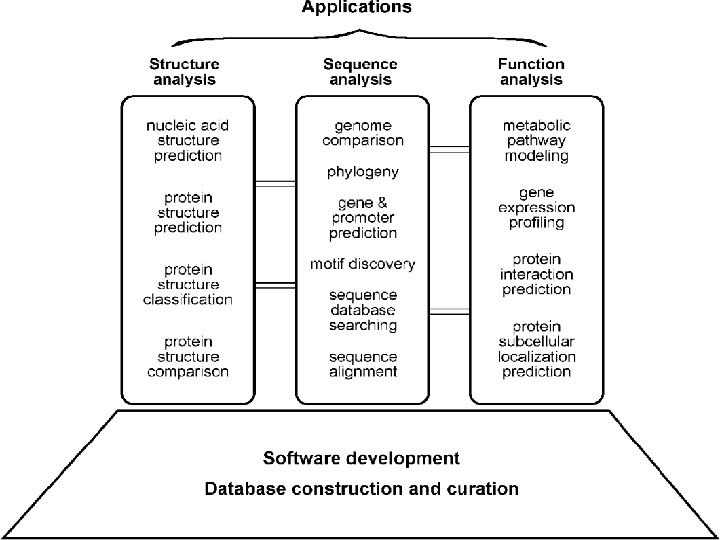

Bioinformatics data There is different data types usually used in bioinformatics The same data may be used in different areas

Data types • DNA sequences • RNA sequences • Expression (microarray) profile • Proteome (x-ray, NMR) profile • Metabolome profile • Haplotype profile • Phenotype profile

1 - DNA Sequences • Simple sequence analysis – Database searching – Pairwise and multiple analysis • • Regulatory regions Gene finding Whole genome annotation Comparative genomics
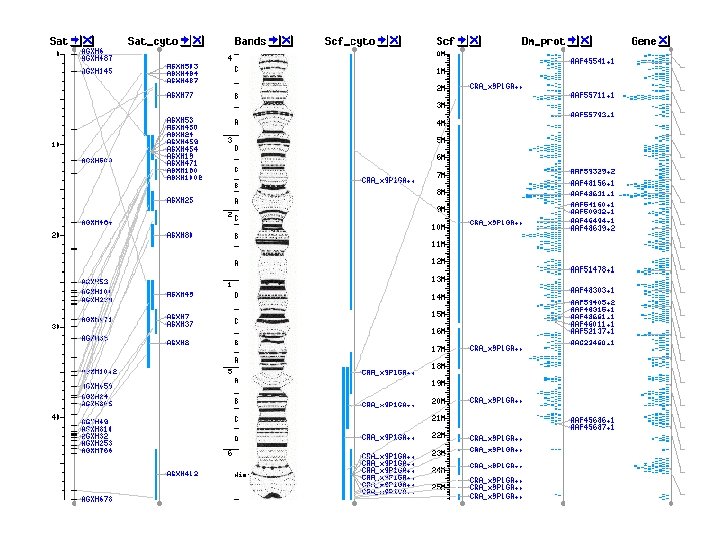

2 - RNAs • • • Splice variants Tissue specific expression 2 D structure 3 D structure Single gene analysis Microarray
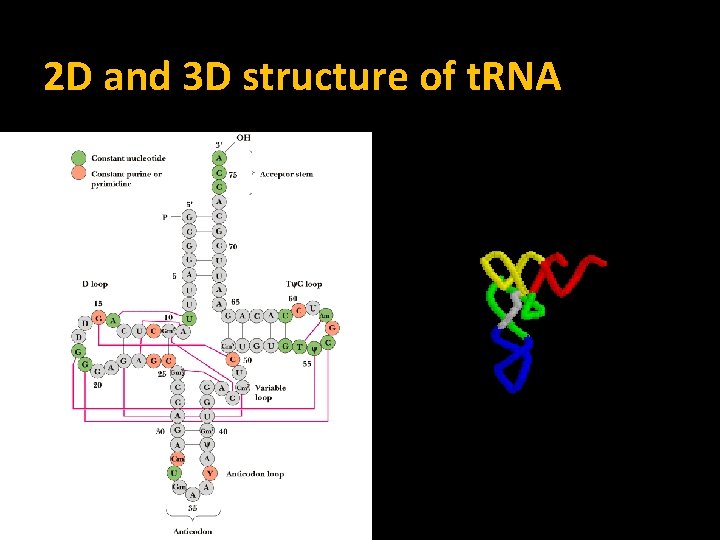
2 D and 3 D structure of t. RNA
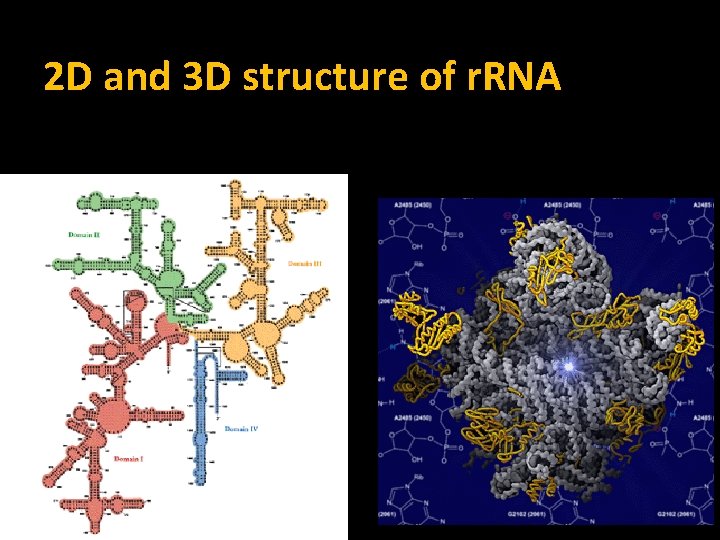
2 D and 3 D structure of r. RNA

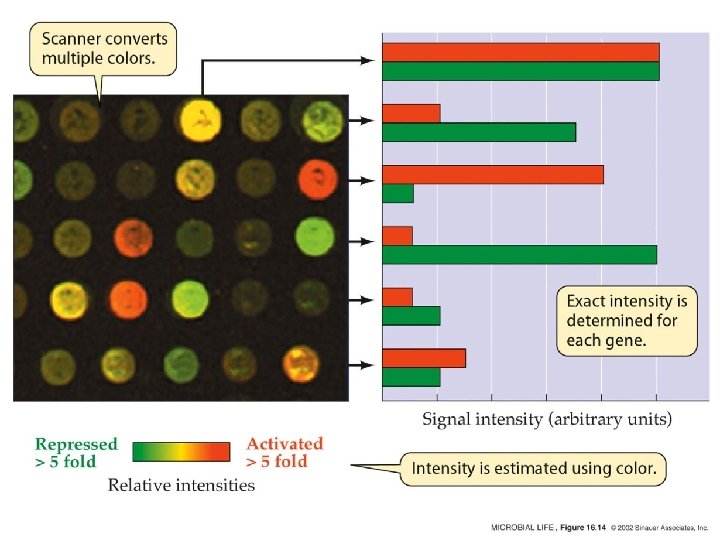
Microarray • 20, 000 to 60, 000 short DNA probes of specified sequences are orderly tethered on a small slide. Each probe corresponds to a particular short section of a gene.

Microarray • DNA microarrays measure the RNA abundance with either 1 channel (one color) or 2 channels (two colors). • Stanford microarrays measure by competitive hybridization the relative expression under a given condition (fluorescent red dye Cy 5) compared to its control (labeled with a green fluorescent dye, Cy 3) (Two channels) • Affymetrix Gene. Chip has 1 channel and use either fluorescent red dye Cy 5 or green fluorescent dye, Cy 3

3 - Proteins • Protein sequences analysis – Database searching – Pairwise and multiple analysis • • 2 D structure 3 D structure Classification of proteins families Protein arrays
3 D structure
Animation

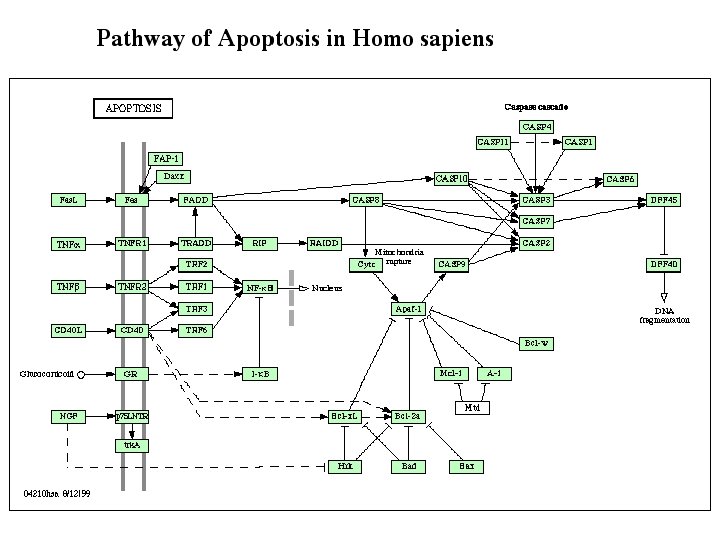
4 - Metabolome and molecular biology • Metabolic pathways • Regulatory networks Helps to understand systems biology

5 - Haplotype • Molecular Markers – – – RFLP RAPD SSR ISSR AFLP DAr. T – SNP – ….
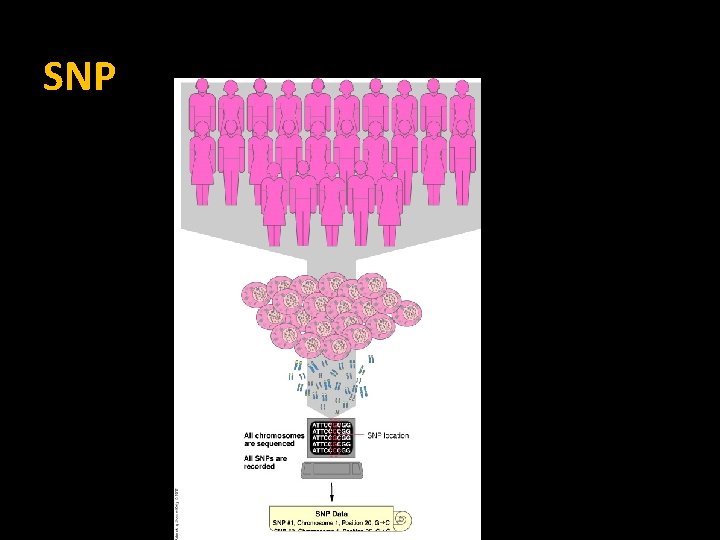
SNP

6 - Phenotype • • Morphological data Physiological data Stresses tolerance Pathogenic infections Diseases resistance Cancers types …. .
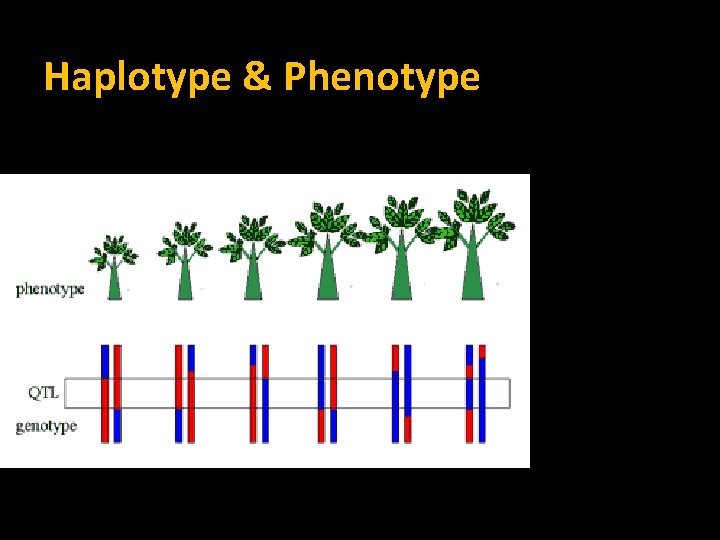
Haplotype & Phenotype
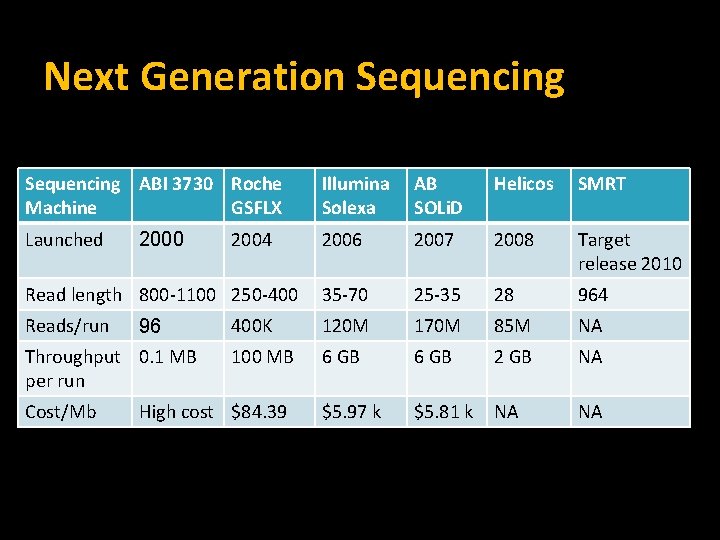
Next Generation Sequencing ABI 3730 Roche Machine GSFLX Illumina Solexa AB SOLi. D Helicos SMRT Launched 2006 2007 2008 Target release 2010 Read length 800 -1100 250 -400 35 -70 25 -35 28 964 Reads/run 400 K 120 M 170 M 85 M NA 100 MB 6 GB 2 GB NA $5. 97 k $5. 81 k NA NA 2000 96 Throughput 0. 1 MB per run Cost/Mb 2004 High cost $84. 39
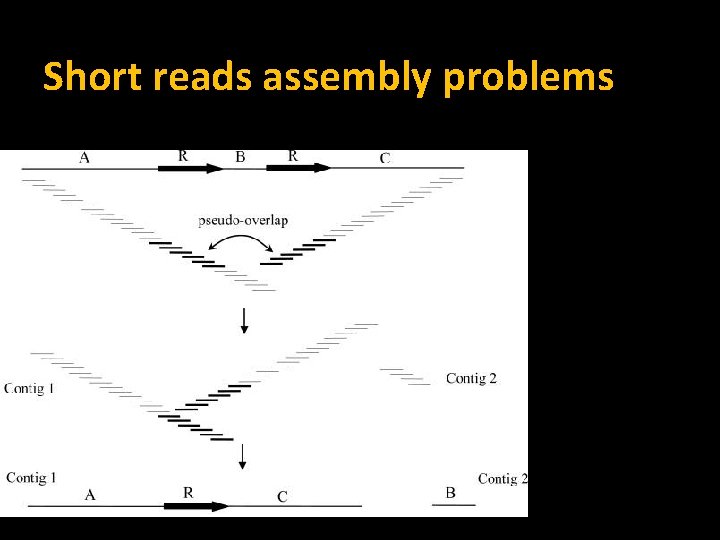
Short reads assembly problems
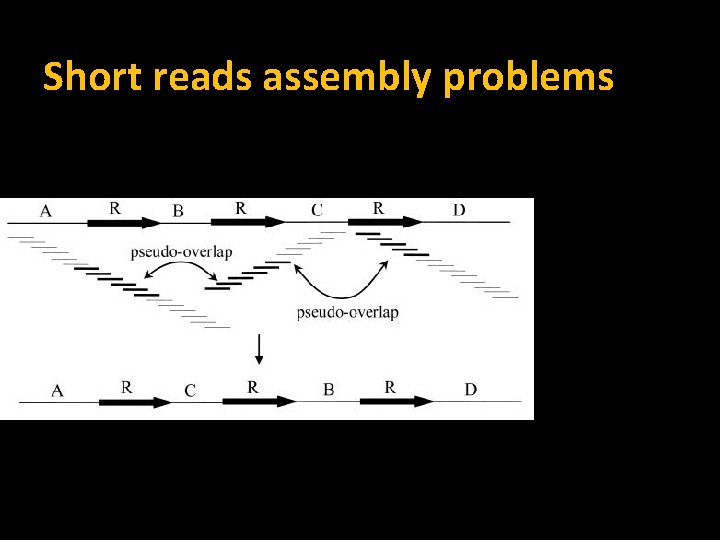
Short reads assembly problems
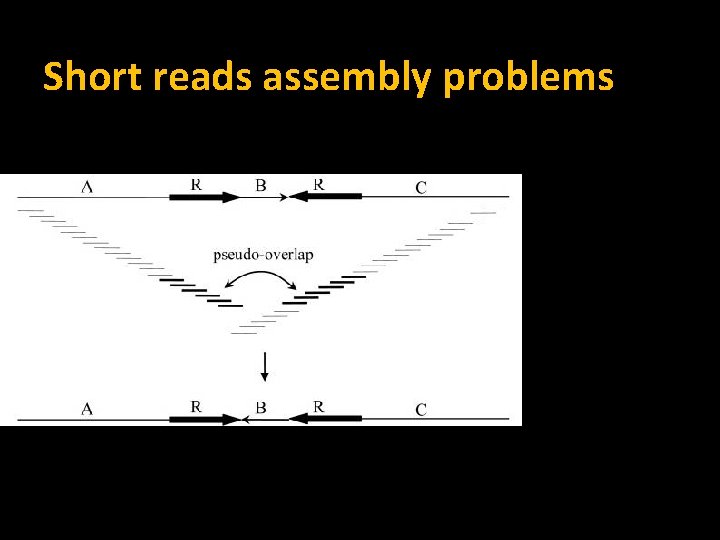
Short reads assembly problems

Algorithms in bioinformatics • String algorithms • Dynamic programming • Machine learning (NN, k-NN, SVM, GA, . . ) • Markov chain models • Hidden Markov models • Markov Chain Monte Carlo (MCMC) algorithms • Stochastic context free grammars • EM algorithms • Gibbs sampling • Clustering • Tree algorithms (suffix trees) • Graph algorithms • Text analysis • Hybrid/combinatorial techniques • ….

Joint international programming initiatives • Bioperl http: //www. bioperl. org/wiki/Main_Page • Biopython http: //www. biopython. org/ • Bio. Tcl http: //wiki. tcl. tk/12367 • Bio. Java www. biojava. org/wiki/Main_Page

Thank You
- Slides: 34