Bioinformatics 2 course Course contents General Introduction Part

Bioinformatics 2η course

Course contents General Introduction ü Part I -- Introduction to algorithms for efficient storage and management of strings and biological data sequences. -Algorithms for exact pattern matching (Boyer-Moore, Knuth-Morris -Pratt, Shift-Or, Multiple pattern matching). -- Introduction to suffix trees and its applications. -- Αlgorithms for approximate pattern matching and Sequence Alignment -Algorithms for searching in Sequence Data Bases (FASTA, BLAST, PROSITE)

ü Part II -- The theoretical base of molecular design. -- Molecular models and biochemical information -- Structure based drug design -- Open problems ü Part III Clustering and categorization techniques for biological data, in order to predict the behavior of biological molecules.

Techniques for Analysing and Comparing Sequences of Biological Data n n Examples of Data Bases with Biological Sequences Basic Definitions The problem of exact pattern matching ¨ Naïve method ¨ Boyer-Moore algorithm ¨ Knuth-Morris-Pratt algorithm ¨ Shift-Or/Shift And Algorithm ¨ Aho-Corasick automaton Applications in Molecular Biology problems

Biological Data Bases q q ü Generalised or Archival biological data bases. We can discriminate them to: - Primary Sequence Databases. They contain nucleic and amino acid genome sequences that have been sequenced completely or -data bases that contain three-dimensional structures of nucleic acid and proteins (GENBANK, EMBL-Bank, DDJB, Swiss-Prot, PIR-PSD) Secondary biological data bases that emerge from analysing data that are stored in archival biological data bases and are distinguished to: Secondary DB DNA sequences and proteins that emerge from the basic DB sequences and contain (α) DB sequences from where we have removed the sequences that have been stored more than once (β) DB that register mutations or variations in DNA sequences and proteins (γ) Genomic DB that cluster neighboring and not fully sequenced genomes or deal with genomes of organisms.

DB that are related to the hierarchical and/or correlations between molecules such as protein families, common motifs of DNA sequences, and proteins: q. Specialized D. B. , a category in which they belong: ü D. B. of microarrays that contain information for gene expression and proteins ü D. B. of metabolic pathways that contain information for chemical reactions that take place in the cell q. Bibliographical Biological Data Bases q. Biological Data Bases of web pages that contain: ü D. B. that contain as records biological bases ü Links between biological data bases. ü

Biological Data Bases(1) n Gen. Bank: NCBI (http: //www. ncbi. nlm. nih. gov) Gen. Bank® is the NIH genetic sequence database, an annotated collection of all publicly available DNA sequences. (National Center of Biotechnology Information) n PIR: Protein Information Resource (http: //pir. georgetown. edu) PIR produces the Protein Sequence Database (PSD) of functionally annotated protein sequences, which grew out of the Atlas of Protein Sequence and Structure (1965 -1978) edited by Margaret Dayhoff and has been incorporated into an integrated knowledge base system of value-added databases and analytical tools. n Swiss-Prot + Tr. EMBL: Swiss-Prot. htm (http: //tw. expasy. org/sprot/) Swiss-Prot is a curated protein sequence database which strives to provide a high level of annotation (such as the description of the function of a protein, its domains structure, post-translational modifications, variants, etc. ), a minimal level of redundancy and high level of integration with other databases Tr. EMBL is a computer-annotated supplement of Swiss-Prot that contains all the translations of EMBL nucleotide sequence entries not yet integrated in Swiss-Prot.

Biological Data Bases(2) n PROSITE: Prosite (http: //tw. expasy. org/prosite/) PROSITE is a database of protein families and domains. It consists of biologically significant sites, patterns and profiles that help to reliably identify to which known protein family (if any) a new sequence belongs. n PDB-Protein Data Bank: PDB (http: //www. rcsb. org/pdb/ The PDB is the single worldwide repository for the processing and distribution of 3 -D structure data of large molecules of proteins and nucleic acids. § SCOP: (http: //scop. berkeley. edu/) Structural Classification of Proteins n PRINTS: ( http: //umber. sbs. man. ac. uk/dbbrowser/PRINTS ) PRINTS is a compendium of protein fingerprints. A fingerprint is a group of conserved motifs used to characterise a protein family; its diagnostic power is refined by iterative scanning of a SWISS-PROT/Tr. EMBL composite.

An example: PDB data entry example
![Basic Definitions(1) n String: x=x[1]x[2]…. . x[n], x[i] Σ & |x|=n x= acgttaaaca, |x|=10 Basic Definitions(1) n String: x=x[1]x[2]…. . x[n], x[i] Σ & |x|=n x= acgttaaaca, |x|=10](http://slidetodoc.com/presentation_image/0292ed3e6e2a3baaa44646ba847053c0/image-10.jpg)
Basic Definitions(1) n String: x=x[1]x[2]…. . x[n], x[i] Σ & |x|=n x= acgttaaaca, |x|=10 & Σ={Α, C, G, T} (Α (adenine ), Τ (thymine ), C (cytosine ), G (guanine ) ) n Σ+: the set of strings that are defined in the alphabet Σ ? How many strings of length « » are defined in Σ={a, c, g, t} n n Empty string: ε Substring w: x=uwv Prefix w: x=wu Suffix w: x=uw

Basic Definitions (2) n n Border of string x: string w that is both prefix and suffix and x. xk=x…x , κ-th power of x κ φορές n n n y=xk, k>1 y is periodical with period x Period y= the smallest such string Primitive string Cover of a string Seed of a string

Exact pattern matching(1) n n Exact matching: we are interested in locating all occurences of a given motif P (“structured” or “nonstructure d”) in a string (biological sequence) Τ. Approximate matching: for a text T, a motif P, a parameter k and a similarity function d( ) , locate all positions i, j in text, so that, d(P, Ti. . j ) >=k.

Pattern matching problems (2) n The procedure of similarity matching between two sequences is based on matrices that score the matches and mismatches betweeb successive symbols. This kind of matrices are: Dayhoff Mutation Data Matrix, BLOSUM κτλ. n Hence comparing sequences, can be categorized in : α) local alignment and β) global alignment. In local alignment we seek of local similarity regions. Known algorithms of this kind are Smith. Waterman (local), Needleman & Wunsch (global). In both cases there more than one alignments. The optimal solution is to minimize the differences between two seauences or differently to maximize the similarity function.

Sequence Alignment n Compare two or more sequences controling for a sequence of atomix symbols that are in the same order with the sequence. n Discover functional, structural and evolutionary information. L G P S S K Q T G K G S – S R I W D N global alignment L N – I T K S A G K G A I M R L G D A -- -- T K G S – -- -- -- -Structural alignment -- -- – -- -- A K G A -- --

Finding repetitions in Biological Sequences n Repetitions in biological sequences are categorized in 3 basic categories : Limited length repetitions that appear in local level and whose function is known, ¨ Limited length repetitions that appear in the whole length f the sequence and whose functionality in not known, ¨ Structured repetitions of large length whose functionality has not been defined. ¨ Εισαγωγή στη Βιοπληροφορική

Example of repetitions n 1 st category: The complementary palindromes in DNA & RNA sequences that regulate the DNA transcription, ¨ The nested complentary palindromes in RNA sequences ¨ n 2 th category: tandem repeats, ¨ DNA- satellite DNA, (micro & mini satellite DNA) ¨ n 3 nd category: ¨ SINE-Short Interspersed Nuclear Sequences (e. g. : Alu family) ¨ LINE-Long Interspersed Nuclear Sequences.

Motifs DNA motif TRANSFAC, JASPAR, SCPD, DBTBS, Regulon. DB n Protein motifs PROSITE, Pfam, Pro. Dom, BLOCKS, TIGRFAM, Interpro n Εισαγωγή στη Βιοπληροφορική Περδικούρη Αικατερίνη, Μακρής Χρήστος

Exact matching (applications) n n n Text processors Utilities (grep στο Unix) Textual Information Retrieval (Medline, Lexis, Nexis) Internet News Readers On-line dictionaries and thesaures Molecular Biology Databases
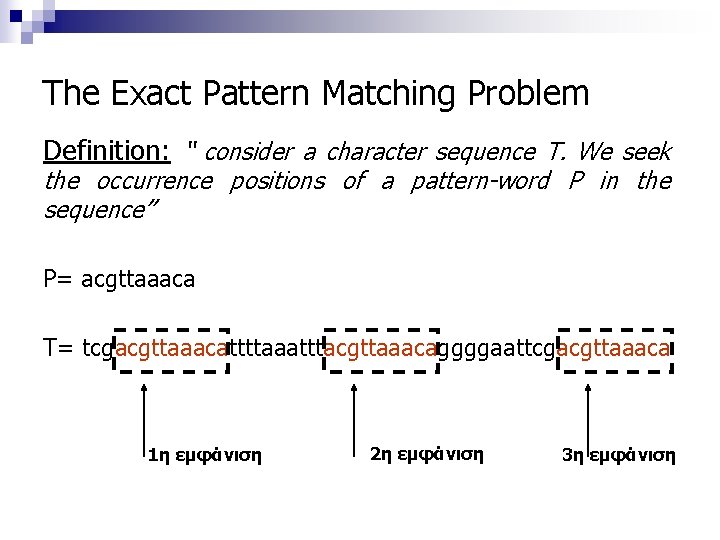
The Exact Pattern Matching Problem Definition: “ consider a character sequence T. We seek the occurrence positions of a pattern-word P in the sequence” P= acgttaaaca T= tcgacgttaaacattttaaatttacgttaaacaggggaattcgacgttaaaca 1η εμφάνιση 2η εμφάνιση 3η εμφάνιση

Naïve Solution Method 1ο step: se align the sequence and the pattern & compare the characters g c a t g c a g a g t 1 2 3 4 5 6 7 8 g c a g a g a t a c a g t a c g 2ο βήμα: In the first mismatch – 4η position we locate the pattern in 1 position g c a t g c a g a g t 1 2 3 4 5 6 7 8 g c a g a g a t a c a g t a c g

Naive Method 3ο step: in every mismatch we shift the pattern to 1 position g c a t g c a g a g t 1 2 3 4 5 6 7 8 g c a g a g a t a c a g t a c g 4ο step: g c a t g c a g a g t 1 2 3 4 5 6 7 8 g c a g a g

Naive Method 5ο step: 1η location of the pattern L{X}={5, …} g c a t g c a g a g t 1 2 3 4 5 6 7 8 g c a g a g a g t a c g a t a c a g t a c g 6ο step: g c a t 1 2 3 4 5 6 7 8 g c a g a g

Code of naïve method void Naïve-Method (char *P, int m, char *T, int n) { int i, j; for (j=0; j<=n-m; ++j) { for (i=0; i<m && P[i]==T[i+j]; ++i); if (i>=m) output(j); } }

Analysing naïve time method n Complexity of method: Ο(n*m), where|T|=n & |P|=m 1. How many relocations are needed? |T|- |P|+1= n –m +1 2. How many comparisons every time? |P|=m 3. Total processing time: (n –m +1)*m How to improve time? 4.

Data scale We will present 3 efficient linear time algorithms , linear to the size of the input sequence Ο(|T|)

Fundamental Preprocessing (1) (D. Gusfield) n n n Ζi(S)= the length of the largest substring of S, that begins at i and matches prefix of S. Z-box at i= the set of characters that begin at i and end at i+Zi(S)-1. For every i, ri symbolizes the rightmost edge of Z-boxes that begins before or at the position i. Differently ri is the largest value of j+Ζj-1 for every 2<j<=i. The leftmost edge(j) is symbolized as li.

Fundamental preprocessing(2) Given Zi for i<=k-1 and r, l για Zk-1: 1. If k>r, find Zk explicitly. If Zk>0, r=k+Zk-1, l=k. 2. If k<=r, S[k. . r] matches S[k’. . Zl] and the substring at k matches a prefix of S of length >=min(Zk’, r-k+1) (k’=kl+1) 2. a. If Zk’ < |S[k. . r]| then Zk=Zk’ , r, l remain unchanged 2. b. Compare the characters starting at r+1 of S with the characters starting at |S[k. . r]| until a mismatch. Say the mismatch occurs at q>=r+1. Then Zk is q-k, r is set to q 1 and l is set to k.

Fundamental preprocessing(3) n n Apply the algorithm for P$T Every value Zi=m, i>m signifies match in location i-m-1. The method can be implemented so that it demands besides the storage space of P, T, O(m) space. It is a method whose complexity is independent of the alphabet size (the same property have algorithms Boyer Moore, Knuth Morris Pratt, but not Shift Or and Aho. Corasick)

Boyer-Moore algorithm 1 st idea: We align the sequence and the pattern, & we compare the characters From right to left g c a t c g c a g a g t 1 2 3 4 5 6 7 8 g c a g a g a t a c a g t a c g 2 nd idea: In every mismatch we shift the pattern more than one positions based on two rules g c a t c g c a g a g t 1 2 3 4 5 6 7 8 g c a g a g a t a c a g t a c g
![α’ rule: “good suffix shift” T b u a u P[i-1] T [j+i-1] P α’ rule: “good suffix shift” T b u a u P[i-1] T [j+i-1] P](http://slidetodoc.com/presentation_image/0292ed3e6e2a3baaa44646ba847053c0/image-30.jpg)
α’ rule: “good suffix shift” T b u a u P[i-1] T [j+i-1] P P[i… m] = T[j+i…j+m] idea: let us align the next appearance of u with the respective segment T[j+i, …, j+m] P c u shift
![β’ rule: “bad character shift” T b u a u P[i-1] T [i+j-1] P β’ rule: “bad character shift” T b u a u P[i-1] T [i+j-1] P](http://slidetodoc.com/presentation_image/0292ed3e6e2a3baaa44646ba847053c0/image-31.jpg)
β’ rule: “bad character shift” T b u a u P[i-1] T [i+j-1] P T[i+j-1] = b idea: let us align the character T[i+j-1] =b with its rightmost appearance – if it exists – in pattern P P b δεν περιέχει b shift

Implementin “bad character shift” n (Simple Shift) Use an array of size m x |Σ| and scan the pattern from left to right n (Extended Shift) Scan the pattern from right to left and for every character create a list of occurences. Scan the proper list when it is located.

Implementing“goof suffix shift” (1) n n Let L(i) be the largest position in P, so that P[i. . m] matches a suffix of P[1. . L[i]] with the extra constraint that the character that is before the suffix should be different from P[i-1]. We define l’(i) the length of the largest suffix of P[i. . m] that is also a prefix of P.

Implementin“goof suffix shift” (2) Let Nj(P) be the length of the largest suffix of P[1. . j] that is a suffix of P. Nj(P)= Zm-j+1(Pr) n From the definition we have that L(i) is the largest value j<m so that Νj(P)=|P[i. . m]| Hence l’[i] is the largest j<=|P[i. . m]|, so that Nj(P)=j

Implementing“goof suffix shift” (3) for i=1 to m do L’(i)=0; for j=1 to m-1 do begin i=m-Nj(P)+1; L(i): = j; end

Analysing Boyer Moore in time n Time Complexity: Ο(n*m), where |T|=n & |P|=m 1. The needed arrays are computed in O(m+σ) time. 2. In real applications we need “ 3 n” comparisons. 3. For a large alphabet |Σ|= (≈|pattern|), we need Ο(n/m) comparisons. Without match time complexity is O(n). There exist variants with worst-case O(n+m) χρόνο 4. 5.
![Algorithm Knuth-Morris-Pratt Τ b P[i+1] T [i+1+j] u P u a P[1… i] = Algorithm Knuth-Morris-Pratt Τ b P[i+1] T [i+1+j] u P u a P[1… i] =](http://slidetodoc.com/presentation_image/0292ed3e6e2a3baaa44646ba847053c0/image-37.jpg)
Algorithm Knuth-Morris-Pratt Τ b P[i+1] T [i+1+j] u P u a P[1… i] = T [j…i+j-1] = u idea: let us align the maximal prefix(P)=v with the respective segment of suffix u of sequence shift P v c

Let us see the specific algorithm 1ο step: we align the sequence and the pattern & we compare the characters from left to right: x y a b c x a d c d q f 1 2 3 4 5 6 7 8 9 a b c x a b c d e e g a g t a c g 2ο step: We shift the pattern for 4 positions x y a b c x a d c d q f 1 2 3 4 5 6 7 8 9 a b c x a b c d e

Implementing KMP (1) n n n We define as shi(P) the length of the largest suffix of P[1. . i] that matches a prefix of P, with the extra condition that the characters P[i+1] and P[shi(P)+1] are different. Position j>1 is mapped to i when i=j+Zj(P)-1. For every i>1 shi(P)=i-j+1 ; where j is the smallest position that maps to i.

Implementing KMP (2) For i=1 to m to shi(P)=0; For j=m downto 2 do begin i=j+Zj(P)-1; shi(P)=Zj(P); end

Real Time KMP n We define as shi(P, x) the length of the largest prefix of P[1. . i] that matches a prefix of P, with the extra condition that character P[shi(P, x)+1] is x. As previously: n For i=1 to m to shi(P)=0; For j=m downto 2 do begin i=j+Zj(P)-1; x=P[Zj(P)+1] shi(P, x)=Zj(P); end

Analyzing time complexity of algorithm Knuth. Morris-Pratt n Method complexity: Ο(n+m), όπου |T|=n & |P|=m 1. In time O(m) we compute, the «period» of the pattern with which we shift the pattern

Algorithm Shift-Or n The algorithm uses arithmetic techniques: 1. Consider for char c Σ, the vector Sc of size m=|P|, that stores the occurrences of c in the pattern (with 0 we designate occurence), 2. The matrix R[mxn]: bit-array where R[i, j] is 0 if and only if the first i characters of P match the i characters that end in the j-th character of T. 3. Rj+1 = Shift (Rj ) ΟR ST[j+1]

Let us see the algorithm 1ο step: we compute the vectors SC for the pattern x=gcagagag Sa Sc Sg St g 1 1 0 1 c 1 0 1 1 a 0 1 1 1 g 1 1 0 1

…Let us see the algorithm in practise 2ο step: we compute the values of the array R, according to the value : Rj+1 = Shift (Rj ) Or SΤ[j+1] 0 1 2 3 4 5 6 7 8 9 10 11 12 g c a t c g c a g a g 0 1 1 0 1 0 1 c 1 0 1 0 1 1 1 2 a 1 1 0 1 1 1 1 1 3 g 1 1 1 1 0 1 1 4 a 1 1 1 1 1 0 1 1 1 5 g 1 1 1 1 1 0 1 1 6 a 1 1 1 0 1 7 g 1 1 1 0

We analyse Shift-Or in time n Complexity of method: Ο(n+m), where |T|=n & |P|=m 1. In time O(m*σ) we compute, the vectors SC
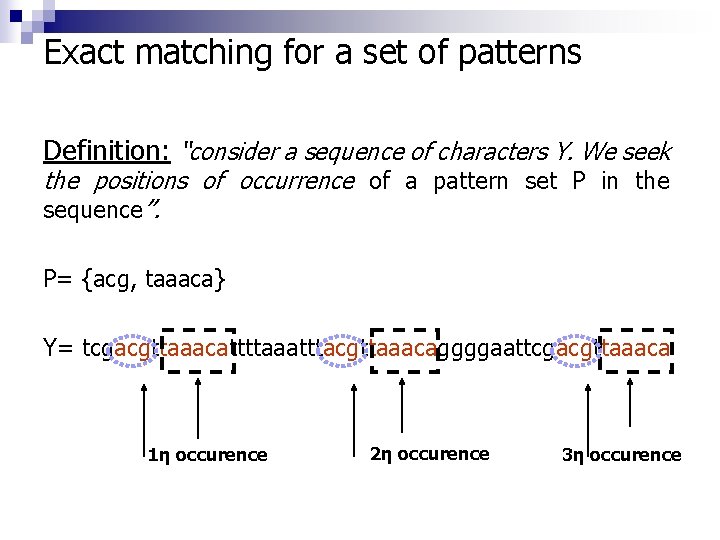
Exact matching for a set of patterns Definition: “consider a sequence of characters Υ. We seek the positions of occurrence of a pattern set Ρ in the sequence”. Ρ= {acg, taaaca} Y= tcgacgttaaacattttaaatttacgttaaacaggggaattcgacgttaaaca 1η occurence 2η occurence 3η occurence
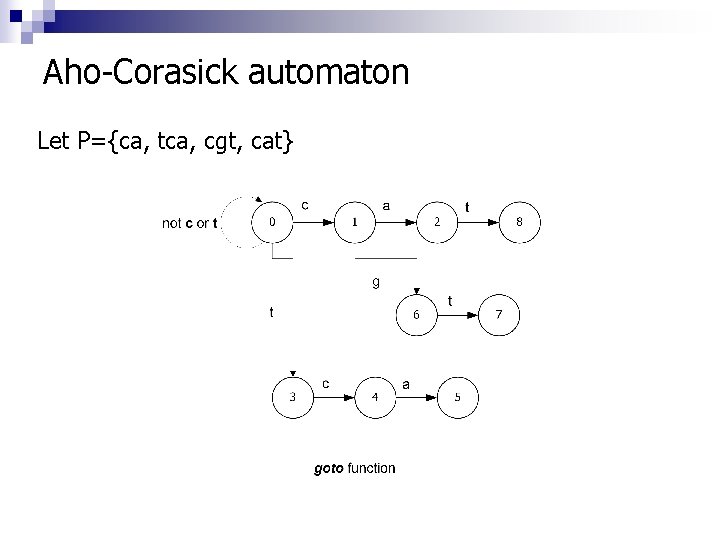
Aho-Corasick automaton Let Ρ={ca, tca, cgt, cat}

The function “goto” n n g(s, α) = s’: the automaton transists to state s’ and the next character of the sequence gets read in the input, g(s, α) = fail, the automaton transits to state s’=f(s) according to the failure function. The search continues to the current state s’ and the input symbol becomes the character that has been read in input α.

The function “failure-function” n n n f(s) = 0, for every state s, of depth 1. In order to compute f(s), for every state s, of depth d, consider all the states r, of depth d-1 : n If g(r, α) = fail, for every α, do nothing, n Differently, for every symbol α, so that g(r, α)= s, then: o Set state = f(r) o Execute the command state f(state), till g(state, α) fail Set f(s)= g(state, α)

Applications of pattern matching in Bioinformatics problems n Searching Sequence-tagged-site (STS) & Expressed Sequence Tags (ESTs) in genome sequences. ¨ STS: τμήματα του DNA μήκους 200 -300 νουκλεοτιδίων ¨ ESTs: segments of m. RNA & c. DNA sequences that represent the coding segment of a protein in a genome sequence. n Searching “regular expressions” (regular expressions) ¨ [ED]-[EN]-L-[SAN]-x-x-[DE]-x-E-L ENLSSEDEEL
- Slides: 51