BIODEGRADATION OF POLYCYCLIC AROMATIC HYDROCARBONS USING BACTERIA ISOLATED

BIODEGRADATION OF POLYCYCLIC AROMATIC HYDROCARBONS USING BACTERIA ISOLATED FROM USED ENGINE OIL CONTAMINATED SOIL ZUBAIRU UMAR DARMA, Ph. D DEPARTMENT OF MICROBIOLOGY FACULTY OF NATURAL AND APPLIED SCIENCES UMARU MUSA YAR’ADUA UNIVERSITY, KATSINA
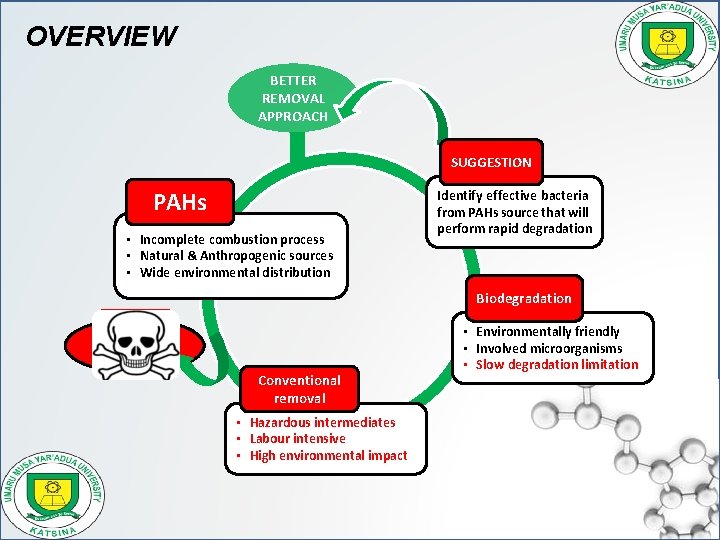
OVERVIEW BETTER REMOVAL APPROACH SUGGESTION PAHs • Incomplete combustion process • Natural & Anthropogenic sources • Wide environmental distribution Identify effective bacteria from PAHs source that will perform rapid degradation Biodegradation Dangerous to living cells Conventional removal • Hazardous intermediates • Labour intensive • High environmental impact • Environmentally friendly • Involved microorganisms • Slow degradation limitation
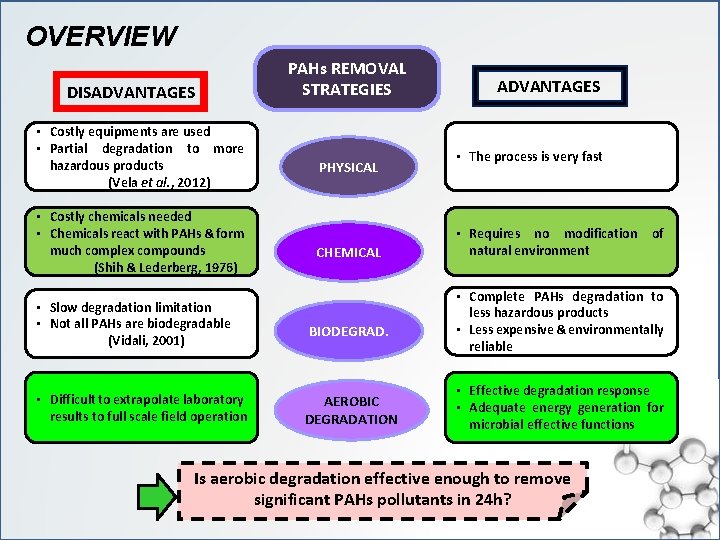
OVERVIEW DISADVANTAGES • Costly equipments are used • Partial degradation to more hazardous products (Vela et al. , 2012) • Costly chemicals needed • Chemicals react with PAHs & form much complex compounds (Shih & Lederberg, 1976) PAHs REMOVAL STRATEGIES PHYSICAL CHEMICAL • Slow degradation limitation • Not all PAHs are biodegradable (Vidali, 2001) BIODEGRAD. • Difficult to extrapolate laboratory results to full scale field operation AEROBIC DEGRADATION ADVANTAGES • The process is very fast • Requires no modification of natural environment • Complete PAHs degradation to less hazardous products • Less expensive & environmentally reliable • Effective degradation response • Adequate energy generation for microbial effective functions Is aerobic degradation effective enough to remove significant PAHs pollutants in 24 h?
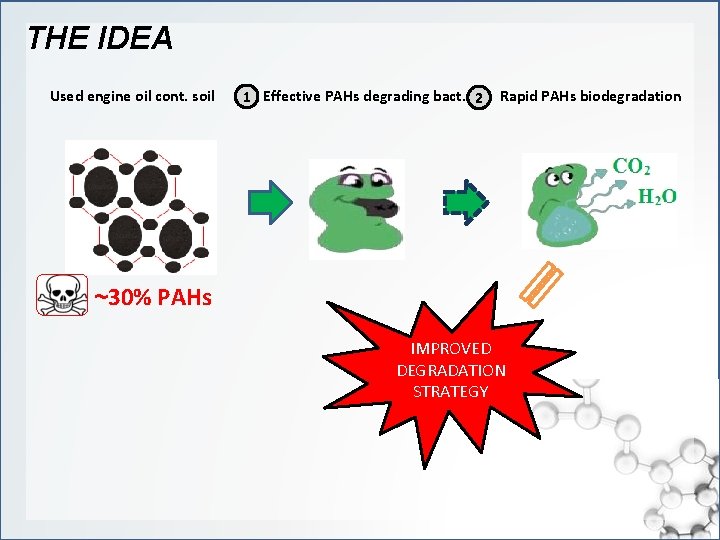
THE IDEA Used engine oil cont. soil 1 Effective PAHs degrading bact. 2 Rapid PAHs biodegradation ~30% PAHs IMPROVED DEGRADATION STRATEGY

OBJECTIVE 1: An attempt to identify effective PAHs degrading bacteria QUESTION ? Ø Does PAHs degrading bacteria exist in used engine oil contaminated soil? OBJECTIVE 1 Ø Identification of PAHs degrading bacteria from used engine oil contaminated soil HOW DO WE ACHIEVE THIS? Ø Enrichment of soil bacteria & screening the best isolates Ø Molecular identification SIGNIFICANT OF THE STUDY Ø Strategizing the best PAHs biodegradation
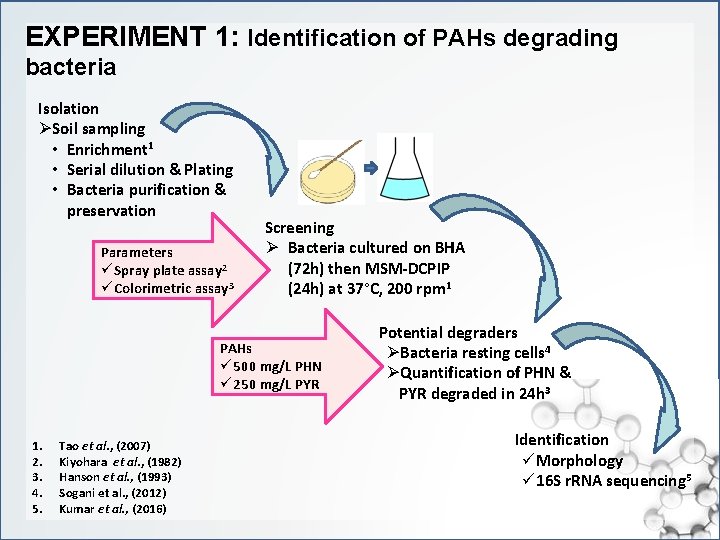
EXPERIMENT 1: Identification of PAHs degrading bacteria Isolation ØSoil sampling • Enrichment 1 • Serial dilution & Plating • Bacteria purification & preservation Parameters ü Spray plate assay 2 ü Colorimetric assay 3 Screening Ø Bacteria cultured on BHA (72 h) then MSM-DCPIP (24 h) at 37°C, 200 rpm 1 PAHs ü 500 mg/L PHN ü 250 mg/L PYR 1. 2. 3. 4. 5. Tao et al. , (2007) Kiyohara et al. , (1982) Hanson et al. , (1993) Sogani et al. , (2012) Kumar et al. , (2016) Potential degraders ØBacteria resting cells 4 ØQuantification of PHN & PYR degraded in 24 h 3 Identification üMorphology ü 16 S r. RNA sequencing 5
OUTCOME 1: Isolation & Screening v Total bacteria isolated : 93 Isolates v 72 h spray plate screened : 53 Isolates 24 h colorimetric rapid response : MM 045 & MM 087 DCPIP
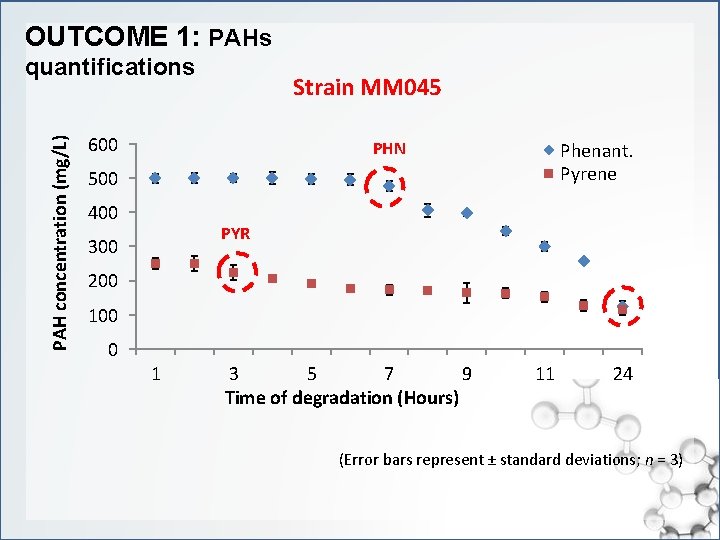
OUTCOME 1: PAHs PAH concentration (mg/L) quantifications Strain MM 045 600 PHN Phenant. Pyrene 500 400 PYR 300 200 100 0 1 3 5 7 9 Time of degradation (Hours) 11 24 (Error bars represent ± standard deviations; n = 3)
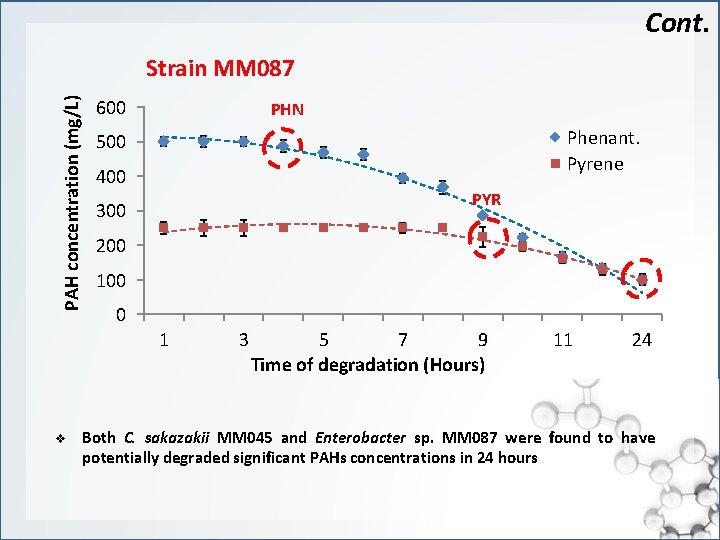
Cont. PAH concentration (mg/L) Strain MM 087 600 PHN Phenant. Pyrene 500 400 PYR 300 200 100 0 1 v 3 5 7 9 Time of degradation (Hours) 11 24 Both C. sakazakii MM 045 and Enterobacter sp. MM 087 were found to have potentially degraded significant PAHs concentrations in 24 hours
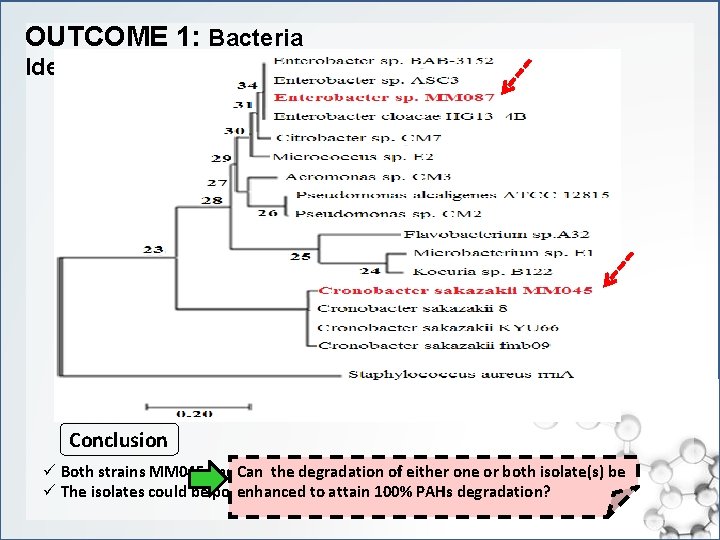
OUTCOME 1: Bacteria Identification Conclusion ü Both strains MM 045 and MM 087 degraded >70% PHN & >50% PYR in 24 hrs Can the degradation of either one or both isolate(s) be ü The isolates could be potential degraders that effectively remove PAHs rapidly enhanced to attain 100% PAHs degradation?
OBJECTIVE 2: Optimization of PAHs biodegradation conditions QUESTION ? v Can C. sakazakii MM 045 & Enterobacter sp. MM 087 perform 100% PAHs degradation in 24 hrs? OBJECTIVE 2 ! v Optimization of phenanthrene and pyrene biodegradation conditions HOW TO ACHIEVE IT ? v Single factor optimization v Response surface methodology SIGNIFICANCE v Enhancing degradation performance of C. sakazakii MM 045 & Enterobacter sp. MM 087
EXPERIMENT 2: Optimization of biodegradation condition Strain MM 045 & MM 087 ü Colorimetric assay 3 ü PHN (500 mg/L) & PYR (250 mg/L) Parameters ü Agitation ü Temp. ü p. H ü Inoc. vol. ü Salinity Single factor optimization 6 Ø Bacteria resting cells preparation, Ø Culturing cells @ variable factors, Ø PAH quantification after 24 hours 3, Ø Selecting significant deg. response Parameters ü SFO selected factors range 6. 7. Djekrif et al. , (2006) Silver & Roberto, (2001) Response surface optimization 7 ü Experimental design ü Modeling ü Optimization
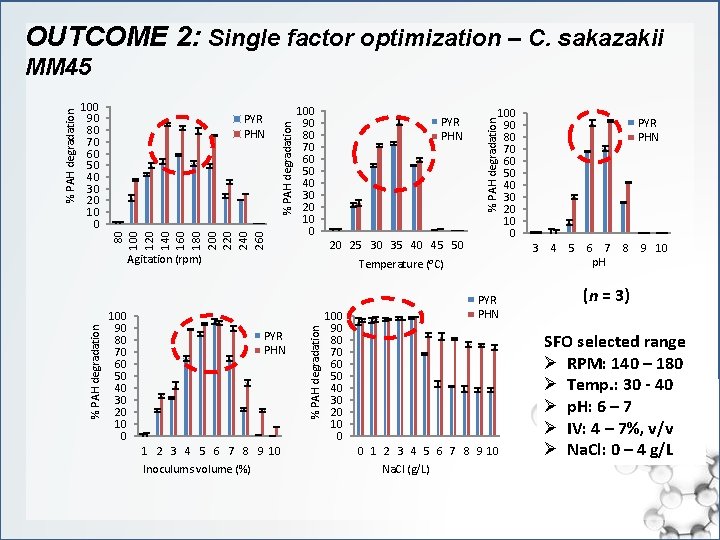
OUTCOME 2: Single factor optimization – C. sakazakii 80 100 120 140 160 180 200 220 240 260 100 90 80 70 60 50 40 30 20 10 0 20 25 30 35 40 45 50 Agitation (rpm) PYR PHN 1 2 3 4 5 6 7 8 9 10 Inoculums volume (%) 100 90 80 70 60 50 40 30 20 10 0 PYR PHN 100 90 80 70 60 50 40 30 20 10 0 0 1 2 3 4 5 6 7 8 9 10 Na. Cl (g/L) PYR PHN 3 4 5 6 7 8 9 10 p. H Temperature (⁰C) % PAH degradation 100 90 80 70 60 50 40 30 20 10 0 PYR PHN % PAH degradation 100 90 80 70 60 50 40 30 20 10 0 % PAH degradation MM 45 (n = 3) SFO selected range Ø RPM: 140 – 180 Ø Temp. : 30 - 40 Ø p. H: 6 – 7 Ø IV: 4 – 7%, v/v Ø Na. Cl: 0 – 4 g/L
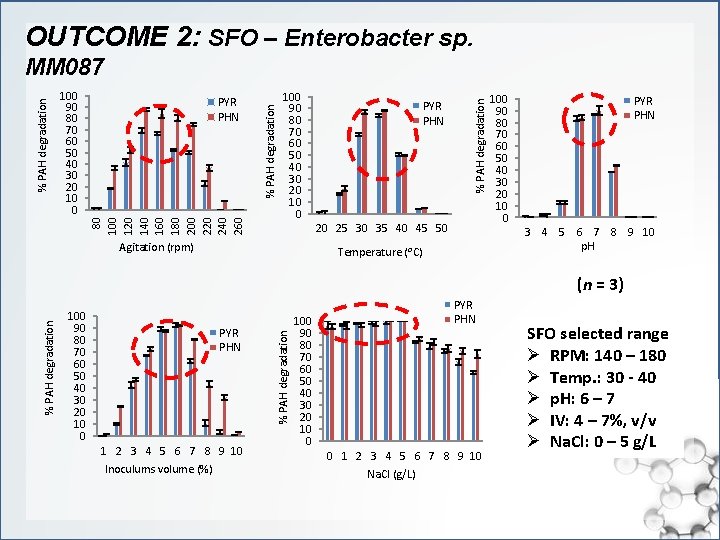
OUTCOME 2: SFO – Enterobacter sp. 100 90 80 70 60 50 40 30 20 10 0 PYR PHN % PAH degradation 100 90 80 70 60 50 40 30 20 10 0 80 100 120 140 160 180 200 220 240 260 % PAH degradation MM 087 20 25 30 35 40 45 50 Agitation (rpm) 100 90 80 70 60 50 40 30 20 10 0 PYR PHN 3 4 5 6 7 8 9 10 p. H Temperature (⁰C) 100 90 80 70 60 50 40 30 20 10 0 PYR PHN 1 2 3 4 5 6 7 8 9 10 Inoculums volume (%) % PAH degradation (n = 3) 100 90 80 70 60 50 40 30 20 10 0 PYR PHN 0 1 2 3 4 5 6 7 8 9 10 Na. Cl (g/L) SFO selected range Ø RPM: 140 – 180 Ø Temp. : 30 - 40 Ø p. H: 6 – 7 Ø IV: 4 – 7%, v/v Ø Na. Cl: 0 – 5 g/L
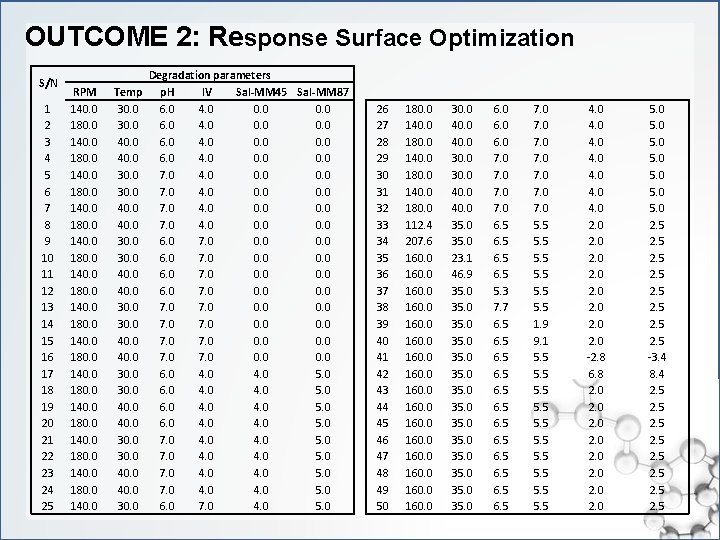
OUTCOME 2: Response Surface Optimization S/N 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 RPM 140. 0 180. 0 140. 0 Temp 30. 0 40. 0 30. 0 Degradation parameters p. H IV Sal-MM 45 Sal-MM 87 6. 0 4. 0 0. 0 7. 0 4. 0 0. 0 6. 0 7. 0 0. 0 6. 0 4. 0 5. 0 7. 0 4. 0 5. 0 6. 0 7. 0 4. 0 5. 0 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 180. 0 140. 0 180. 0 112. 4 207. 6 160. 0 160. 0 30. 0 40. 0 35. 0 23. 1 46. 9 35. 0 35. 0 6. 0 7. 0 6. 5 5. 3 7. 7 6. 5 6. 5 7. 0 7. 0 5. 5 5. 5 1. 9 9. 1 5. 5 5. 5 4. 0 4. 0 2. 0 -2. 8 6. 8 2. 0 2. 0 5. 0 2. 5 2. 5 -3. 4 8. 4 2. 5 2. 5
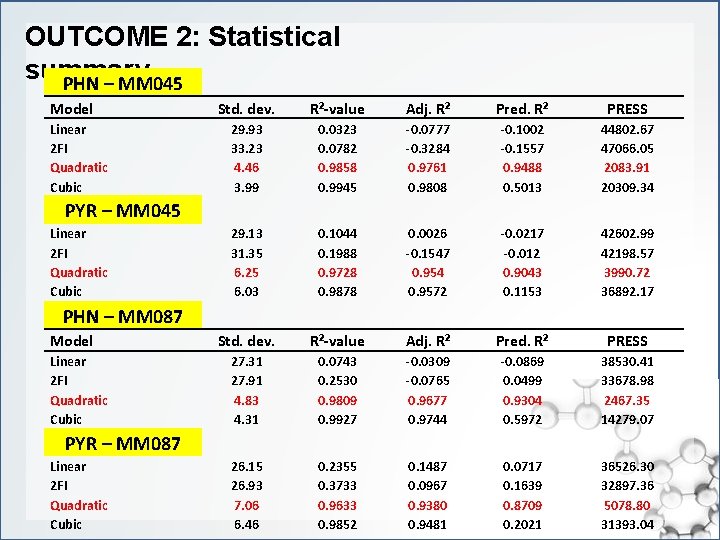
OUTCOME 2: Statistical summary PHN – MM 045 Model Linear 2 FI Quadratic Cubic Std. dev. R²-value Adj. R² Pred. R² PRESS 29. 93 33. 23 4. 46 3. 99 0. 0323 0. 0782 0. 9858 0. 9945 -0. 0777 -0. 3284 0. 9761 0. 9808 -0. 1002 -0. 1557 0. 9488 0. 5013 44802. 67 47066. 05 2083. 91 20309. 34 29. 13 31. 35 6. 25 6. 03 0. 1044 0. 1988 0. 9728 0. 9878 0. 0026 -0. 1547 0. 954 0. 9572 -0. 0217 -0. 012 0. 9043 0. 1153 42602. 99 42198. 57 3990. 72 36892. 17 Std. dev. R²-value Adj. R² Pred. R² PRESS 27. 31 27. 91 4. 83 4. 31 0. 0743 0. 2530 0. 9809 0. 9927 -0. 0309 -0. 0765 0. 9677 0. 9744 -0. 0869 0. 0499 0. 9304 0. 5972 38530. 41 33678. 98 2467. 35 14279. 07 26. 15 26. 93 7. 06 6. 46 0. 2355 0. 3733 0. 9633 0. 9852 0. 1487 0. 0967 0. 9380 0. 9481 0. 0717 0. 1639 0. 8709 0. 2021 36526. 30 32897. 36 5078. 80 31393. 04 PYR – MM 045 Linear 2 FI Quadratic Cubic PHN – MM 087 Model Linear 2 FI Quadratic Cubic PYR – MM 087 Linear 2 FI Quadratic Cubic
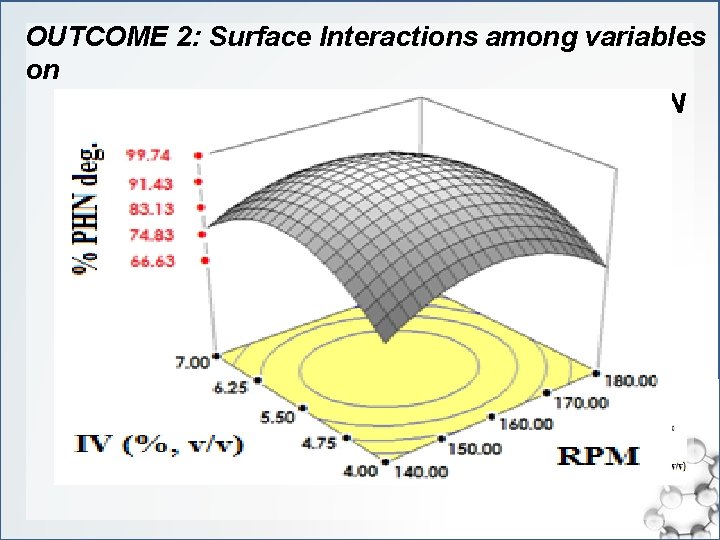
OUTCOME 2: Surface Interactions among variables on C. sakazakii MM 045 response to PHN
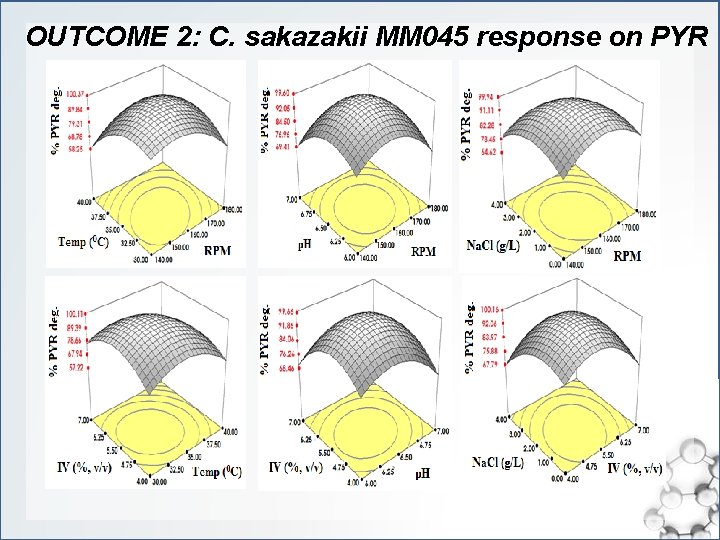
OUTCOME 2: C. sakazakii MM 045 response on PYR
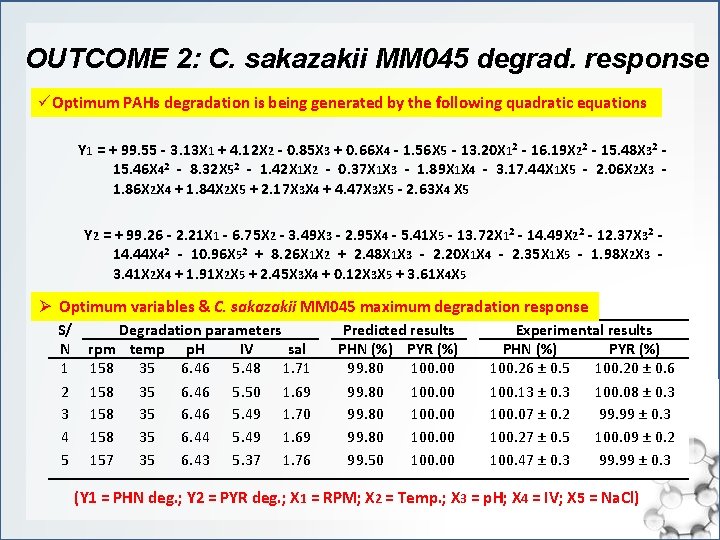
OUTCOME 2: C. sakazakii MM 045 degrad. response üOptimum PAHs degradation is being generated by the following quadratic equations Y 1 = + 99. 55 - 3. 13 X 1 + 4. 12 X 2 - 0. 85 X 3 + 0. 66 X 4 - 1. 56 X 5 - 13. 20 X 12 - 16. 19 X 22 - 15. 48 X 32 - 15. 46 X 42 - 8. 32 X 52 - 1. 42 X 1 X 2 - 0. 37 X 1 X 3 - 1. 89 X 1 X 4 - 3. 17. 44 X 1 X 5 - 2. 06 X 2 X 3 - 1. 86 X 2 X 4 + 1. 84 X 2 X 5 + 2. 17 X 3 X 4 + 4. 47 X 3 X 5 - 2. 63 X 4 X 5 Y 2 = + 99. 26 - 2. 21 X 1 - 6. 75 X 2 - 3. 49 X 3 - 2. 95 X 4 - 5. 41 X 5 - 13. 72 X 12 - 14. 49 X 22 - 12. 37 X 32 - 14. 44 X 42 - 10. 96 X 52 + 8. 26 X 1 X 2 + 2. 48 X 1 X 3 - 2. 20 X 1 X 4 - 2. 35 X 1 X 5 - 1. 98 X 2 X 3 - 3. 41 X 2 X 4 + 1. 91 X 2 X 5 + 2. 45 X 3 X 4 + 0. 12 X 3 X 5 + 3. 61 X 4 X 5 Ø Optimum variables & C. sakazakii MM 045 maximum degradation response S/ N 1 2 3 4 5 Degradation parameters rpm temp p. H IV sal 158 35 6. 46 5. 48 1. 71 158 35 6. 46 5. 50 1. 69 158 35 6. 46 5. 49 1. 70 158 35 6. 44 5. 49 1. 69 157 35 6. 43 5. 37 1. 76 Predicted results PHN (%) PYR (%) 99. 80 100. 00 99. 50 100. 00 Experimental results PHN (%) PYR (%) 100. 26 ± 0. 5 100. 20 ± 0. 6 100. 13 ± 0. 3 100. 08 ± 0. 3 100. 07 ± 0. 2 99. 99 ± 0. 3 100. 27 ± 0. 5 100. 09 ± 0. 2 100. 47 ± 0. 3 99. 99 ± 0. 3 (Y 1 = PHN deg. ; Y 2 = PYR deg. ; X 1 = RPM; X 2 = Temp. ; X 3 = p. H; X 4 = IV; X 5 = Na. Cl)
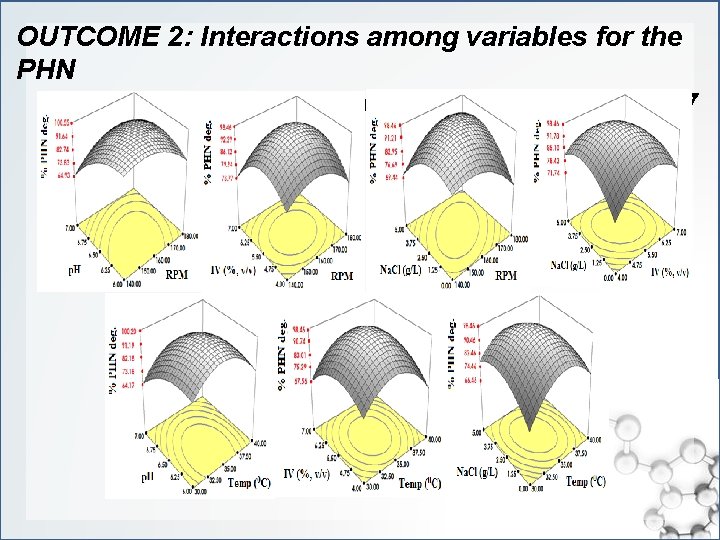
OUTCOME 2: Interactions among variables for the PHN degradation by Enterobacter sp. MM 87
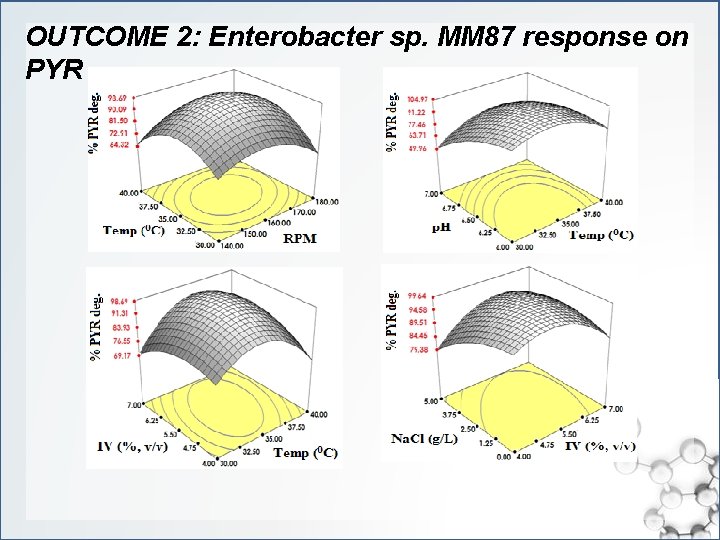
OUTCOME 2: Enterobacter sp. MM 87 response on PYR
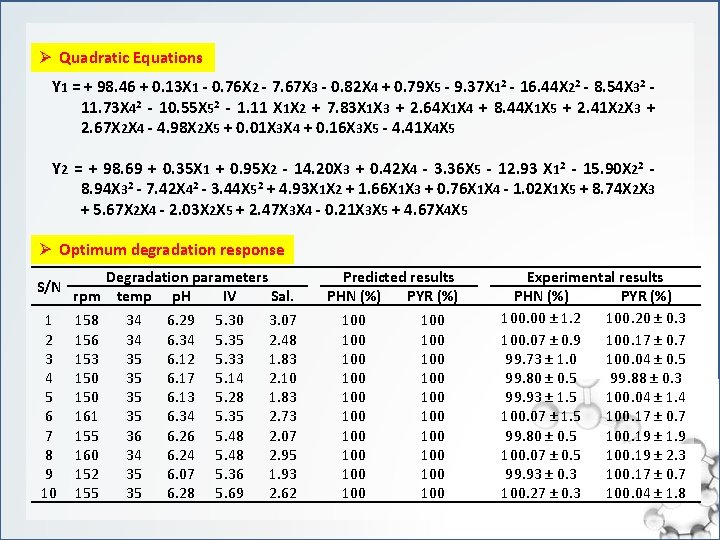
Ø Quadratic Equations Y 1 = + 98. 46 + 0. 13 X 1 - 0. 76 X 2 - 7. 67 X 3 - 0. 82 X 4 + 0. 79 X 5 - 9. 37 X 12 - 16. 44 X 22 - 8. 54 X 32 - 11. 73 X 42 - 10. 55 X 52 - 1. 11 X 1 X 2 + 7. 83 X 1 X 3 + 2. 64 X 1 X 4 + 8. 44 X 1 X 5 + 2. 41 X 2 X 3 + 2. 67 X 2 X 4 - 4. 98 X 2 X 5 + 0. 01 X 3 X 4 + 0. 16 X 3 X 5 - 4. 41 X 4 X 5 Y 2 = + 98. 69 + 0. 35 X 1 + 0. 95 X 2 - 14. 20 X 3 + 0. 42 X 4 - 3. 36 X 5 - 12. 93 X 12 - 15. 90 X 22 - 8. 94 X 32 - 7. 42 X 42 - 3. 44 X 52 + 4. 93 X 1 X 2 + 1. 66 X 1 X 3 + 0. 76 X 1 X 4 - 1. 02 X 1 X 5 + 8. 74 X 2 X 3 + 5. 67 X 2 X 4 - 2. 03 X 2 X 5 + 2. 47 X 3 X 4 - 0. 21 X 3 X 5 + 4. 67 X 4 X 5 Ø Optimum degradation response S/N Degradation parameters rpm temp p. H IV Sal. 1 2 3 4 5 6 7 8 9 10 158 156 153 150 161 155 160 152 155 34 34 35 35 36 34 35 35 6. 29 6. 34 6. 12 6. 17 6. 13 6. 34 6. 26 6. 24 6. 07 6. 28 5. 30 5. 35 5. 33 5. 14 5. 28 5. 35 5. 48 5. 36 5. 69 3. 07 2. 48 1. 83 2. 10 1. 83 2. 73 2. 07 2. 95 1. 93 2. 62 Predicted results PHN (%) PYR (%) 100 100 100 100 100 Experimental results PHN (%) PYR (%) 100. 00 ± 1. 2 100. 20 ± 0. 3 100. 07 ± 0. 9 100. 17 ± 0. 7 99. 73 ± 1. 0 100. 04 ± 0. 5 99. 80 ± 0. 5 99. 88 ± 0. 3 99. 93 ± 1. 5 100. 04 ± 1. 4 100. 07 ± 1. 5 100. 17 ± 0. 7 99. 80 ± 0. 5 100. 19 ± 1. 9 100. 07 ± 0. 5 100. 19 ± 2. 3 99. 93 ± 0. 3 100. 17 ± 0. 7 100. 27 ± 0. 3 100. 04 ± 1. 8
Objective 2: Conclusion v RSM optimization significantly enhanced PAHs biodegradation responses of C. sakazakii MM 045 and Enterobacter sp. MM 087 from < 70% degradations to 100% within 24 hours How do we confirm such effective degradation responses?

OBJECTIVE 3: Identification of PHN & PYR degrading intermediates QUESTION ? v What strategy can be applied to ascertain the effective PAHs degradation by MM 045 & MM 087 ? OBJECTIVE 3 ! v Identification of phenanthrene and pyrene biodegradation metabolites HOW TO ACHIEVE IT? v Gas chromatography-mass spectrophotometer (GC-MS) analyses SIGNIFICANT v To establish PAHs degradation pathway followed by C. sakazakii MM 045 & Enterobacter sp. MM 087
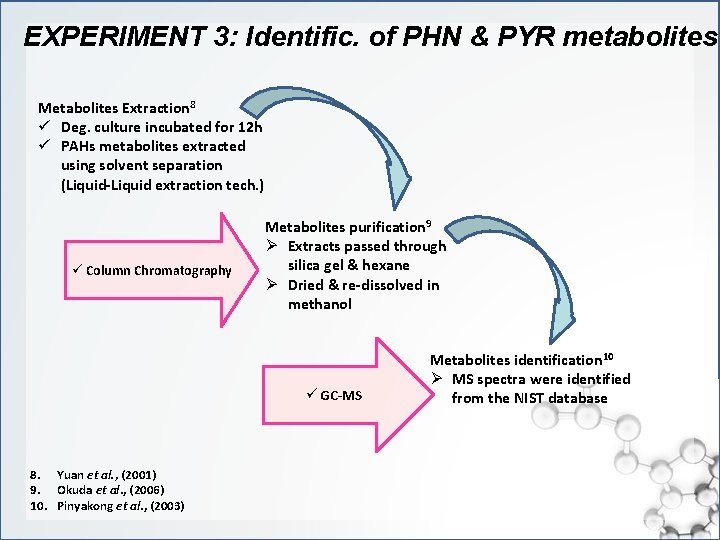
EXPERIMENT 3: Identific. of PHN & PYR metabolites Metabolites Extraction 8 ü Deg. culture incubated for 12 h ü PAHs metabolites extracted using solvent separation (Liquid-Liquid extraction tech. ) ü Column Chromatography Metabolites purification 9 Ø Extracts passed through silica gel & hexane Ø Dried & re-dissolved in methanol ü GC-MS 8. Yuan et al. , (2001) 9. Okuda et al. , (2006) 10. Pinyakong et al. , (2003) Metabolites identification 10 Ø MS spectra were identified from the NIST database
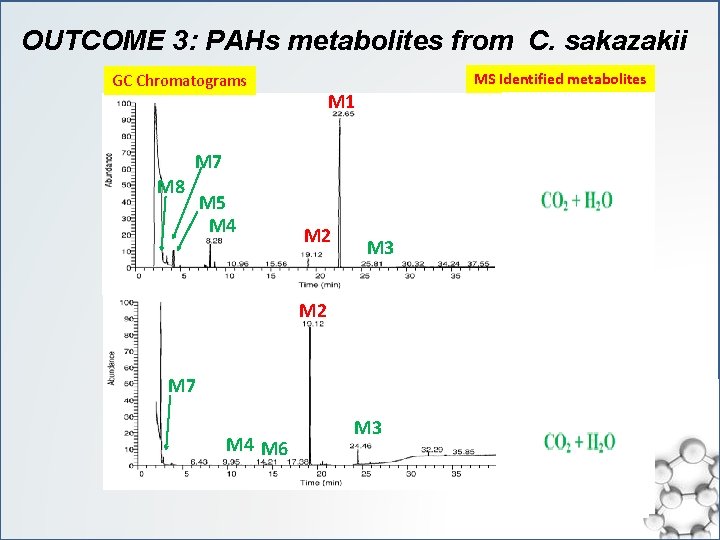
OUTCOME 3: PAHs metabolites from C. sakazakii MS Identified metabolites GC Chromatograms M 1 M 7 M 8 M 5 M 4 M 2 M 3 M 2 M 7 M 4 M 6 M 3
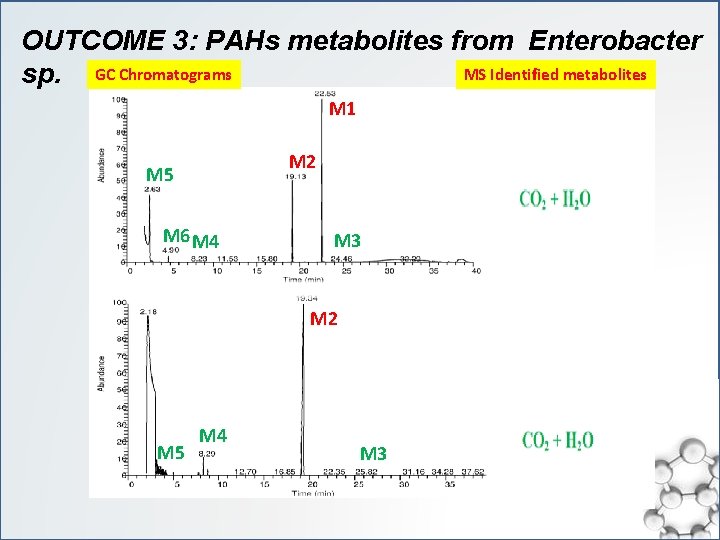
OUTCOME 3: PAHs metabolites from Enterobacter MS Identified metabolites sp. GC Chromatograms M 1 M 2 M 5 M 6 M 4 M 3 M 2 M 5 M 4 M 3
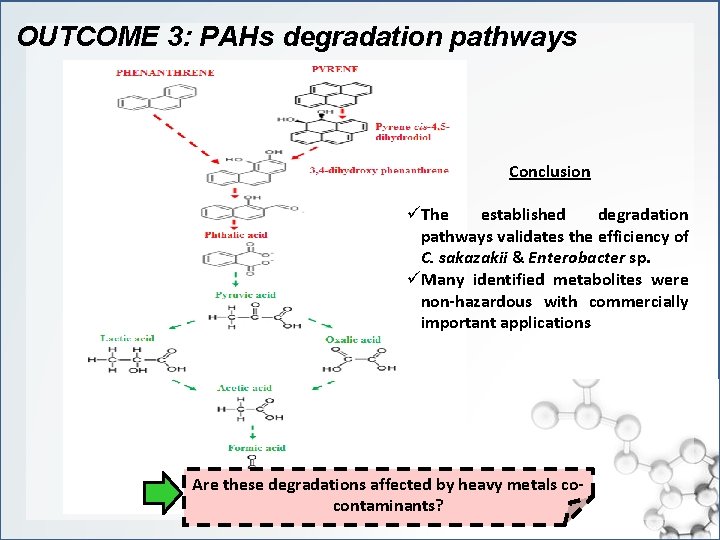
OUTCOME 3: PAHs degradation pathways Conclusion üThe established degradation pathways validates the efficiency of C. sakazakii & Enterobacter sp. üMany identified metabolites were non-hazardous with commercially important applications Are these degradations affected by heavy metals cocontaminants?
OBJECTIVE 4: Effects of heavy metals on PAHs degrading consortium QUESTION ? v What impact does heavy metals co-contaminants have on the degradation efficiency of MM 045 & MM 087? OBJECTIVE 4 ! v To determine the effect of heavy metals co-contaminants on PHN & PYR biodegrading consortium. HOW DO WE ACHIEVE THIS? v Formulating bacteria consortium using MM 045 & MM 087 v PAHs deg. effects of heavy metals on bacteria consortium SIGNIFICANT v To confirm heavy metals resistance during PAHs degradation by C. sakazakii MM 45 & Enterobacter sp. MM 87
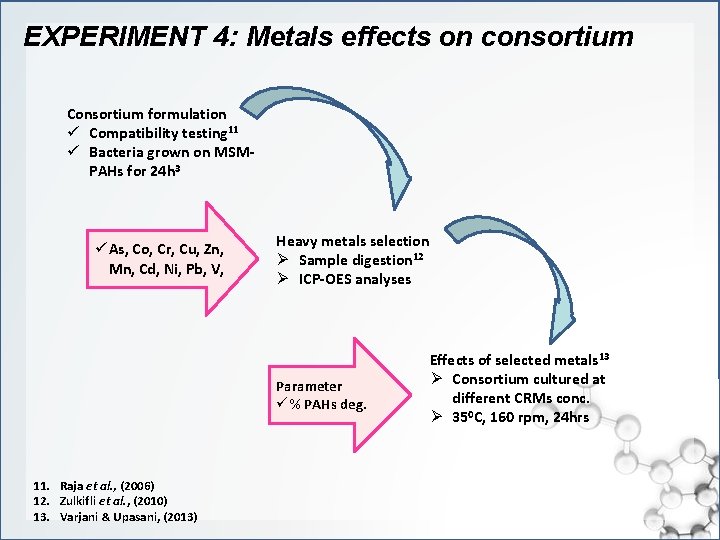
EXPERIMENT 4: Metals effects on consortium Consortium formulation ü Compatibility testing 11 ü Bacteria grown on MSMPAHs for 24 h 3 ü As, Co, Cr, Cu, Zn, Mn, Cd, Ni, Pb, V, Heavy metals selection Ø Sample digestion 12 Ø ICP-OES analyses Parameter ü % PAHs deg. 11. Raja et al. , (2006) 12. Zulkifli et al. , (2010) 13. Varjani & Upasani, (2013) Effects of selected metals 13 Ø Consortium cultured at different CRMs conc. Ø 350 C, 160 rpm, 24 hrs

OUTCOME 4: Consortium selection S/N 1 2 3 4 5 6* 7 8 9 10 Consortium (%, v/v) 1: 4 2: 3 3: 2 4: 1 1: 3 *2 : 2 3: 1 1: 2 2: 1 1: 1 PHN deg. (%) 101. 0 ± 1. 40 100. 2 ± 0. 72 100. 5 ± 0. 50 100. 2 ± 0. 31 98. 3 ± 0. 90 100. 1 ± 0. 81 98. 9 ± 0. 70 49. 9 ± 0. 90 47. 9 ± 0. 50 0. 00 PYR deg. (%) 100. 7 ± 0. 71 100. 0 ± 9. 2 100. 5 ± 1. 38 100. 2 ± 1. 17 94. 3 ± 1. 50 100. 0 ± 0. 92 94. 9 ± 2. 12 63. 9 ± 1. 67 52. 7 ± 3. 34 10. 4 ± 1. 43 Ø C. sakazakii MM 045 (2%, v/v) + Enterobacter sp. MM 087 (2%, v/v)
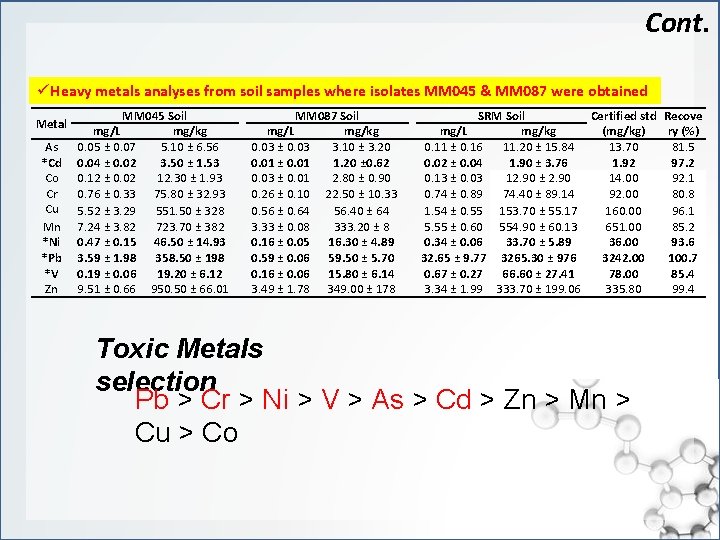
Cont. ü Heavy metals analyses from soil samples where isolates MM 045 & MM 087 were obtained Metal As *Cd Co Cr Cu Mn *Ni *Pb *V Zn MM 045 Soil mg/L mg/kg 0. 05 ± 0. 07 5. 10 ± 6. 56 0. 04 ± 0. 02 3. 50 ± 1. 53 0. 12 ± 0. 02 12. 30 ± 1. 93 0. 76 ± 0. 33 75. 80 ± 32. 93 5. 52 ± 3. 29 551. 50 ± 328 7. 24 ± 3. 82 723. 70 ± 382 0. 47 ± 0. 15 46. 50 ± 14. 93 3. 59 ± 1. 98 358. 50 ± 198 0. 19 ± 0. 06 19. 20 ± 6. 12 9. 51 ± 0. 66 950. 50 ± 66. 01 MM 087 Soil mg/L mg/kg 0. 03 ± 0. 03 3. 10 ± 3. 20 0. 01 ± 0. 01 1. 20 ± 0. 62 0. 03 ± 0. 01 2. 80 ± 0. 90 0. 26 ± 0. 10 22. 50 ± 10. 33 0. 56 ± 0. 64 56. 40 ± 64 3. 33 ± 0. 08 333. 20 ± 8 0. 16 ± 0. 05 16. 30 ± 4. 89 0. 59 ± 0. 06 59. 50 ± 5. 70 0. 16 ± 0. 06 15. 80 ± 6. 14 3. 49 ± 1. 78 349. 00 ± 178 SRM Soil Certified std Recove mg/L mg/kg (mg/kg) ry (%) 0. 11 ± 0. 16 11. 20 ± 15. 84 13. 70 81. 5 0. 02 ± 0. 04 1. 90 ± 3. 76 1. 92 97. 2 0. 13 ± 0. 03 12. 90 ± 2. 90 14. 00 92. 1 0. 74 ± 0. 89 74. 40 ± 89. 14 92. 00 80. 8 1. 54 ± 0. 55 153. 70 ± 55. 17 160. 00 96. 1 5. 55 ± 0. 60 554. 90 ± 60. 13 651. 00 85. 2 0. 34 ± 0. 06 33. 70 ± 5. 89 36. 00 93. 6 32. 65 ± 9. 77 3265. 30 ± 976 3242. 00 100. 7 0. 67 ± 0. 27 66. 60 ± 27. 41 78. 00 85. 4 3. 34 ± 1. 99 333. 70 ± 199. 06 335. 80 99. 4 Toxic Metals selection Pb > Cr > Ni > V > As > Cd > Zn > Mn > Cu > Co
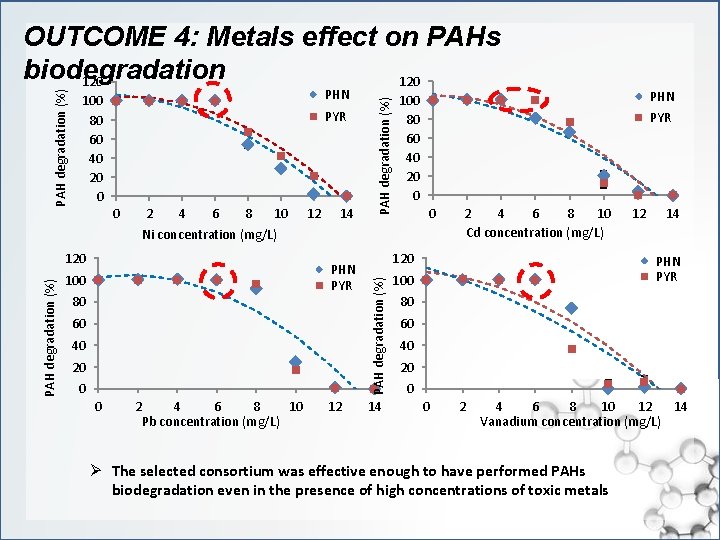
PHN 100 80 60 40 20 0 PYR 0 2 4 6 8 10 12 14 PAH degradation (%) OUTCOME 4: Metals effect on PAHs biodegradation 120 0 Ni concentration (mg/L) PHN PYR 100 80 60 40 20 0 0 2 4 6 8 10 Pb concentration (mg/L) 12 2 4 6 8 10 Cd concentration (mg/L) 120 PAH degradation (%) 120 14 PHN PYR 100 80 60 40 20 0 12 14 PHN PYR 100 80 60 40 20 0 0 2 4 6 8 10 12 14 Vanadium concentration (mg/L) Ø The selected consortium was effective enough to have performed PAHs biodegradation even in the presence of high concentrations of toxic metals

Conclusio n v Effective PAHs degrading bacteria were identified as C. sakazakii MM 045 and Enterobacter sp. MM 087 v The degradation effectiveness of both bacteria was significantly enhanced by SFO and RSM v Both bacteria degraded 100% of 500 mg/L PHN & 250 mg/L PYR in just 24 h v Such degradation response was successfully validated by establishing the bacteria degradation pathway. v The bacteria responses were further complimented with heavy metals resistance during PAHs biodegradation C. Sakazakii MM 045 and Enterobacter sp. MM 087 could be used for effective PAHs bioremoval

PUBLICATIONS 1. Umar, Z. D. , Aziz, N. A. A. , Zulkifli, S. Z. , & Muskhazli, M. (2016): Identification of phenanthrene and pyrene degrading bacteria from used engine oil contaminated soil. International Journal of Scientific & Engineering Research, 7, 680 -686. 2. Umar, Z. D. , Aziz, N. A. A. , Zulkifli, S. Z. , & Muskhazli, M. (2017): Rapid biodegradation of polycyclic aromatic hydrocarbons using effective Cronobacter sakazakii MM 045 (KT 933253). Methods. X, 4, 104 -117. 3. Umar, Z. D. , Aziz, N. A. A. , Zulkifli, S. Z. , & Muskhazli, M. (in press): Effective phenanthrene and pyrene biodegradation using Enterobacter sp. MM 087 (KT 933254) isolated from used engine oil contaminated soil. Egyptian Journal of Petroleum, vol. 27. doi: 10. 1016/j. ejpe. 2017. 06. 001 4. Umar, Z. D. , Aziz, N. A. A. , Zulkifli, S. Z. , & Muskhazli, M. (submitted): Efficiency of effective polycyclic aromatic hydrocarbon (PAHs) degrading consortium in resisting toxic heavy metals during PAHs biodegradation (submitted)

The Research was conducted at the: • Universiti Putra Malaysia (2014 to 2017) Under the Supervisions of • Associate Prof. Muskhazli Mustafa (Main Super. ) • Associate Prof. Nur Azwady Abd. Aziz (Co Super • Dr Syaizwan Zahmir Zulkifli (Co Super. )

Acknowledgements v The Management of UMYUK for the Ph. D Sponsorship under the Staff Development Program v The Putra Grant (GP-IPS/2016/9489800) for Funding the research

Reference s • Tao, X. Q. , Lu, G. N. , Dang, Z. , Yang, C. , & Yi, X. Y. (2007): A phenanthrene-degrading strain Sphingomonas sp. GY 2 B isolated from contaminated soils. Process Biochemistry, 42, 401 -408. • Kiyohara, H. , Nagao, K. , & Yana, K. (1982): Rapid screen for bacteria degrading water-insoluble, solid hydrocarbons on agar plates. Applied & Environmental Microbiology , 43, 454 -457. • Hanson, K. G. , Desai, J. D. , & Desai, A. J. (1993): A rapid and simple screening technique for potential crude oil degrading microorganisms. Biotechnology Techniques, 7, 745 -748 • Sogani, M. , Mathur, N. , Sharma, P. , & Bhatnagar, P. (2012): Comparison of immobilized whole resting cells in different matrices vis-avis free cells of Bacillus megaterium for acyltransferase activity. Journal of Environmental Research and Development, 6 , 695 -701. • Kumar, S. , Stecher, G. , & Tamura, K. (2016): MEGA 7: Molecular Evolutionary Genetics Analysis version 7. 0 for bigger datasets. Molecular Biology & Evolution, 33, 1870 -1874. • Djekrif-Dakhmouche, S. , Gheribi-Aoulmi, Z. , Meraihi, Z. , & Bennamoun, L. (2006): Application of a statistical design to the optimization of culture medium for α-amylase production by Aspergillus niger ATCC 16404 grown on orange waste powder. Journal of Food Engineering, 73, 190– 197. • Silva, C. J. S. M. , & Roberto, I. C. , (2001): Optimization of xylitol production by Candida guilliermondii FTI 20037 using response surface methodology. Process Biochemistry, 361, 119 -124. • Yuan, S. Y. , Chang, J. S. , Yen, J. H. , & Chang, B. V. (2001): Biodegradation of phenanthrene in river sediment. Chemosphere, 43, 273 -278. • Okuda, T. , Naoi, D. , Tenmoku, M. , Tanaka, S. , He, K. , Ma, Y. & Zhang, D. (2006): Polycyclic aromatic hydrcarbns in the aerosol in Beijing, China, measured by aminopropylsilane chemically-bonded stationary-phase column chromatography and HPLC/fluorescence detection. Chemosphere, 65, 427 -435. • Pinyakong, O. , Habe, H. , Yoshida, T. , Nojiri, H. , & Omori, T. (2003): Identification of three novel salicylate 1 -hydroxylases involved in the phenanthrene degradation of Sphingobium sp. strain P 2. Biochemical and Biophysical Research Communications , 301, 350 -357. • Raja, P. , Uma, S. , Gopal, H. , & Govindarajan, K. (2006): Impact of bio inoculants consortium on rice root exudates, biological nitrogen fixation and plant growth. Journal Biological Science, 6, 815 -823. • Zulkifli, S. Z. , Ismail, A. , Mohamat-Yusuff, F. , Arai, T. , & Miyazaki, N. (2010): Johor Strait as a hotspot for trace elements contamination in Peninsular Malaysia. Bulletin of Environmental Contamination and Toxicology , 84, 568 -573. • Varjani, S. J. , & Upasani, V. N. (2013): Comparative studies on bacterial consortia for hydrocarbon degradation. International Journal of Innovative Research in Science, Engineering and Technology , 2, 10. • Vela, N. , Martínez-Menchón, M. , Navarro, G. , Pérez-Lucas, G. , & Navarro, S. (2012): Removal of polycyclic aromatic hydrocarbons from groundwater by heterogeneous photocatalysis under natural sunlight. Journal of Photochemistry and Photobiology A: Chemistry , 232, 32 -40.

• Vidali, M. (2001): Bioremediation; an overview. Pure & Applied Chemistry, 73, 1163 -1172. • Shih, K. L. , & Lederberg, J. (1976): Chloramine mutagenesis in Bacillus subtilis. Science, 192, 1141 -1143. • Brinda, L. M. , Muthukumar, K. , & Velan, M. (2013): Optimization of minimal salt medium for efficient phenanthrene biodegradation by Mycoplana sp. MVMB 2 isolated from petroleum contaminated soil using factorial design experiments. CLEAN–Soil, Air, Water, 41, 51 -59. • Cycoń, M. , Żmijowska, A. , Wójcik, M. , & Piotrowska-Seget, Z. (2013): Biodegradation and bioremediation potential of diazinondegrading Serratia marcescens to remove other organophosphorus pesticides from soils. Journal of Environmental Management, 117, 7 -16. • Gillespie, S. H. , & Hawkey, P. M. (2006): Principles and Practice of Clinical Bacteriology, second edition , John Wiley Publisher, UK. • Kandhai, M. C. , Reij, M. W. , Gorris, L. G. , Guillaume-Gentil, O. , & Schothorst, M. (2004): Occurrence of Enterobacter sakazakii in food production environments and households. Lancet, 363, 39 -40.
- Slides: 39