Biochemistry Macromolecules Proteins Amino Acids Introduction to Biochemistry

Biochemistry Macromolecules, Proteins, Amino Acids

Introduction to Biochemistry Most biologically important macromolecules are: 1) proteins 2) polysaccharides (carbohydrates) 3) nucleic acids 4) Oils and fats All are polymers made via condensation

Amino Acids

Amino acids They can act as either an acid or a base They are Zwitterions that have both a positive and a negative charge Because of their ionic nature they have extremely high melting temperatures and so are crystalline solids, (all are colourless) As they are ionically charged they are soluble in
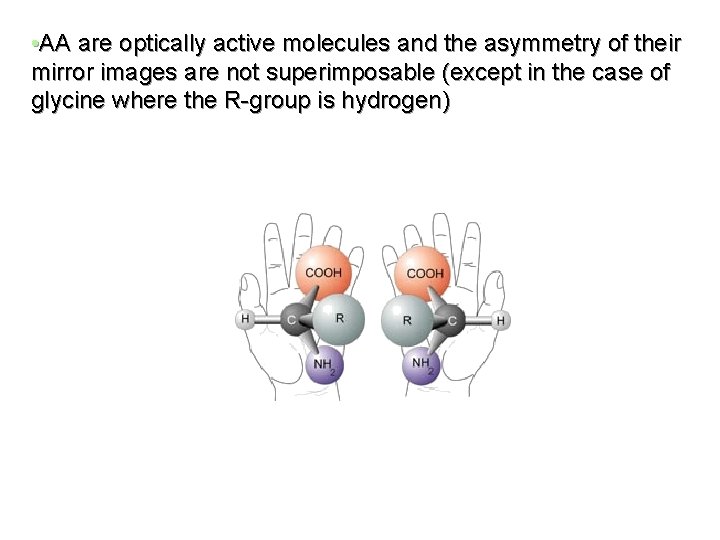
• AA are optically active molecules and the asymmetry of their mirror images are not superimposable (except in the case of glycine where the R-group is hydrogen)
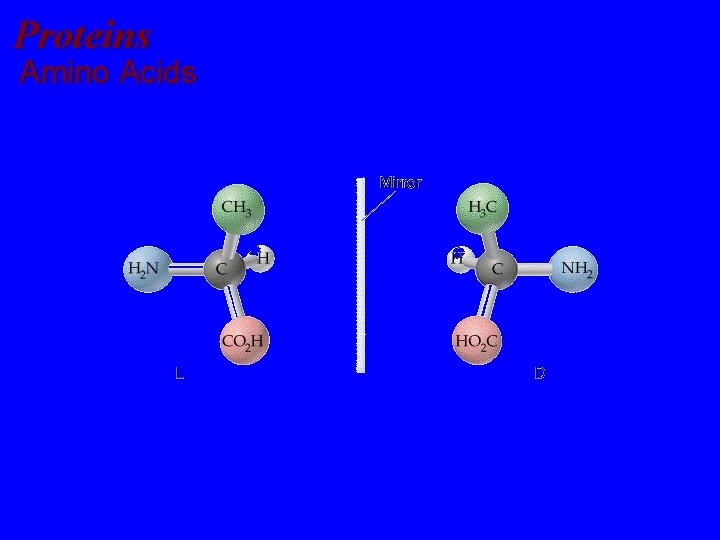
Proteins Amino Acids
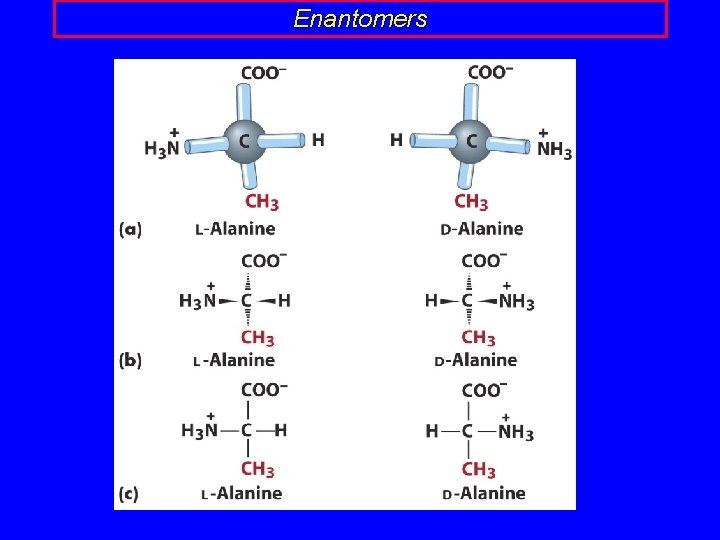
Enantomers
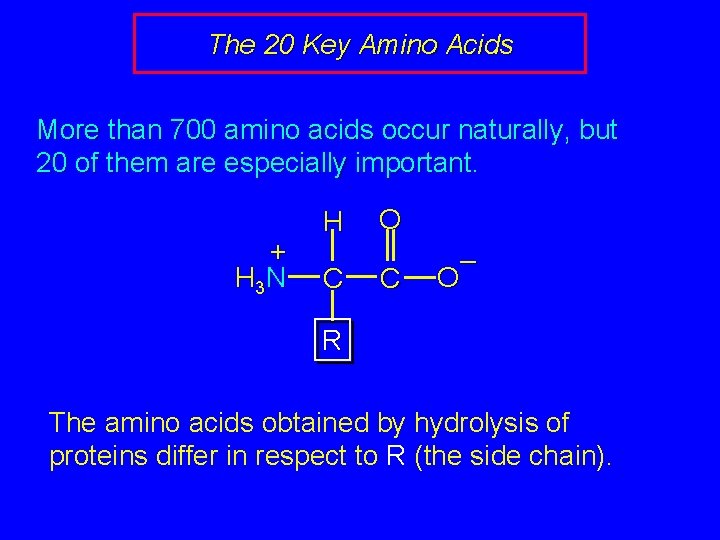
The 20 Key Amino Acids More than 700 amino acids occur naturally, but 20 of them are especially important. + H 3 N H C O C – O R The amino acids obtained by hydrolysis of proteins differ in respect to R (the side chain).
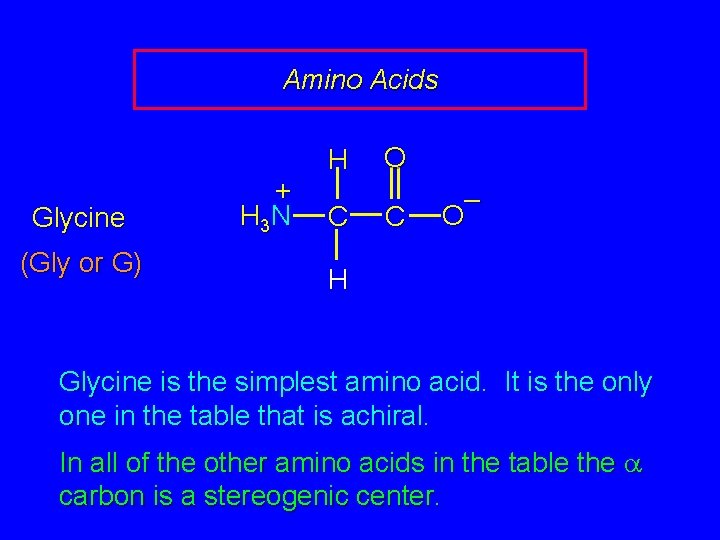
Amino Acids Glycine (Gly or G) + H 3 N H C O C – O H Glycine is the simplest amino acid. It is the only one in the table that is achiral. In all of the other amino acids in the table the carbon is a stereogenic center.
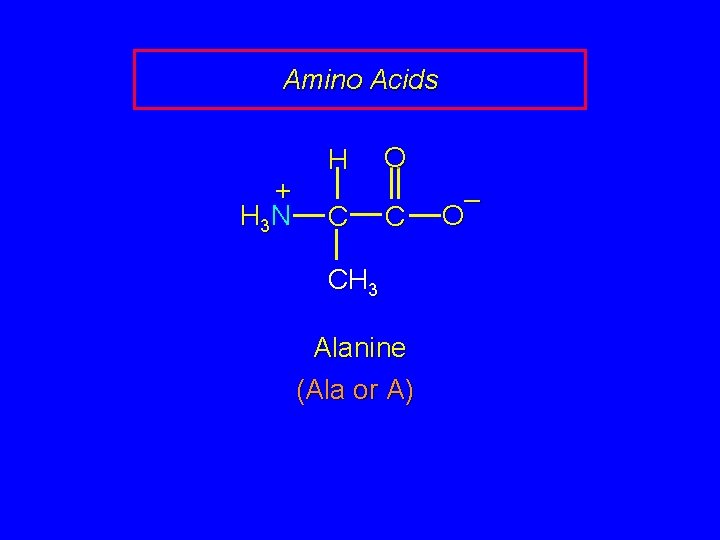
Amino Acids + H 3 N H C O C CH 3 Alanine (Ala or A) – O
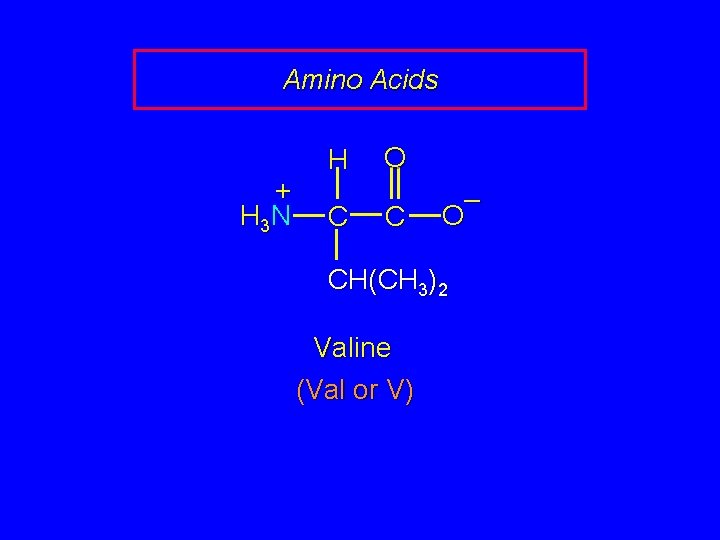
Amino Acids + H 3 N H C O C – O CH(CH 3)2 Valine (Val or V)
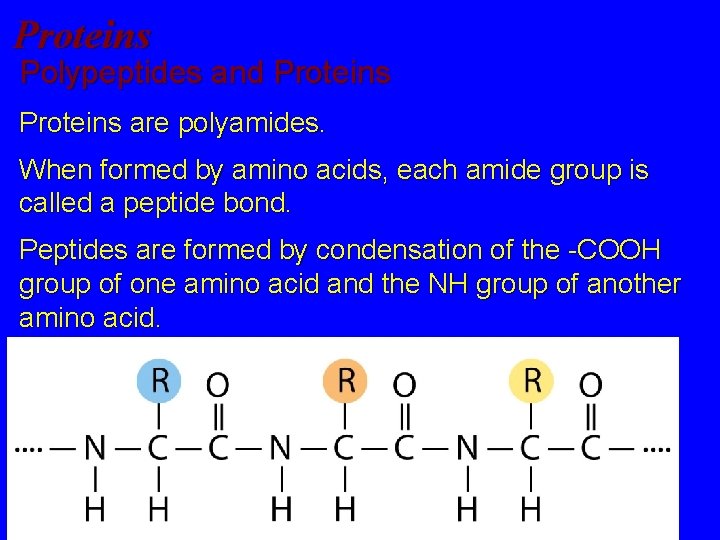
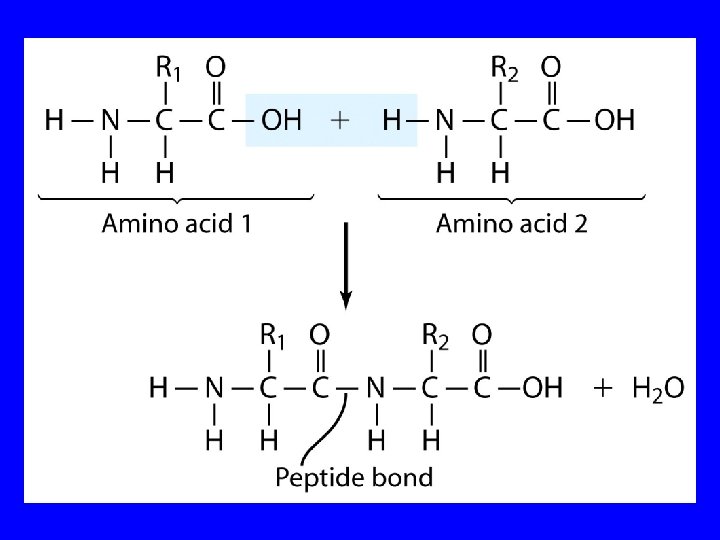
Proteins Polypeptides and Proteins are polyamides. When formed by amino acids, each amide group is called a peptide bond. Peptides are formed by condensation of the -COOH group of one amino acid and the NH group of another amino acid. .

Acid-Base Behavior of Amino Acids

Amino Acids While their name implies that amino acids are compounds that contain an —NH 2 group and a —CO 2 H group, these groups are actually present as —NH 3+ and —CO 2– respectively. How do we know this?

Properties of Glycine The properties of glycine: high melting point (when heated to 233°C it decomposes before it melts) solubility: soluble in water; not soluble in nonpolar solvent more consistent with this than this • • O • • + H 3 NCH 2 C • • O • • • – O • • • H 2 NCH 2 C • • OH • •

Properties of Glycine The properties of glycine: high melting point (when heated to 233°C it decomposes before it melts) solubility: soluble in water; not soluble in nonpolar solvent more consistent with this • • O • • + H 3 NCH 2 C • • • – O • • • Thus we know it is a zwitterion or dipolar ion

Acid-Base Properties of Glycine The zwitterionic structure of glycine also follows from considering its acid-base properties. A good way to think about this is to start with the structure of glycine in strongly acidic solution, say p. H = 1. At p. H = 1, glycine exists in its protonated form (a monocation). • • O • • + H 3 NCH 2 C • • OH • •
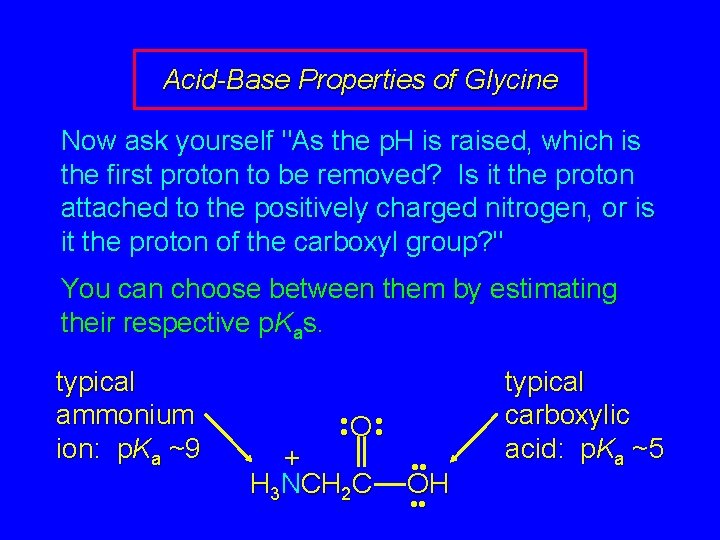
Acid-Base Properties of Glycine Now ask yourself "As the p. H is raised, which is the first proton to be removed? Is it the proton attached to the positively charged nitrogen, or is it the proton of the carboxyl group? " You can choose between them by estimating their respective p. Kas. typical ammonium ion: p. Ka ~9 • • O • • + H 3 NCH 2 C • • OH • • typical carboxylic acid: p. Ka ~5
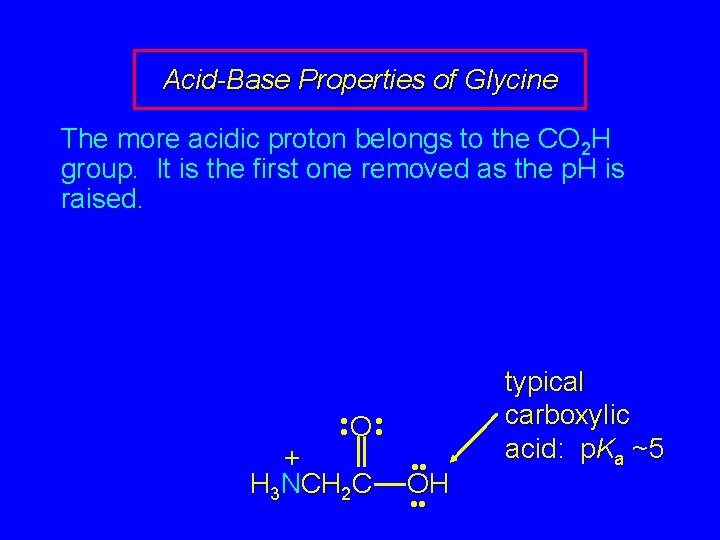
Acid-Base Properties of Glycine The more acidic proton belongs to the CO 2 H group. It is the first one removed as the p. H is raised. • • O • • + H 3 NCH 2 C • • OH • • typical carboxylic acid: p. Ka ~5
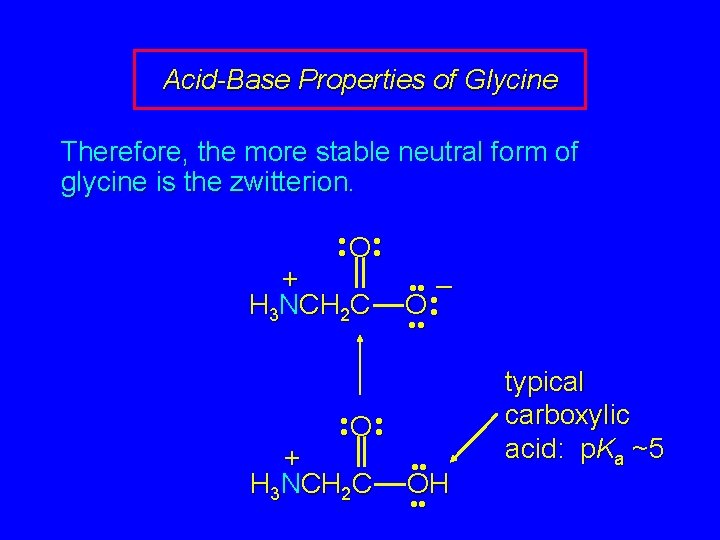
Acid-Base Properties of Glycine Therefore, the more stable neutral form of glycine is the zwitterion. • • O • • + H 3 NCH 2 C • • • – O • • • O • • + H 3 NCH 2 C • • OH • • typical carboxylic acid: p. Ka ~5
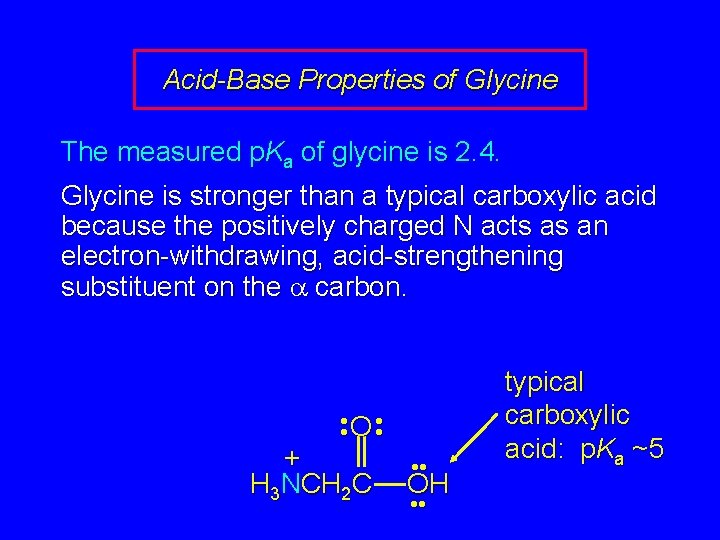
Acid-Base Properties of Glycine The measured p. Ka of glycine is 2. 4. Glycine is stronger than a typical carboxylic acid because the positively charged N acts as an electron-withdrawing, acid-strengthening substituent on the carbon. • • O • • + H 3 NCH 2 C • • OH • • typical carboxylic acid: p. Ka ~5
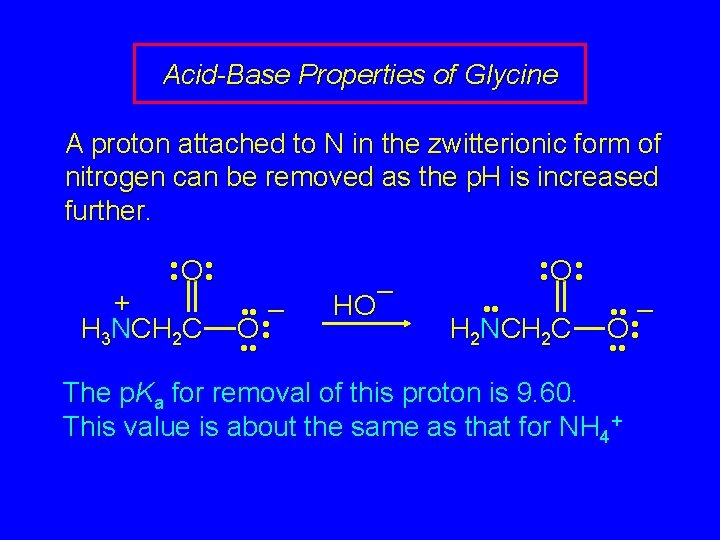
Acid-Base Properties of Glycine A proton attached to N in the zwitterionic form of nitrogen can be removed as the p. H is increased further. • • O • • + H 3 NCH 2 C • • • – O • • • HO – • • O • • H 2 NCH 2 C • • • – O • • • The p. Ka for removal of this proton is 9. 60. This value is about the same as that for NH 4+
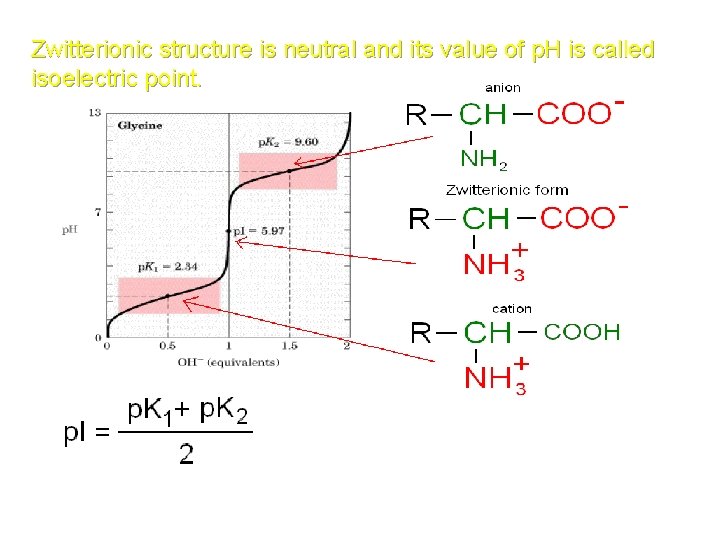
Zwitterionic structure is neutral and its value of p. H is called isoelectric point.
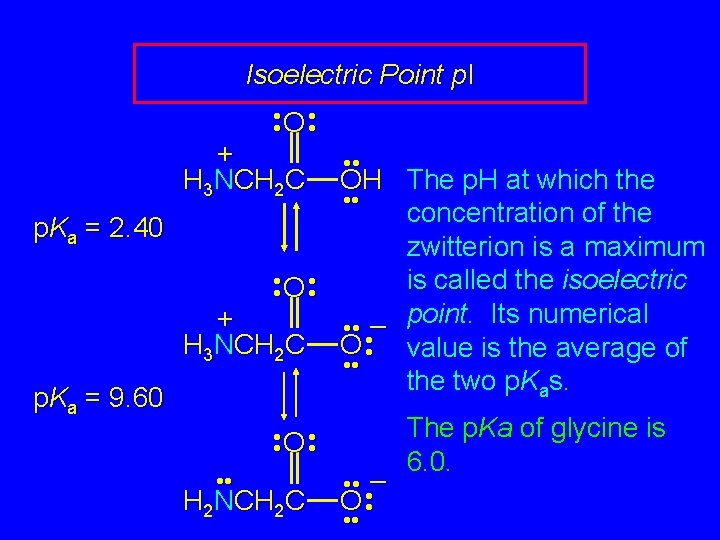
Isoelectric Point p. I • • O • • + H 3 NCH 2 C • • OH The p. H at which the • • concentration of the p. Ka = 2. 40 zwitterion is a maximum • • O • • is called the isoelectric point. Its numerical + – • • • H 3 NCH 2 C O • value is the average of • • the two p. Kas. p. Ka = 9. 60 The p. Ka of glycine is • • O • • 6. 0. • • • – H 2 NCH 2 C O • • •
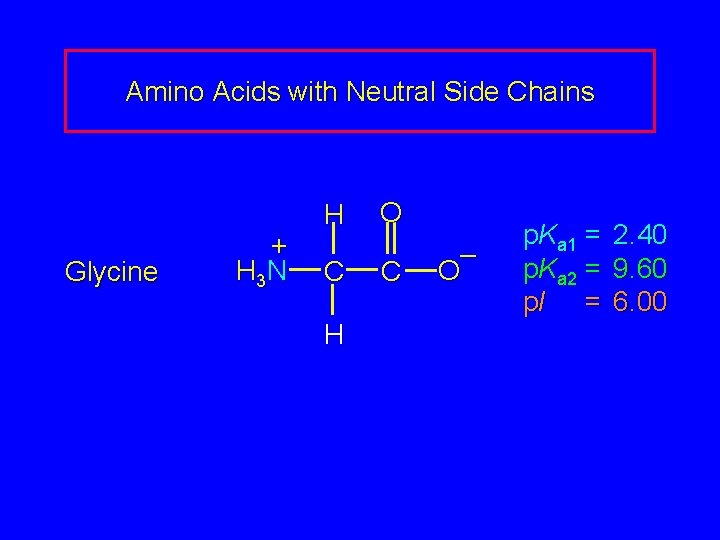
Amino Acids with Neutral Side Chains Glycine + H 3 N H C H O C – O p. Ka 1 = p. Ka 2 = p. I = 2. 40 9. 60 6. 00

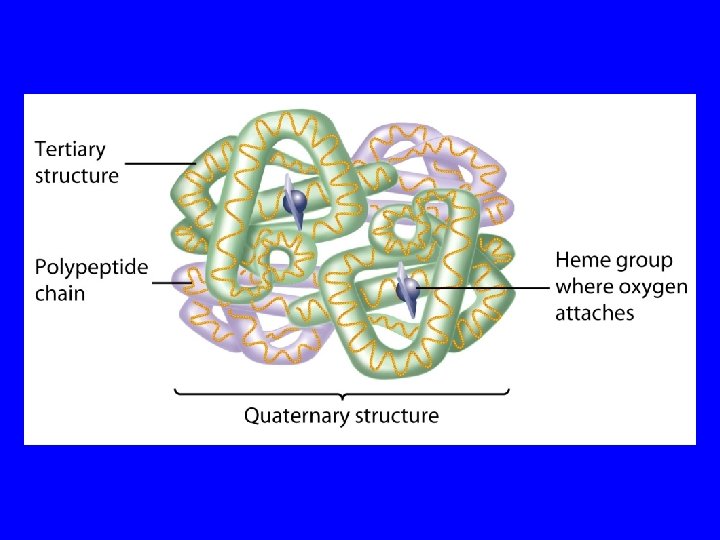
Protein Structure Primary structure is the amino acid sequence. Secondary structure is how the amino acids in sequence fold up locally. Commons examples are -helixes and -strands. Tertiary structure is the 3 -dimensional folding of the secondary structural elements and connecting loops in space. Quaternary structure is the association of multiple subunits, each with a tertiary structure and each a separate gene product.

1 O Structure Consider a peptide with two amino acids AA 1 20 AA 1 AA 2 x 20 AA 2 = 400 different molecules AA 3 20 x 20 (or 203 ) = 8000 different molecules For 100 amino acid protein the # of possibilities are: The total number of atoms in the universe is

Proteins Polypeptides and Proteins Glycylalanine is abbreviated gly-ala. Polypeptides are formed with a large number of amino acids (usually result in proteins with molecular weights between 6000 and 50 million amu). Protein Structure Primary structure is the sequence of the amino acids in the protein. A change in one amino acid can alter the biochemical behavior of the protein.

Protein Structure 1 o : The linear sequence of amino acids and disulfide bonds eg. ARDV: Ala. Arg. Asp. Val. 2 o : Local structures which include, folds, turns, helices and -sheets held in place by hydrogen bonds. 3 o : 3 -D arrangement of all atoms in a single polypeptide chain. 4 o : Arrangement of polypeptide chains into a functional protein, eg. hemoglobin.
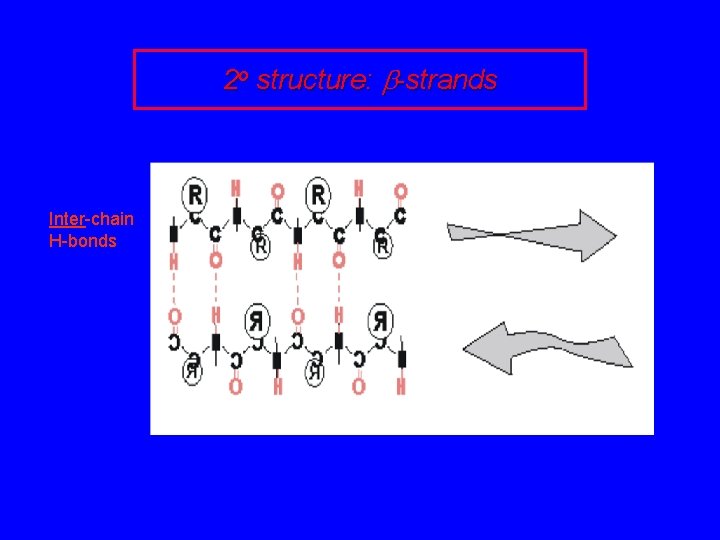
2 o structure: b-strands Inter-chain H-bonds
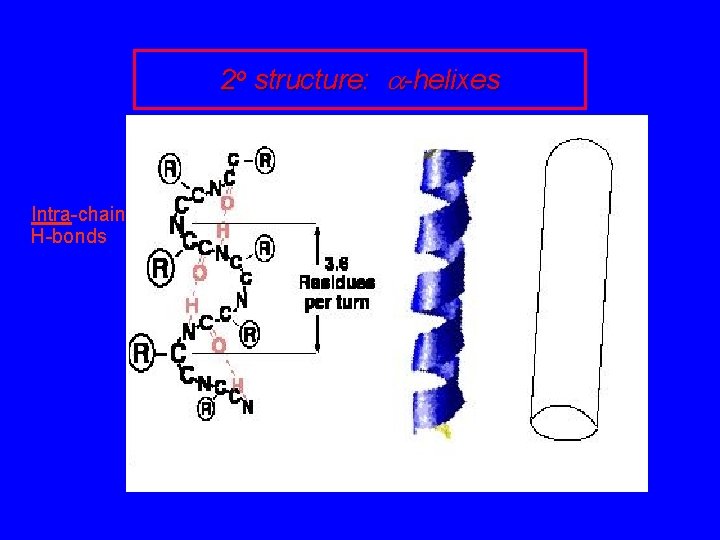
2 o structure: a-helixes Intra-chain H-bonds
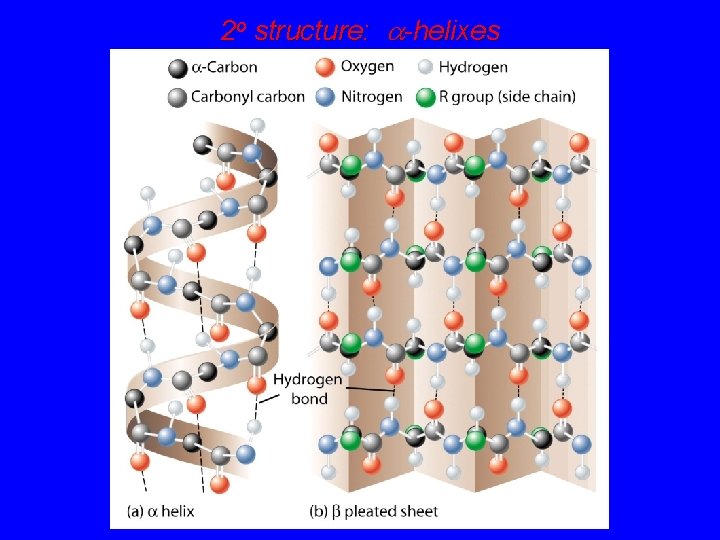
2 o structure: a-helixes
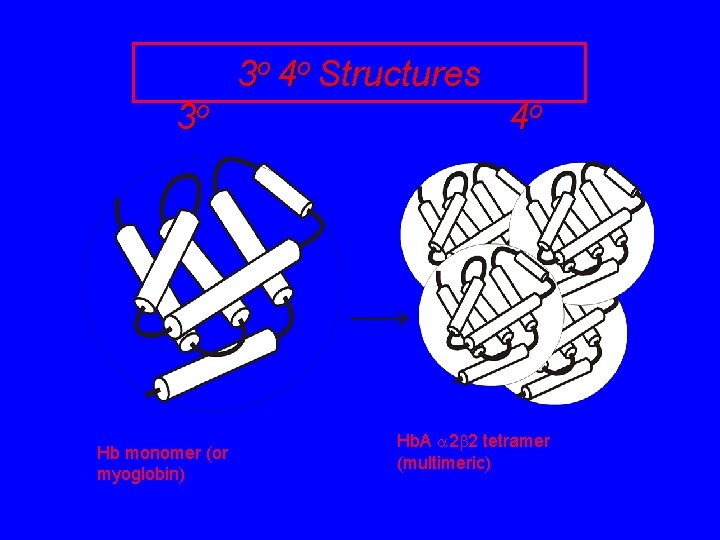
3 o 4 o Structures 3 o Hb monomer (or myoglobin) 4 o Hb. A 2 2 tetramer (multimeric)
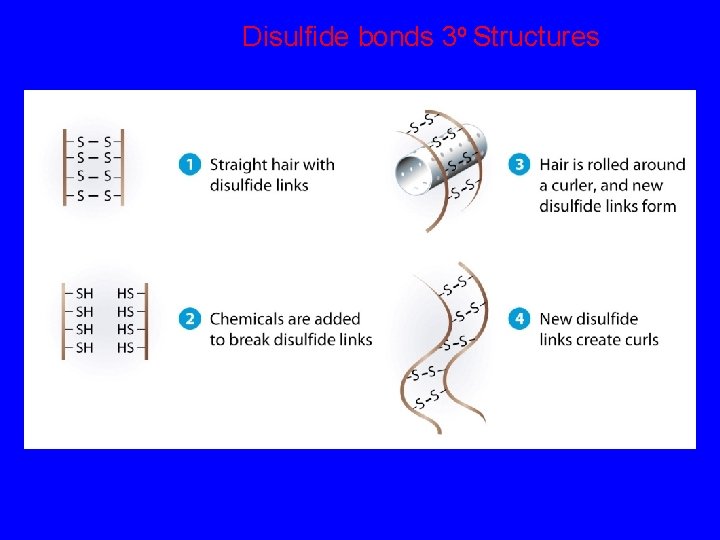
Disulfide bonds 3 o Structures
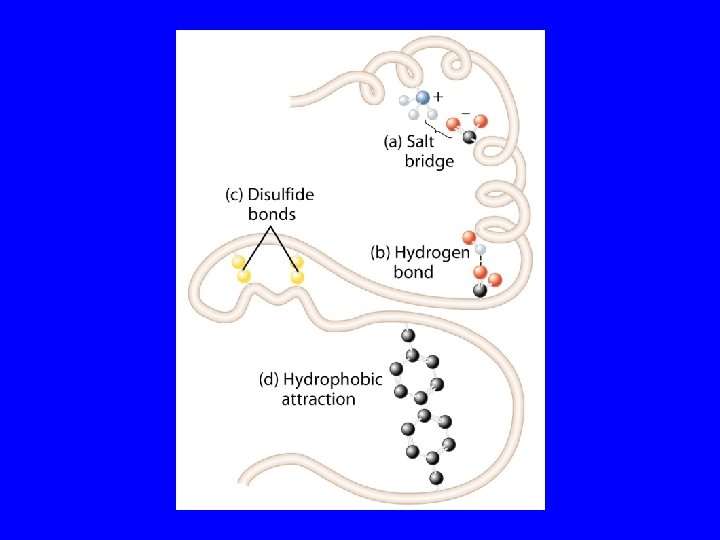

Stabilization of Protein Structure Electrostatic interactions involve the interaction of (+) and (-) charged (side) groups. Hydrogen bonds involve sharing of a hydrogen atom between two eletronegative atoms (e. g. , O, N). Van der Waal’s forces are weak forces based on optimal overlap of adjacent electronic orbitals. Hydrophobic interactions are, by far, the most powerful force stabilizing protein structure. Basis of force is entropy gain realized by burying hydrophobic residues.


Enzymes are proteins which act as biological catalysts. Over 1500 have been isolated. Human genome project scientists estimate that there about 30, 000 enzymes in a human. Active (catalytic) site is a crevice which binds a substrate. Lock & key metaphore. . but, protein can change conformation. The active site is evolutionarily conserved.
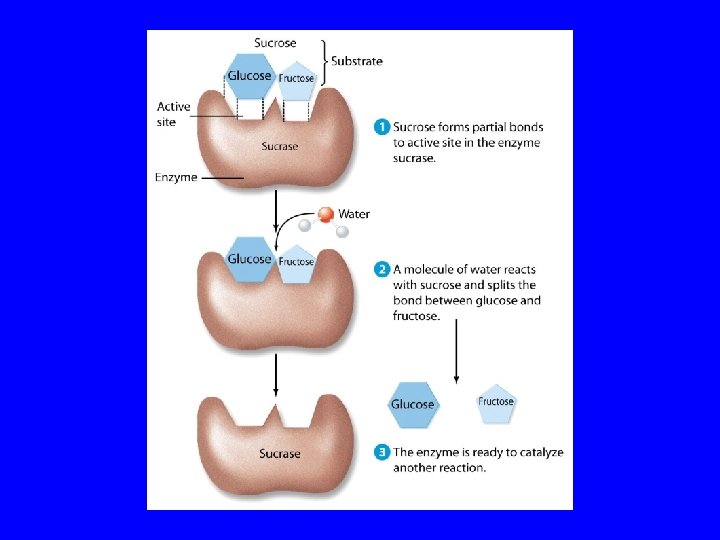

Enzyme Inhibitors / Effectors Michaelis-Minton Kinetics E = Enzyme; S = Substrate Enzyme Activity is reduced by inhibitors. Four types of inhibitors: Reversible, Irreversible, Competitive, Non-competitive Equilibrium Constant & Free Energy K[ES]eq = 10 -2 to 10 -6 ; Free Energies -3 to -12 kcal/mol vs. covalent bonds -50 to -110 kcal/mol Effectors increase enzyme activity.
- Slides: 42