Biochemistry 2e Garrett Grisham Chapter 33 Protein Synthesis

Biochemistry 2/e - Garrett & Grisham Chapter 33 Protein Synthesis and Degradation to accompany Biochemistry, 2/e by Reginald Garrett and Charles Grisham All rights reserved. Requests for permission to make copies of any part of the work should be mailed to: Permissions Department, Harcourt Brace & Company, 6277 Sea Harbor Drive, Orlando, Florida 32887 -6777 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Outline • • 33. 1 Ribosome Structure and Assembly 33. 2 Mechanics of Protein Synthesis 33. 3 Protein Synthesis in Eukaryotes 33. 4 Inhibitors of Protein Synthesis 33. 5 Protein Folding 33. 6 Post-Translational Processing of Proteins 33. 7 Protein Degradation 2 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Ribosome Structure and Assembly • E. coli ribosome is 25 nm diameter, 2520 k. D in mass, and consists of two unequal subunits that dissociate at < 1 m. M Mg 2+ • 30 S subunit is 930 k. D with 21 proteins and a 16 S r. RNA • 50 S subunit is 1590 k. D with 31 proteins and two r. RNAs: 23 S r. RNA and 5 S r. RNA • These ribosomes and others are roughly 2/3 RNA • 20, 000 ribosomes in a cell, 20% of cell's mass 3 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham 4 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Ribosomal Proteins • One of each per ribosome, except L 7/L 12 with 4 • L 7/L 12 identical except for extent of acetylation at N-terminus • Four L 7/L 12 plus L 10 makes "L 8" • Only one protein is common to large and small subunits: S 20 = L 26 • Variety of structures, still being characterized 5 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Ribosome Assembly/Structure • If individual proteins and r. RNAs are mixed, functional ribosomes will assemble • Gross structures of large and small subunits are known - see Figure 33. 3 • A tunnel runs through the large subunit • Growing peptide chain is thought to thread through the tunnel during protein synthesis 6 Copyright © 1999 by Harcourt Brace & Company
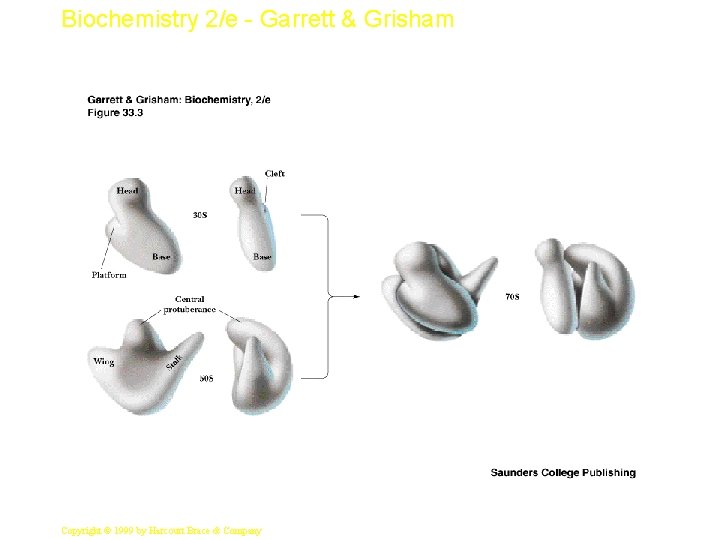
Biochemistry 2/e - Garrett & Grisham 7 Copyright © 1999 by Harcourt Brace & Company
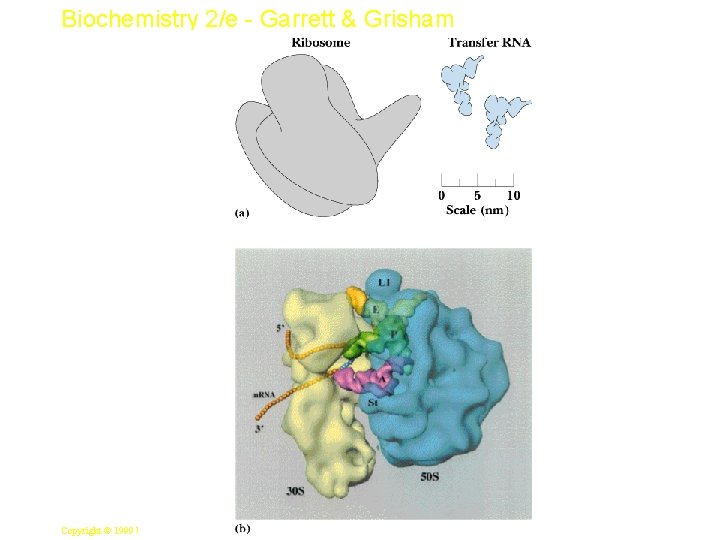
Biochemistry 2/e - Garrett & Grisham 8 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Eukaryotic Ribosomes • Mitochondrial and chloroplast ribosomes are quite similar to prokaryotic ribosomes, reflecting their supposed prokaryotic origin • Cytoplasmic ribosomes are larger and more complex, but many of the structural and functional properties are similar • See Table 33. 2 for properties 9 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Mechanics of Protein Synthesis • All protein synthesis involves three phases: initiation, elongation, termination • Initiation involves binding of m. RNA and initiator aminoacyl-t. RNA to small subunit, followed by binding of large subunit • Elongation: synthesis of all peptide bonds - with t. RNAs bound to acceptor (A) and peptidyl (P) sites. See Figure 33. 5 • Termination occurs when "stop codon" reached 10 Copyright © 1999 by Harcourt Brace & Company
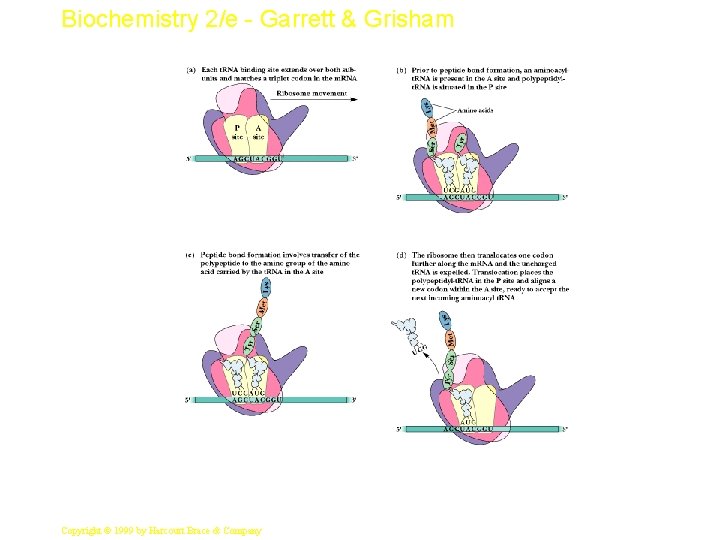
Biochemistry 2/e - Garrett & Grisham 11 Copyright © 1999 by Harcourt Brace & Company
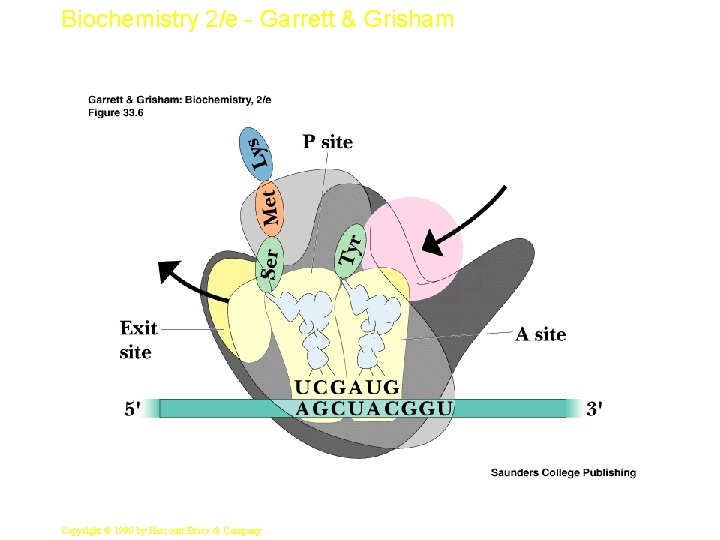
Biochemistry 2/e - Garrett & Grisham 12 Copyright © 1999 by Harcourt Brace & Company

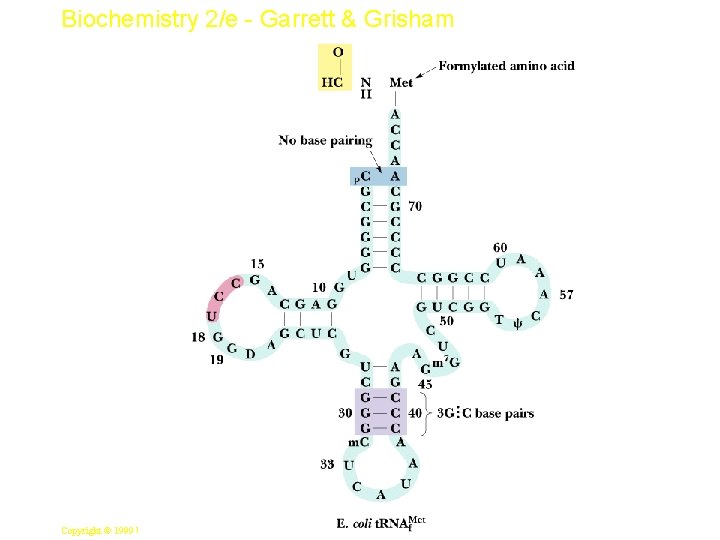
Biochemistry 2/e - Garrett & Grisham Prokaryotic Initiation • The initiator t. RNA is one with a formylated methionine: f-Met-t. RNAf. Met • It is only used for initiation, and regular Met -t. RNAm. Met is used instead for Met addition • N-formyl methionine is first aa of all E. coli proteins, but this is cleaved in about half • A formyl transferase adds the formyl group (see Figure 33. 8) 13 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 14 Copyright © 1999 by Harcourt Brace & Company

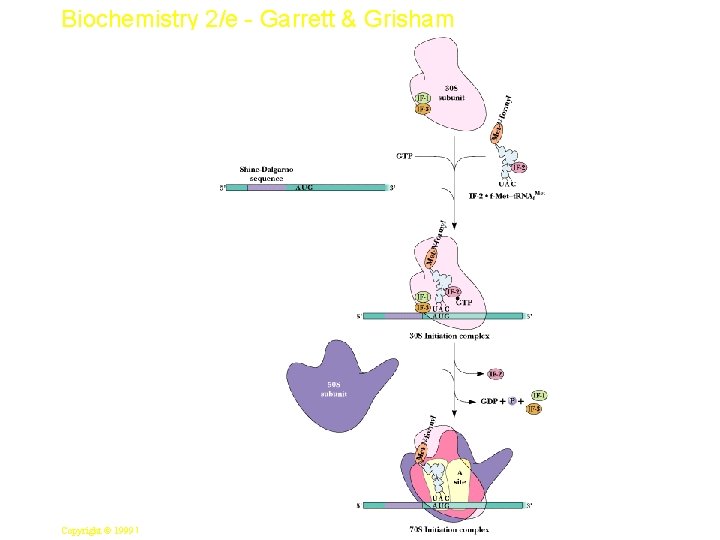
Biochemistry 2/e - Garrett & Grisham More Initiation • Correct registration of m. RNA on ribosome requires alignment of a pyrimidine-rich sequence on 3'-end of 16 S RNA with a purine-rich part of 5'-end of m. RNA • The purine-rich segment - the ribosomebinding site - is known as the Shine-Dalgarno sequence (see Figure 33. 9) • Initiation factor proteins, GTP, N-formyl-Mett. RNAf. Met, m. RNA and 30 S ribosome form the 30 S initiation complex 15 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham 16 Copyright © 1999 by Harcourt Brace & Company

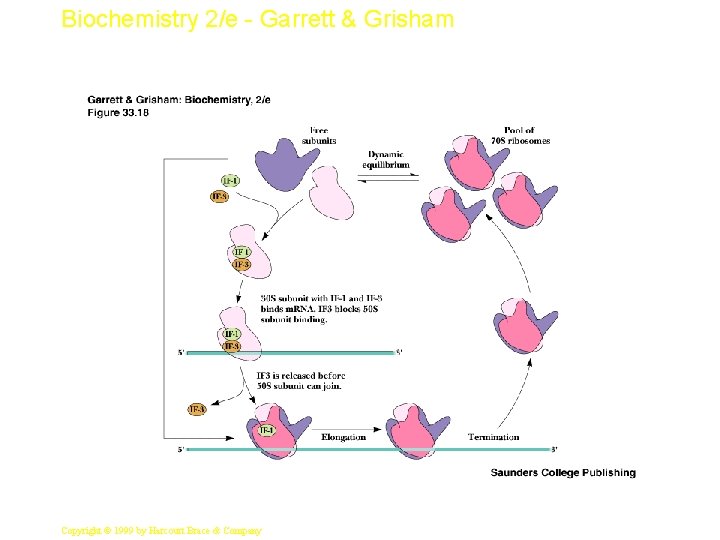
Biochemistry 2/e - Garrett & Grisham Events of Initiation • 30 S subunit with IF-1 and IF-3 binds m. RNA, IF-2, GTP and f-Met-t. RNAf. Met (Figure 33. 10) • IF-2 delivers the initiator t. RNA in a GTPdependent process • Loss of the initiation factors leads to binding of 50 S subunit • Note that the "acceptor site" is now poised to accept an incoming aminoacyl-t. RNA 17 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 18 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 19 Copyright © 1999 by Harcourt Brace & Company

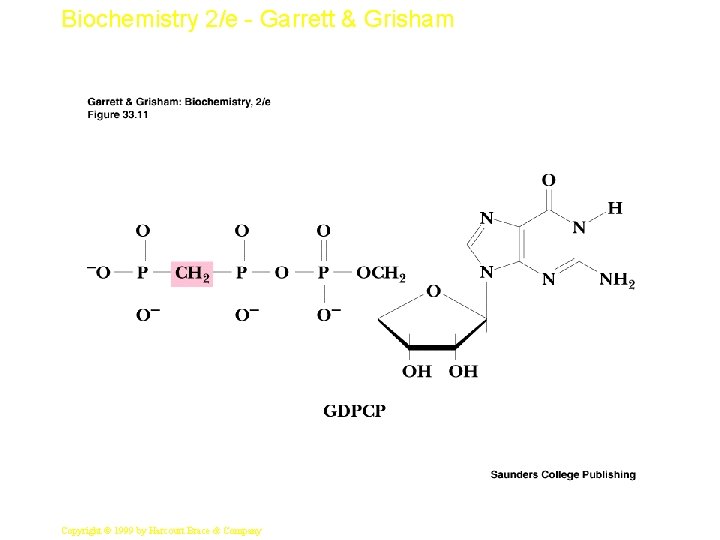
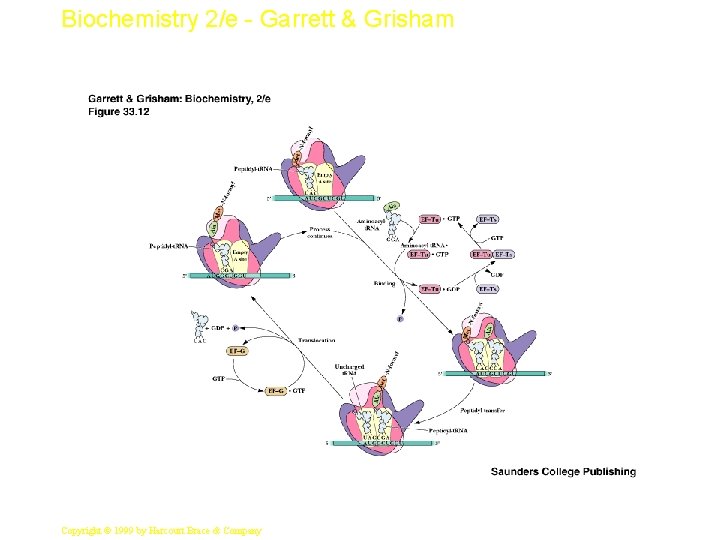
Biochemistry 2/e - Garrett & Grisham The Elongation Cycle • The elongation factors are vital to cell function, so they are present in significant quantities (EFTu is 5% of total protein in E. coli • EF-Tu binds aminoacyl-t. RNA and GTP • Aminoacyl-t. RNA binds to A site of ribosome as a complex with 2 EF-Tu and 2 GTP • GTP is then hydrolyzed and EF-Tu: GDP complexes dissociate • EF-Ts recycles EF-Tu by exchanging GTP for GDP 20 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 21 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham 22 Copyright © 1999 by Harcourt Brace & Company

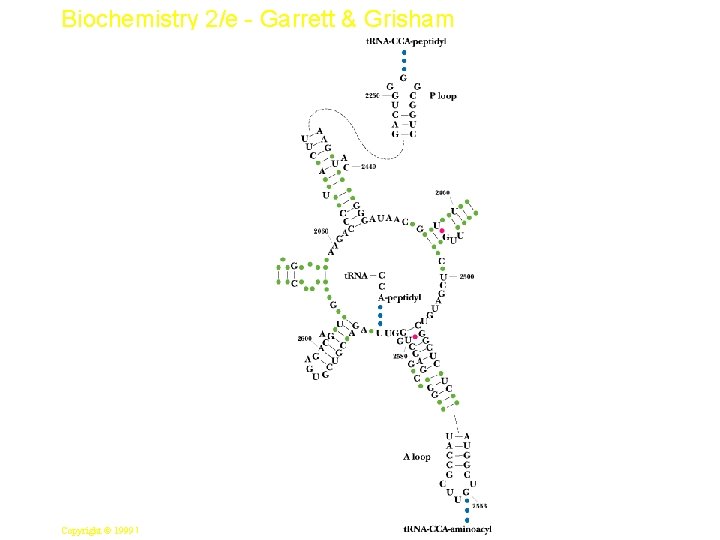
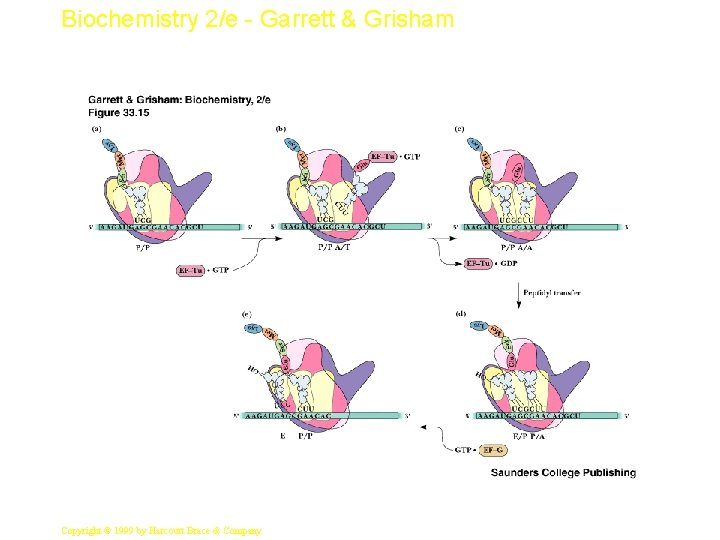
Biochemistry 2/e - Garrett & Grisham Peptidyl Transferase • This is the central reaction of protein synthesis • 23 S r. RNA is the peptidyl transferase! • The "reaction center" of 23 S r. RNA is shown in Figure 33. 14 - these bases are among the most highly conserved in all of biology. • Translocation of peptidyl-t. RNA from the A site to the P site follows (see Figures 33. 12 & 33. 15 ) catalyzed by EF-G 23 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 24 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 25 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham 26 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 27 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham The Role of GTP Hydrolysis • Three GTPs are hydrolyzed for each amino acid incorporated into peptide. • Hydrolysis drives essential conformation changes • Total of five high-energy phosphate bonds are expended per amino acid residue added - three GTP here and two in amino acid activation via aminoacylt. RNA synthesis 28 Copyright © 1999 by Harcourt Brace & Company

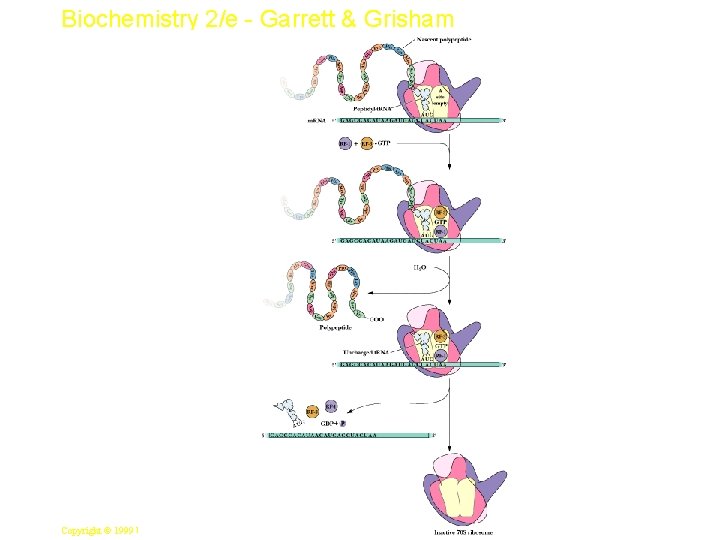
Biochemistry 2/e - Garrett & Grisham Peptide Chain Termination • Proteins known as "release factors" recognize the stop codon at the A site • Presence of release factors with a nonsense codon at A site transforms the peptidyl transferase into a hydrolase, which cleaves the peptidyl chain from the t. RNA carrier 29 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 30 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 31 Copyright © 1999 by Harcourt Brace & Company

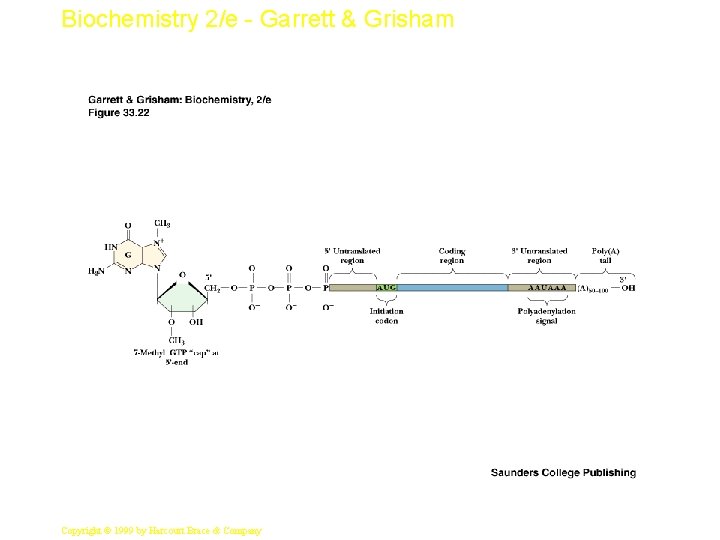
Biochemistry 2/e - Garrett & Grisham Eukaryotic Protein Synthesis See Figure 33. 22 for the structure of the typical m. RNA transcript • Note the 5'-methyl-GTP cap and the poly A tail • Initiation of protein synthesis in eukaryotes involves a family of at least 11 eukaryotic initiation factors • The initiator t. RNA is a special one that carries only Met and functions only in initiation - it is called t. RNAi. Met but it is not formylated 32 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 33 Copyright © 1999 by Harcourt Brace & Company

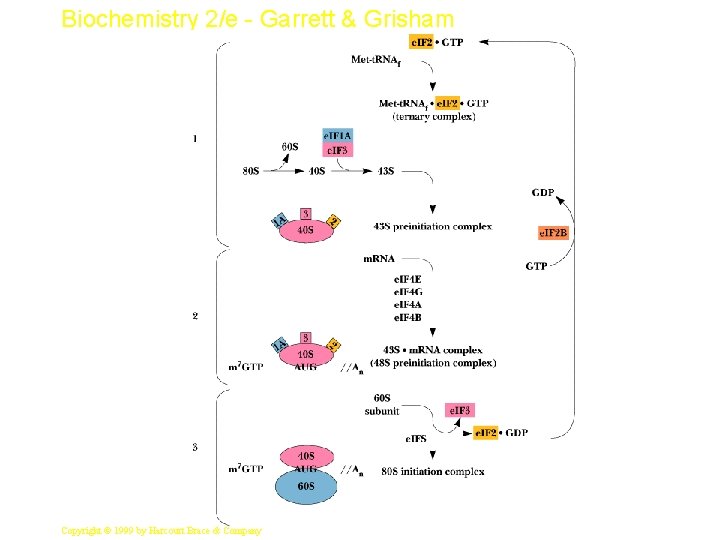
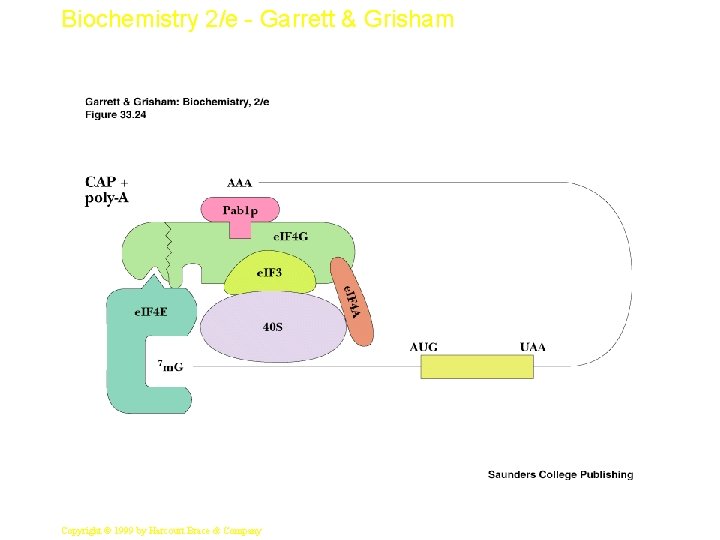
Biochemistry 2/e - Garrett & Grisham Eukaryotic Initiation • Begins with formation of ternary complex of e. IF-2, GTP and Met-t. RNAi. Met • 1) This binds to 40 S ribosomal subunit: e. IF-3: e. IF 1 A complex to form the 43 S preinitiation complex – Note no m. RNA yet, so no codon association with Mett. RNAi. Met • 2) m. RNA then adds with several other factors, forming the 48 S initiation complex (Fig. 33. 23) – 48 S initiation complex scans to find the first AUG (start) codon • 3) At AUG, 60 S subunit adds to make 80 S initiation complex (GTP is hydrolyzed) 34 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 35 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 36 Copyright © 1999 by Harcourt Brace & Company

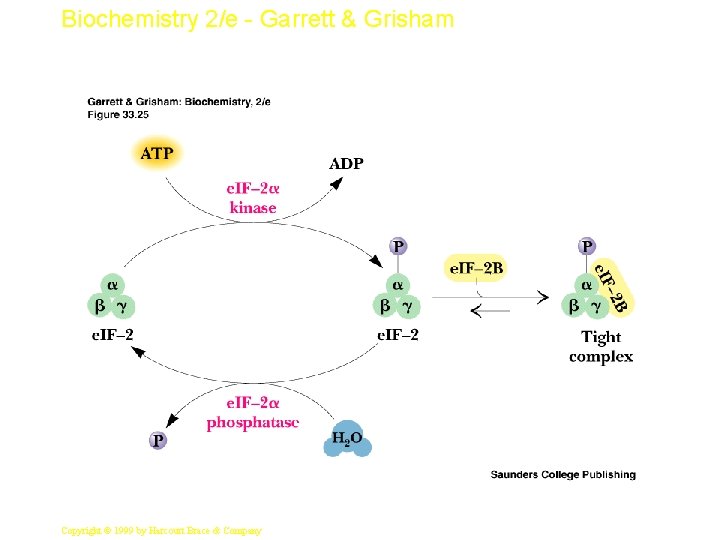
Biochemistry 2/e - Garrett & Grisham Regulation of Initiation Phosphorylation is the key, as usual • At least two proteins involved in initiation (Ribosomal protein S 6 and e. IF-4 F) are activated by phosphorylation • But phosphorylation of e. IF-2 a causes it to bind all available e. IF-2 B and sequesters it 37 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 38 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Elongation and Termination • Elongation is similar to procaryotic elongation: – EF 1 A homolog to EF-Tu, EF 1 B homolog to EF-Ts, EF 2 homolog to EF-G • Termination even simpler: only one RF, binds with GTP at the termination codon 39 Copyright © 1999 by Harcourt Brace & Company

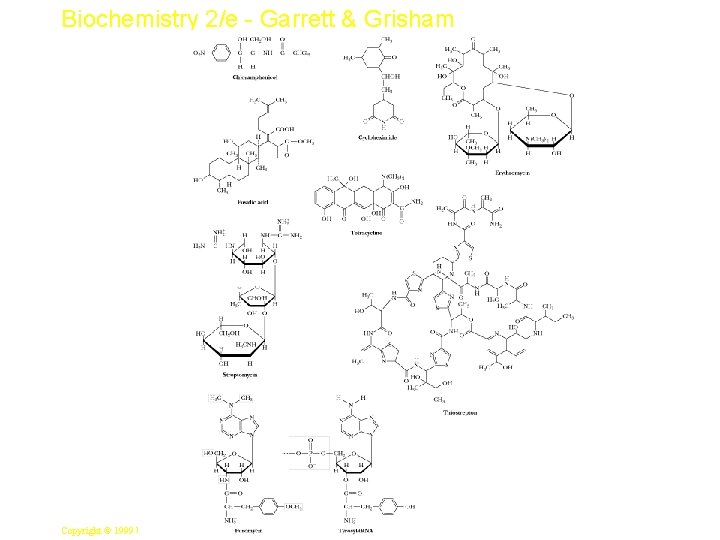
Biochemistry 2/e - Garrett & Grisham Inhibitors of Protein Synthesis • • Two important purposes to biochemists These inhibitors (Figure 33. 26) have helped unravel the mechanism of protein synthesis Those that affect prokaryotic but not eukaryotic protein synthesis are effective antibiotics Streptomycin - an aminoglycoside antibiotic - induces m. RNA misreading. Resulting mutant proteins slow the rate of bacterial growth Puromycin - binds at the A site of both prokaryotic and eukaryotic ribosomes, accepting the peptide chain from the P site, and terminating protein synthesis 40 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 41 Copyright © 1999 by Harcourt Brace & Company

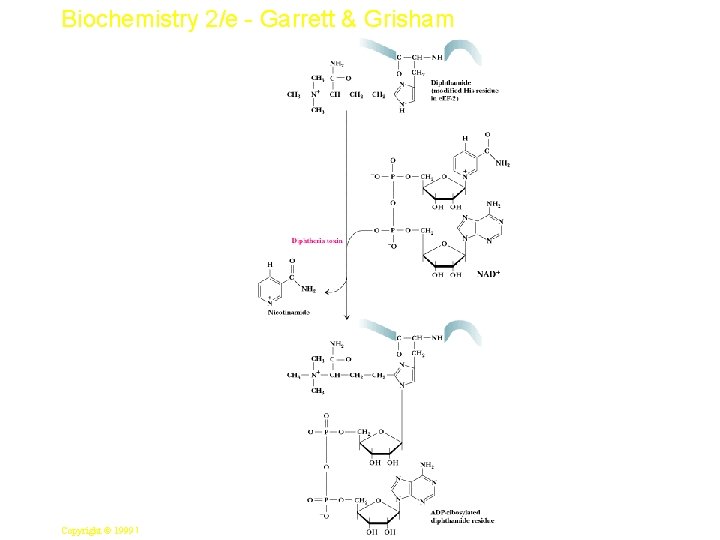
Biochemistry 2/e - Garrett & Grisham Diphtheria Toxin • • An NAD+-dependent ADP ribosylase One target of this enzyme is EF 2 has a diphthamide (see Figure 33. 27) Toxin-mediated ADP-ribosylation of EF 2 allows it to bind GTP but makes it inactive in protein synthesis One toxin molecule ADP-ribosylates many EF 2 s, so just a little is lethal! 42 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 43 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Ricin • • from Ricinus communis (castor bean) One of the most deadly substances known A glycoprotein that is a disulfide-linked heterodimer of 30 k. D subunits The B subunit is a lectin (a class of proteins that binds specifically to glycoproteins & glycolipids) Endocytosis followed by disulfide reduction releases A subunit, which catalytically inactivates the large subunit of ribosomes 44 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Ricin A subunit mechanism • Ricin A chain specifically attacks a single, highly conserved adenosine near position 4324 in eukaryotic 28 S RNA • N-glycosidase activity of A chain removes the adenosine base • Removal of this A (without cleaving the RNA chain) inactivates the large subunit of the ribosome • One ricin molecules can inactivate 50, 000 ribosomes, killing the eukaryotic cell! 45 Copyright © 1999 by Harcourt Brace & Company

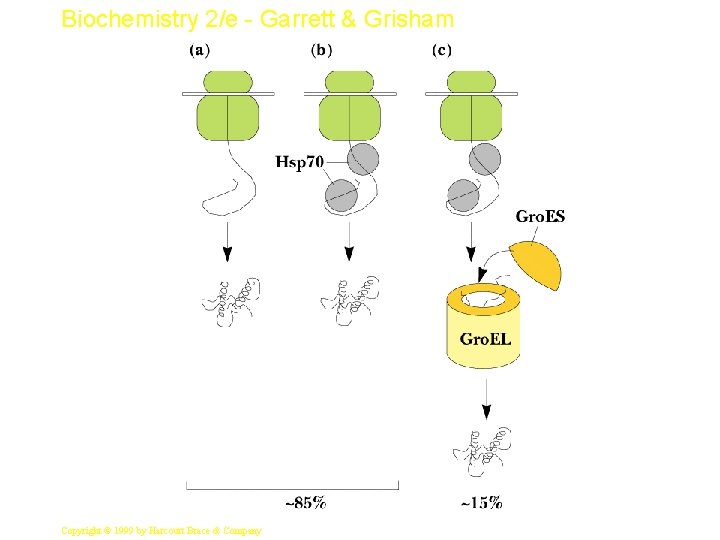
Biochemistry 2/e - Garrett & Grisham Protein Folding • Proteins are assisted in folding by molecular chaperones • Hsp 60 (chaperonins) and Hsp 70 are two main classes • Hsp 70 recognizes exposed, unfolded regions of new protein chains - especially hydrophobic regions • It binds to these regions, protecting them until productive folding reactions can occur 46 Copyright © 1999 by Harcourt Brace & Company
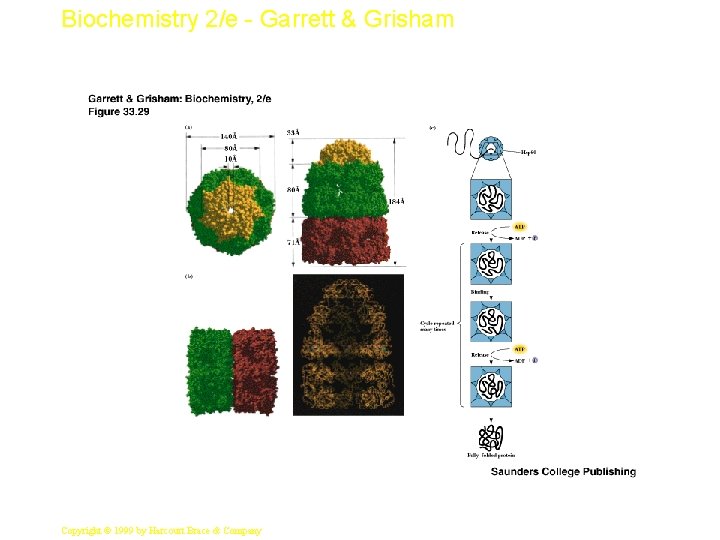
Biochemistry 2/e - Garrett & Grisham 47 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham The Gro. ES-Gro. EL Complex • The principal chaperonin in E. coli • Gro. EL forms two stacked 7 -membered rings of 60 k. D subunits; Gro. ES is a dome on the top • Nascent protein apparently binds reversibly many times to the walls of the donut structure, each time driven by ATP hydrolysis, eventually adopting its folded structure, then being released from the Gro. ES-Gro. EL complex • Rhodanese (as one example) requires 48 hydrolysis of 130 ATP to reach fully folded state Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 49 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Protein Translocation An essential process for membrane proteins and secretory proteins • Such proteins are synthesized with a "leader peptide", aka a "signal sequence" of about 16 -26 amino acids • The signal sequence has a basic N-terminus, a central domain of 7 -13 hydrophobic residues, and a nonhelical C-terminus • The signal sequence directs the newly synthesized protein to its proper destination 50 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Protein Translocation II • • Four common features Proteins are made as preproteins containing domains that act as sorting signals Membranes involved in protein translocation have specific receptors on their cytosolic faces Translocases catalyze the movement of the proteins across the membrane with metabolic energy (ATP, GTP, ion gradients) essential Preproteins bind to chaperones to stay loosely folded 51 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Prokaryotic Protein Transport All non-cytoplasmic proteins must be translocated • The leader peptide retards the folding of the protein so that molecular chaperone proteins can interact with it and direct its folding • The leader peptide also provides recognition signals for the translocation machinery • A leader peptidase removes the leader sequence when folding and targeting are assured 52 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Eukaryotic Protein Sorting Eukaryotic cells contain many membrane-bounded compartments • Most (but not all) targeting sequences are Nterminal, cleaveable presequences • Charge distribution, polarity and secondary structure of the signal sequence, rather than a particular sequence, appears to target to particular organelles and membranes • Synthesis of secretory and membrane proteins is coupled to translocation across ER membrane 53 Copyright © 1999 by Harcourt Brace & Company

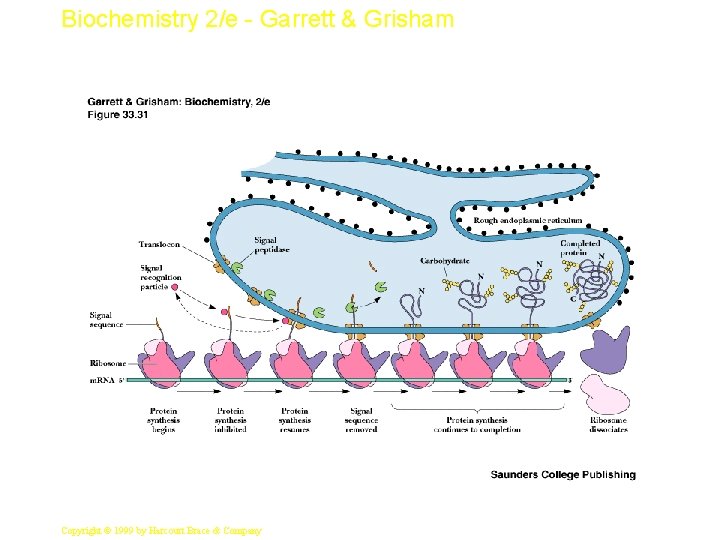
Biochemistry 2/e - Garrett & Grisham Events at the ER Membrane • As the signal sequence emerges from the ribosome, a signal recognition particle (SRP) finds it and escorts it to the ER membrane • There it docks with a docking protein or SRP receptor - see Figure 33. 31 • SRP dissociates in a GTP-dependent process • Protein synthesis resumes and protein passes into ER or into ER membrane; signal is cleaved 54 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 55 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Mitochondrial protein import • Mitochondria have two membranes, and two spaces in between the membranes • Signal sequences are N-terminal, positively charged regions of 10 -70 aa • Form amphiphilic a-helices, positive on one side and uncharged, hydrophobic on the other • Mitochondrial receptor will bind to preprotein 56 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham Protein Degradation • Some protein degradation pathways are nonspecific - randomly cleaved proteins seem to be rapidly degraded • However, there is also a selective, ATPdependent pathway for degradation - the ubiquitin-mediated pathway • Ubiquitin is a highly-conserved, 76 residue (8. 5 k. D) protein found widely in eukaryotes • Proteins are committed to degradation by conjugation with ubiquitin 57 Copyright © 1999 by Harcourt Brace & Company

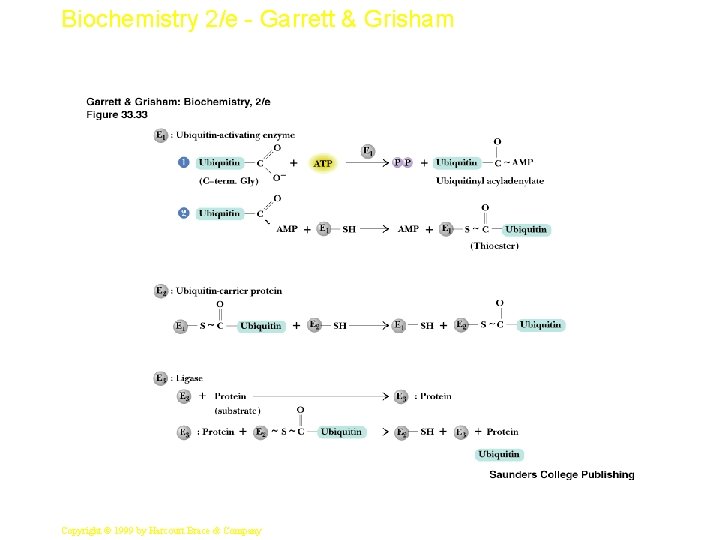
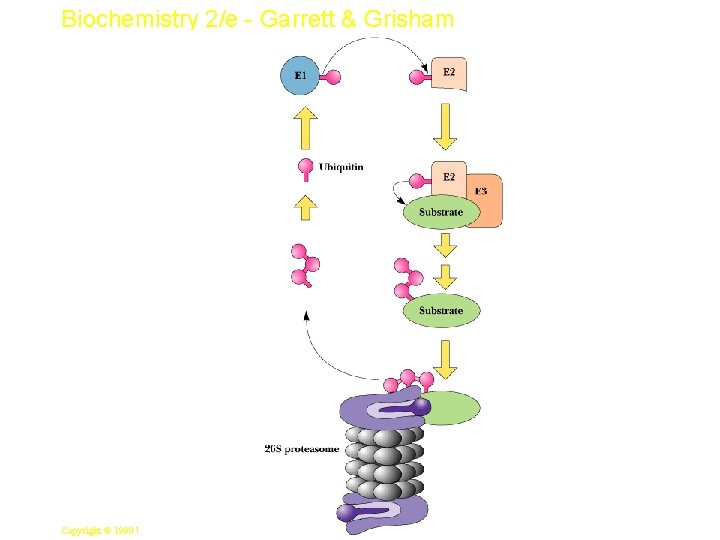
Biochemistry 2/e - Garrett & Grisham Ubiquitin and Degradation • • Three proteins involved: E 1, E 2 and E 3 E 1 is the ubiquitin-activating enzyme - it forms a thioester bond with C-terminal Gly of ubiquitin Ubiquitin is then transferred to a Cys-thiol of E 2, the ubiquitin-carrier protein Ligase (E 3) selects proteins for degradation. the E 2 -S~ubiquitin complex transfers ubiquitin to these selected proteins More than one ubiquitin may be attached to a protein target 58 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 59 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 60 Copyright © 1999 by Harcourt Brace & Company

Biochemistry 2/e - Garrett & Grisham 61 Copyright © 1999 by Harcourt Brace & Company
Biochemistry 2/e - Garrett & Grisham 62 Copyright © 1999 by Harcourt Brace & Company
- Slides: 62