Bio2 The Geant 4 DNA project http geant

Bio-2 The Geant 4 -DNA project http: //geant 4 -dna. org Takashi Sasaki – KEK, Japan Sébastien Incerti – IN 2 P 3/CENBG, France on behalf of the Bio-2 team TYL-FKPPL Joint Workshop, Seoul, June 4 -6, 2013

Outline • Context – Development of a simulation platform dedicated to the modelling biological effects of ionising radiation down to the cellular & sub-cellular scales – the Geant 4 -DNA project • Proposed workplan for 2013 1. Validation of theoretical cross section models in biomolecules through dedicated measurements 2. Development of realistic cellular geometries 3. Efforts toward Geant 4 -DNA computation speedup 4. Application in targeted radiotherapy • Collaboration matters 2

Modelling biological effects of ionising radiation remains a major scientific challenge Chronic exposure Fukushima Diagnosis Space missions http: //rcwww. kek. jp/norm/index-e. html Space exploration ISS Proton & hadrontherapy « A major challenge lies in providing a sound mechanistic understanding of low-dose radiation carcinogenesis » L. Mullenders et al. Mars Assessing cancer risks of low-dose radiation Nature Reviews Cancer (2009) NCC 3

The Monte Carlo approach • Can « reproduce » with accuracy the stochastic nature of particle-matter interactions • Many Monte Carlo codes are already available today in radiobiology for the simulation of track structures at the molecular scale in biological medium – E. g. PARTRAC, TRIOL, PHITS, KURBUC, NOREC… – Include physics & physico-chemistry processes, detailed geometrical descriptions of biological targets down to the DNA size, DNA and chromosome damage simulation and even repair mechanisms (PARTRAC)… • Usually designed for very specific applications • Not always easily accessible – Is it possible to access the source code ? – Are they adapted to recent OSs ? – Are they extendable by the user ? « To expand accessibility and avoid ‘reinventing the wheel’, track structure codes should be made available to all users via the internet from a central data bank» H. Nikjoo, IJRB 73, 355 (1998)
ATLAS, CMS, LHCb, ALICE @ CERN PET Scan (GATE) Ba. Bar, ILC… Brachytherapy Medical linac Earth magnetosphere GLAST/FERMI (NASA) GAIA Physics-Biology DICOM dosimetry ISS Hadrontherapy 5

Geant 4 ? • Can we try to extend Geant 4 to model biological effects of radiation ? • Strong limitations prevent its usage for the modelling of biological effects of ionising radiation at the sub-cellular & DNA scale – Condensed-history approach • No step-by-step transport on small distances, a key requirement for micro/nano-dosimetry – Low-energy limit applicability of EM physics models is limited • ~250 e. V for Livermore Low Energy EM models • ~100 e. V for Penelope Low Energy EM models – No description of target molecular properties • Liquid water, DNA nucleotides – Only physical particle-matter interactions • Physical interactions are NOT the dominant processes for DNA damage at low LET. . . 6

See Int. J. Model. Simul. Sci. Comput. 1 (2010) 157– 178 The Geant 4 -DNA project • Initiated in 2001 by Dr Petteri Nieminen at the European Space Agency/ESTEC • Main objective: extension of the general purpose Geant 4 Monte Carlo toolkit for the simulation of interactions of radiation with biological systems at the cellular and DNA level in order to predict early DNA damages in the context of manned space exploration missions ( « bottom-up » approach) • Providing an open source access to the scientific community that can be easily upgraded & improved • First prototypes of physics models were added to Geant 4 in 2007 • It is now a full independent sub-category of the electromagnetic category of Geant 4 – $G 4 INSTALL/source/processes/electromagnetic/dna • Usable from external codes such as GATE since 2012 • Currently an on-going interdisciplinary activity of the Geant 4 collaboration « low energy electromagnetic physics » working group – Coordinated by CNRS/IN 2 P 3 since 2008 – Supported by several funding agencies : ESA (AO 6041, AO 7146), ANR, INSERM. . . 7
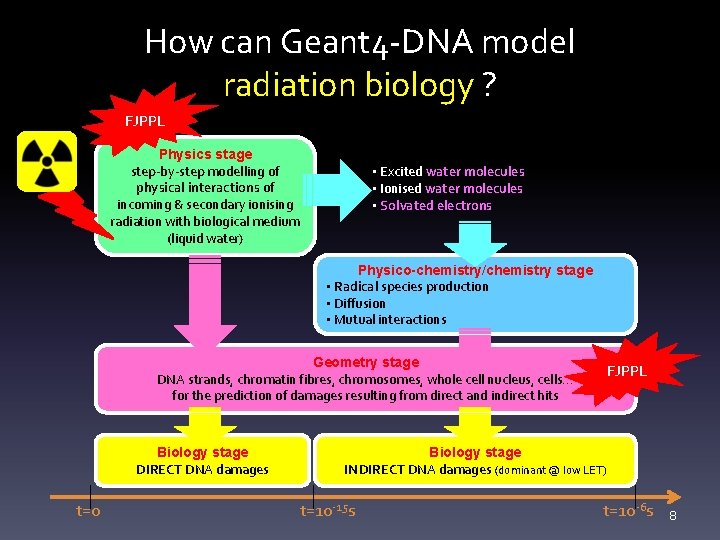
How can Geant 4 -DNA model radiation biology ? FJPPL Physics stage step-by-step modelling of physical interactions of incoming & secondary ionising radiation with biological medium (liquid water) • Excited water molecules • Ionised water molecules • Solvated electrons Physico-chemistry/chemistry stage • Radical species production • Diffusion • Mutual interactions Geometry stage DNA strands, chromatin fibres, chromosomes, whole cell nucleus, cells… for the prediction of damages resulting from direct and indirect hits Biology stage DIRECT DNA damages t=0 FJPPL Biology stage INDIRECT DNA damages (dominant @ low LET) t=10 -15 s t=10 -6 s 8

a) Physics stage: improving Physics

Physics models available in Geant 4 9. 6+P 01 (Spring 2013) • Geant 4 -DNA physics models are applicable to liquid water, the main component of biological matter • • • They can reach the very low energy domain down to electron thermalization – Compatible with molecular description of interactions (5 excitation & ionisation levels of the water molecule) – Sub-excitation electrons (below ~9 e. V ) can undergo vibrational excitation, attachment and elastic scattering Purely discrete – Simulate all elementary interactions on an event-by-event basis – No condensed history approximation Models can be purely analytical and/or use interpolated data tables – • For eg. computation of integral cross sections They use the same software design as all electromagnetic models available in Geant 4 ( « standard » and « low energy » EM models and processes) – Allows the combination of models and processes 10

Overview of physics models for liquid water • Protons & H – • Excitation • Electrons – energies and Born and Bethe theories above 500 ke. V Elastic scattering • Screened Rutherford and Brenner-Zaider below 200 e. V • Partial wave framework model, – Ionisation • 5 levels for H 2 O • Dielectric formalism & FBA using Heller optical data up to 1 Me. V, and low energy corrections – ke. V (relativistic + Fermi density) Ionisation • – 5 levels for H 2 O • Dielectric formalism & FBA using Heller optical data and • – Michaud et al. xs measurements in amorphous ice • Factor 2 to account for phase effect Speed and effective charge scaling from protons by Dingfelder et al. , – Charge change • • Dissociative attachment • Excitation and ionisation • Vibrational excitation • Analytical parametrizations by Dingfelder et al. He 0, He+, He 2+ – semi-empirical low energy corrections – Charge change • Excitation • Rudd semi-empirical approach by Dingfelder et al. and Born and Bethe theories & dielectric formalism above 500 3 contributions to the interaction potential – Miller & Green speed scaling of e- excitation at low C, N, O, Fe – Melton et al. xs measurements Semi-empirical models from Dingfelder et al. Ionisation • Speed scaling and global effective charge by Booth and Grant See Med. Phys. 37 (2010) 4692 -4708 and Appl. Radiat. Isot. 69 (2011) 220 -226 • Photons – from EM « standard » and « low energy » 11
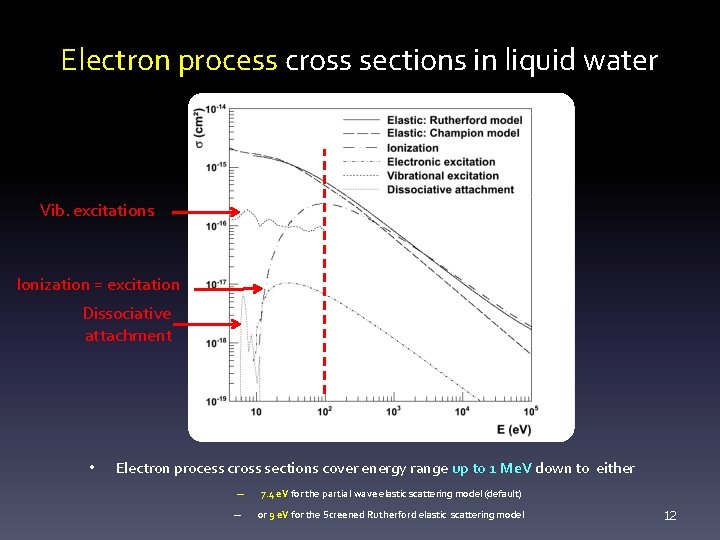
Electron process cross sections in liquid water Vib. excitations Ionization = excitation Dissociative attachment • Electron process cross sections cover energy range up to 1 Me. V down to either – 7. 4 e. V for the partial wave elastic scattering model (default) – or 9 e. V for the Screened Rutherford elastic scattering model 12

Extension to DNA material • We have implemented physics processes and models for the modeling of proton and neutral hydrogen atom interactions with – liquid water – the four DNA nucleobases : Adenine (A), Thymine (T), Guanine (G) and Cytosine (C) • within a Classical Trajectory Monte Carlo approach and taking into account specific energetic criteria • Mean energy transfers (potential and kinematical) during interactions were computed from quantum mechanics and are tabulated • First time that a general purpose Monte Carlo toolkit is equipped with such functionnalities • Dedicated Physics constructor: G 4 Em. DNACTMCPhysics • Expected to be released in Geant 4 X 13
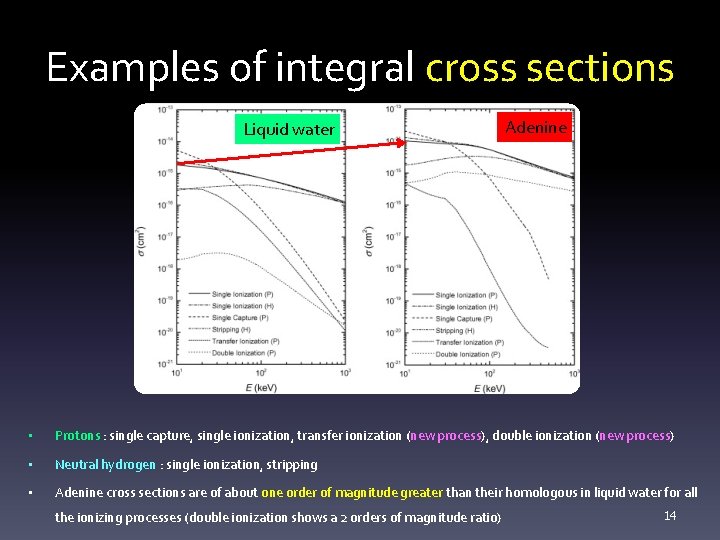
Examples of integral cross sections Liquid water Adenine • Protons : single capture, single ionization, transfer ionization (new process), double ionization (new process) • Neutral hydrogen : single ionization, stripping • Adenine cross sections are of about one order of magnitude greater than their homologous in liquid water for all the ionizing processes (double ionization shows a 2 orders of magnitude ratio) 14
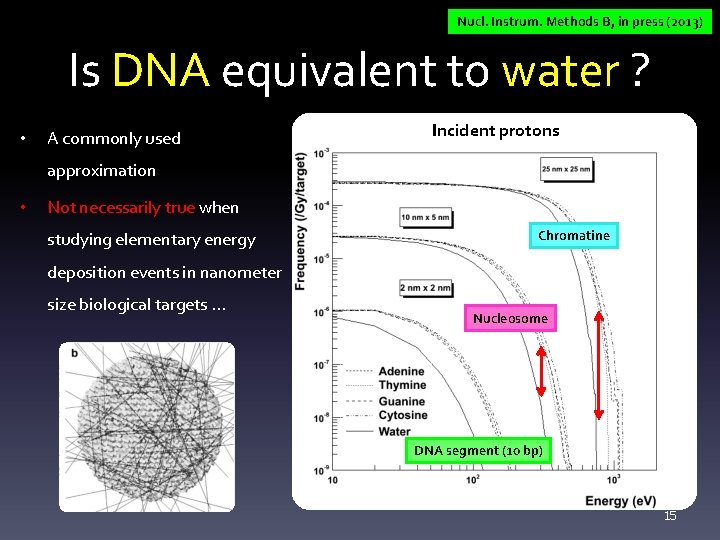
Nucl. Instrum. Methods B, in press (2013) Is DNA equivalent to water ? • A commonly used Incident protons approximation • Not necessarily true when studying elementary energy Chromatine deposition events in nanometer size biological targets. . . Nucleosome DNA segment (10 bp) 15

Need for experimental validation of theoretical cross section models • Experimental measurement of cross section in biomolecules remain very scarce today • Pr A. Itoh, Kyoto University, joined this proposal in order to propose an experimental workplan dedicated to the measurement of cross sections in biomolecules – Ion induced collisions on DNA & RNA components • Ionisation and electronic capture processes – Measurement of multi-differential and total cross sections • Will allow the validation Geant 4 -DNA theoretical models 16
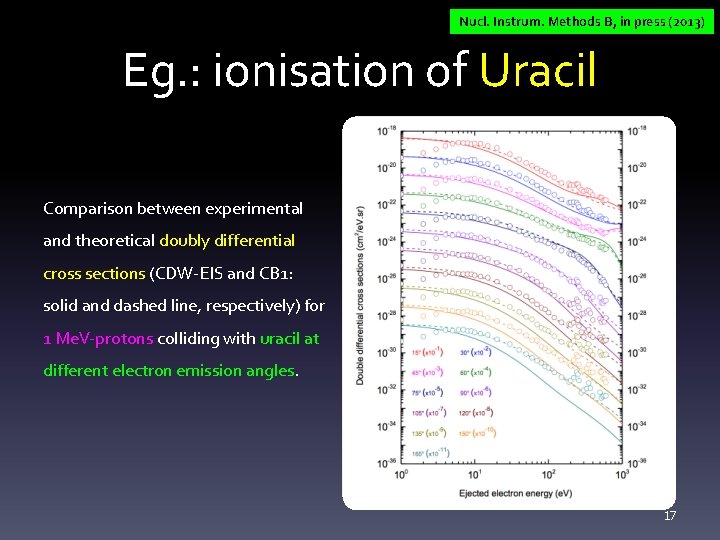
Nucl. Instrum. Methods B, in press (2013) Eg. : ionisation of Uracil Comparison between experimental and theoretical doubly differential cross sections (CDW-EIS and CB 1: solid and dashed line, respectively) for 1 Me. V-protons colliding with uracil at different electron emission angles. 17

Experimental setup • A beam of 1 Me. V H+ is extracted from a Van de Graff accelerator at Kyoto University • Beam was well collimated to about 1 x 3 mm 2 in size and was introduced into a double mumetal shielded collision chamber. • A typical beam current of about 50 n. A measured by a Faraday cup was used in this work. • An effusive molecular beam target of uracil was produced by heating crystalline uracil powder (99 % purity) in a stainless steel oven at a temperature of 473 K. • The energy and angular distributions of secondary electrons ejected from uracil molecules were measured by a 45° parallel plate electrostatic spectrometer. • The measurements were carried out for electron energies from 1 e. V to 1 ke. V and emission angles from 15° to 165°. • Experimental errors on the obtained single and doubly differential cross sections are 8 -13%. 18

b) Geometry stage: improving geometries
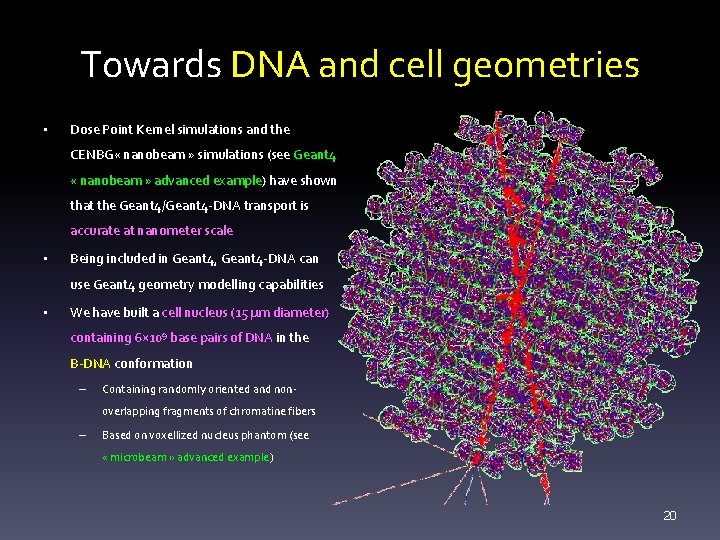
Towards DNA and cell geometries • Dose Point Kernel simulations and the CENBG « nanobeam » simulations (see Geant 4 « nanobeam » advanced example) have shown that the Geant 4/Geant 4 -DNA transport is accurate at nanometer scale • Being included in Geant 4, Geant 4 -DNA can use Geant 4 geometry modelling capabilities • We have built a cell nucleus (15 μm diameter) containing 6× 109 base pairs of DNA in the B-DNA conformation – Containing randomly oriented and nonoverlapping fragments of chromatine fibers – Based on voxellized nucleus phantom (see « microbeam » advanced example) 20
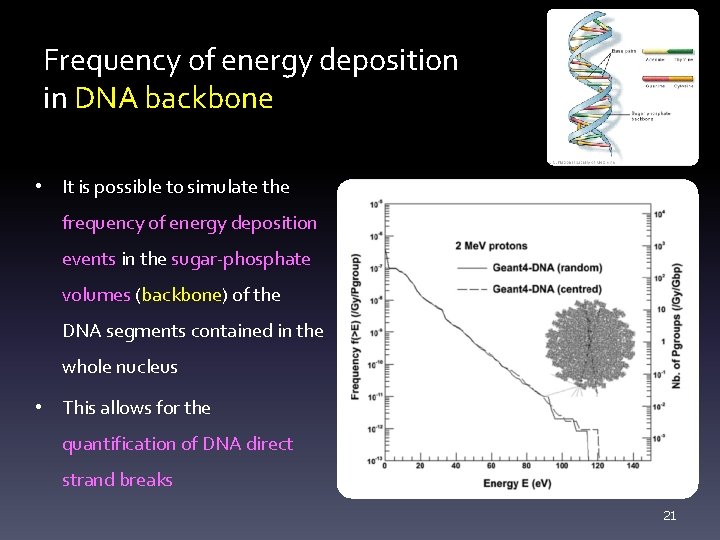
Frequency of energy deposition in DNA backbone • It is possible to simulate the frequency of energy deposition events in the sugar-phosphate volumes (backbone) of the DNA segments contained in the whole nucleus • This allows for the quantification of DNA direct strand breaks 21
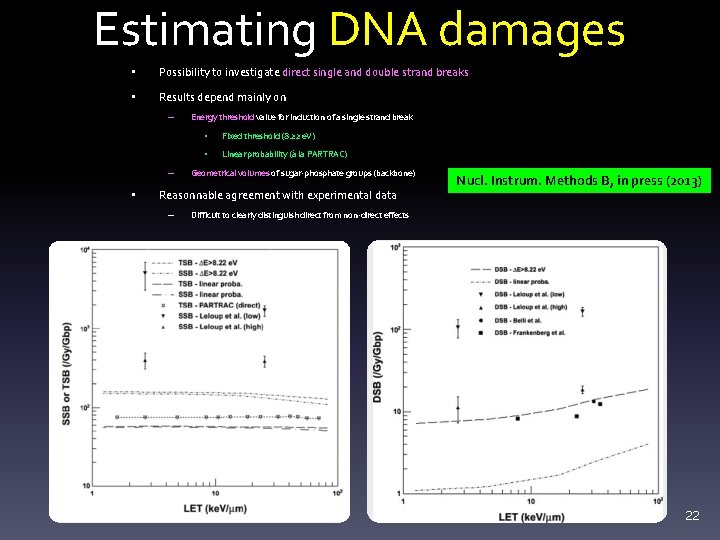
Estimating DNA damages • Possibility to investigate direct single and double strand breaks • Results depend mainly on – – • Energy threshold value for induction of a single strand break • Fixed threshold (8. 22 e. V) • Linear probability (à la PARTRAC) Geometrical volumes of sugar-phosphate groups (backbone) Reasonnable agreement with experimental data – Nucl. Instrum. Methods B, in press (2013) Difficult to clearly distinguish direct from non-direct effects 22
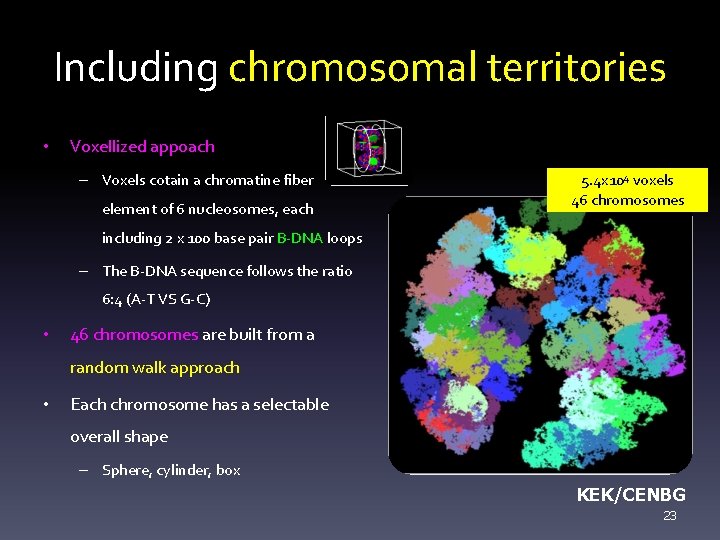
Including chromosomal territories • Voxellized appoach – Voxels cotain a chromatine fiber element of 6 nucleosomes, each 5. 4 x 104 voxels 46 chromosomes including 2 x 100 base pair B-DNA loops – The B-DNA sequence follows the ratio 6: 4 (A-T VS G-C) • 46 chromosomes are built from a random walk approach • Each chromosome has a selectable overall shape – Sphere, cylinder, box KEK/CENBG 23
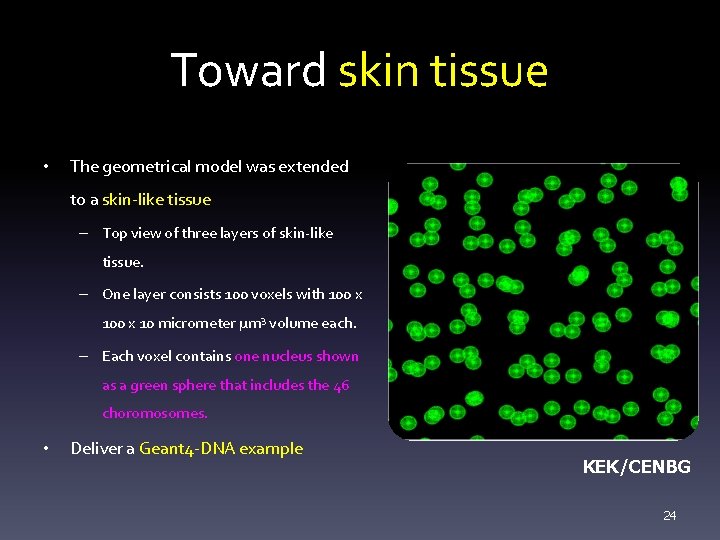
Toward skin tissue • The geometrical model was extended to a skin-like tissue – Top view of three layers of skin-like tissue. – One layer consists 100 voxels with 100 x 10 micrometer µm 3 volume each. – Each voxel contains one nucleus shown as a green sphere that includes the 46 choromosomes. • Deliver a Geant 4 -DNA example KEK/CENBG 24

c) Software performance
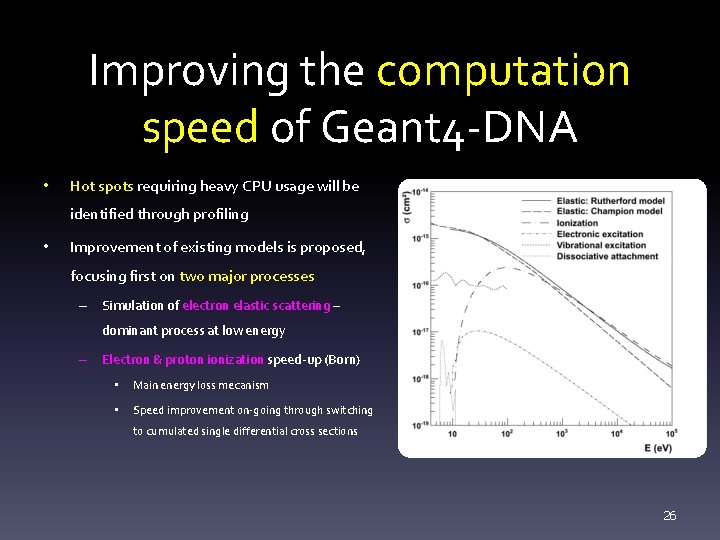
Improving the computation speed of Geant 4 -DNA • Hot spots requiring heavy CPU usage will be identified through profiling • Improvement of existing models is proposed, focusing first on two major processes – Simulation of electron elastic scattering – dominant process at low energy – Electron & proton ionization speed-up (Born) • Main energy loss mecanism • Speed improvement on-going through switching to cumulated single differential cross sections 26

d) Application: targeted radiotherapy
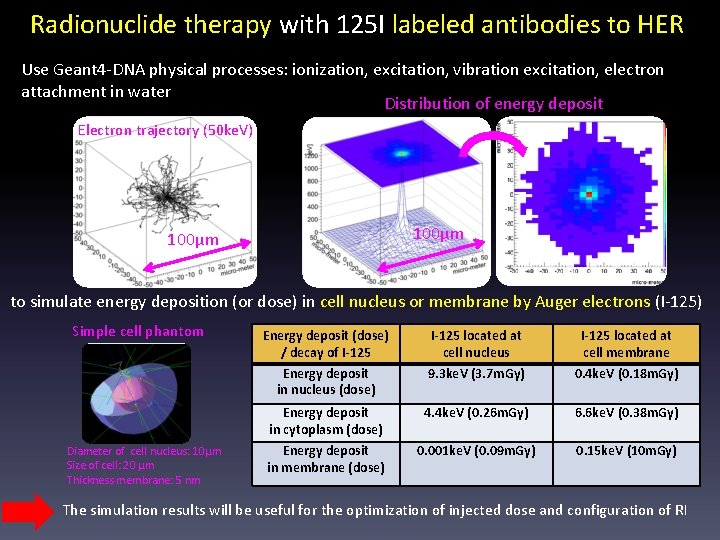
Radionuclide therapy with 125 I labeled antibodies to HER Use Geant 4 -DNA physical processes: ionization, excitation, vibration excitation, electron attachment in water Distribution of energy deposit Electron trajectory (50 ke. V) 100μm to simulate energy deposition (or dose) in cell nucleus or membrane by Auger electrons (I-125) Simple cell phantom Diameter of cell nucleus: 10μm Size of cell: 20 μm Thickness membrane: 5 nm Energy deposit (dose) / decay of I-125 Energy deposit in nucleus (dose) I-125 located at cell nucleus 9. 3 ke. V (3. 7 m. Gy) I-125 located at cell membrane 0. 4 ke. V (0. 18 m. Gy) Energy deposit in cytoplasm (dose) 4. 4 ke. V (0. 26 m. Gy) 6. 6 ke. V (0. 38 m. Gy) Energy deposit in membrane (dose) 0. 001 ke. V (0. 09 m. Gy) 0. 15 ke. V (10 m. Gy) The simulation results will be useful for the optimization of injected dose and configuration of RI
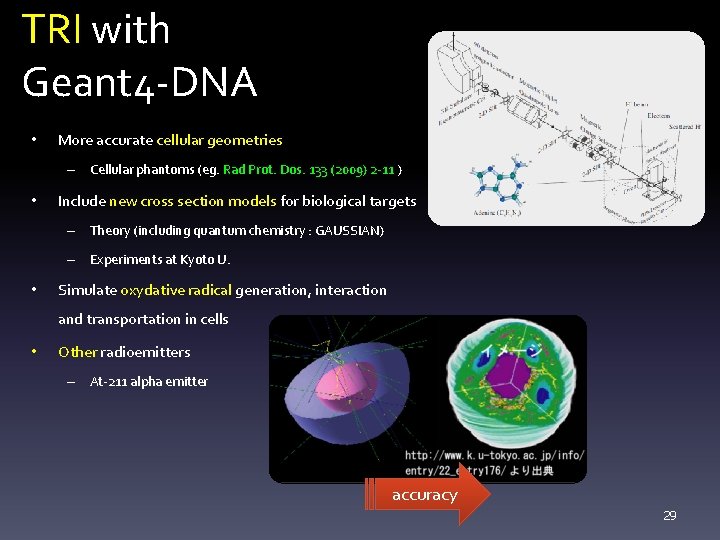
TRI with Geant 4 -DNA • More accurate cellular geometries – Cellular phantoms (eg. Rad Prot. Dos. 133 (2009) 2 -11 ) • Include new cross section models for biological targets – Theory (including quantum chemistry : GAUSSIAN) – Experiments at Kyoto U. • Simulate oxydative radical generation, interaction and transportation in cells • Other radioemitters – At-211 alpha emitter accuracy 29

Bio-2 collaboration matters
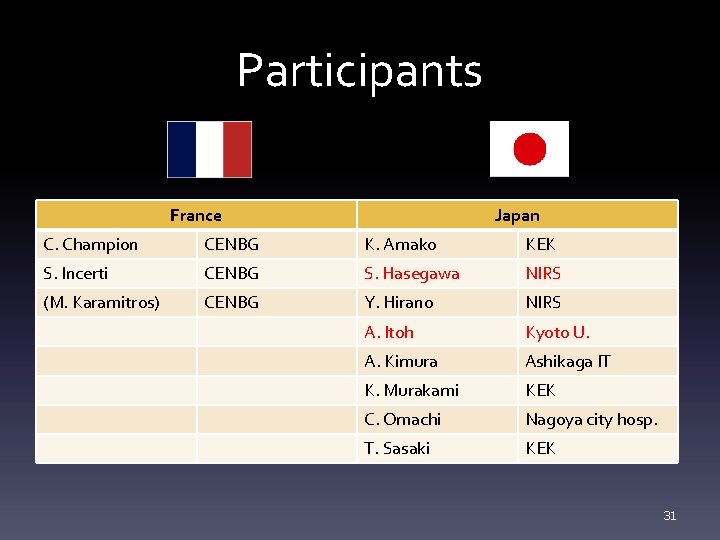
Participants France Japan C. Champion CENBG K. Amako KEK S. Incerti CENBG S. Hasegawa NIRS (M. Karamitros) CENBG Y. Hirano NIRS A. Itoh Kyoto U. A. Kimura Ashikaga IT K. Murakami KEK C. Omachi Nagoya city hosp. T. Sasaki KEK 31

Budget request for 2013 • Request from France – 2 french visitors to KEK & Kyoto U. • Participation to experimental program • (possibly participate to a Geant 4 Japanese tutorial) • Request from Japan – 3 japanese visitors to CENBG for a week • Note: we do not befenit from any other funding for Japan-France collaboration around Geant 4 -DNA & Geant 4 32
Geant 4 -DNA from the Internet A unique web site for Geant 4 -DNA: http: //geant 4 -dna. org Fully included in Geant 4 33

Thank you very much
- Slides: 34