Bio Sci D 145 Lecture 3 Bruce Blumberg
Bio. Sci D 145 Lecture #3 • Bruce Blumberg (blumberg@uci. edu) – 4103 Nat Sci 2 - office hours Tu, Th 3: 30 -5: 00 (or by appointment) – phone 824 -8573 • TA – Angela Kuo (akuo 4@uci. edu) – 4311 Nat Sci 2– office hours W 10 -12 – Phone 824 -6873 • check e-mail regularly for announcements, etc. . • Lectures will be posted in advance (without answers) • Updated lectures (with answers) will be posted after lecture – http: //blumberg-lab. bio. uci. edu/biod 145 -w 20120 • Don’t forget to discuss term paper topics with me in office hours or by email • Last year’s midterm is posted Bio. Sci D 145 lecture 1 page 1 ©copyright Bruce Blumberg 2020. All rights reserved

Term paper specific aims • Title of your proposal • A paragraph introducing your topic and explaining why it is important; i. e. , what impact will the knowledge gained have. – Why should any funding agency give you money to pursue this research? • NIH now requires a statement of human health relevance for all grant applications • NSF wants to know what is the intellectual merit of your proposed research and what broader impacts of your proposed research • Present your hypothesis – A supposition or conjecture put forth to account for known facts; esp. in the sciences, a provisional supposition from which to draw conclusions that shall be in accordance with known facts, and which serves as a starting-point for further investigation by which it may be proved or disproved and the true theory arrived at. • Enumerate 2 -3 specific aims in the form of questions that test your hypothesis – At least one of these aims needs to have a strong “whole genome” component – This is not a review article – propose something new. Bio. Sci D 145 lecture 4 page 2 ©copyright Bruce Blumberg 2004 -2016. All rights reserved
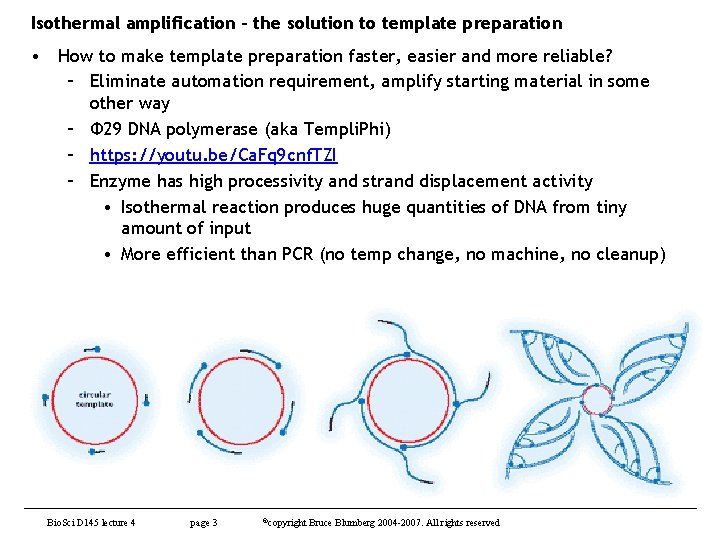
Isothermal amplification – the solution to template preparation • How to make template preparation faster, easier and more reliable? – Eliminate automation requirement, amplify starting material in some other way – Φ 29 DNA polymerase (aka Templi. Phi) – https: //youtu. be/Ca. Fq 9 cnf. TZI – Enzyme has high processivity and strand displacement activity • Isothermal reaction produces huge quantities of DNA from tiny amount of input • More efficient than PCR (no temp change, no machine, no cleanup) Bio. Sci D 145 lecture 4 page 3 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

Modern DNA sequence analysis • Cycle sequencing – Virtually all routine DNA sequencing today is done by cycle sequencing with fluorescent dd. NTPs • ABI Big Dye chemistry – Template preparation still tedious for small scale • Templi. PHi used in genome centers (no need for most automation) – Capillary sequencers predominant for small scale sequencing • Retrogen and similar companies • But, next generation sequencing has already rapidly displaced old technology in genome centers. – 454 sequencing (Roche) – Solexa (Illumina) *dominant player at the moment* – So. LID (Applied Biosystems) (dead technology due to poor support) • 3 rd generation sequencing (individual DNA molecule) now available – e. g. , Pacific Biosciences (sequence reads of 1, 000 -10 K bases) – Oxford Biosciences Nanopore (read length 5 kb— 200 kb) Bio. Sci D 145 lecture 4 page 4 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

DNA sequence analysis • Landmarks in DNA sequencing – Sanger, Nicklen and Coulson. Sequencing with chain terminating inhibitors. Proc. Natl. Acad. Sci. 74, 5463 -5467 (1977). – Sanger, F. et al. The nucleotide sequence of bacteriophage ΦX 174. J Mol Biol 125, 225 -46. (1978). – Sutcliffe, J. G. Complete nucleotide sequence of the Escherichia coli plasmid p. BR 322. Cold Spring Harb Symp Quant Biol 43, 77 -90. (1979). – Sanger et al. , Nucleotide sequence of bacteriophage lambda DNA. J Mol Biol 162, 729 -73. (1982). – Messing, J. , Crea, R. & Seeburg, P. H. A system for shotgun DNA sequencing. Nucl. Acids Res 9, 309 -21 (1981). – Anderson, S. et al. Sequence and organization of the human mitochondrial genome. Nature 290, 457 -65 (1981). – Deininger, P. L. Random subcloning of sonicated DNA: application to shotgun DNA sequence analysis. Anal Biochem 129, 216 -23. (1983). – Baer et al. DNA sequence and expression of the B 95 -8 Epstein-Barr virus genome. Nature 310, 207 -11. (1984). (189 kb) – Innis et al. DNA sequencing with Taq DNA polymerase and direct sequencing of PCR-amplified DNA Proc. Natl. Acad. Sci. 85, 9436 -9440 (1988) Bio. Sci D 145 lecture 4 page 5 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

DNA sequence analysis (contd) • Landmarks in DNA sequencing (contd). – 1995 - Haemophilus influenzae (1. 83 Mb) • first bacterium sequenced, human pathogen – 1995 - Mycoplasma genitalium (0. 58 Mb) • smallest free living organism – 1996 - Saccharomyces cerevisiae genome (13 Mb) – 1996 - Methanococcus jannaschii (1. 66 Mb) • first Archaebacterium – 1997 - Escherichia coli (4. 6 Mb) – 1997 - Bacillus subtilis (4. 2 Mb) – 1997 - Borrelia burgdorferi (1. 44 Mb) • Lyme disease – 1997 - Archaeoglobus fulgidus (2. 18 Mb) • first sulfur metabolizing bacterium – 1997 - Helicobacter pylori (1. 66 Mb) • first bacterium proven to cause cancer Bio. Sci D 145 lecture 4 page 6 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

DNA sequence analysis (contd) • Landmarks in DNA sequencing (contd) – 1998 - Treponema pallidum (1. 14 Mb) – 1998 - Caenorhabditis elegans genome (97 Mb) – 1999 - Deinococcus radiodurans (3. 28 Mb) • resistant to radiation, starvation, ox stress – 2000 - Drosophila melanogaster (120 Mb) – 2000 - Arabidopsis thaliana (115 Mb) – 2001 - Escherichia coli O 157: H 7 (4. 1 Mb) • Pathogenic variant of E. coli – 2001 – draft Human “genome” – 2002 – mouse genome – 2002 – Ciona intestinalis – – – • Primitive chordate 2003 – “complete “human genome 2004 – rat genome 2006 – Human “genome” complete sequence of all chromosomes 2010 – Neanderthal genome sequenced 2012 – Denisovan genome sequenced Bio. Sci D 145 lecture 4 page 7 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

DNA Sequence analysis • Complete DNA sequence (all nts both strands, no gaps) – complete sequence is desirable but takes time • how long depends on size and strategy employed – which strategy to use depends on various factors • how large is the clone? – c. DNA – genomic • How fast is sequence required? • sequencing strategies – primer walking – cloning and sequencing of restriction fragments – progressive deletions • Bidirectional, unidirectional – Shotgun sequencing • whole genome • with mapping – map first (C. elegans) – map as you go (many) Bio. Sci D 145 lecture 4 page 8 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
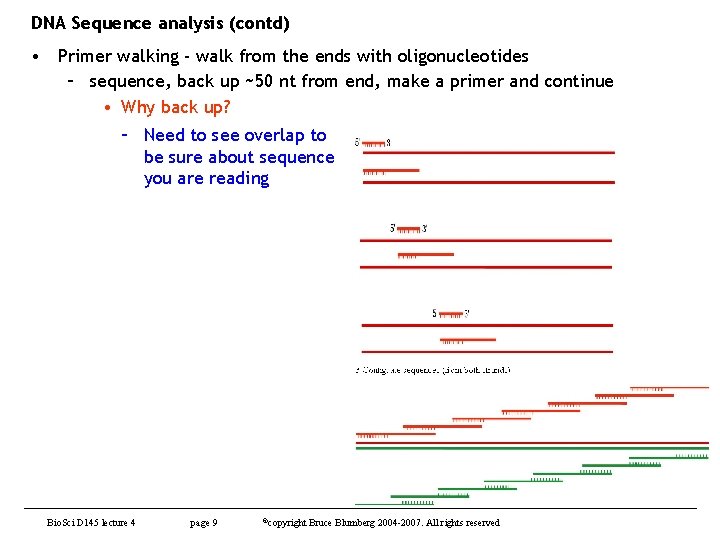
DNA Sequence analysis (contd) • Primer walking - walk from the ends with oligonucleotides – sequence, back up ~50 nt from end, make a primer and continue • Why back up? – Need to see overlap to be sure about sequence you are reading Bio. Sci D 145 lecture 4 page 9 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

DNA Sequence analysis (contd) • Primer walking (contd) – advantages • very simple • no possibility to lose bits of DNA – restriction mapping – deletion methods • no restriction map needed • best choice for short DNA – disadvantages • slowest method – about a week between sequencing runs • oligos are not free (and not reusable) • not feasible for large sequences – applications • c. DNA sequencing when time is not critical • targeted sequencing – verification – closing gaps in sequences Bio. Sci D 145 lecture 4 page 10 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

DNA Sequence analysis (contd) • Cloning and sequencing of restriction fragments – once the most popular method • make a restriction map, subclone fragments • sequence – advantages • straightforward • directed approach • can go quickly • cloned fragments often useful otherwise – RNase protection, nuclease mapping, in situ hybridization – disadvantages • possible to lose small fragments – must run high quality analytical gels • depends on quality of restriction map – mistaken mapping -> wrong sequence • restriction site availability – applications • sequencing small c. DNAs • isolating regions to close gaps Bio. Sci D 145 lecture 4 page 11 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

DNA Sequence analysis (contd) • nested deletion strategies - sequential deletions from one end of the clone – cut, close and sequence • Approach – make restriction map – use enzymes that cut in polylinker and insert – Religate, sequence from end with restriction site – repeat until finished, filling in gaps with oligos • advantages – Fast, simple, efficient • disadvantages – limited by restriction site availability in vector and insert – need to make a restriction map Bio. Sci D 145 lecture 4 page 12 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
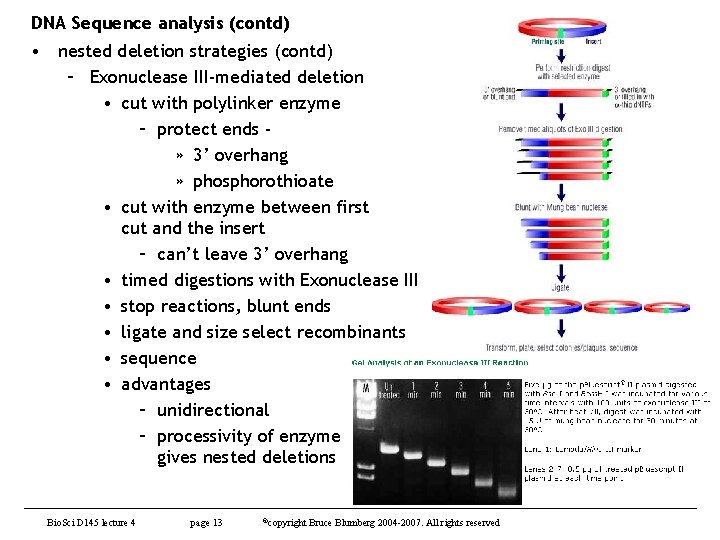
DNA Sequence analysis (contd) • nested deletion strategies (contd) – Exonuclease III-mediated deletion • cut with polylinker enzyme – protect ends » 3’ overhang » phosphorothioate • cut with enzyme between first cut and the insert – can’t leave 3’ overhang • timed digestions with Exonuclease III • stop reactions, blunt ends • ligate and size select recombinants • sequence • advantages – unidirectional – processivity of enzyme gives nested deletions Bio. Sci D 145 lecture 4 page 13 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

DNA Sequence analysis (contd) • Nested deletion strategies – Exonuclease III-mediated deletion (contd) • disadvantages – need two unique restriction sites flanking insert on each side – best used successively to get > 10 kb total deletions – may not get complete overlaps of sequences » fill in with restriction fragments or oligos • applications – method of choice for moderate size sequencing projects » c. DNAs » genomic clones – good for closing larger gaps • Small-scale sequence analysis – how is it practiced today? – Primer walking – Exo. III-mediated deletion with primer walking Bio. Sci D 145 lecture 4 page 14 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

Genome sequencing • The problem – Genome sizes for most eukaryotes are large (108 -109 bp) – High quality sequences only about 600 -800 bp per run (Sanger) – Nextgen sequences ~150 bp/read • The solution – Break genome into lots of bits and sequence them all – Reassemble with computer • The benefit – Rapid increase in information about genome size, gene comparisons, etc • The cost – 3 x 109 bp(human haploid genome) ÷ 600 bp/reaction = 5 x 106 reactions for 1 x coverage! – Need both strands (x 2), need overlaps and need to be sure of sequences – ~107 -108 reactions/runs required for a human-sized genome – About $1 -2 per reaction these days, ~$8 commercially. Bio. Sci D 145 lecture 4 page 15 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
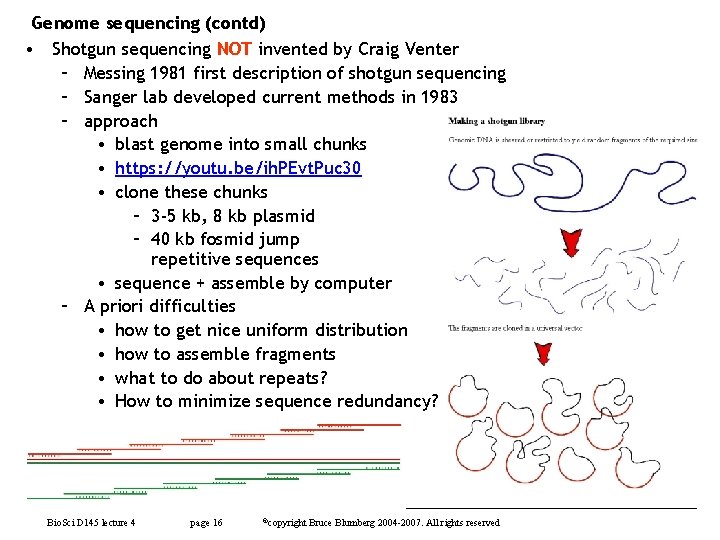
Genome sequencing (contd) • Shotgun sequencing NOT invented by Craig Venter – Messing 1981 first description of shotgun sequencing – Sanger lab developed current methods in 1983 – approach • blast genome into small chunks • https: //youtu. be/ih. PEvt. Puc 30 • clone these chunks – 3 -5 kb, 8 kb plasmid – 40 kb fosmid jump repetitive sequences • sequence + assemble by computer – A priori difficulties • how to get nice uniform distribution • how to assemble fragments • what to do about repeats? • How to minimize sequence redundancy? Bio. Sci D 145 lecture 4 page 16 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
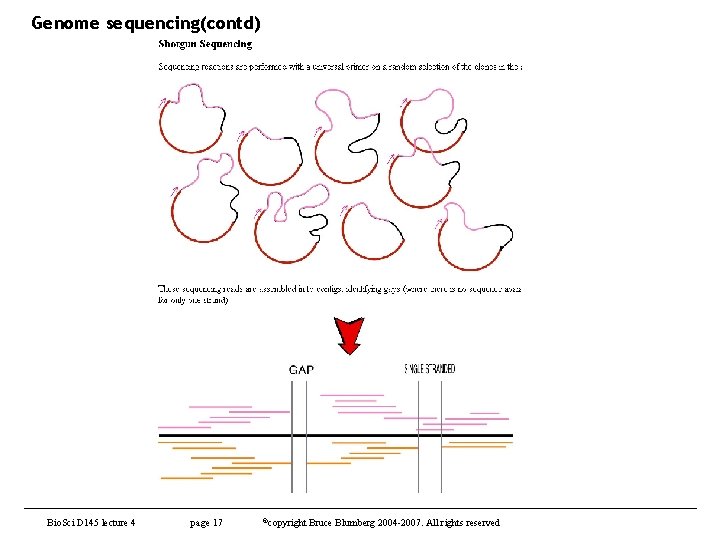
Genome sequencing(contd) Bio. Sci D 145 lecture 4 page 17 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
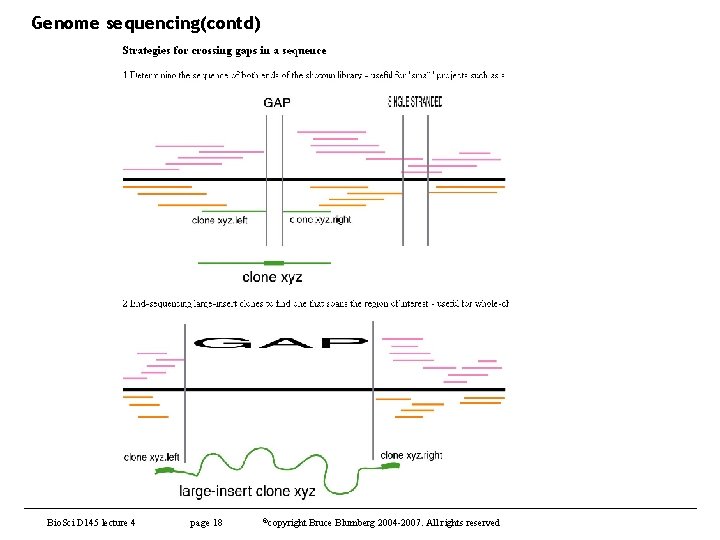
Genome sequencing(contd) Bio. Sci D 145 lecture 4 page 18 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

Genome sequencing (contd) • Shotgun sequencing (contd) – How to minimize sequence redundancy? • Best way to minimize redundancy is map before you start – C. elegans was done this way - when the sequence was finished, it was FINISHED » mapping took almost 10 years – mapping much too tedious and nonprofitable for Celera » who cares about redundancy, let’s sequence and make $$ » There is scientific value to draft genomes, too. • why does redundancy matter? – Finished sequence today costs about $0. 50/base – Note that 10 x, 99. 995% coverage leaves at least 150 kb unsequenced Bio. Sci D 145 lecture 4 page 19 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
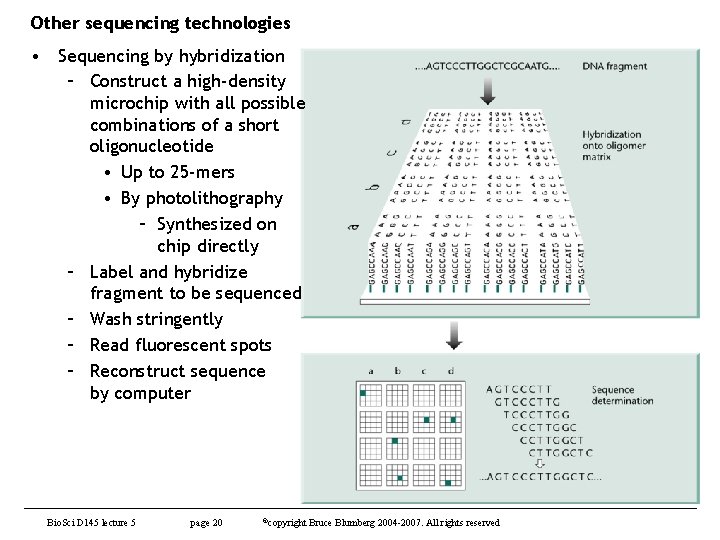
Other sequencing technologies • Sequencing by hybridization – Construct a high-density microchip with all possible combinations of a short oligonucleotide • Up to 25 -mers • By photolithography – Synthesized on chip directly – Label and hybridize fragment to be sequenced – Wash stringently – Read fluorescent spots – Reconstruct sequence by computer Bio. Sci D 145 lecture 5 page 20 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
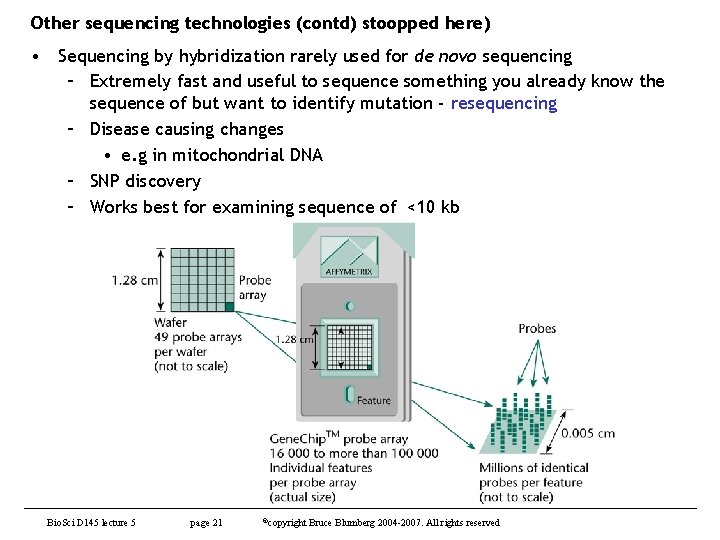
Other sequencing technologies (contd) stoopped here) • Sequencing by hybridization rarely used for de novo sequencing – Extremely fast and useful to sequence something you already know the sequence of but want to identify mutation - resequencing – Disease causing changes • e. g in mitochondrial DNA – SNP discovery – Works best for examining sequence of <10 kb Bio. Sci D 145 lecture 5 page 21 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

Other sequencing technologies (contd) • https: //www. thermofisher. com/us/en/home/life-science/microarrayanalysis/affymetrix. html? nav. Mode=35810&a. Id=products. Nav • SNP discovery – Photo shows mitochondrial chip – Right panel shows pairs of normal (top) vs disease (bottom) (Leber’s Hereditary Optic Neuropathy) • Top 3 disease mutations • Bottom control with no change Bio. Sci D 145 lecture 5 page 22 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

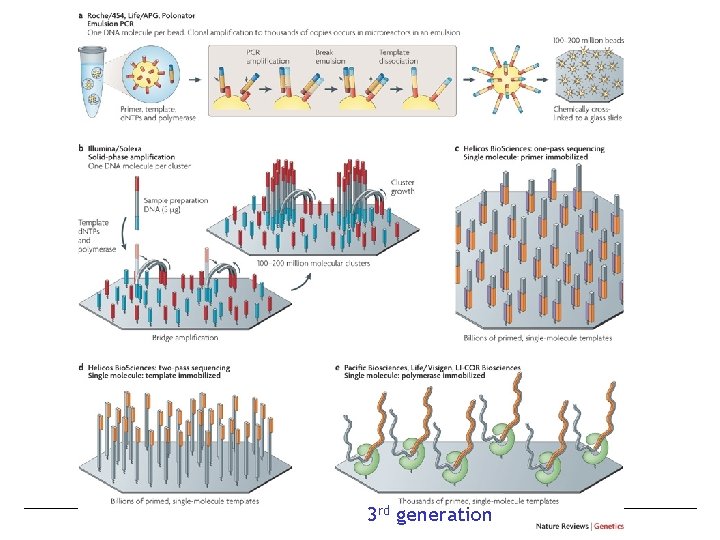
Other sequencing technologies – Next Generation sequencing • 2 nd generation = high throughput, short sequences • 3 rd generation = single molecule sequencing • Small number of sequence templates (thousands) but very long reads (~105 bp) • What is the immediate implication of this technology for genome assembly? We should now be able to completely sequence large insert clones directly and avoid fragmentation by repetitive elements! • See Metzger, M. L. (2010) Sequencing technologies — the next generation, Nature Reviews Genetics 11, 31 -46. Bio. Sci D 145 lecture 5 page 23 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
3 rd generation

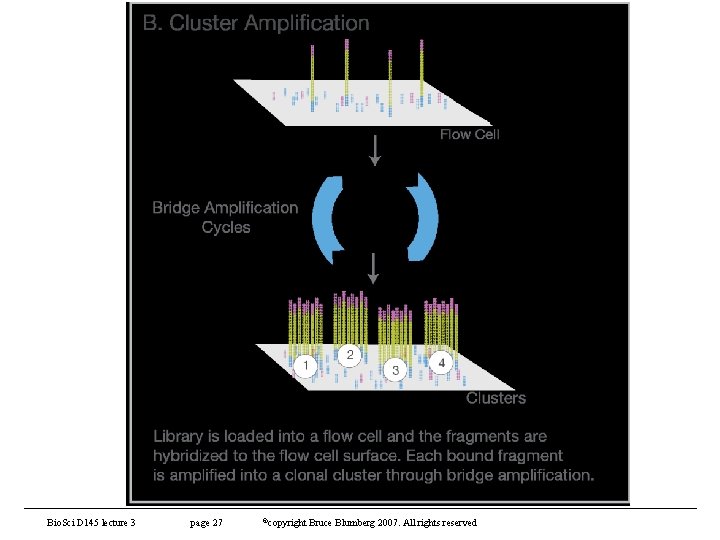
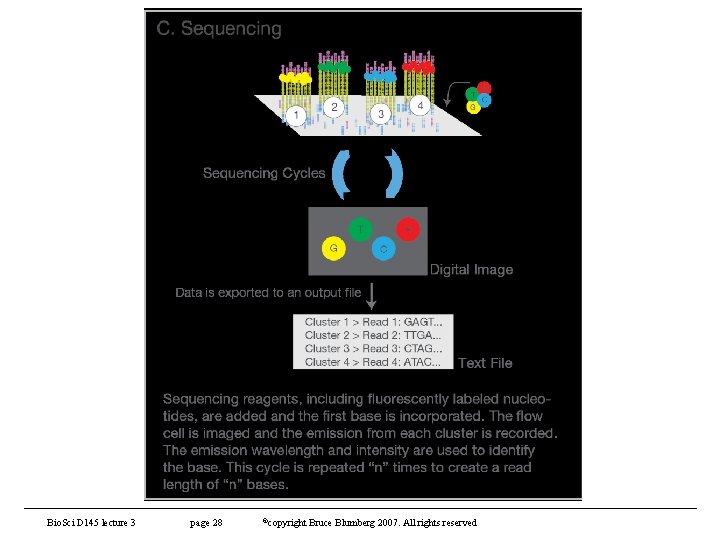
Other sequencing technologies (contd) • Illumina (Solexa) sequencing – https: //www. illumina. com/content/dam/illuminamarketing/documents/products/illumina_sequencing_introduction. pdf – Based on synthesis of complementary strand to a template (like Sanger) • Detection of elongation with labeled terminators – Steps • Library generation - fragment genome to appropriate size (depends on application) and adapters to each end • Cluster generation – capture fragments on lawn of oligos and amplify • Sequencing – reversible terminator • Data analysis – – align reads to reference genome – Analysis of reads Bio. Sci D 145 lecture 5 page 25 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

Other sequencing technologies (contd) • Illumina sequencing (contd) – Library preparation – fragment target and adapters. • Can multiplex to gain additional capacity • That is, Hiseq-X can generate 1. 8 Tb sequence per run, but we don’t need this much for most applications so use different adapters and “bar-code” samples. • This way, you can get many sequences from one run and then deconvolute them • also has advantage of removing batch effects – Can directly compare all sequences with each other because they come from same run of machine. Bio. Sci D 145 lecture 5 page 26 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
Bio. Sci D 145 lecture 3 page 27 ©copyright Bruce Blumberg 2007. All rights reserved
Bio. Sci D 145 lecture 3 page 28 ©copyright Bruce Blumberg 2007. All rights reserved

Bio. Sci D 145 lecture 3 page 29 ©copyright Bruce Blumberg 2007. All rights reserved
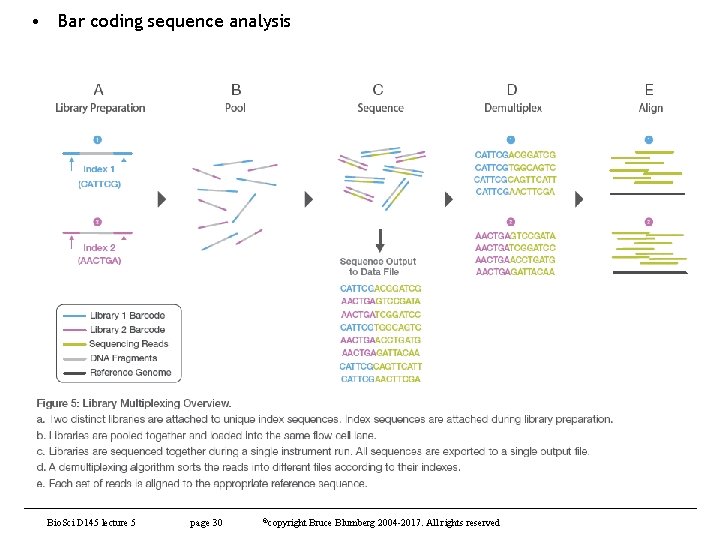
• Bar coding sequence analysis Bio. Sci D 145 lecture 5 page 30 ©copyright Bruce Blumberg 2004 -2017. All rights reserved
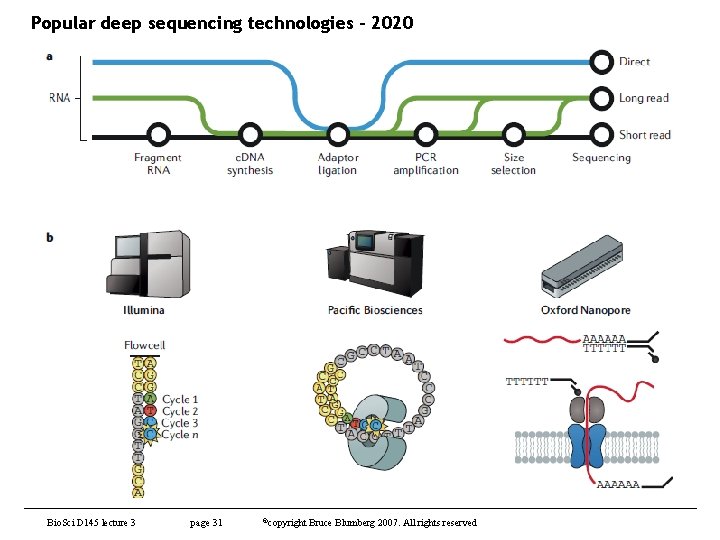
Popular deep sequencing technologies - 2020 Bio. Sci D 145 lecture 3 page 31 ©copyright Bruce Blumberg 2007. All rights reserved
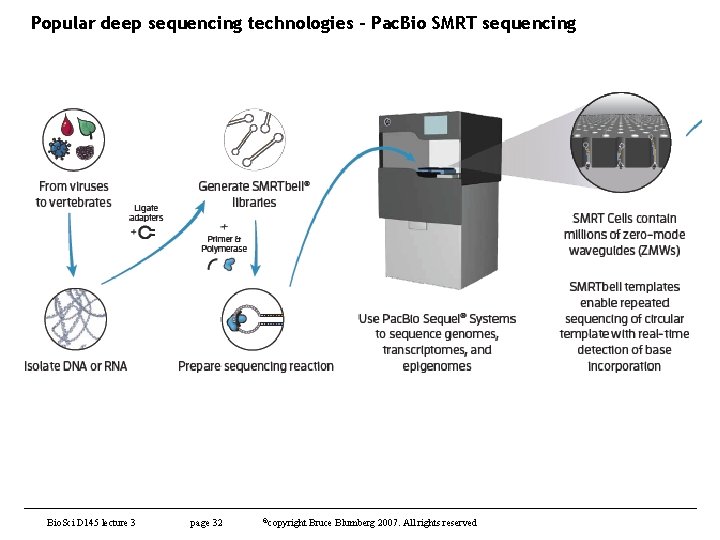
Popular deep sequencing technologies – Pac. Bio SMRT sequencing Bio. Sci D 145 lecture 3 page 32 ©copyright Bruce Blumberg 2007. All rights reserved
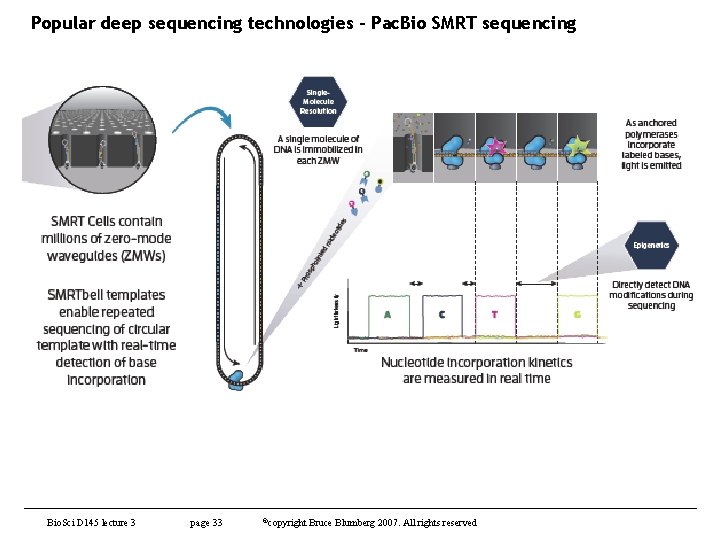
Popular deep sequencing technologies – Pac. Bio SMRT sequencing Bio. Sci D 145 lecture 3 page 33 ©copyright Bruce Blumberg 2007. All rights reserved
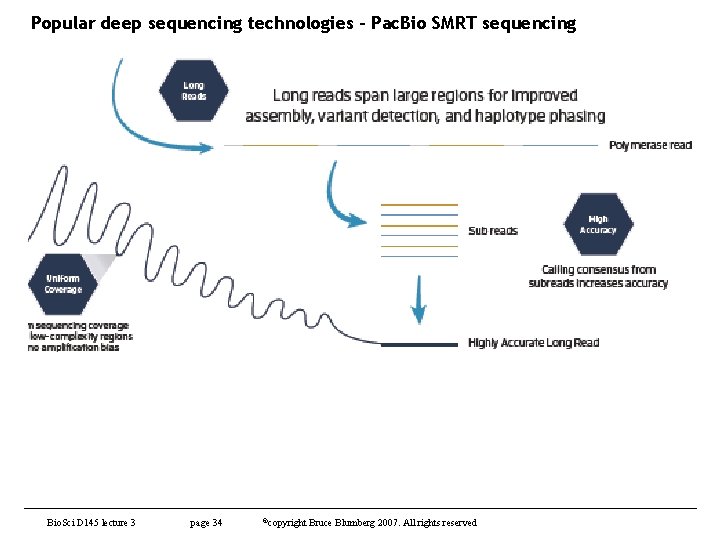
Popular deep sequencing technologies – Pac. Bio SMRT sequencing Bio. Sci D 145 lecture 3 page 34 ©copyright Bruce Blumberg 2007. All rights reserved
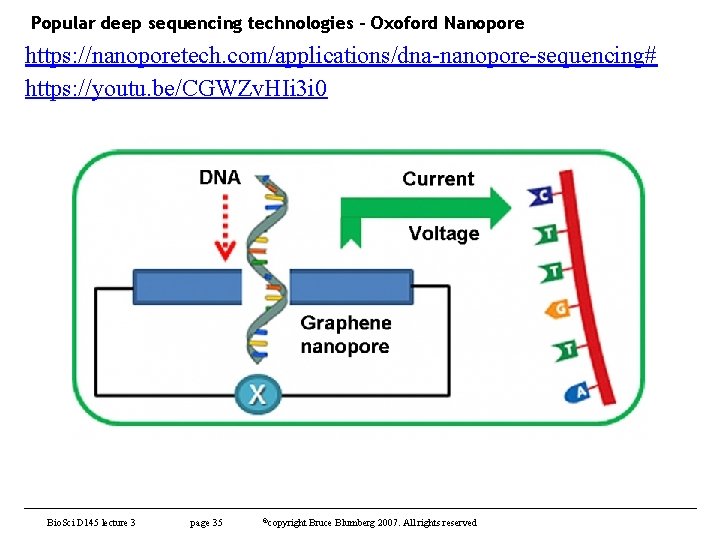
Popular deep sequencing technologies – Oxoford Nanopore https: //nanoporetech. com/applications/dna-nanopore-sequencing# https: //youtu. be/CGWZv. HIi 3 i 0 Bio. Sci D 145 lecture 3 page 35 ©copyright Bruce Blumberg 2007. All rights reserved

Other sequencing technologies (contd) • Deep sequencing - what is the point? – Can generate huge number of reads in parallel • i. Seq 100 – 1. 2 Gb (4 million reads/run, 2 x 150 bp) • Miniseq – 7. 5 Gb (25 million reads/run, 2 x 150 bp) • Mi. Seq – 15 gb (15 million reads/run, 2 x 300 bp) • Next. Seq – 120 Gb (400 million reads/run, 2 x 150 bp) • Hi. Seq – 1. 5 Tb (5 billion/run, 2 x 150 bp) • Hiseq. X – 1. 8 Tb (6 billion/run, 2 x 150 bp) • Novaseq – 6. 0 Tb (20 billion/run, 2 x 150 bp) • What is massively parallel sequencing good for? – Rapid sequencing of genomes, or resequencing of known sequences – Ancient DNA (even dinosaurs? ) § Probably not, 1. 5 million years appears to be upper limit – Ch. IP-sequencing – Sequencing ESTs or other tags – Determining microbial diversity in field samples – Transcriptome sequencing – Single cell sequencing – Identification of infrequent variants in large populations (e. g. , viruses) Bio. Sci D 145 lecture 5 page 36 ©copyright Bruce Blumberg 2004 -2007. All rights reserved
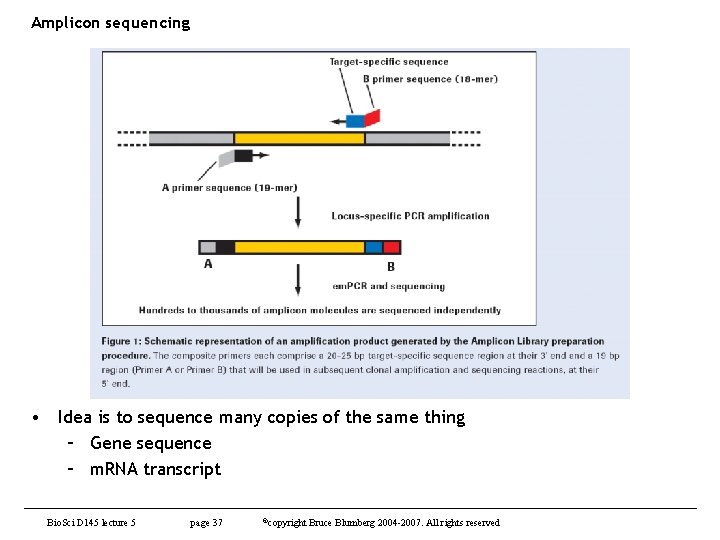
Amplicon sequencing • Idea is to sequence many copies of the same thing – Gene sequence – m. RNA transcript Bio. Sci D 145 lecture 5 page 37 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

Amplicon sequencing (contd) • What is amplicon sequencing good for? – Discovery of rare somatic mutations in complex samples (e. g. , cancerous tumors - mixed with germline DNA) based on ultra-deep sequencing of amplicons – Sequencing collections of exons from populations of individuals to identify diversity – Sequencing collections of human exons from populations of individuals for the identification of rare alleles associated with disease – Analysis of viral quasi-species present within infected populations in the context of epidemiological studies (find virulent mutations in population) – Evolutionary biology in populations Bio. Sci D 145 lecture 5 page 38 ©copyright Bruce Blumberg 2004 -2007. All rights reserved

The human genome • In Feb 12 2001, Celera and Human Genome project published “draft” human genome sequencs – Celera -> 39114 – Ensembl -> 29691 – Consensus from all sources ~30 K • Number of genes – C. elegans – 19, 000 – Arabidopsis - 25, 000 • Predictions had been from 50 -140 k human genes – What’s up with that? – Are we only slightly more complicated than a weed? – How can we possibly get a human with less than 2 x the number of genes as C. elegans – Implications? • UNRAVELING THE DNA MYTH: The spurious foundation of genetic engineering, Barry Commoner, Harpers Magazine Feb, 2002 Bio. Sci D 145 lecture 4 page 39 ©copyright Bruce Blumberg 2004 -2016. All rights reserved

The human genome • The answer – Gene sets don’t overlap completely (duh) – Floor is 42 K – 130029 build #236 Uni. Gene Clusters (from EST and m. RNA sequencing) – http: //www. ncbi. nlm. nih. gov/unigene – Up from 123, 459 in 2013 (85, 793, 105, 680, 128, 826, 123, 891 previous years) (“final” count • Important questions to be answered about what constitutes a “gene” – Crick genes? DNA-RNA-protein – How about RNAs? – mi. RNAs? – Antisense transcripts? – lnc. RNAs? Bio. Sci D 145 lecture 4 page 40 ©copyright = 42113 Bruce Blumberg 2004 -2016. All rights reserved

Genome sequencing(contd) – Whole genome shotgun sequencing (Celera) • premise is that rapid generation of draft sequence is valuable • why bother trying to clone and sequence difficult regions? – Basically just forget regions of repetitive DNA - not cost effective • using this approach, genomes rarely are completely finished – rule of thumb is that it takes at least as long to finish the last 5% as it took to get the first 95% • problems – sequence may never be complete as is C. elegans – much redundant sequence with many sparse regions and lots of gaps. – Fragment assembly for regions of highly repetitive DNA is dubious at best – “Finished” fly and human genomes lack more than a few already characterized genes Bio. Sci D 145 lecture 4 page 41 ©copyright Bruce Blumberg 2004 -2016. All rights reserved

Genome sequencing (contd) • Knowing what we know – how to approach a large new genome? – Xenopus tropicalis 1. 7 Gb (about ½ human) – BAC end sequencing – Whole genome shotgun – HAPPY mapping and radiation hybrid mapping to order scaffolds – Gaps closed with BACs – 8. 5 x coverage (but > 9000 scaffolds for 18 chromosomes) • 2019 update – now version 10. 0 – FINALLY integrated BAC end sequences and genetic map – 99. 86% of genome mapped to chromosomes • 167 scaffolds, ~150 Mbp, 10 chromosomes – ~45 k protein coding genes • Xenopus laevis – v 9. 2 • >90% of genome in chromosomal scaffolds • 2 “subgenomes” fully characterized. Bio. Sci D 145 lecture 4 page 42 ©copyright Bruce Blumberg 2004 -2016. All rights reserved
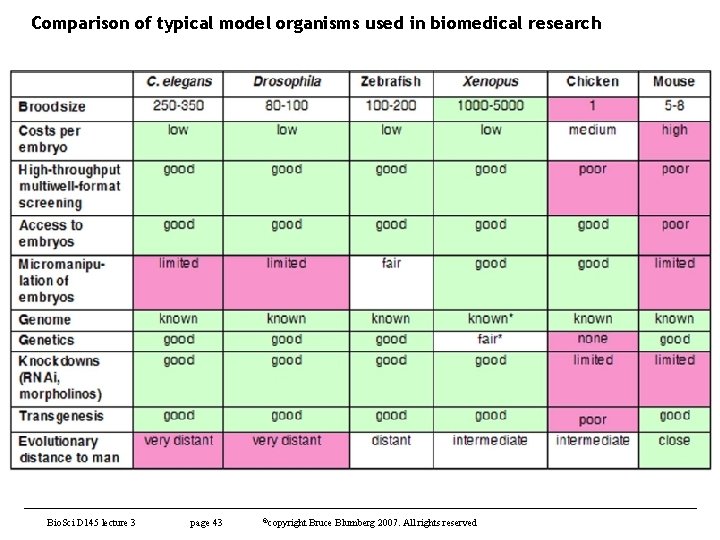
Comparison of typical model organisms used in biomedical research Bio. Sci D 145 lecture 3 page 43 ©copyright Bruce Blumberg 2007. All rights reserved
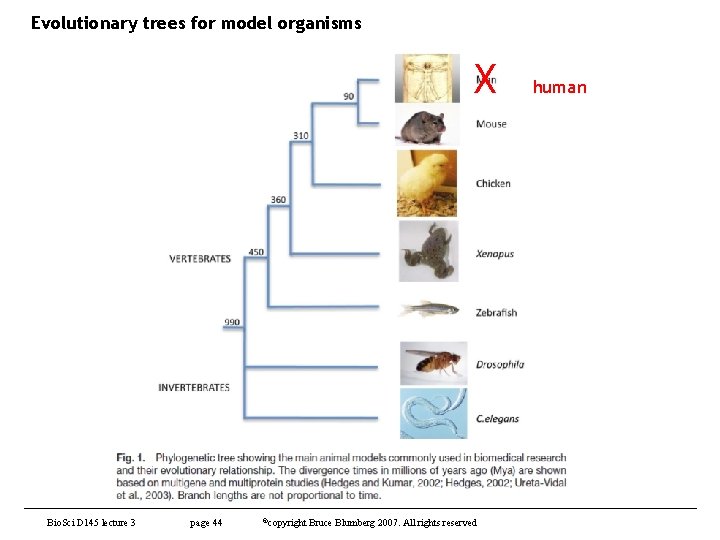
Evolutionary trees for model organisms X Bio. Sci D 145 lecture 3 page 44 ©copyright Bruce Blumberg 2007. All rights reserved human
- Slides: 44