Bio Sci D 145 Lecture 1 Bruce Blumberg

Bio. Sci D 145 Lecture #1 • Bruce Blumberg (blumberg@uci. edu) – 4103 Nat Sci 2 - office hours Tu, Th 3: 30 -5: 00 (or by appointment) – phone 824 -8573 • TA – Angela Kuo (akuo 4@uci. edu) – 4311 Nat Sci 2– office hours TBA – Phone 824 -6873 • check e-mail regularly for announcements, etc. . • Lectures will be posted in advance (without answers) • Updated lectures (with answers) will be posted after lecture – http: //blumberg-lab. bio. uci. edu/biod 145 -w 20120 Bio. Sci D 145 lecture 1 page 1 ©copyright Bruce Blumberg 2020. All rights reserved

Introductions and Goals • Let’s introduce each other – Name – Major – Favorite thing about UCI – Least favorite thing about UCI Bio. Sci D 145 lecture 1 page 2 ©copyright Bruce Blumberg 2020. All rights reserved
Class requirements • Grading Midterm Final exam Presentation Term paper Participation 35% 10% 10% (attendance, class discussion) • How are grades determined? • 20 minute presentation and discussion of a journal article is required • These will be randomly assigned – Angela will schedule yours • Presentations will be done as teams for most papers (depending on class size) • Volunteers for 1/16 ? See Angela. • Attendance and participation is important • Please come to class having read assigned material • Final examination will not be cumulative, however, understanding of concepts and techniques from first part of course is required. Bio. Sci D 145 lecture 1 page 3 ©copyright Bruce Blumberg 2020. All rights reserved

General comments • Overall philosophy – This class is about understanding genomic and proteomic (i. e. whole genome) approaches to problems of biological interest • Focus will be on research problems – Intended to be informative and cutting edge but also interesting and relevant, even fun. – Office hours are after class but I am always around – Questions are welcome • Please stop me and ask questions if something is unclear – I am going to ask you questions • Answers get participation credit • Memorizing vs. understanding – I am not concerned with your memory – This course is about problem solving – how to address interesting biological problems using modern, whole-genome approaches Bio. Sci D 145 lecture 1 page 4 ©copyright Bruce Blumberg 2020. All rights reserved

General comments • Letters of recommendation – If you want a letter from me, I need to know you as more than a student number and grade • come to office hours • participate in class discussions • make your interest in the subject apparent Bio. Sci D 145 lecture 1 page 5 ©copyright Bruce Blumberg 2020. All rights reserved

About the texts • Bookstore vs. online? • Neither text book is absolutely required – Brown has lots of introductory material that will help to fill in background between Bio. Sci 99 and this class • Reading noted in text books are intended to supplement lecture material • Source of material for this class will be lectures and assigned papers. Bio. Sci D 145 lecture 1 page 6 ©copyright Bruce Blumberg 2020. All rights reserved

Requirements for the term paper • Goals – Analytical thinking – Improved writing • Select a topic of interest to you and then propose a whole genome approach to address the problem (not necessarily your 199 research!) – Talk with me about your topic (so that I can help you focus it on something do-able and rewarding to you) • Write a short paper (5 pages) in the style of a pre-doctoral fellowship proposal describing how you will attack this problem (examples posted). – Specific aims (~1/2 page) • Hypotheses to be tested • How will you test hypotheses? – Background and significance (1 -2 pages) • What is known, what remains to be learned • why should someone give you money to study this problem? – Research plan (~3 pages) • specific experiments to answer the questions posed in specific aims • How will you handle expected vs. unexpected results Bio. Sci D 145 lecture 1 page 7 ©copyright Bruce Blumberg 2020. All rights reserved

Requirements for the term paper (contd) • Outline (due Friday February 7 (24: 00) – Title and topic – Introductory paragraph telling why the problem is important – What is the hypothesis that your proposed research will address? – Enumerate 1 -3 specific aims in the form of questions that will test aspects of your hypothesis • Topic can be changed later, if necessary • What is a hypothesis? – A supposition or conjecture put forth to account for known facts; esp. in the sciences, a provisional supposition from which to draw conclusions that shall be in accordance with known facts, and which serves as a starting-point for further investigation by which it may be proved or disproved and the true theory arrived at. • What is a theory ? – An analytical framework that explains a set of observations – A comprehensive explanation of an important feature of nature that is supported by facts that have been repeatedly confirmed through observation and experiment Bio. Sci D 145 lecture 1 page 8 ©copyright Bruce Blumberg 2020. All rights reserved

Requirements for the oral presentation • Goal – again to get you to think more analytically – Exposure to literature (classic and current) – Learn critical reading – Discuss practical applications of what we are learning • Powerpoint (“journal club”) presentation – as a presenter – 15 -20 minutes with time allowed for discussion (max of 15 – 20 slides) – Frame the problem – what are the big picture questions? • What was known before they started? What was unknown? • Present background (not more than 5 slides) – What are specific questions asked or hypotheses tested • Discuss figures – What is the question being asked in each figure or panel? – What experiments did the authors do to answer questions? – Do the data support the conclusions drawn? • What did they conclude overall? • What could have been improved? – Point out a few papers for further reading (reviews, follow-ups, etc) – Summarize main points and key techniques used in the last slide Bio. Sci D 145 lecture 1 page 9 ©copyright Bruce Blumberg 2020. All rights reserved

Requirements for the oral presentation (contd) • Powerpoint presentation – as a listener – READ THE PAPERS – you are responsible for the material covered – Study the figures • What points don’t you understand? – Make notations, ask the speaker to clarify these – Listen to the speaker • If presentation is unclear, ask the speaker to elaborate • Always feel free to ask questions – we want an open discussion • Papers are posted on the web sites listed • Logistics – Prepare presentation in Powerpoint or PDF (not Keynote) and either email to me or bring it on a USB stick. Bio. Sci D 145 lecture 1 page 10 ©copyright Bruce Blumberg 2020. All rights reserved
Presentation schedule • • • • • Week 1 papers – Dear and Cook, 1993, Jiang et al, 2011 (Angela) Week 2 papers – (1) Geisler et al. , 1999 (2) Redon et al. , 2006 (3) Venter et al. , 2004 Week 3 papers – (4) Bentley et al. , 2008 (5) Lindblad-Toh et al. , 2011 (6) Sessions et al. , 2016 Week 4 papers – (7) Kapranov et al. , 2007 (8) Morrison et al. , 2017 (9) Owens et al. , 2016 Week 5 papers - (10) Cheng et al. , 2019 (11) Chen et al. , 2012 (12) Silvert et al. , 2019 Week 6 papers –Midterm, no presentations Week 7 papers – (13) Flyamer et al. , 2017 (14) Buenrostro et al. , 2012 (15) Argelaguet et al. , 2019 Week 8 papers – (16) Gilbert et al. , 2014 (17) Anzalone et al. , 2019 (18) Luo et al. , 2009 Week 9 papers – (19) Ito et al. , 2001 (20) Dejardin and Kingston, 2009 (21) Gavin et al. , 2002 Week 10 papers - (22) David et al. , 2014 (23) Rampelli et al. , 2015 (24) Tang et al. , 2019 Bio. Sci D 145 lecture 1 page 11 ©copyright Bruce Blumberg 2020. All rights reserved

Lecture Outline – Organization and Structure of Genomes • Today’s topics – Genome complexity – Implications of split genes for protein diversity – Repetitive elements and gene evolution • The big picture for the next 2 lectures – How are genomes similar and different? – How do we find out this information? – Why do we care? • What is genomics? Proteomics? – ‘omics is the study of a property using “whole genome” approach – Genomics – study of genes and gene function – Proteomics – study of all the proteins Bio. Sci D 145 lecture 1 page 12 ©copyright Bruce Blumberg 2020. All rights reserved

The rise of -omics • The -omics revolution of science – http: //www. genomicglossaries. com/content/omes. asp • What does it all mean? – Transcriptomics – large scale profiling of gene expression – Proteomics – study of complement of expressed proteins – Functional genomics – vague term, typically encompasses many others – Structural genomics – prediction of structure and interactions from sequence (Rick Lathrop, Pierre Baldi) – Pharmacogenomics – transcriptional profiling of response to drug treatment – often looking for genetic basis of differences – Toxicogenomics – transcriptional profiling of response to toxicants (often includes pharmacogenomics • Seeks mechanistic understanding of toxic response – Metabolomics – analysis of total metabolite pool ("metabolome") to reveal novel aspects of cellular metabolism and global regulation – Interactomics – genome wide study of macromolecular interactions, physical and genetic are included – Bibliomics – identifying words that occur together in papers Sadly, usually just abstracts Bio. Sci D 145 lecture 1 page 13 ©copyright Bruce Blumberg 2020 All rights reserved

Organization and Structure of Genomes (contd) • Genome size – i. e. total number of DNA bp – Varies widely - WHY? C- paradox – i. e. , what is the source of the differences? • Do the number of genes required vary so much? unlikely — (how many “phyla” are represented at the right? ) Mixed bag Bio. Sci D 145 lecture 1 page 14 ©copyright Bruce Blumberg 2020. All rights reserved Phylum Chordata Phylum Arthropoda

Organization and Structure of Genomes (contd) • How to measure genome complexity? – Hybridization kinetics – Shear and melt DNA – Allow to hybridize and measure doublestranded vs. single-stranded by spectrophotometry • Cot½ - measures genome size and complexity – What does a large value (longer to hybridize) mean? • k is smaller (rate constant slower) • Longer to hybridize – more unique sequences, larger genome – Much of what we knew about genome size and complexity (until advent of genome sequencing) comes from these studies Bio. Sci D 145 lecture 1 page 15 ©copyright Bruce Blumberg 2020. All rights reserved
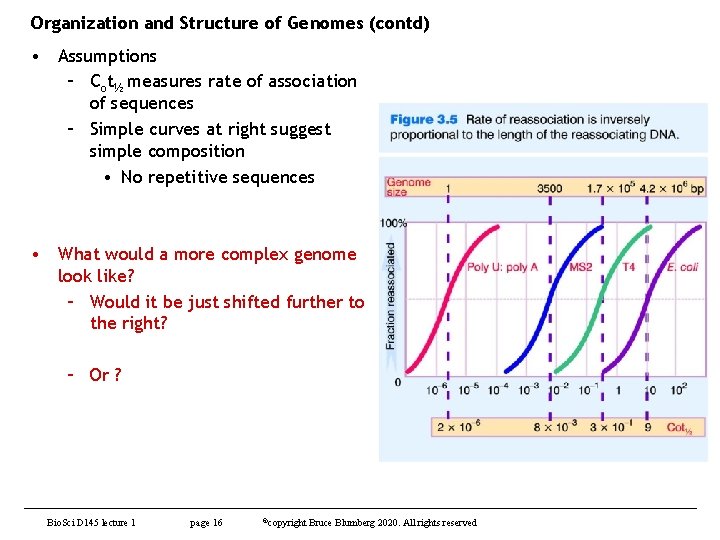
Organization and Structure of Genomes (contd) • Assumptions – Cot½ measures rate of association of sequences – Simple curves at right suggest simple composition • No repetitive sequences • What would a more complex genome look like? – Would it be just shifted further to the right? – Or ? Bio. Sci D 145 lecture 1 page 16 ©copyright Bruce Blumberg 2020. All rights reserved

Organization and Structure of Genomes (contd) • Measure eukaryotic DNA – Multiple components – Can calculate more than 1 Cot½ value – Either means starting material is not pure (i. e. , multiple types of DNA) – Or means different frequency classes of DNA • Highly repetitive • Moderately repetitive • Unique – Very big surprise Bio. Sci D 145 lecture 1 page 17 ©copyright Bruce Blumberg 2020. All rights reserved

Organization and Structure of Genomes (contd) • What can we conclude from great variation in genome size ? Genetic complexity is not directly proportional to genome size! • Increase in C is not always accompanied by proportional increase in number of genes — Gene number is controversial — Depends on what is a “gene” — Are we no more complicated than a weed (Arabidopsis) ? Bio. Sci D 145 lecture 1 page 18 ©copyright Bruce Blumberg 2020. All rights reserved
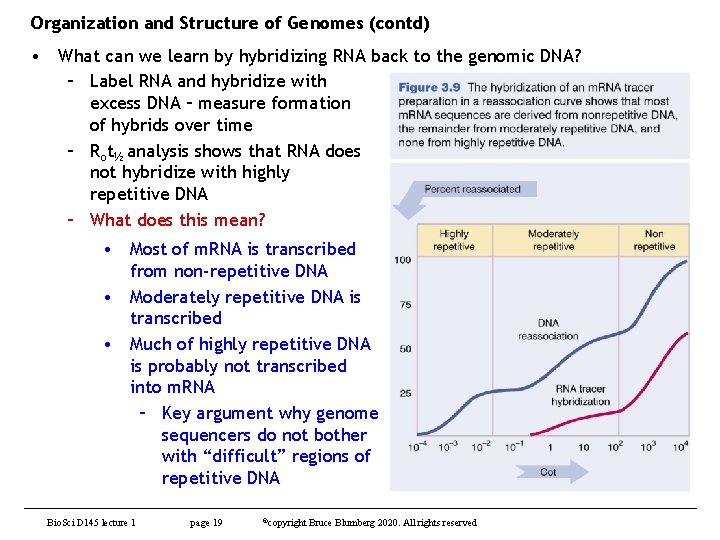
Organization and Structure of Genomes (contd) • What can we learn by hybridizing RNA back to the genomic DNA? – Label RNA and hybridize with excess DNA – measure formation of hybrids over time – Rot½ analysis shows that RNA does not hybridize with highly repetitive DNA – What does this mean? • Most of m. RNA is transcribed from non-repetitive DNA • Moderately repetitive DNA is transcribed • Much of highly repetitive DNA is probably not transcribed into m. RNA – Key argument why genome sequencers do not bother with “difficult” regions of repetitive DNA Bio. Sci D 145 lecture 1 page 19 ©copyright Bruce Blumberg 2020. All rights reserved
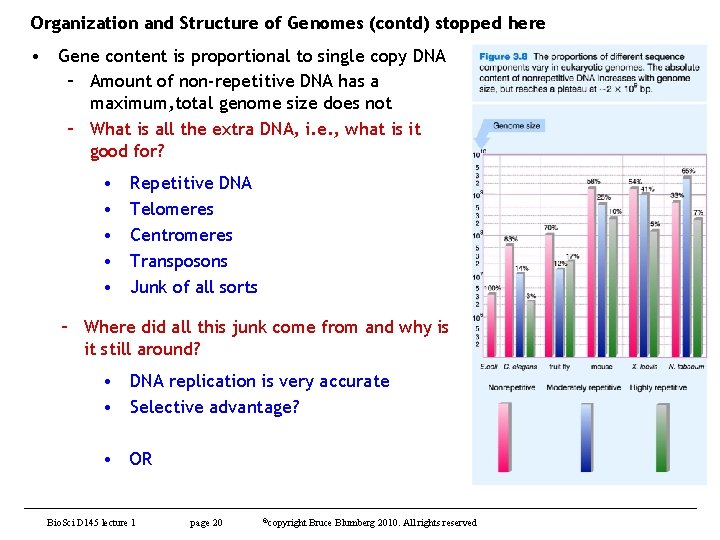
Organization and Structure of Genomes (contd) stopped here • Gene content is proportional to single copy DNA – Amount of non-repetitive DNA has a maximum, total genome size does not – What is all the extra DNA, i. e. , what is it good for? • • • Repetitive DNA Telomeres Centromeres Transposons Junk of all sorts – Where did all this junk come from and why is it still around? • DNA replication is very accurate • Selective advantage? • OR Bio. Sci D 145 lecture 1 page 20 ©copyright Bruce Blumberg 2010. All rights reserved

Organization and Structure of Genomes (contd) • What is this highly repetitive DNA? • Selfish DNA? – Parasitic sequences that exist solely to replicate themselves? • Or evolutionary relics? – Produced by recombination, duplication, unequal crossing over • Probably both – Transposons exemplify “selfish DNA” • Akin to viruses? – Crossing over and other recombination lead to large scale duplications • ENCODE (encyclopedia of DNA elements) considers > 80% of genome to be functional. − But see Grauer et al, 2013. Genome Biol Evol 5: 578 -590. On the immortality of television sets: “function” in the human genome according to the evolution-free gospel of ENCODE Bio. Sci D 145 lecture 1 page 21 ©copyright Bruce Blumberg 2010. All rights reserved

Transcription of Prokaryotic vs Eukaryotic genomes (stopped here) • Prokaryotic genes are expressed in linear order on chromosome – m. RNA corresponds directly to g. DNA • Most eukaryotic genes are interrupted by non-coding sequences – Introns (Gilbert 1978) – These are spliced out after transcription and prior to transport out of nucleus – Post-transcriptional processing in an important feature of eukaryotic gene regulation • Why do eukaryotes have introns, i. e. , what are they good for? • Main function may be to generate protein diversity • Harbor regulatory sequences Bio. Sci D 145 lecture 1 page 22 ©copyright Bruce Blumberg 2010. All rights reserved
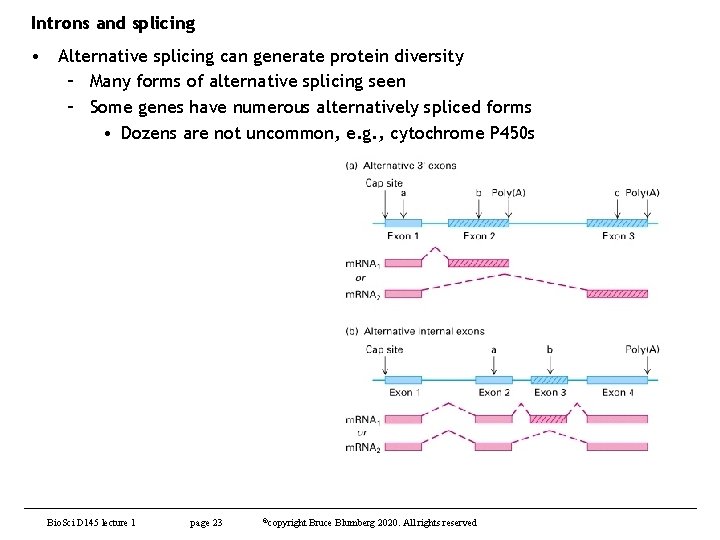
Introns and splicing • Alternative splicing can generate protein diversity – Many forms of alternative splicing seen – Some genes have numerous alternatively spliced forms • Dozens are not uncommon, e. g. , cytochrome P 450 s Bio. Sci D 145 lecture 1 page 23 ©copyright Bruce Blumberg 2020. All rights reserved
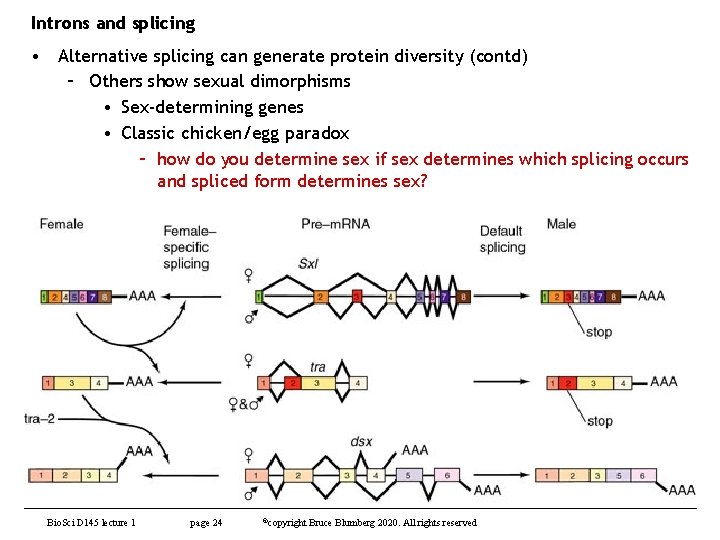
Introns and splicing • Alternative splicing can generate protein diversity (contd) – Others show sexual dimorphisms • Sex-determining genes • Classic chicken/egg paradox – how do you determine sex if sex determines which splicing occurs and spliced form determines sex? Bio. Sci D 145 lecture 1 page 24 ©copyright Bruce Blumberg 2020. All rights reserved
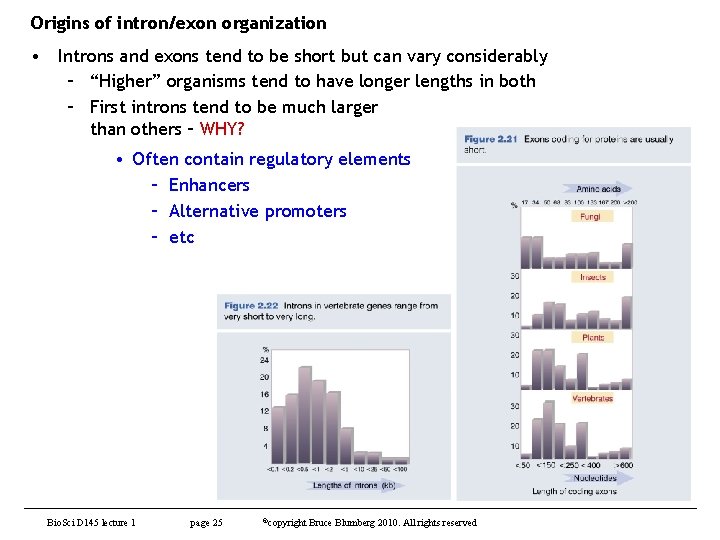
Origins of intron/exon organization • Introns and exons tend to be short but can vary considerably – “Higher” organisms tend to have longer lengths in both – First introns tend to be much larger than others – WHY? • Often contain regulatory elements – Enhancers – Alternative promoters – etc Bio. Sci D 145 lecture 1 page 25 ©copyright Bruce Blumberg 2010. All rights reserved
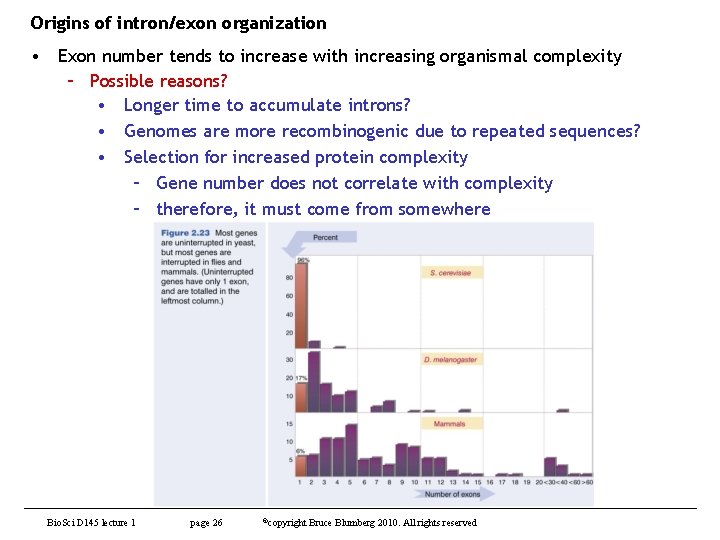
Origins of intron/exon organization • Exon number tends to increase with increasing organismal complexity – Possible reasons? • Longer time to accumulate introns? • Genomes are more recombinogenic due to repeated sequences? • Selection for increased protein complexity – Gene number does not correlate with complexity – therefore, it must come from somewhere Bio. Sci D 145 lecture 1 page 26 ©copyright Bruce Blumberg 2010. All rights reserved

Origins of intron/exon organization • When did introns arise – Introns early – Walter Gilbert • There from the beginning, lost in bacteria and many simpler organisms – Introns late – Cavalier-Smith, Ford Doolittle, Russell Doolittle • Introns acquired over time as a result of transposable elements, aberrant splicing, etc • If introns benefit protein evolution – why would they be lost? – Which is it? Actin • Introns “late” (at the moment) • But late = ~580 million years ago • What is common factor among animals that share intron locations? All deuterostomes (echinoderms, chordates, hemichordates, xenoturbellids – diverged about 580 x 106 years ago Bio. Sci D 145 lecture 1 page 27 ©copyright Bruce Blumberg 2010. All rights reserved

Evolution of gene clusters • Many genes occur as multigene families (e. g. , actin, tubulin, globins, Hox) – Inference is that they evolved from a common ancestor – Families can be • clustered - nearby on chromosomes (α-globins, Hox. A) • Dispersed – on various chromosomes (actin, tubulin) • Both – related clusters on different chromosomes (α, β-globins, Hox. A, B, C, D) – Members of clusters may show stage or tissue-specific expression • Implies means for coregulation as well as individual regulation • Much recent evidence showing that gene clusters occur on loops of chromatin that are bounded by particular structural protein (cohesins, CTCF) Bio. Sci D 145 lecture 1 page 28 ©copyright Bruce Blumberg 2010. All rights reserved
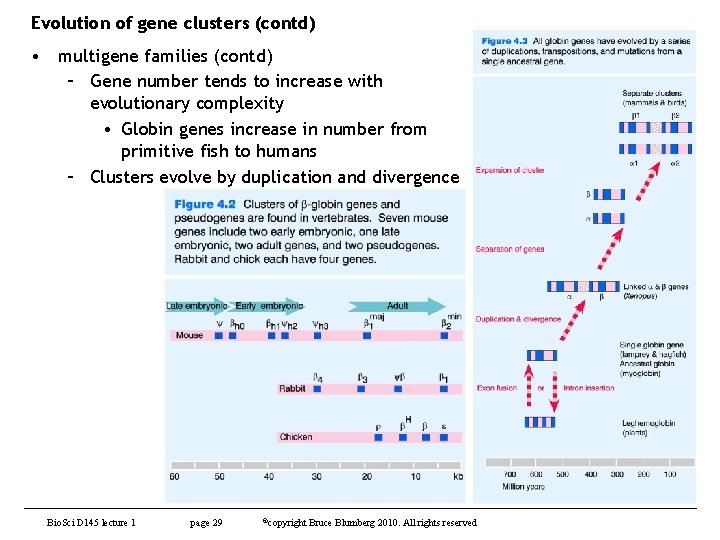
Evolution of gene clusters (contd) • multigene families (contd) – Gene number tends to increase with evolutionary complexity • Globin genes increase in number from primitive fish to humans – Clusters evolve by duplication and divergence Bio. Sci D 145 lecture 1 page 29 ©copyright Bruce Blumberg 2010. All rights reserved
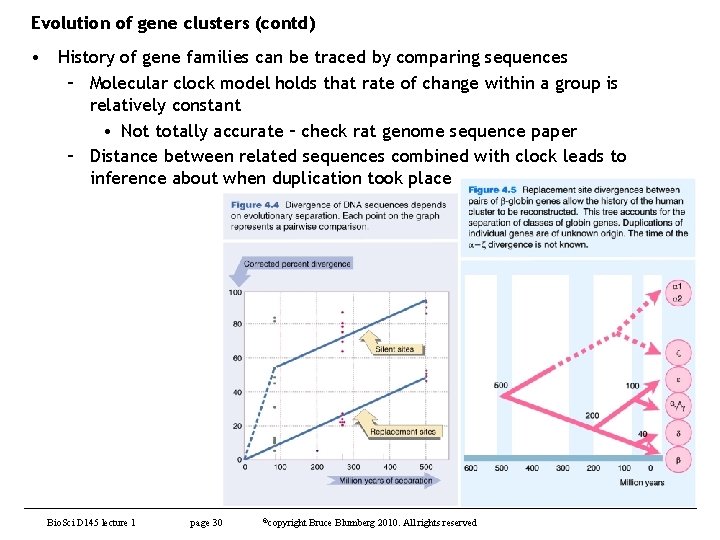
Evolution of gene clusters (contd) • History of gene families can be traced by comparing sequences – Molecular clock model holds that rate of change within a group is relatively constant • Not totally accurate – check rat genome sequence paper – Distance between related sequences combined with clock leads to inference about when duplication took place Bio. Sci D 145 lecture 1 page 30 ©copyright Bruce Blumberg 2010. All rights reserved

Types and origin of repetitive elements • DNA sequences are not random – genes, restriction sites, methylation sites • Repeated sequences are not random either – Some occur as tandemly repeated sequences – Usually generated by unequal crossing over during meiosis – These resolve in ultracentrifuge into satellite bands because GC content differs from majority of DNA – This “satellite” DNA is highly variable • Between species • And among individuals within a population • Can be useful for mapping genotyping, etc – Much highly repetitive DNA is in heterochromatin (highly condensed regions) • Centromeres are one such place Bio. Sci D 145 lecture 2 page 31 ©copyright Bruce Blumberg 2007. All rights reserved
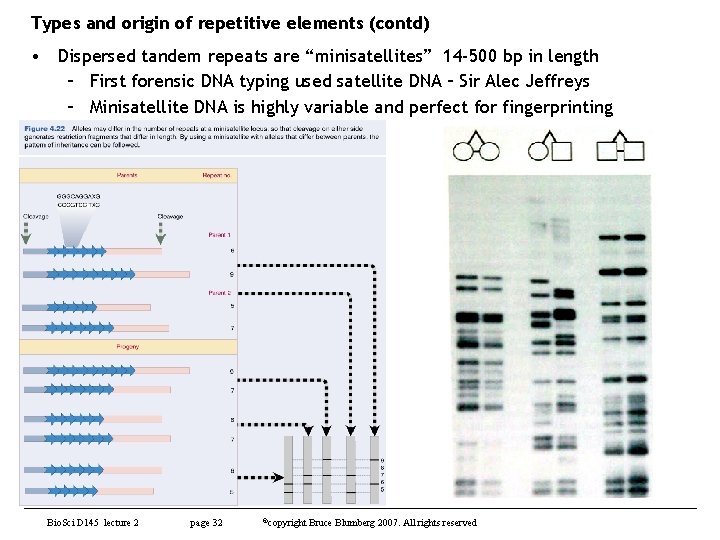
Types and origin of repetitive elements (contd) • Dispersed tandem repeats are “minisatellites” 14 -500 bp in length – First forensic DNA typing used satellite DNA – Sir Alec Jeffreys – Minisatellite DNA is highly variable and perfect for fingerprinting Bio. Sci D 145 lecture 2 page 32 ©copyright Bruce Blumberg 2007. All rights reserved
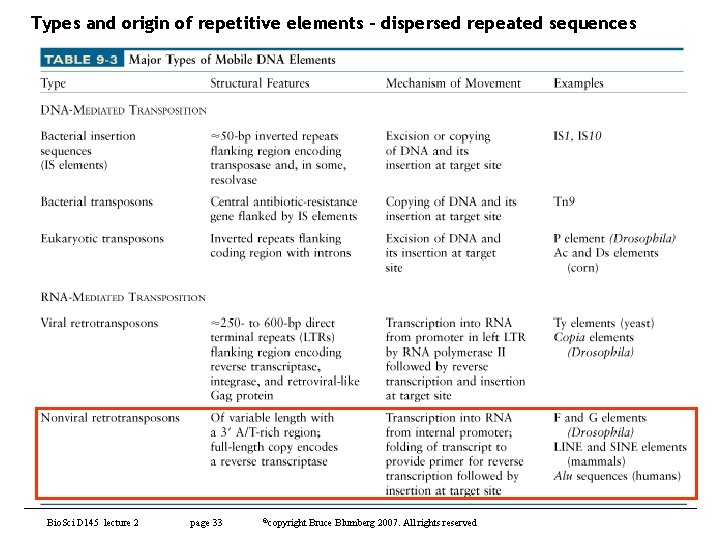
Types and origin of repetitive elements – dispersed repeated sequences Bio. Sci D 145 lecture 2 page 33 ©copyright Bruce Blumberg 2007. All rights reserved
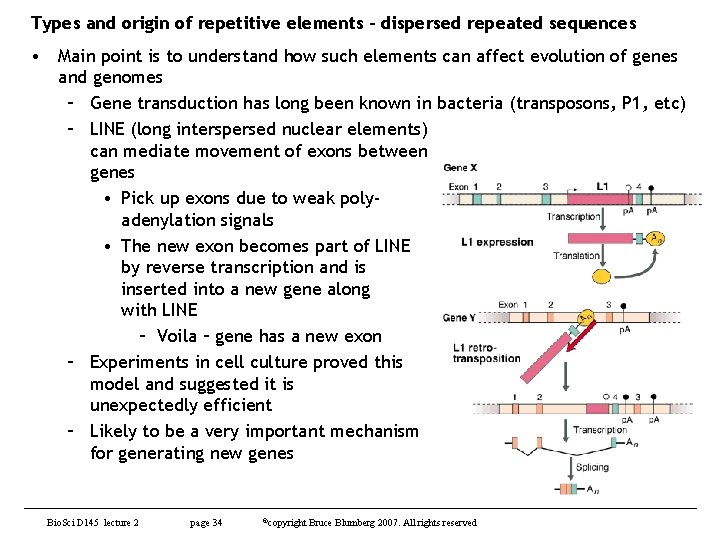
Types and origin of repetitive elements – dispersed repeated sequences • Main point is to understand how such elements can affect evolution of genes and genomes – Gene transduction has long been known in bacteria (transposons, P 1, etc) – LINE (long interspersed nuclear elements) can mediate movement of exons between genes • Pick up exons due to weak polyadenylation signals • The new exon becomes part of LINE by reverse transcription and is inserted into a new gene along with LINE – Voila – gene has a new exon – Experiments in cell culture proved this model and suggested it is unexpectedly efficient – Likely to be a very important mechanism for generating new genes Bio. Sci D 145 lecture 2 page 34 ©copyright Bruce Blumberg 2007. All rights reserved
- Slides: 34