Bio Sci 145 B Lecture 3 4202004 Bruce

Bio. Sci 145 B Lecture #3 4/20/2004 • Bruce Blumberg – 2113 E Mc. Gaugh Hall - office hours Wed 12 -1 PM (or by appointment) – phone 824 -8573 – blumberg@uci. edu • TA – Curtis Daly – 2113 Mc. Gaugh Hall, 924 -6873, 3116 – Office hours Tuesday 11 -12 (or by appointment) • lectures will be posted on web pages after lecture – http: //eee. uci. edu/04 s/05705/ - link only here – http: //blumberg-serv. bio. uci. edu/bio 145 b-sp 2004 – http: //blumberg. bio. uci. edu/bio 145 b-sp 2004 Bio. Sci 145 B lecture 3 page 1 ©copyright Bruce Blumberg 2004. All rights reserved
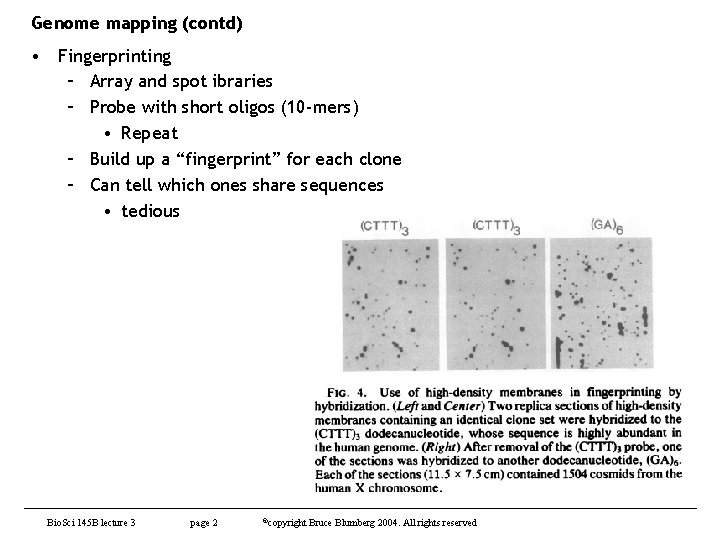
Genome mapping (contd) • Fingerprinting – Array and spot ibraries – Probe with short oligos (10 -mers) • Repeat – Build up a “fingerprint” for each clone – Can tell which ones share sequences • tedious Bio. Sci 145 B lecture 3 page 2 ©copyright Bruce Blumberg 2004. All rights reserved
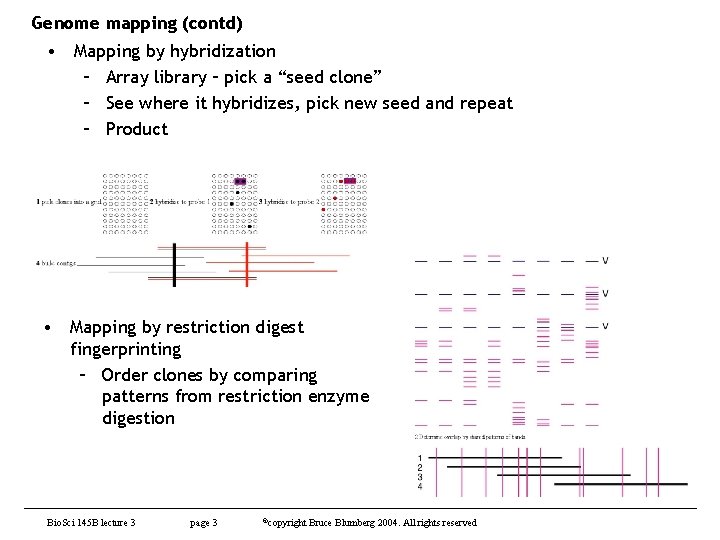
Genome mapping (contd) • Mapping by hybridization – Array library – pick a “seed clone” – See where it hybridizes, pick new seed and repeat – Product • Mapping by restriction digest fingerprinting – Order clones by comparing patterns from restriction enzyme digestion Bio. Sci 145 B lecture 3 page 3 ©copyright Bruce Blumberg 2004. All rights reserved
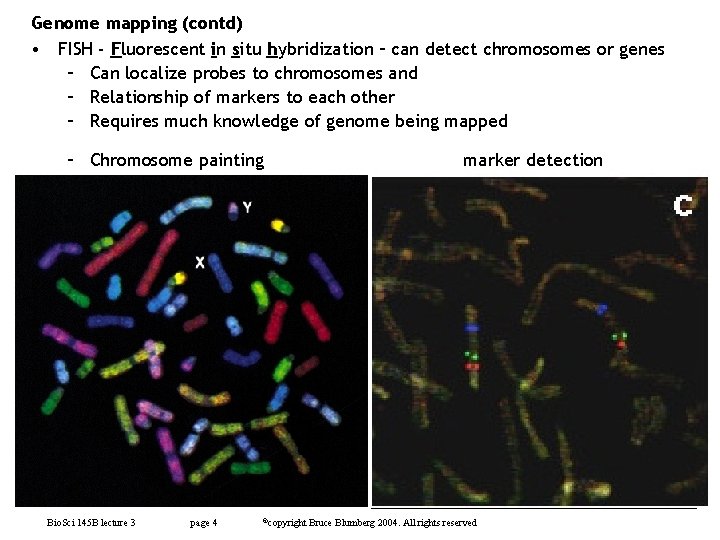
Genome mapping (contd) • FISH - Fluorescent in situ hybridization – can detect chromosomes or genes – Can localize probes to chromosomes and – Relationship of markers to each other – Requires much knowledge of genome being mapped – Chromosome painting Bio. Sci 145 B lecture 3 page 4 ©copyright marker detection Bruce Blumberg 2004. All rights reserved

Genome mapping (contd) • Radiation hybrid mapping – Old but very useful technique • Lethally irradiate cells with X-rays • Fuse with cells of another species, e. g. , blast human cells then fuse with hamster cells – Chunks of human DNA will remain in mouse cells • Expand colonies of cells to get a collection of cell lines, each containing a single chunk of human c. DNA • Collection = RH panel – Now map markers onto these RH panels • Can identify which of any type of markers map together – STS, EST (very commonly used), etc • Can then map others by linkage to the ones you have mapped – Compare RH panel with other maps • Utility – great for cloning gaps in other maps Bio. Sci 145 B lecture 3 page 5 ©copyright Bruce Blumberg 2004. All rights reserved

Genome mapping (contd) • How should maps be made with current knowledge? – All methods have strengths and weaknesses – must integrate data for useful map • e. g, RH panel, BAC maps, STS, ESTs – Size and complexity of genome is important • More complex genomes require more markers and time mapping – Breakpoints and markers are mapped relative to each other – Maps need to be defined by markers (cities, lakes, roads in analogy) – Key part of making a finely detailed map is construction of genomic libraries and cell lines for common use • Efforts by many groups increase resolution and utility of maps • Current strategies – BAC end sequencing – Whole genome shotgun sequencing – EST sequencing – Mapping of above to RH panels Bio. Sci 145 B lecture 3 page 6 ©copyright Bruce Blumberg 2004. All rights reserved

Sequence stability in E. coli • How can we be sure that our genomic and c. DNA libraries do not change during growth and screening? • What are the sorts of factors that might modulate whether a sequence can be stably propagated in E. coli? – 1 – 2 – 3 Bio. Sci 145 B lecture 3 page 7 ©copyright Bruce Blumberg 2004. All rights reserved

Sequence stability in E. coli • toxicity – sequence may lead to the production of a toxic product or toxic levels of an otherwise innocuous product – more problematic with c. DNA than genomic clones • restriction - Raleigh 1987 Meth. Enzymol. 152, 130 -141 – virtually all microorganisms have systems to destroy non-endogenous DNA host range restriction • four classes of restriction endonucleases – very important for cloning purposes are recently discovered systems that degrade DNA containing 5 -methyl cytosine or 6 -methyl adenine. – If you are cloning genomic DNA, or hemimethylated c. DNA these are very important! • virtually all eukaryotic DNA contains 5 -methyl cytosine and/or 6 methyl adenine – mcr. A, B, C - methylcytosine – mrr - methyl adenine Bio. Sci 145 B lecture 3 page 8 ©copyright Bruce Blumberg 2004. All rights reserved

Sequence stability in E. coli (contd) • Restriction (contd) – foreign DNA escapes restriction 1/105 for Eco. K and Eco. B, 1/10 for mcr. A. – one needs to be conscious of the mcr and mrr restriction status of strains and packaging extracts to be used. • Recombination - Wyman and Wertman (1987) Meth Enzymol 152, 173 -180 – genomic DNA contains lots of repeated sequences • direct repeats • inverted repeats • interspersed repeats (e. g. Alu) – repeated sequences unstable in recombination proficient E. coli if in: • lambda • plasmid • cosmid – seems not to apply to single copy vectors such as BAC, PAC, fosmid • What does this imply? – ~30% of the human genome is unstable in plasmid or phage clones • phages with such sequences either don’t grow at all or get shorter with time Bio. Sci 145 B lecture 3 page 9 ©copyright Bruce Blumberg 2004. All rights reserved

Sequence stability in E. coli (contd) • Recombination (contd) – E. coli has a variety of recombination pathways. These are the major players in causing sequence underrepresentation • rec. A required for all pathways • rec. BCD - major recombination pathway • sbc. B, C - suppressor of B, C • minor pathways – rec. E – rec. F – rec. J • rule of thumb - the more recombination pathways mutated, the sicker the cells and the slower they grow – major players for inverted repeats are rec. BCD and sbc – rec. A is most important for stabilizing direct repeats and preventing plasmid concatamerization Bio. Sci 145 B lecture 3 page 10 ©copyright Bruce Blumberg 2004. All rights reserved

Sequence stability in E. coli (contd) • Plating a genomic library – whenever possible, select a cell type that is rec. A, rec. D, sbc. B and deficient in all restriction systems. • Conveniently, Eco. K, mcr. B, C and mrr are all linked and often deleted together in strains • can get more than 100 fold difference in numbers of phage between wild type and recombination deficient – rec. D is preferred over rec. B, C because rec. D promotes rolling circle replication in lambda which improves yields Bio. Sci 145 B lecture 3 page 11 ©copyright Bruce Blumberg 2004. All rights reserved

What do I need to know about E. coli genetics? • You look in a supplier’s catalog and see lots of E. coli with different genotypes of the following general form: – F’{lac. Iq Tn 10 (Tet. R)} mcr. A, Δ(mrr-hsd. RMS-mcr. BC), Φ 80 lac. ZΔM 15, Δlac. X 74, deo. R, rec. A 1, ara. D 139, Δ(ara-leu)7697, gal. U, gal. K, rps. L(Str. R), end. A 1, nup. G • Does this make any difference for your experiments? – Or should you simply follow the supplier’s instructions? – Or just use whatever people in the next lab are using without thinking about it? Bio. Sci 145 B lecture 3 page 12 ©copyright Bruce Blumberg 2004. All rights reserved

What do I need to know about E. coli genetics? • F’{lac. Iq Tn 10 (Tet. R)} mcr. A, Δ(mrr-hsd. RMS-mcr. BC), Φ 80 lac. ZΔM 15, Δlac. X 74, deo. R, rec. A 1, ara. D 139, Δ(ara-leu)7697, gal. U, gal. K, rps. L(Str. R), end. A 1, nup. G • restriction systems – mcr. A - cuts Cm 5 CGG – mcr. B, C - complex cuts at Gm 5 C – mrr - restricts 6 -methyl adenine containing DNA – Why are these important? – hsd. RMS - Eco. K restriction system • R cuts 5'-AAC(N)6 GTGC-3’ • M/S methylates A residues in this sequence • for stability of long repeated sequences – rec. A 1 - deficient in general recombination – rec. D - deficiency in Exonuclease V – sbc. B, C - Exonuclease I – deo. R - allows uptake of large DNA Bio. Sci 145 B lecture 3 page 13 ©copyright Bruce Blumberg 2004. All rights reserved

What do I need to know about E. coli genetics? (contd) • for lac color selection – lac. Z ΔM 15 either on F’ or on Φ 80 prophage – lac. Iq - constitutive expression of lac repressor. Prevents leaky expression of promoters containing lac operator • for high quality DNA preps – rec. A 1 - deficient in general recombination – end. A 1 - deficient in endonuclease I • if you buy ESTs from Research Genetics (In. Vitrogen) or Open. Biosystems – ton. A - resistant to bacteriophage T 1 • for recombinant protein expression – lon - protease deficiency – Omp. T - protease found in periplasmic space – most important protease inhibitor for E. coli protein preps is pepstatin A Bio. Sci 145 B lecture 3 page 14 ©copyright Bruce Blumberg 2004. All rights reserved

What do I need to know about E. coli genetics? (contd) • suppressors – sup. E - inserts glutamine at UAG (amber) codons – sup. F - inserts tyrosine at UAG (amber) codons • many older phages have S 100 am which can only be suppressed by sup. F – λZAP, λgt 11, λZip. LOX, Bio. Sci 145 B lecture 3 page 15 ©copyright Bruce Blumberg 2004. All rights reserved

Construction of c. DNA libraries • What is a c. DNA library? • What are they good for? Bio. Sci 145 B lecture 3 page 16 ©copyright Bruce Blumberg 2004. All rights reserved

Determinants of library quality • What constitutes a full-length c. DNA? – Strictly it is an exact copy of the m. RNA – full-length protein coding sequence considered acceptable for most purposes • m. RNA – full-length, capped m. RNAs are critical to making full-length libraries – cytoplasmic m. RNAs are best – WHY? • 1 st strand synthesis – complete first strand needs to be synthesized – issues about enzymes • 2 nd strand synthesis – thought to be less difficult than 1 st strand (probably not) • choice of vector – plasmids are best for EST sequencing – phages are best for manual screening • how will library quality be evaluated – test with 2, 4, 6, 8 kb probes to ensure that these are well represented Bio. Sci 145 B lecture 3 page 17 ©copyright Bruce Blumberg 2004. All rights reserved

c. DNA synthesis • Scheme – m. RNA is isolated from source of interest – 1 -10 μg are denatured annealed to primer containing d(T)n. V – reverse transcriptase copies m. RNA into c. DNA – DNA polymerase I and Rnase H convert remaining m. RNA into DNA – c. DNA is rendered blunt ended – linkers or adapters are added for cloning – c. DNA is ligated into a suitable vector – vector is introduced into bacteria • Caveats – there is lots of bad information out there • much is derived from vendors who want to increase sales of their enzymes or kits – all manufacturers do not make equality enzymes – most kits are optimized for speed at the expense of quality – small points can make a big difference in the final outcome Bio. Sci 145 B lecture 3 page 18 ©copyright Bruce Blumberg 2004. All rights reserved

c. DNA synthesis (contd) • Preparation of m. RNA – want minimum of non poly A+ m. RNAs – affinity chromatography on oligo d(T) or (U) – Oligo d(T)30 latex (Nippon Roche) works best overall (a. k. a. Oligo. Tex Qiagen) – 2 successive runs gives ~90% pure A+ m. RNA • denaturation of m. RNA – critical step – most protocols use heat denaturation • Heat RNA in the presence of metal ions = chemical cleavage! – CH 3 Hg. OH is method of choice for best libraries • Potent, reversible denaturant • But VERY TOXIC! Bio. Sci 145 B lecture 3 page 19 ©copyright Bruce Blumberg 2004. All rights reserved

c. DNA synthesis (contd) • First strand synthesis - lots of misinformation about enzymes – reverse transcriptase contains 2 subunits • polymerase • RNase H - critical for processivity of the enzyme! – What is processivity? – Manufacturers prefer to sell MMLV RNase H- RT – cloned and cheap – best enzyme for 1 st strand synthesis is AMV RT from Seikagaku America • But not best overall – thought that 1 st strand is main failure point in c. DNA synthesis - NOT – addition of 0. 6 M trehalose to AMV reactions increases yield • allows rxns to run at ~60° C – Betaine is very big help for MMLV RT Bio. Sci 145 B lecture 3 page 20 ©copyright Bruce Blumberg 2004. All rights reserved
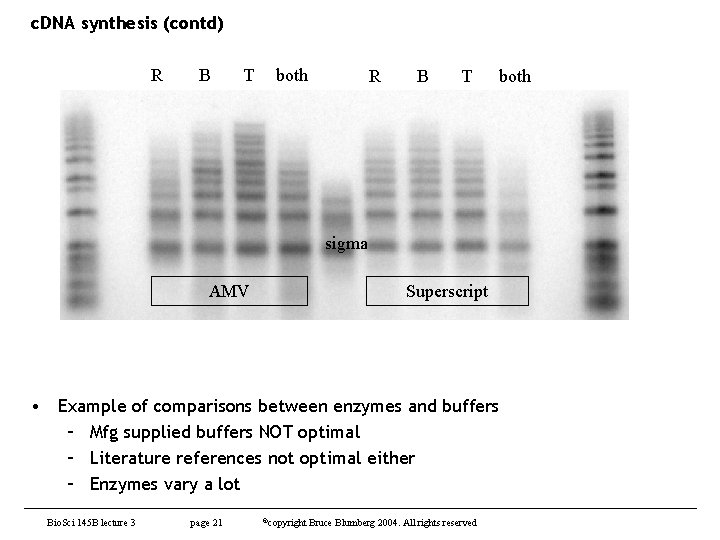
c. DNA synthesis (contd) R B T both sigma AMV Superscript • Example of comparisons between enzymes and buffers – Mfg supplied buffers NOT optimal – Literature references not optimal either – Enzymes vary a lot Bio. Sci 145 B lecture 3 page 21 ©copyright Bruce Blumberg 2004. All rights reserved

c. DNA synthesis (contd) • 2 nd strand – must remove m. RNA – best way is with RNAse H so that fragments serve as primers for DNA pol I – Gubler and Hoffman (1983) Gene 25, 263 – in my experience, 2 nd strand synthesis is the point of failure in c. DNA • virtually all kits shortcut this step (1 -2 hrs) • should be overnight • recent improvement is to use thermostable RNAse H, DNA ligase and DNA polymerase to maximize production of 2 nd strand. Bio. Sci 145 B lecture 3 page 22 ©copyright Bruce Blumberg 2004. All rights reserved
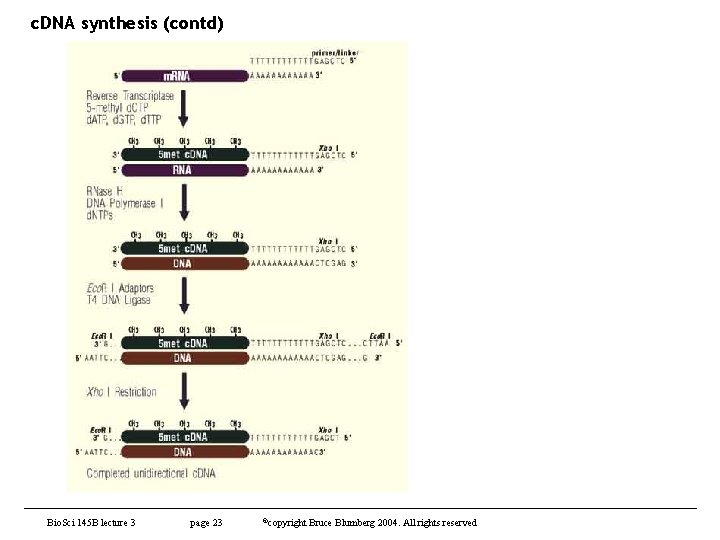
c. DNA synthesis (contd) Bio. Sci 145 B lecture 3 page 23 ©copyright Bruce Blumberg 2004. All rights reserved

c. DNA synthesis (contd) • Cloning – after 2 nd strand is made, the ends must be blunted and linkers or adapters added • usually T 4 DNA polymerase WHY? – perfect c. DNAs will retain 2 -20 bp of RNA at the 5’ end. • Linkers can not be added to this by any DNA ligase! • But T 4 RNA ligase can ligate DNA-RNA and stimulates blunt end ligation 10 x • no commercial products use T 4 RNA ligase so it is no wonder that fulllength c. DNAs are lost – if internal restriction sites have not been protected, they need to be methylated now before linkers are added. • Most methylase preps are not clean Bio. Sci 145 B lecture 3 page 24 ©copyright Bruce Blumberg 2004. All rights reserved
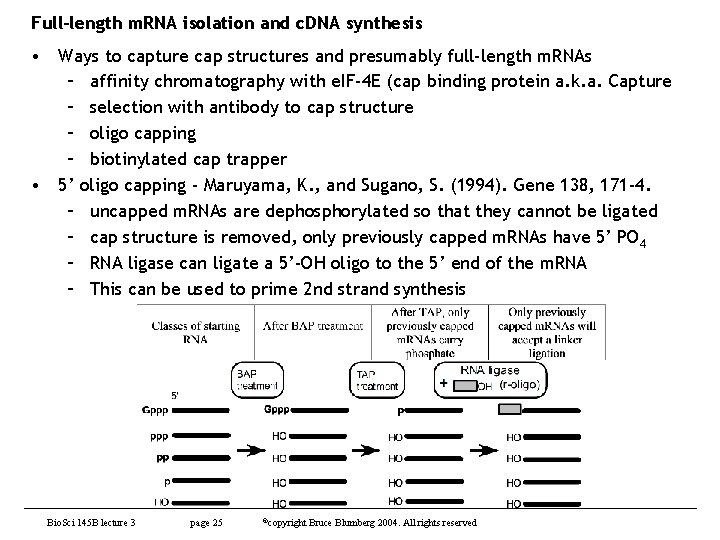
Full-length m. RNA isolation and c. DNA synthesis • Ways to capture cap structures and presumably full-length m. RNAs – affinity chromatography with e. IF-4 E (cap binding protein a. k. a. Capture – selection with antibody to cap structure – oligo capping – biotinylated cap trapper • 5’ oligo capping - Maruyama, K. , and Sugano, S. (1994). Gene 138, 171 -4. – uncapped m. RNAs are dephosphorylated so that they cannot be ligated – cap structure is removed, only previously capped m. RNAs have 5’ PO 4 – RNA ligase can ligate a 5’-OH oligo to the 5’ end of the m. RNA – This can be used to prime 2 nd strand synthesis Bio. Sci 145 B lecture 3 page 25 ©copyright Bruce Blumberg 2004. All rights reserved

Full-length m. RNA isolation and c. DNA synthesis (contd) • 5’ oligo capping (contd) – advantages • very simple • no homopolymeric regions to worry about • can put arbitrary sequence at 5’ end. – Enables custom vector construction – also enables PCR to make driver for normalization – disadvantages • cap trapper paper claims this method only gives 70% full-length c. DNAs • high quality TAP is not easy to find • original paper used PCR between 5’ and 3’ primer to make c. DNAs – PCR => bias! Bio. Sci 145 B lecture 3 page 26 ©copyright Bruce Blumberg 2004. All rights reserved
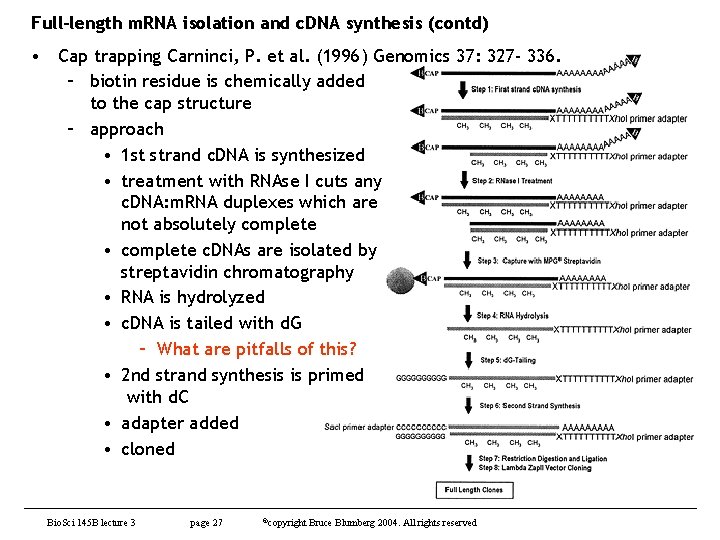
Full-length m. RNA isolation and c. DNA synthesis (contd) • Cap trapping Carninci, P. et al. (1996) Genomics 37: 327 - 336. – biotin residue is chemically added to the cap structure – approach • 1 st strand c. DNA is synthesized • treatment with RNAse I cuts any c. DNA: m. RNA duplexes which are not absolutely complete • complete c. DNAs are isolated by streptavidin chromatography • RNA is hydrolyzed • c. DNA is tailed with d. G – What are pitfalls of this? • 2 nd strand synthesis is primed with d. C • adapter added • cloned Bio. Sci 145 B lecture 3 page 27 ©copyright Bruce Blumberg 2004. All rights reserved

Full-length m. RNA isolation and c. DNA synthesis (contd) • Cap trapping (contd) – advantages • claimed to give 90% recovery of full-length c. DNAs • lots of history at RIKEN – disadvantages • homopolymeric region • many steps -> points of failure Bio. Sci 145 B lecture 3 page 28 ©copyright Bruce Blumberg 2004. All rights reserved
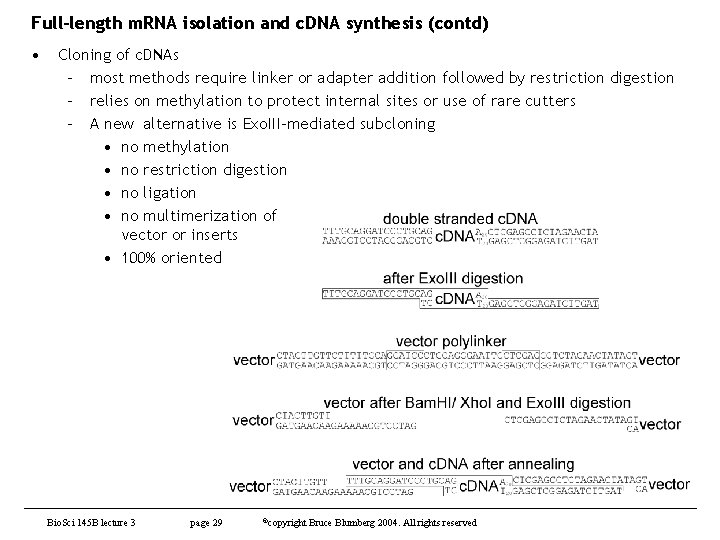
Full-length m. RNA isolation and c. DNA synthesis (contd) • Cloning of c. DNAs – most methods require linker or adapter addition followed by restriction digestion – relies on methylation to protect internal sites or use of rare cutters – A new alternative is Exo. III-mediated subcloning • no methylation • no restriction digestion • no ligation • no multimerization of vector or inserts • 100% oriented Bio. Sci 145 B lecture 3 page 29 ©copyright Bruce Blumberg 2004. All rights reserved

Vectors for c. DNA cloning • Plasmids vs phage – phage preferred for high density manual screening – plasmids are better for functional screening • microinjection • transfection • panning – phage packaging and infection more efficient than electroporation • 10 -100 x better than best transformation frequency • what will the library be used for ? – Consider the intended use as well as other contemplated uses • will the library go to an EST project? – Plasmid • will it be screened manually – phage • or arrayed and screened on high density filters – plasmid • will we normalize it? – Probably plasmid Bio. Sci 145 B lecture 3 page 30 ©copyright Bruce Blumberg 2004. All rights reserved

Vectors for c. DNA cloning (contd) • Analysis of c. DNAs obtained – rate limiting step in clone analysis is getting them into a usable form • usually a plasmid – cloning is tedious, particularly if one has many positives • some tricks can be used but this is still the bottleneck • in about 1985 or so, Stratagene introduced lambda ZAP – phage with an embedded plasmid and M 13 packaging signals – plasmid can be automatically excised by adding a helper phage • gene II protein replicates plasmid into ss phagemid which is secreted – this was a major advance and many phage libraries today are made in ZAP or its derivatives – early protocols had problems with helper phage but this has been overcome • later, others developed a Cre-lox based system – instead of M 13 used lox. P sites. – When Cre recombinase is added, recombination between the lox. P sites excises a plasmid • both methods work very well and make analysis of many clones very straightforward Bio. Sci 145 B lecture 3 page 31 ©copyright Bruce Blumberg 2004. All rights reserved
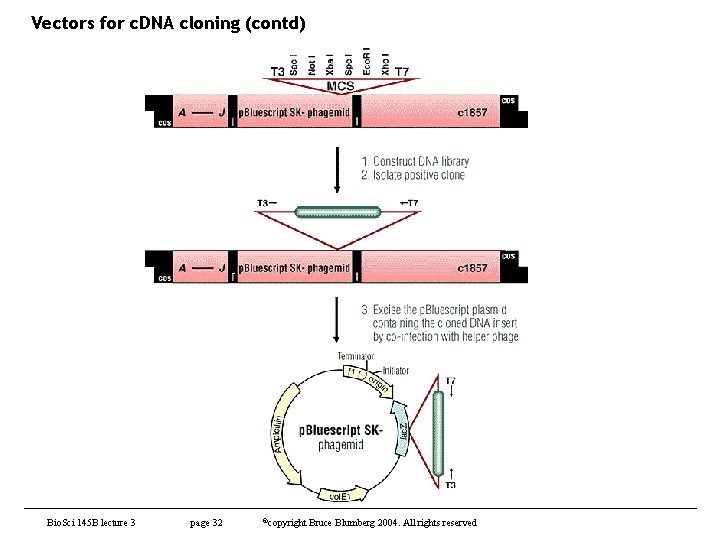
Vectors for c. DNA cloning (contd) Bio. Sci 145 B lecture 3 page 32 ©copyright Bruce Blumberg 2004. All rights reserved
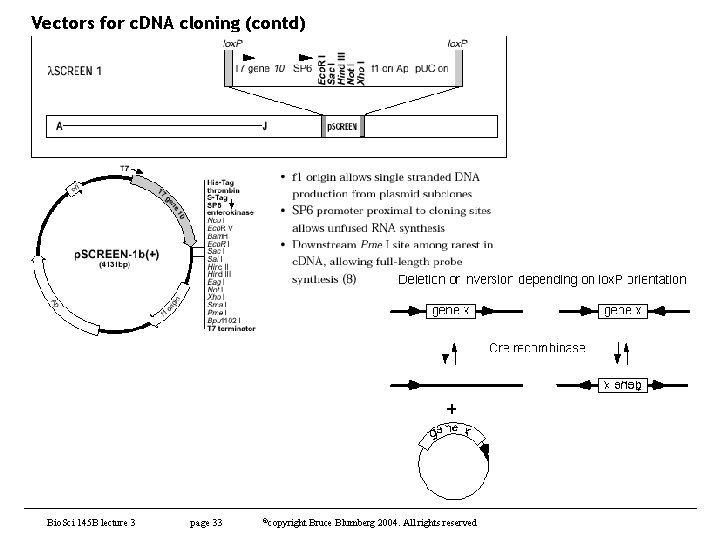
Vectors for c. DNA cloning (contd) Bio. Sci 145 B lecture 3 page 33 ©copyright Bruce Blumberg 2004. All rights reserved

m. RNA frequency and cloning • m. RNA frequency classes – classic references • Bishop et al. , 1974 Nature 250, 199 -204 • Davidson and Britten, 1979 Science 204, 1052 -1059 – abundant • 10 -15 m. RNAs that together represent 10 -20% of the total RNA mass • > 0. 2% – intermediate • 1, 000 -2, 000 m. RNAs together comprising 40 -45% of the total • 0. 05 -0. 2% abundance – rare • 15, 000 -20, 000 m. RNAs comprising 40 -45% of the total • abundance of each is less than 0. 05% of the total • some of these might only occur at a few copies per cell • How does one go about identifying genes that might only occur at a few copies per cell? Bio. Sci 145 B lecture 3 page 34 ©copyright Bruce Blumberg 2004. All rights reserved

Normalization and subtraction • How to identify genes that might only occur at a few copies per cell? – alter the representation of the c. DNAs in a library or probe – Normalization - process of reducing the frequency of abundant and increasing the frequency of rare m. RNAs • Bonaldo et al. , 1996 Genome Research 6, 791 -806 – Subtraction - removing c. DNAs (m. RNAs) expressed in two populations leaving only differentially expressed • Sagerström et al. (1997) Ann Rev. Biochem 66, 751 -783 Bio. Sci 145 B lecture 3 page 35 ©copyright Bruce Blumberg 2004. All rights reserved

Normalization and subtraction • Normalization - reducing abundant, increase rare m. RNAs – normalization should bring c. DNA abundunce to within 10 x • rarely works this well • Typically, abundant genes reduced 10 x, rare ones increased 3 -10 x • Intermediate class genes do not change much at all – Approach • make a population of c. DNAs single stranded - tester • hybridize with a large excess of c. DNA or m. RNA to Cot =5. 5 – driver • Cot value is critical for success of normalization – 5 -10 optimal higher values NOT better Bio. Sci 145 B lecture 3 page 36 ©copyright Bruce Blumberg 2004. All rights reserved

Normalization and subtraction (contd) – Approach (contd) • various approaches to make driver – use m. RNA - may not be easy to get – make ss. RNA by transcribing library – ss. DNA from gene II/Exo. III treating inserts from plasmid library – PCR amplification of library • best approach is to use driver derived from the same library by PCR – rapid, simple and effective – other approaches each have various technical difficulties – see the Bonaldo review for details. Bio. Sci 145 B lecture 3 page 37 ©copyright Bruce Blumberg 2004. All rights reserved

Normalization and subtraction (contd) – What are normalized libraries good for? • EST sequencing • gene identification – biggest use is to reduce the number of c. DNAs that must be screened – good general purpose target to screen » subtracted libraries are useful but limited in utility – Drawbacks • Not trivial to make • Size distribution of library changes – Longer c. DNAs lost Bio. Sci 145 B lecture 3 page 38 ©copyright Bruce Blumberg 2004. All rights reserved
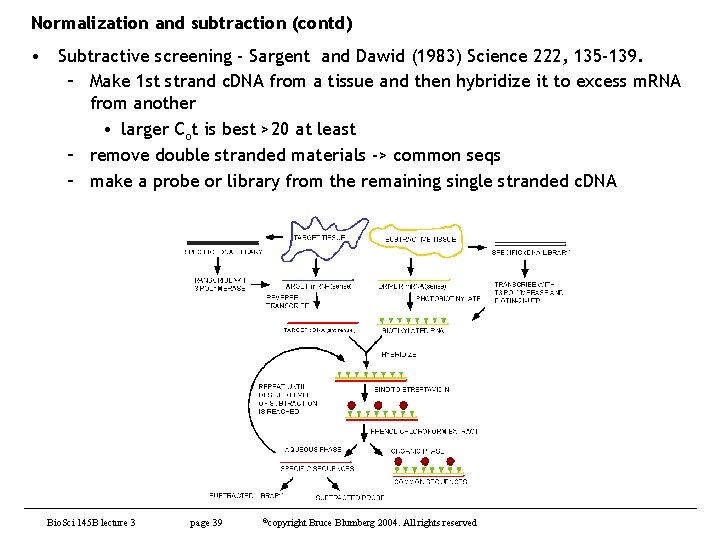
Normalization and subtraction (contd) • Subtractive screening - Sargent and Dawid (1983) Science 222, 135 -139. – Make 1 st strand c. DNA from a tissue and then hybridize it to excess m. RNA from another • larger Cot is best >20 at least – remove double stranded materials -> common seqs – make a probe or library from the remaining single stranded c. DNA Bio. Sci 145 B lecture 3 page 39 ©copyright Bruce Blumberg 2004. All rights reserved

Normalization and subtraction (contd) • Subtractive screening (contd) – benefits • sensitive • can simultaneously identify all c. DNAs that are differentially present in a population • good choice for identifying unknown, tissue specific genes – drawbacks • easy to have abundant housekeeping genes slip through – multistage subtraction is best – in effect normalize first, then subtract • libraries have limited applications – may not be useful for multiple purposes Bio. Sci 145 B lecture 3 page 40 ©copyright Bruce Blumberg 2004. All rights reserved

Normalization and subtraction (contd) – rule of thumb • make a high quality representative library from a tissue of interest • save subtraction and other fancy manipulations for making probes to screen such libraries with – unlimited screening – easy to use libraries for different purposes, e. g. the liver library » hepatocarcinoma » cirrhosis » regeneration specific genes Bio. Sci 145 B lecture 3 page 41 ©copyright Bruce Blumberg 2004. All rights reserved
How to identify your gene of interest • Screening methods depend on what type of information you have in hand. – Related gene from another species? • Low stringency hybridization – A piece of genomic DNA? • Hybridization – A mutant • Complementation • Positional cloning – A functional assay? • Expression screening – An antibody? • Expression library screening – A partial amino acid sequence? • Oligonucleotide screening – A DNA element required for expression of an interesting gene? • Various binding protein strategies – An interacting protein? • Interaction screening – A specific tissue or embryonic stage? • Subtracted screening Bio. Sci 145 B lecture 3 page 42 ©copyright Bruce Blumberg 2004. All rights reserved

How to identify your gene of interest (contd) • What is the most important piece of information you need to clone a c. DNA? – Information on where the m. RNA is expressed • either what tissue or • what time during development – such information is indispensable!! • First step in any hybridization based method (high or low stringency) is to get information on expression – high stringency homologous screening - Northern analysis – cross species screening requires more care • perform a genomic Southern to identify hybridization and washing conditions that identify a small number of hybridizing fragments – standard conditions - 1 M Na+, 43% formamide, 37° C – begin washing at RT in 2 x SSC and expose – increase stringency until signal/noise ratio is acceptable – use these conditions for Northern. • If Northern is unsuccessful - obtain a genomic clone and repeat the screening at high stringency – this approach will never fail to identify a homologous gene Bio. Sci 145 B lecture 3 page 43 ©copyright Bruce Blumberg 2004. All rights reserved
- Slides: 43