Bio Sci 145 B Lecture 10 672005 Bruce

Bio. Sci 145 B Lecture #10 6/7/2005 • Bruce Blumberg – 2113 E Mc. Gaugh Hall - office hours Wed 10 -11 AM (or by appointment) – phone 824 -8573 – blumberg@uci. edu • TA – Suman Verma sverma@uci. edu – 2113 Mc. Gaugh Hall, 924 -6873, 3116 copyright 2000 Bruce Blumberg, all rights reserved
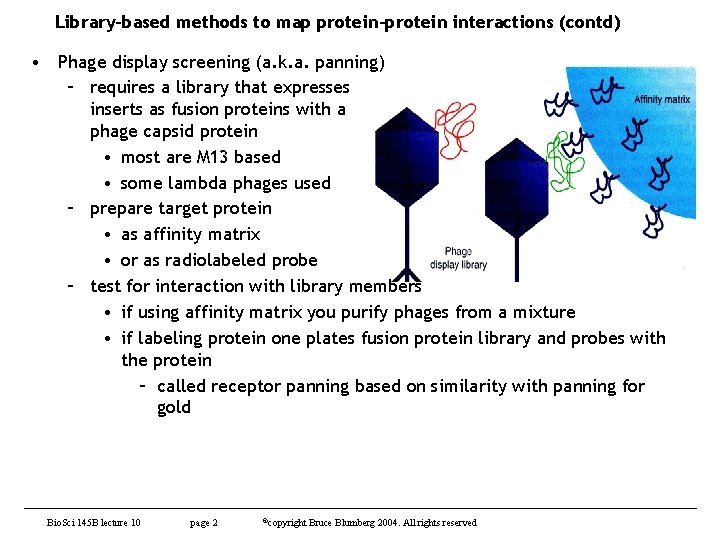
Library-based methods to map protein-protein interactions (contd) • Phage display screening (a. k. a. panning) – requires a library that expresses inserts as fusion proteins with a phage capsid protein • most are M 13 based • some lambda phages used – prepare target protein • as affinity matrix • or as radiolabeled probe – test for interaction with library members • if using affinity matrix you purify phages from a mixture • if labeling protein one plates fusion protein library and probes with the protein – called receptor panning based on similarity with panning for gold Bio. Sci 145 B lecture 10 page 2 ©copyright Bruce Blumberg 2004. All rights reserved

Library-based methods to map protein-protein interactions (contd) • Phage display screening (a. k. a. panning) (contd) – advantages • stringency can be manipulated • if the affinity matrix approach works the cloning could go rapidly – disadvantages • Fusion proteins bias the screen against full-length c. DNAs • Multiple attempts required to optimize binding • Limited targets possible • may not work for heterodimers • unlikely to work for complexes • panning can take many months for each screen – Greg Weiss in Chemistry is local expert Bio. Sci 145 B lecture 10 page 3 ©copyright Bruce Blumberg 2004. All rights reserved
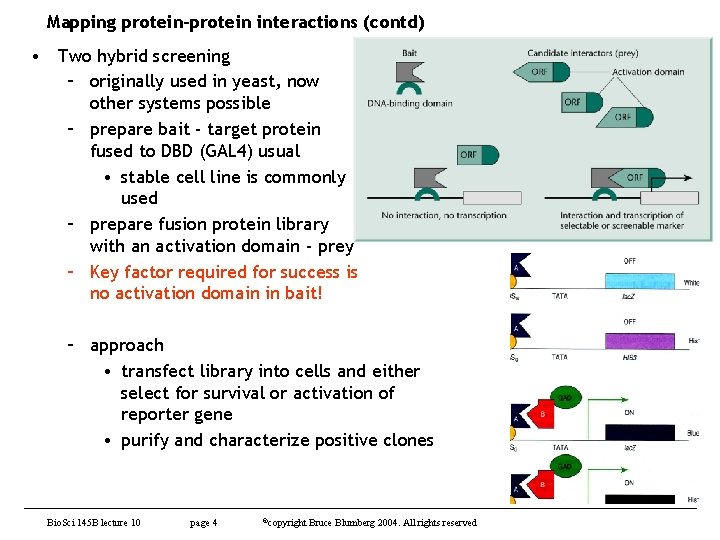
Mapping protein-protein interactions (contd) • Two hybrid screening – originally used in yeast, now other systems possible – prepare bait - target protein fused to DBD (GAL 4) usual • stable cell line is commonly used – prepare fusion protein library with an activation domain - prey – Key factor required for success is no activation domain in bait! – approach • transfect library into cells and either select for survival or activation of reporter gene • purify and characterize positive clones Bio. Sci 145 B lecture 10 page 4 ©copyright Bruce Blumberg 2004. All rights reserved
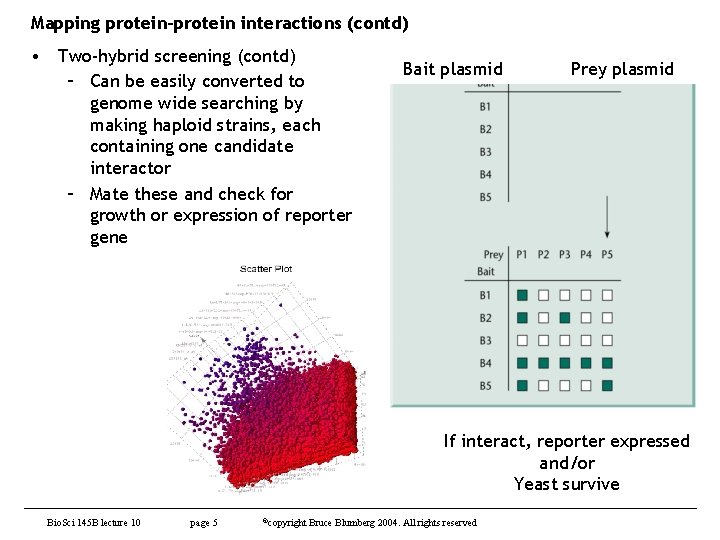
Mapping protein-protein interactions (contd) • Two-hybrid screening (contd) – Can be easily converted to genome wide searching by making haploid strains, each containing one candidate interactor – Mate these and check for growth or expression of reporter gene Bait plasmid Prey plasmid If interact, reporter expressed and/or Yeast survive Bio. Sci 145 B lecture 10 page 5 ©copyright Bruce Blumberg 2004. All rights reserved
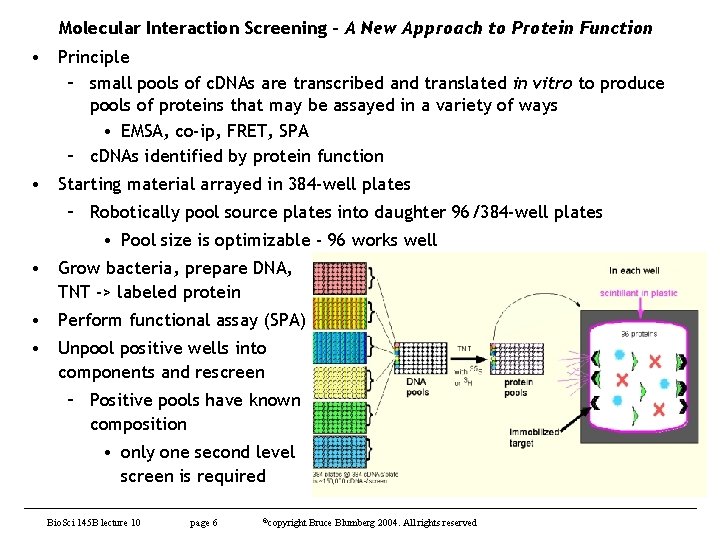
Molecular Interaction Screening - A New Approach to Protein Function • Principle – small pools of c. DNAs are transcribed and translated in vitro to produce pools of proteins that may be assayed in a variety of ways • EMSA, co-ip, FRET, SPA – c. DNAs identified by protein function • Starting material arrayed in 384 -well plates – Robotically pool source plates into daughter 96/384 -well plates • Pool size is optimizable - 96 works well • Grow bacteria, prepare DNA, TNT -> labeled protein • Perform functional assay (SPA) • Unpool positive wells into components and rescreen – Positive pools have known composition • only one second level screen is required Bio. Sci 145 B lecture 10 page 6 ©copyright Bruce Blumberg 2004. All rights reserved

Automated Molecular Interaction Screening • Why do it this way? – arbitrary size and complexity of target is possible – Normalized c. DNA pool -> representation of rare messages – numerous possible endpoint assays • radioactive, fluorescent, luminescent – saturation screening of genome is feasible – two screening steps to pure c. DNA of interest in ~2 weeks Bio. Sci 145 B lecture 10 page 7 ©copyright Bruce Blumberg 2004. All rights reserved
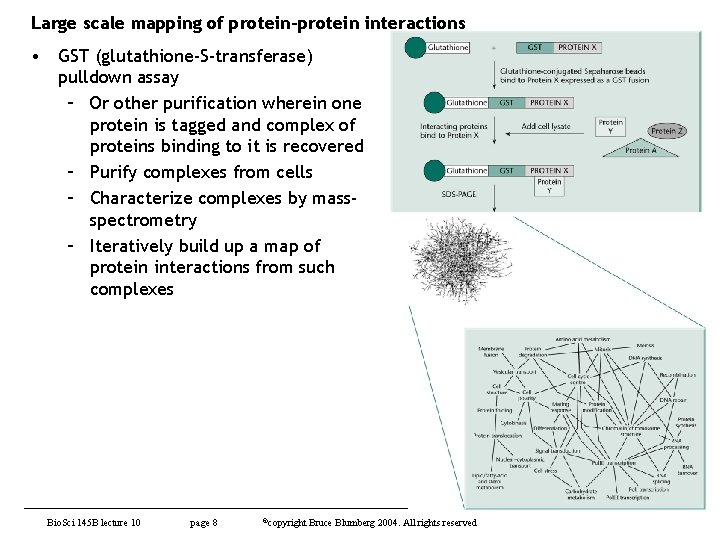
Large scale mapping of protein-protein interactions • GST (glutathione-S-transferase) pulldown assay – Or other purification wherein one protein is tagged and complex of proteins binding to it is recovered – Purify complexes from cells – Characterize complexes by massspectrometry – Iteratively build up a map of protein interactions from such complexes Bio. Sci 145 B lecture 10 page 8 ©copyright Bruce Blumberg 2004. All rights reserved
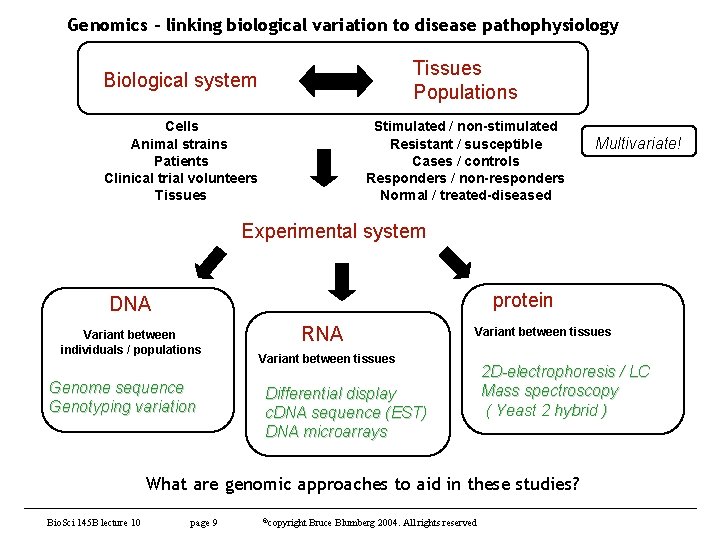
Genomics - linking biological variation to disease pathophysiology Biological system Tissues Populations Cells Animal strains Patients Clinical trial volunteers Tissues Stimulated / non-stimulated Resistant / susceptible Cases / controls Responders / non-responders Normal / treated-diseased Multivariate! Experimental system protein DNA Variant between individuals / populations Genome sequence Genotyping variation RNA Variant between tissues Differential display c. DNA sequence (EST) DNA microarrays 2 D-electrophoresis / LC Mass spectroscopy ( Yeast 2 hybrid ) What are genomic approaches to aid in these studies? Bio. Sci 145 B lecture 10 page 9 ©copyright Bruce Blumberg 2004. All rights reserved

The rise of -omics • The -omics revolution of science – http: //www. genomicglossaries. com/content/omes. asp • What does it all mean? – Transcriptomics – large scale gene profiling (usually microarray) – Proteomics – study of complement of expressed proteins – Functional genomics – very vague term, typically encompasses many others – Structural genomics – prediction of structure and interactions from sequence – Pharmacogenomics – transcriptional profiling of response to drug treatment – often looking for genetic basis of differences – Toxicogenomics – transcriptional profiling of response to toxicants (often includes pharmacogenomics • Seeks mechanistic understanding of toxic response – Metabolomics – analysis of total metabolite pool ("metabolome") to reveal novel aspects of cellular metabolism and global regulation – Interactomics – genome wide study of macromolecular interactions, physical and genetic are included. Bio. Sci 145 B lecture 10 page 10 ©copyright Bruce Blumberg 2004. All rights reserved
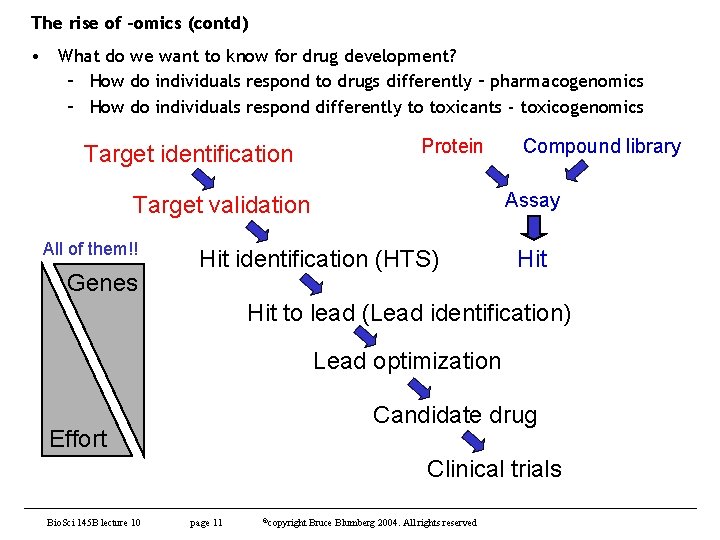
The rise of –omics (contd) • What do we want to know for drug development? – How do individuals respond to drugs differently – pharmacogenomics – How do individuals respond differently to toxicants - toxicogenomics Protein Target identification Assay Target validation All of them!! Genes Compound library Hit identification (HTS) Hit to lead (Lead identification) Lead optimization Candidate drug Effort Clinical trials Bio. Sci 145 B lecture 10 page 11 ©copyright Bruce Blumberg 2004. All rights reserved

Toxicogenomics • Lump pharmacogenomics and toxicogenomics together in the context of drug development • Toxicology is the study of effects of toxicant exposure – Traditional toxicology focuses on exposure, dose, effect – “dose makes the poison” – overly simplistic and probably incorrect • Mechanistic Toxicology (academic and regulatory) – Investigative toxicology • Hypothesis generation for grants and studies – Risk assessment • Understanding the mechanism of toxicity at the molecular level • EPA and NIEHS very concerned with this • Predictive toxicology – Compound avoidance • Elimination of liabilities (pharma, chemical industry) – Compound selection • Select compound with least toxic liability from a series (pharma) – Compound management • Tailor conventional studies and perform timely investigational toxicology studies Bio. Sci 145 B lecture 10 page 12 ©copyright Bruce Blumberg 2004. All rights reserved
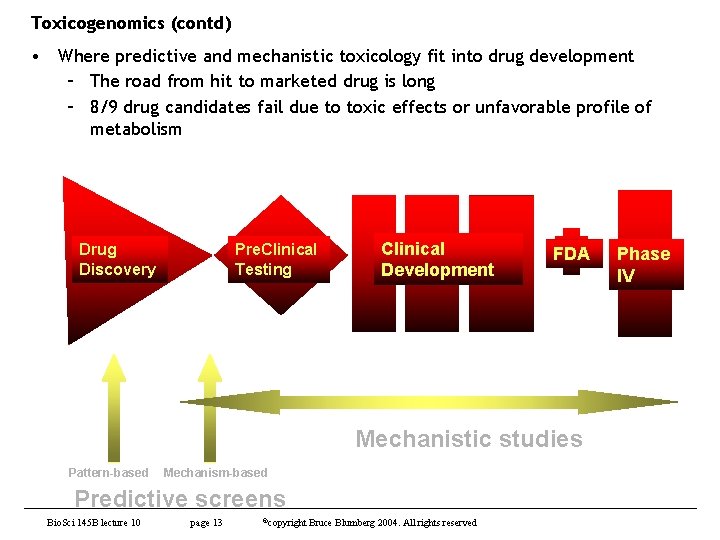
Toxicogenomics (contd) • Where predictive and mechanistic toxicology fit into drug development – The road from hit to marketed drug is long – 8/9 drug candidates fail due to toxic effects or unfavorable profile of metabolism Drug Discovery Pre. Clinical Testing Clinical Development FDA Mechanistic studies Pattern-based Mechanism-based Predictive screens Bio. Sci 145 B lecture 10 page 13 ©copyright Bruce Blumberg 2004. All rights reserved Phase IV
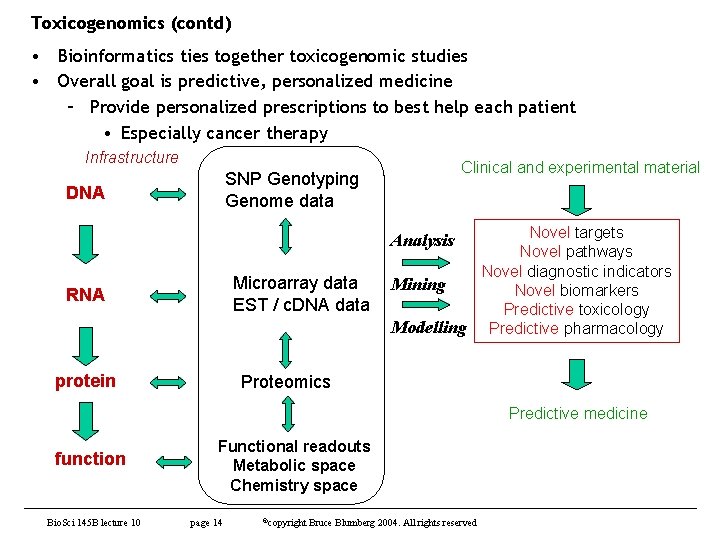
Toxicogenomics (contd) • Bioinformatics ties together toxicogenomic studies • Overall goal is predictive, personalized medicine – Provide personalized prescriptions to best help each patient • Especially cancer therapy Infrastructure Clinical and experimental material SNP Genotyping Genome data DNA Novel targets Novel pathways Novel diagnostic indicators Mining Novel biomarkers Predictive toxicology Modelling Predictive pharmacology Analysis Microarray data EST / c. DNA data RNA protein Proteomics Predictive medicine function Bio. Sci 145 B lecture 10 Functional readouts Metabolic space Chemistry space page 14 ©copyright Bruce Blumberg 2004. All rights reserved
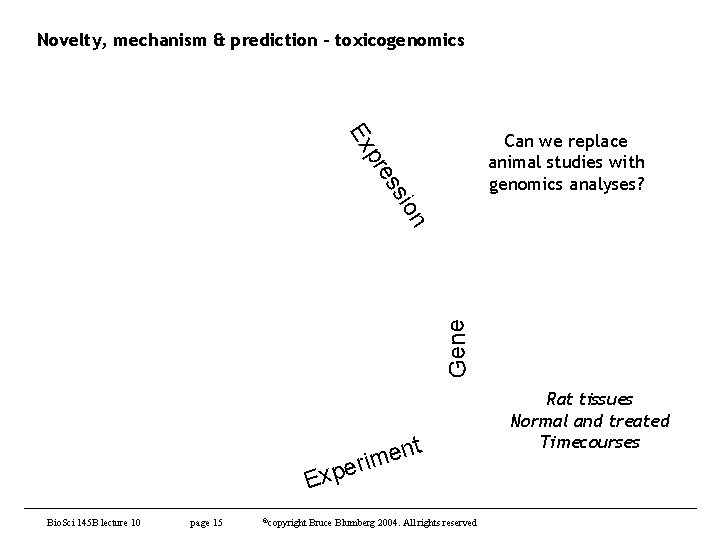
Novelty, mechanism & prediction - toxicogenomics Gene ion ss pre Ex Can we replace animal studies with genomics analyses? t n e m peri Ex Bio. Sci 145 B lecture 10 page 15 ©copyright Bruce Blumberg 2004. All rights reserved Rat tissues Normal and treated Timecourses

Toxicogenomics (contd) • What is toxicogenomics good for? – Obtaining a high level view of a biological system – Rapid generation of response profiles to • Unravel mechanisms • Discriminate among compounds – Signature of exposures? – Probably not a single method to identify toxicity • Problems that must be solved – Interlab variation – different labs use slightly different methods and get results that may not be strictly applicable • Japanese solution is to designate a single lab for entire country – Most genes change expression at high doses of exposure • Relevant? Bio. Sci 145 B lecture 10 page 16 ©copyright Bruce Blumberg 2004. All rights reserved

Genomic technology - implications • Genetics and reverse genetics – gene transfer and selection technology speeds up genetic analysis by orders of magnitude – virtually all conceivable experiments are now possible • all questions are askable • BUT should all questions be asked? – much more straightforward to understand gene function using knockouts and transgenics • gene sequences are coming at an unprecedented rate from the genome projects • Knockouts and transgenics remain very expensive to practice – other yet undiscovered technologies will be required to understand gene function. Bio. Sci 145 B lecture 10 page 17 ©copyright Bruce Blumberg 2004. All rights reserved

Genomic technology – implications (contd) • Clinical genetics – Molecular diagnostics are becoming very widespread as genes are matched with diseases • huge growth area for the future • big pharma is dumping billions into diagnostics – room for great benefit and widespread abuse • diagnostics will enable early identification and treatment of diseases • but insurance companies will want access to these data to maximize profits Bio. Sci 145 B lecture 10 page 18 ©copyright Bruce Blumberg 2004. All rights reserved

Genomic technology – implications (contd) • gene therapy – new viral vector technology is making this a reality • efficient transfer and reasonable regulation possible – long lag time from laboratory to clinic, still working with old technology in many cases – The Biotech Death of Jesse Gelsinger. Sheryl Gay Stolberg, NY Times, Sunday Magazine, 28 Nov 99 • http: //www. frenchanderson. org/history/biotech. pdf • protein engineering – not as widely appreciated as more glamorous techniques such as gene therapy and transgenic crops – better drugs, e. g. , more stable insulin, TPA for heart attacks and strokes, etc. – more efficient enzymes (e. g. subtilisin in detergents) – safe and effective vaccines • just produce antigenic proteins rather than using inactivated or attenuated organisms to reduce undesirable side effects Bio. Sci 145 B lecture 10 page 19 ©copyright Bruce Blumberg 2004. All rights reserved

Genomic technology – implications (contd) • metabolite engineering – enhanced microbial synthesis of valuable products • eg indigo (jeans) • vitamin C – generation of entirely new small molecules • transfer of antibiotic producing genes to related species yields new antibiotics (badly needed) – reduction of undesirable side reactions • faster more efficient production of beer • plants as producers of specialty chemicals – underutilized because plant technology lags behind techniques in animals • But regulations are strict (Monsanto) – plants as factories to produce materials more cheaply and efficiently • especially replacements for petrochemicals – plants and herbs are the original source of many pharmaceutical products • engineer them to overproduce desirable substances Bio. Sci 145 B lecture 10 page 20 ©copyright Bruce Blumberg 2004. All rights reserved

Genomic technology – implications (contd) • transgenic food – gene transfer techniques have allowed the creation of desirable mutations into animals and crops of commercial value • disease resistance (various viruses) • pest resistance (Bt cotton) • Pesticide, herbicide and fungicide resistance • growth hormone and milk production – effective but necessary? – negative implications – “Frankenfoods” • pesticide and herbicide resistance lead to much higher use of toxic compounds • results are not predictable due to small datasets • at least one herbicide (bromoxynil) for which resistance was engineered has since been banned • Atrazine is becoming highly controversial Bio. Sci 145 B lecture 10 page 21 ©copyright Bruce Blumberg 2004. All rights reserved

2004 Final Examination 1. (10 points) Remember our old friend Mars burroughsii from the midterm? The pathogenic eukaryotic microorganism that escaped into Newport Beach and is now spreading along the Pacific Coast? Recall also that we have generated a full genomic sequence and a full set of ESTs. We have learned that a protein on the M. burroughsii surface binds to the human Cad. F protein in order to gain entry into the cell and that polymorphisms in the Cad. F gene confer a degree of resistance to M. burroughsii infection in a subset of the population. It is now time to figure out just how M. burroughsii exerts its pathogenic effects in the cell. A postdoc in your lab hypothesizes that additional M. burroughsii proteins interact with human cellular proteins to cause pathogenesis. Comparisons of different M. burroughsii isolates have identified 50 potential genes that are involved in pathogenecity. a) (5 points) How would you identify the full complement of human proteins in intestinal cells that the 50 M. burroughsii proteins interact with? b) (5 points) Assume that one of the human proteins identified in a) is quite different between humans and mice. Recall that M. burroughsii seems only to infect humans and not mice. How could you go about creating a mouse model for studying M. burroughsii pathogenesis? Assume that M. burroughsii can infect mouse cells by binding to Cad. F just fine, but that they are apparently not pathogenic due to this single protein difference between mice and humans. Bio. Sci 145 B lecture 10 page 22 ©copyright Bruce Blumberg 2004. All rights reserved

a) Possible answers are a proteomic approach or large-scale two hybrid. For the proteomic approach, it would be necessary to tag the 50 proteins so that they could be identified, put these back into the cells, purify complexes and identify the components. For the two-hybrid approach, one would need to make suitable bait from each of the 50 proteins in one yeast mating type strain and then cross these with prey from another strain and look for interactions. b) You would want to use targeted mutagenesis to create a mouse that expresses the human protein in place of the corresponding mouse protein. Alternatively, you could simply make a transgenic mouse expressing the human protein but this might not work if there is interference between the human and mouse versions of the protein. Bio. Sci 145 B lecture 10 page 23 ©copyright Bruce Blumberg 2004. All rights reserved

2. (5 points) OK, you now have a mouse model for M. burroughsii pathogenesis that can be used to study the molecular mechanisms involved and 50 candidate M. burroughsii proteins that are involved in pathogenecity. How would you go about systematically disrupting these genes one by one to determine where each fit into the pathogenesis cascade? The best approach would be to create M. burroughsii that are deficient in the proteins using homologous recombination. Alternatively, you could try something like si. RNA expressed under the control of a ubiquitously active promoter to knock down the proteins and test the effects of these proteins. A proteomic approach wherein you mutate the proteins and test the effect on the complexes identified in question 1 a could also be employed. Bio. Sci 145 B lecture 10 page 24 ©copyright Bruce Blumberg 2004. All rights reserved

3. (5 points) Great, now you have identified which of the M. burroughsii proteins act where to induce a pathogenic response in infected cells. These proteins will be good candidate drug targets. Several companies have identified candidate drugs for testing by screening compound libraries against these targets but first wish to determine the profiles of gene expression in response to these drugs in mice and humans. How would you go about determining these response profiles and how they differ in hepatocytes from different human donors and wild type mice vs. your mouse model from 1 b above? The best approach would be to treat human hepatocytes from different donors with the candidate drugs and evaluate the profile of gene expression using microarray analysis. The profiles could then be compared with each other to develop a sense of how humans vary in response to the drugs and whether the profile of gene expression corresponds to known toxic profiles obtained from toxicogenomic studies. A similar approach using wild-type and your model mouse would give the desired results. Bio. Sci 145 B lecture 10 page 25 ©copyright Bruce Blumberg 2004. All rights reserved

4. (10 points) You are a molecular biologist who really loves eating shrimp after work as much as you enjoy working in the lab. To combine your two loves, you have decided to create a biotech company based on producing the largest shrimp anywhere. You took an invertebrate biology course at UCI and learned that shrimp and related animals express an unidentified growth inhibitory gene when cultured at high density. In answering the questions below, assume that all of the standard sorts of methods we discussed in class will work in shrimp (except targeted gene disruption). Assume that shrimp heterozygous for disruption of the growth inhibitory gene will be large enough to be detected but not big enough for your company. a) (4 points) Describe an approach that will allow you to create mutations in the growth inhibitory gene b) (4 points) Describe how you will clone and/or identify the gene that has been mutated? c) (2 points) How will you figure out where the gene is expressed to begin to understand how it works? Bio. Sci 145 B lecture 10 page 26 ©copyright Bruce Blumberg 2004. All rights reserved

a) you don’t have any genome sequence available from this organism therefore si. RNA or targeted gene disruption will not work. You could try chemical mutagenesis but that will not put you in a position to answer b and c. The best way is to use gene trapping to identify a line that has enhanced growth abilities. b) if you created a gene trap line in a) then all you have to do is clone the region surrounding the insertion and sequence it to identify the disrupted gene c) once you have established the gene trap line, the expression patterns of the reporter gene should tell you where it is expressed. Alternatively, you could use in situ hybridization but this might not be sensitive enough to detect expression everywhere. Bio. Sci 145 B lecture 10 page 27 ©copyright Bruce Blumberg 2004. All rights reserved

5. (5 points) In answering question 4 above, you have identified a chunk of genomic sequence that appears to correspond to the gene responsible for growth inhibition. You identify a large BAC clone containing 150 kb of sequence and sequence it completely. a) (3 points) Describe how you will identify where the introns, exons, splicing signals, and the putative promoter are in this gene. b) (2 points) How will you determine how many m. RNAs are expressed from this region of genomic DNA? a) A bioinformatics approach to identify introns, exons, splicing signals, etc might be good enough. You could also use the genomic sequence to isolate c. DNAs that could then be compared with the genomic sequence to identify the introns, exons and putative promoter b) you could use the genomic DNA to hybridize to northern blots or microarrays or a bioinformatics approach to identify predicted ORFs Bio. Sci 145 B lecture 10 page 28 ©copyright Bruce Blumberg 2004. All rights reserved

6. (5 points) The jumbo shrimp that your company is selling are bringing in serious $ in profits. The CEO decides that improving the product even further would be a wise investment and commissions an EST project to be performed in your laboratory. 15, 000 different transcripts are identified and glass slide microarrays are prepared. To more fully characterize the crowding response, your lab is contracted to identify all of the genes that are upregulated and downregulated by crowding in aquaculture. Describe generally how you will accomplish this. One good approach would be to use the glass slide microarrays to characterize upregulated and downregulated transcripts. You would prepare m. RNA from shrimp that are grown in a crowded environment and from a non -crowded environment. These m. RNAS would be used to prepare fluorescently labeled probes that would be hybridized to the microarrays. Analysis of the microarray data would identify candidate upregulated and downregulated genes that could be tested by QRT-PCR to distinguish which ones are truly regulated by crowding and which are not. Bio. Sci 145 B lecture 10 page 29 ©copyright Bruce Blumberg 2004. All rights reserved
- Slides: 29