Bio Image Analysis course 2020 Cell Profiler handson
Bio. Image Analysis course 2020 Cell. Profiler hands-on workshop Vincent Dumont 1 & Lassi Paavolainen 2 1 Biomedicum 2 High Imaging Unit, University of Helsinki Content Imaging and Analysis unit, Institute for Molecular Medicine Finland, University of Helsinki May 2020

Plan of the workshop • Content of this “general” presentation: • General presentation of Cell. Profiler • Its basic organization and functions • Presentation of the fixed modules common to all pipelines • Content of the other presentations for this course: • Task 1: assemble grayscale images into a merged RGB (red-green-blue) image • Concept of segmentation and image pre-processing • From raw high-content image data to individual cell measurements • Task 2: Illumination correction (pre-processing) • Task 3: Artificial image generation (pre-processing) • Task 4: Segmentation and feature (=numbers) extraction

High-Content Screening (HCS) • Screening = studying biological processes/effects under large amounts of different conditions (typically chemical or genetic perturbations) • High-Content = Using imaging High-Content Screening relies on • Multi-well plates (typically 96 - or 384 -well plate) • Automation: sample preparation, imaging, analysis • Rigorous quality control (QC): On level of an experiment, batch, plate, well, image in some cases even on a single-cell level Need for specialized software to handle these requirements

Example of data amounts from a single experiment Cell Painting assay* for screening drugs • 525 drugs in 5 concentrations => • 8 x 384 -well plates including controls • Imaging in 2 D • 5 channels in Cell Painting assay • 9 fields-of-view from each well using 20 x objective • Single image = 9 MB and includes 100 cells • Output of the experiment • 8*384*9*5 = 138 240 images • 1. 2 TB of data • 2. 7 million cells *Bray et al. (2016). Nature Protocols 11(9), 1757 -1774.
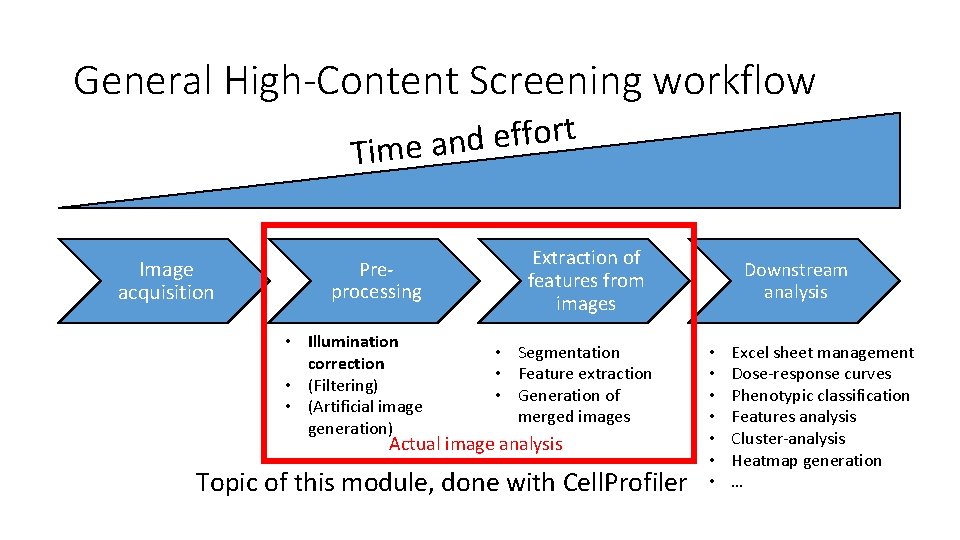
General High-Content Screening workflow t r o f f e d Time an Image acquisition Preprocessing • Illumination correction • (Filtering) • (Artificial image generation) Extraction of features from images • Segmentation • Feature extraction • Generation of merged images Actual image analysis Topic of this module, done with Cell. Profiler Downstream analysis • • Excel sheet management Dose-response curves Phenotypic classification Features analysis Cluster-analysis Heatmap generation …
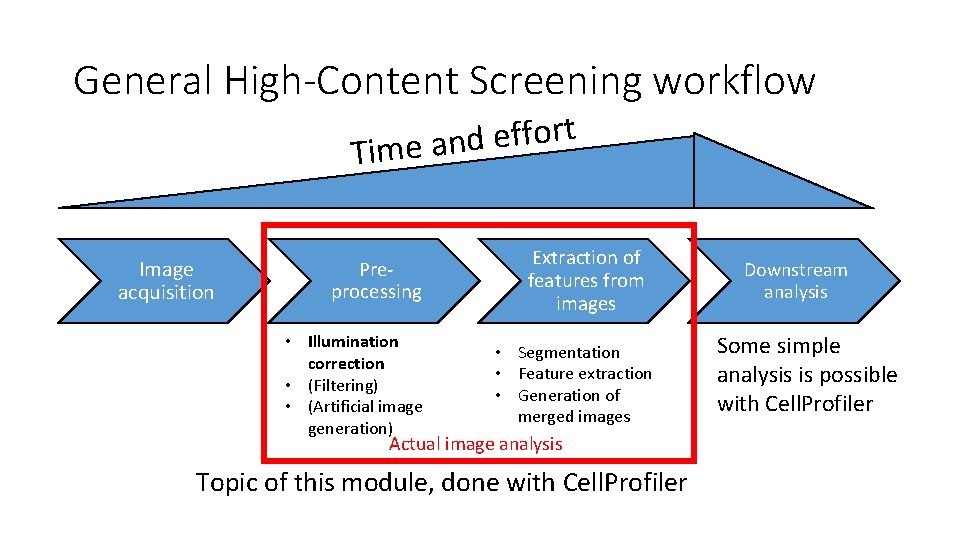
General High-Content Screening workflow t r o f f e d Time an Image acquisition Preprocessing • Illumination correction • (Filtering) • (Artificial image generation) Extraction of features from images • Segmentation • Feature extraction • Generation of merged images Actual image analysis Topic of this module, done with Cell. Profiler Downstream analysis Some simple analysis is possible with Cell. Profiler
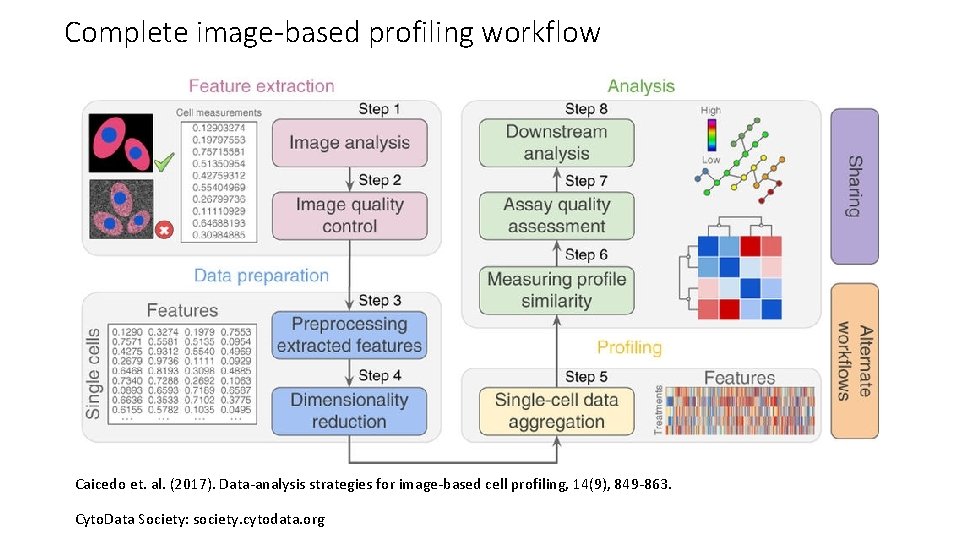
Complete image-based profiling workflow Caicedo et. al. (2017). Data-analysis strategies for image-based cell profiling, 14(9), 849 -863. Cyto. Data Society: society. cytodata. org

Cell. Profiler: generalities • Free and open-source software developed by Carpenter lab, Imaging Platform, Broad Institute, USA • Aims to help biologists without computer science background to analyze big datasets by processing the images in an automated manner • Based on “pipelines” through which image datasets are run • Pipelines can be used • to apply simple modifications to many images (image correction, background subtraction, merging, …) • to extract quantitative information from individual images, cells, organelles or subcellular spaces
Cell. Profiler: generalities • Pipelines are made-up from modules (similar as an electrical circuit made of components) • Each module will perform one step of the overall process Modules • The modules needed in the pipeline depend on the goal of the pipeline • Here, we will build 4 different pipelines for different purposes. But first let’s have a look to the software itself. pipeline

Cell. Profiler: generalities • Please note that this module has been produced with the 3. 1. 8 version of Cell. Profiler. • Using a different version may alter the outlook of the software and the compatibility of the resulting pipelines that are present in the folders • Note that in this course we also use Image. J/Fiji to see how the output images look
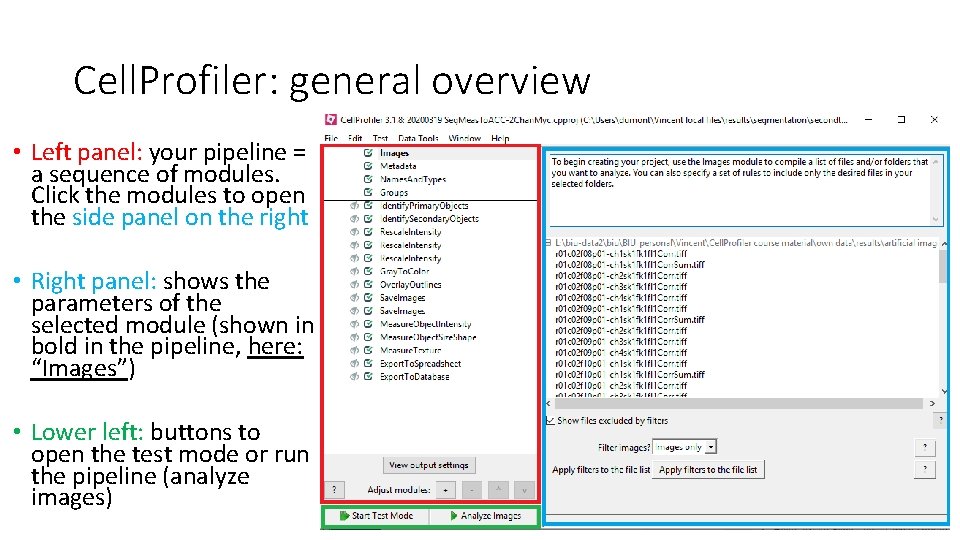
Cell. Profiler: general overview • Left panel: your pipeline = a sequence of modules. Click the modules to open the side panel on the right • Right panel: shows the parameters of the selected module (shown in bold in the pipeline, here: “Images”) • Lower left: buttons to open the test mode or run the pipeline (analyze images)
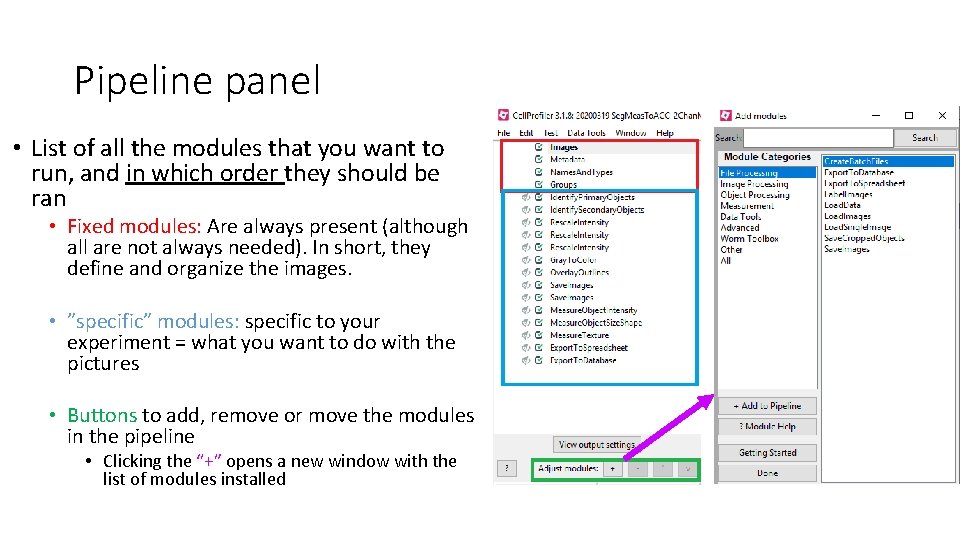
Pipeline panel • List of all the modules that you want to run, and in which order they should be ran • Fixed modules: Are always present (although all are not always needed). In short, they define and organize the images. • ”specific” modules: specific to your experiment = what you want to do with the pictures • Buttons to add, remove or move the modules in the pipeline • Clicking the “+” opens a new window with the list of modules installed
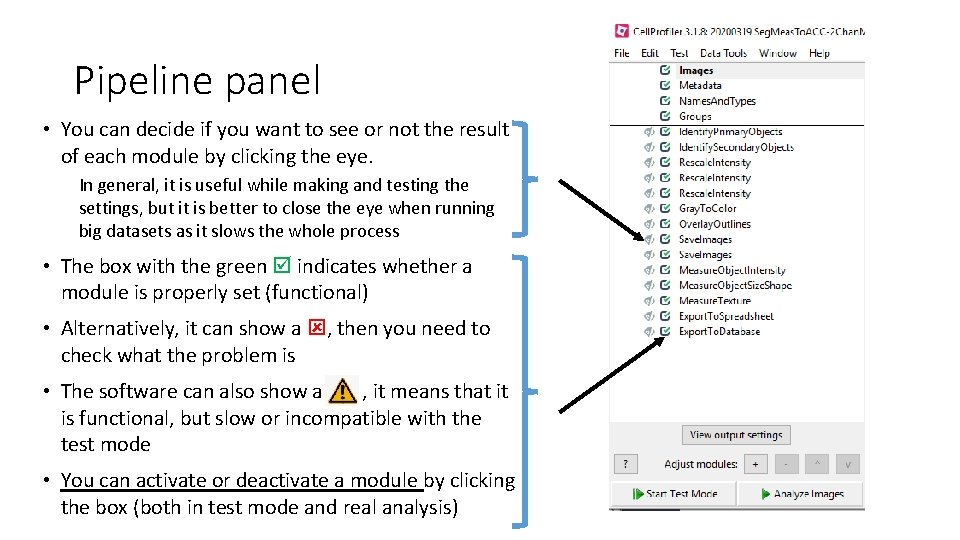
Pipeline panel • You can decide if you want to see or not the result of each module by clicking the eye. In general, it is useful while making and testing the settings, but it is better to close the eye when running big datasets as it slows the whole process • The box with the green indicates whether a module is properly set (functional) • Alternatively, it can show a , then you need to check what the problem is • The software can also show a , it means that it is functional, but slow or incompatible with the test mode • You can activate or deactivate a module by clicking the box (both in test mode and real analysis)
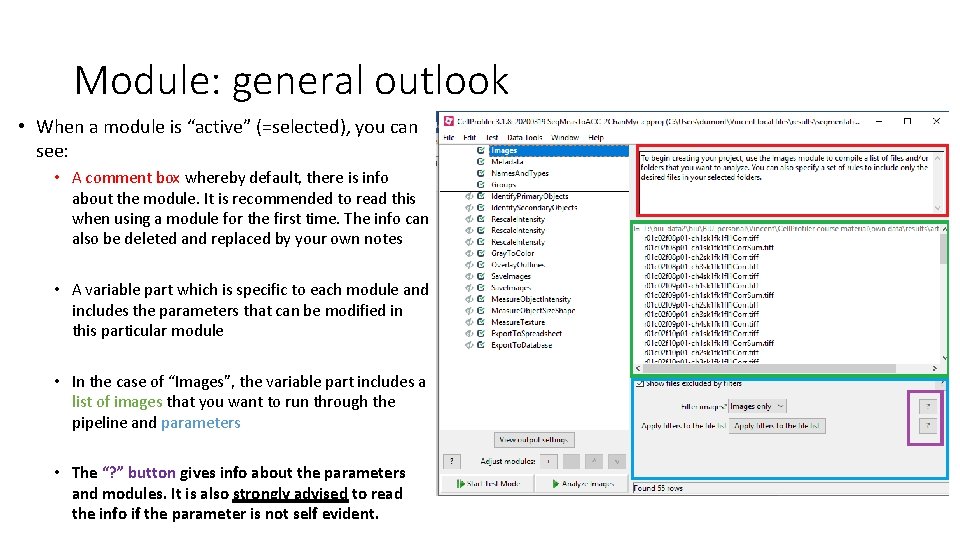
Module: general outlook • When a module is “active” (=selected), you can see: • A comment box whereby default, there is info about the module. It is recommended to read this when using a module for the first time. The info can also be deleted and replaced by your own notes • A variable part which is specific to each module and includes the parameters that can be modified in this particular module • In the case of “Images”, the variable part includes a list of images that you want to run through the pipeline and parameters • The “? ” button gives info about the parameters and modules. It is also strongly advised to read the info if the parameter is not self evident.

Test mode • The test mode can be activated by clicking the “Start Test Mode” button • It is highly useful when building up the pipeline and tuning the parameters • It tests the modules on one “set” of images (=for example an image consisting of 3 channels) • “Run” passes the set of images through the whole pipeline • “Step” passes the set of images through the selected module only (in bold, here the first “Rescale. Intensity”) • “Next Image Set” move to the next image set… (so that you can check whether your settings works with other image sets) Note: Opening the “eye” is necessary to see the results of the module

Test mode • In general, when using test mode, it is good to load few images randomly selected from the whole data set: • the staining quality may vary throughout the set (different wells, coverslips, …) • The phenotype and treatment (drug, gene silencing, …) may alter the cells, which in turn may alter the quality of the image processing

Fixed modules: Always included in the pipeline • Include: Images, Metadata, Names. And. Types, Groups • They will be used to define image set meta information • For example, if you have several channels, which ones should be put together by the software as one image set. • More groups include: planes of z-stacks, time series, etc…
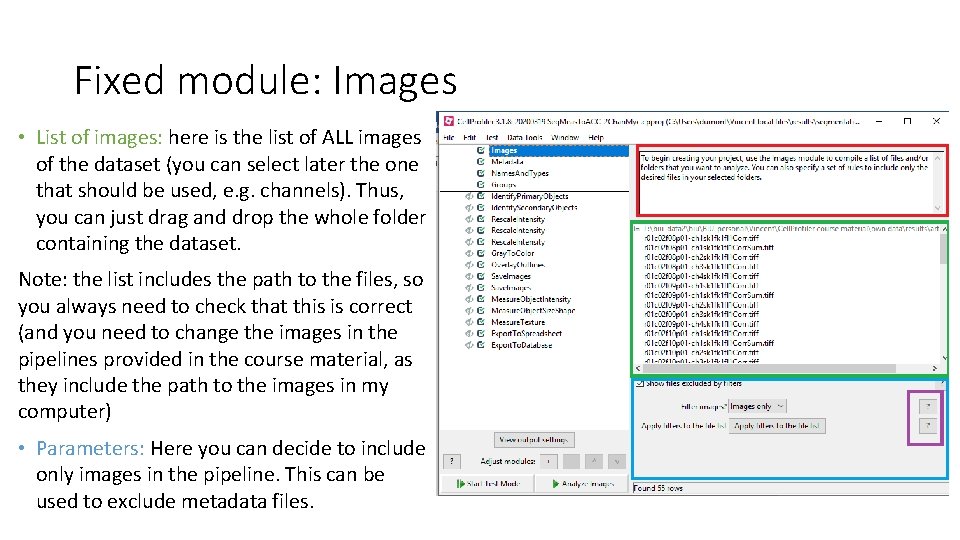
Fixed module: Images • List of images: here is the list of ALL images of the dataset (you can select later the one that should be used, e. g. channels). Thus, you can just drag and drop the whole folder containing the dataset. Note: the list includes the path to the files, so you always need to check that this is correct (and you need to change the images in the pipelines provided in the course material, as they include the path to the images in my computer) • Parameters: Here you can decide to include only images in the pipeline. This can be used to exclude metadata files.
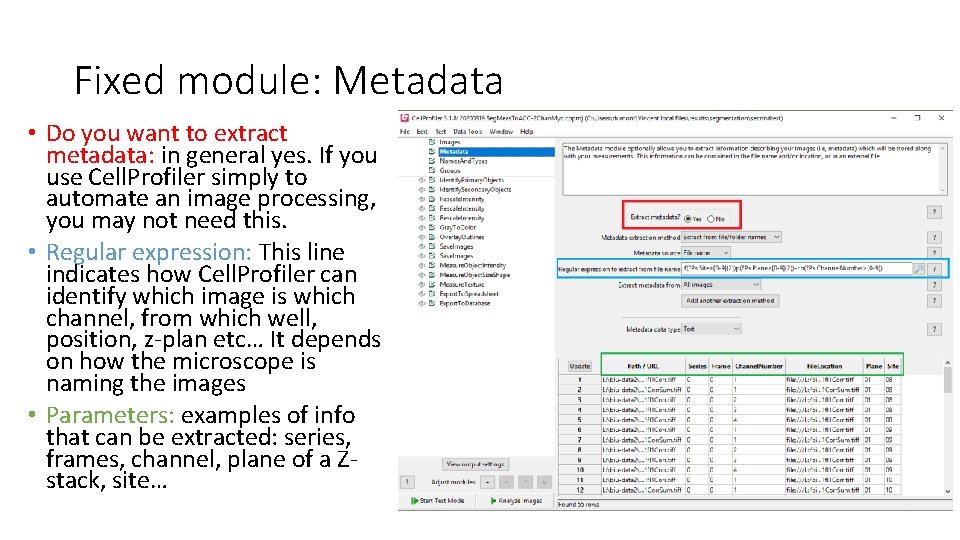
Fixed module: Metadata • Do you want to extract metadata: in general yes. If you use Cell. Profiler simply to automate an image processing, you may not need this. • Regular expression: This line indicates how Cell. Profiler can identify which image is which channel, from which well, position, z-plan etc… It depends on how the microscope is naming the images • Parameters: examples of info that can be extracted: series, frames, channel, plane of a Zstack, site…
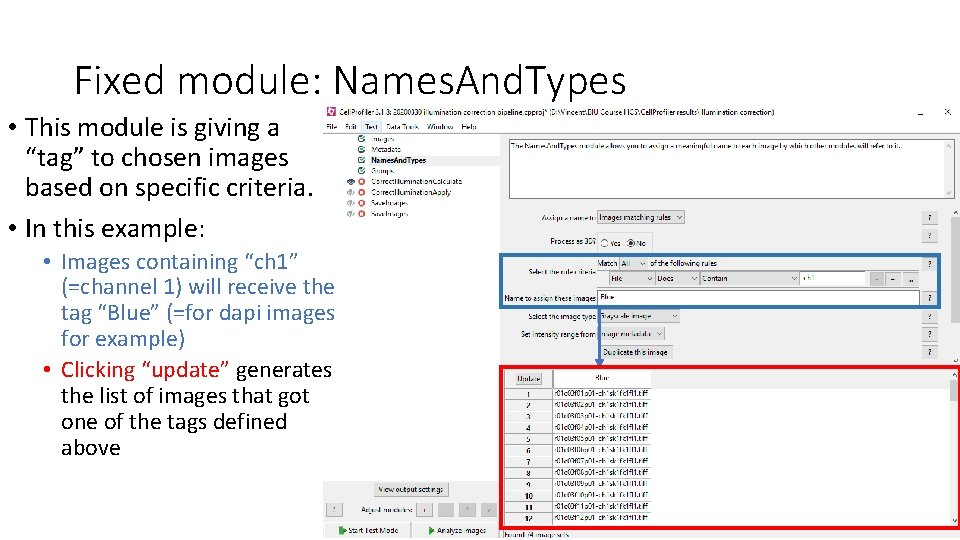
Fixed module: Names. And. Types • This module is giving a “tag” to chosen images based on specific criteria. • In this example: • Images containing “ch 1” (=channel 1) will receive the tag “Blue” (=for dapi images for example) • Clicking “update” generates the list of images that got one of the tags defined above
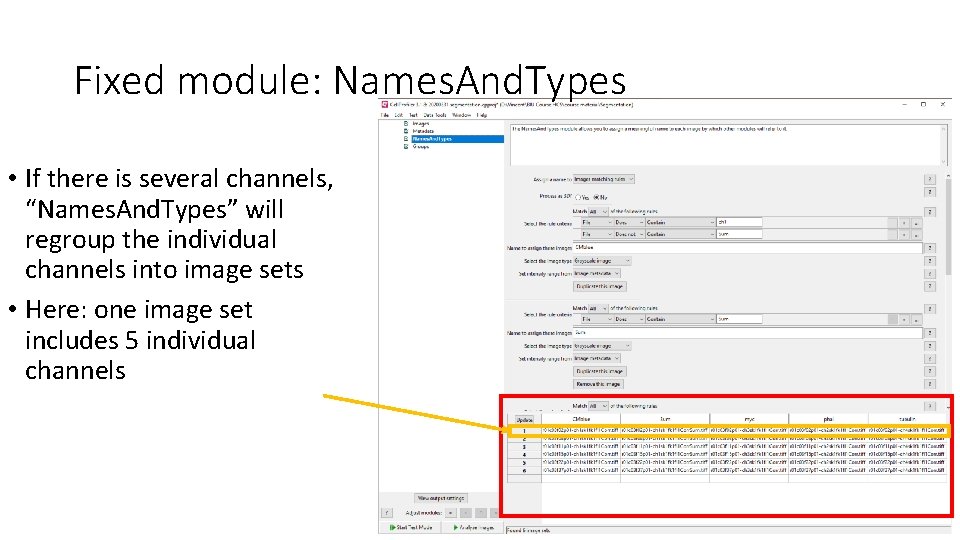
Fixed module: Names. And. Types • If there is several channels, “Names. And. Types” will regroup the individual channels into image sets • Here: one image set includes 5 individual channels
Fixed module: Groups • This allows to subdivide larger datasets. We will not use it in this course

The specific modules: Depends on your needs We will check these in more detail in the tasks that we will perform. Each task is regrouped in a subfolder containing a Power. Point presentation, input images, the final pipeline produced and the output images or spreadsheets. Note that the final pipelines that I included in the folder contain the path to the images in MY computer. You need to delete the images in the “Images” module and add the images from your own computer to make the pipeline functional I have also placed additional pipelines used in real experiments similar as what we do in the tasks.

Task 1: Grayscale to RGB As a start, we will build a pipeline to assemble 16 -bits TIFF images into low information 3 -color PNG images Open the Task 1 folder, and switch to the “task 1 Grayscaleto. RGB” presentation.
- Slides: 24