Bio Curation WG Variant Curation Steven Harrison June

Bio. Curation WG Variant Curation Steven Harrison June 8 2017 sharrison@bwh. harvard. edu
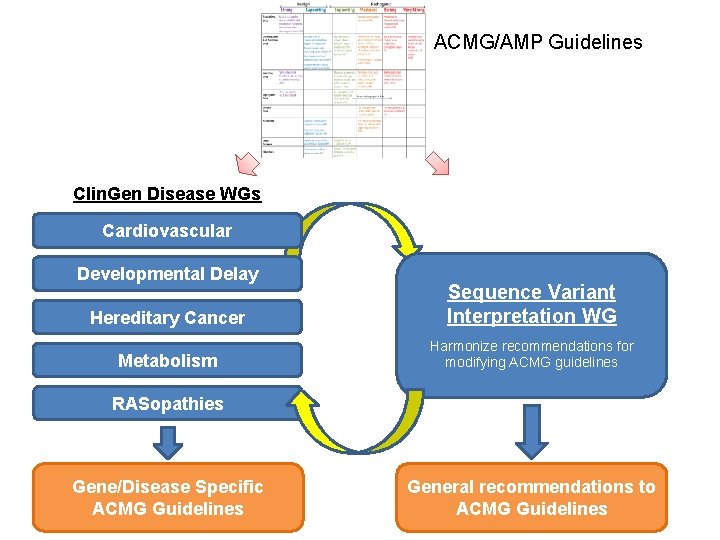
ACMG/AMP Guidelines Clin. Gen Disease WGs Cardiovascular Developmental Delay Hereditary Cancer Sequence Variant Interpretation WG Metabolism Harmonize recommendations for modifying ACMG guidelines RASopathies Gene/Disease Specific ACMG Guidelines General recommendations to ACMG Guidelines
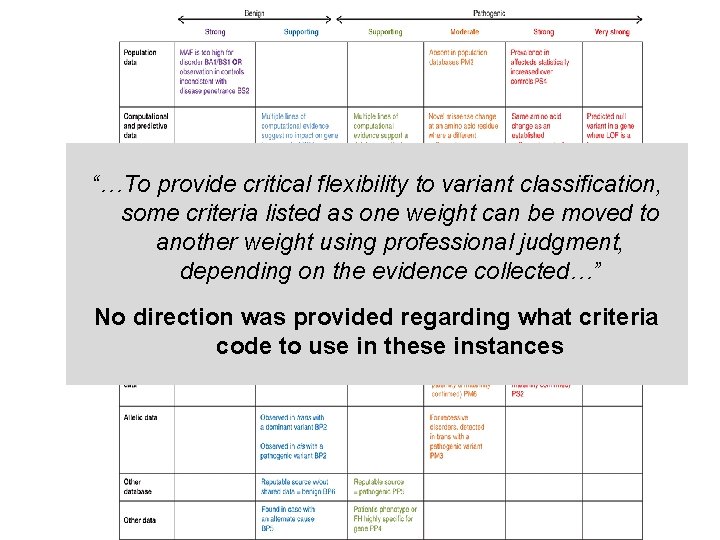
“…To provide critical flexibility to variant classification, some criteria listed as one weight can be moved to another weight using professional judgment, depending on the evidence collected…” No direction was provided regarding what criteria code to use in these instances
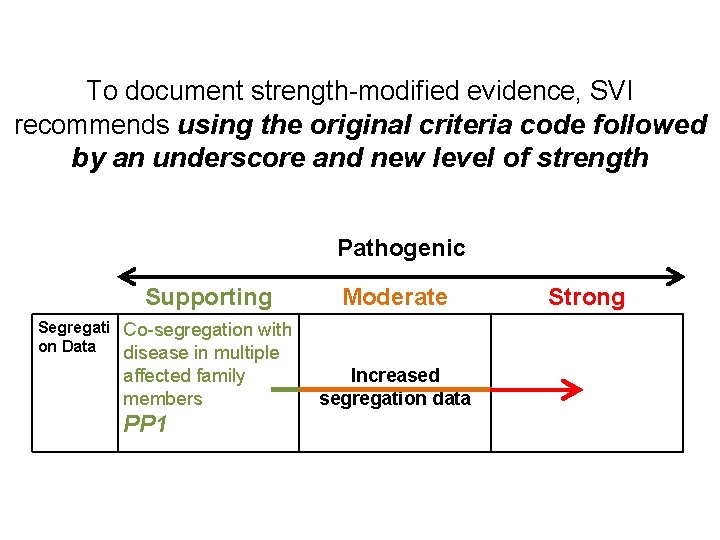
To document strength-modified evidence, SVI recommends using the original criteria code followed by an underscore and new level of strength Pathogenic Supporting Moderate Segregati Co-segregation with on Data disease in multiple affected family members PP 1 Increased segregation data Strong
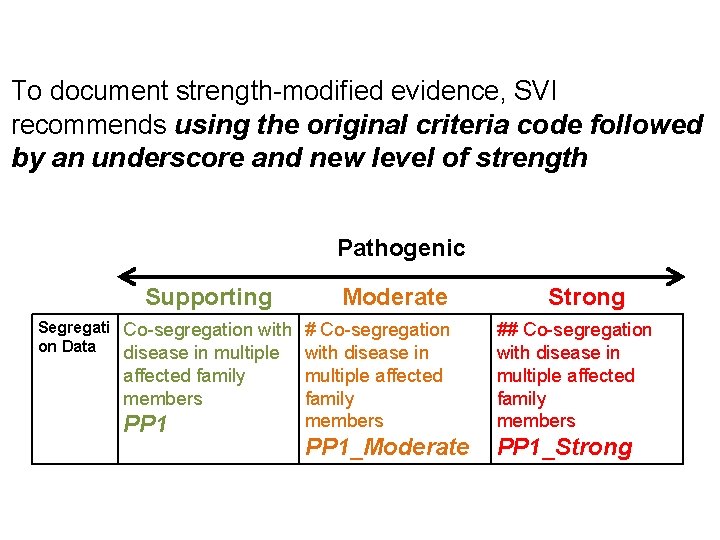
To document strength-modified evidence, SVI recommends using the original criteria code followed by an underscore and new level of strength Pathogenic Supporting Moderate Segregati Co-segregation with # Co-segregation on Data disease in multiple with disease in affected family members PP 1 Strong multiple affected family members ## Co-segregation with disease in multiple affected family members PP 1_Moderate PP 1_Strong
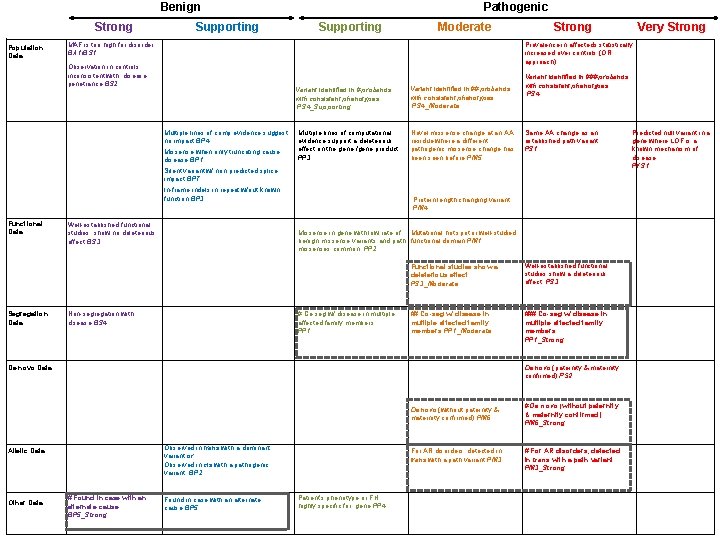
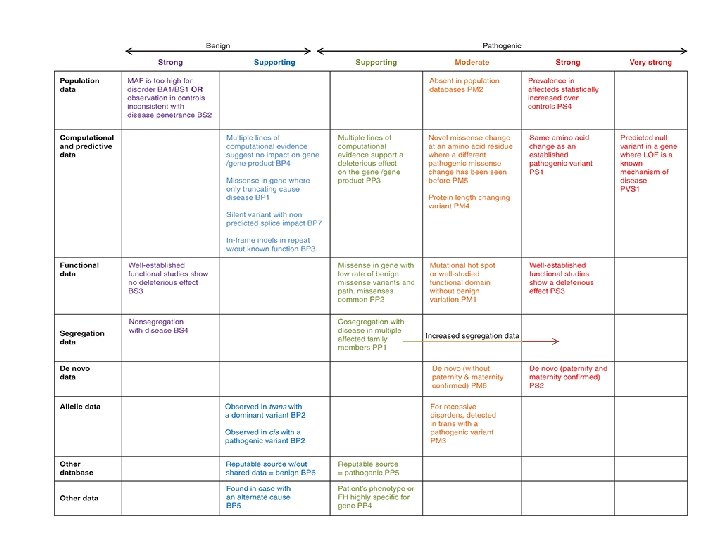
Benign Strong Population Data Supporting Pathogenic Supporting Moderate MAF is too high for disorder BA 1/BS 1 Strong Prevalence in affecteds statistically increased over controls (OR approach) Observation in controls inconsistent with disease penetrance BS 2 Variant identified in # probands with consistent phenotypes PS 4_Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Variant identified in ## probands with consistent phenotypes PS 4_Moderate Novel missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 Variant identified in ### probands with consistent phenotypes PS 4 Same AA change as an established path variant PS 1 Silent variant w/ non predicted splice impact BP 7 In-frame indels in repeat w/out known function BP 3 Functional Data Segregation Data Well-established functional studies show no deleterious effect BS 3 Protein length changing variant PM 4 Missense in gene with low rate of Mutational hot spot or well-studied benign missense variants and path. functional domain PM 1 missenses common PP 2 Non-segregation with disease BS 4 # Co-seg w/ disease in multiple affected family members PP 1 Functional studies show a deleterious effect PS 3_Moderate Well-established functional studies show a deleterious effect PS 3 ## Co-seg w/ disease in multiple affected family members PP 1_Moderate ### Co-seg w/ disease in multiple affected family members PP 1_Strong De novo Data De novo (paternity & maternity confirmed) PS 2 Observed in trans with a dominant variant or Observed in cis with a pathogenic variant BP 2 Allelic Data Other Data # Found in case with an alternate cause BP 5_Strong Very Strong Found in case with an alternate cause BP 5 Patient’s phenotype or FH highly specific for gene PP 4 De novo (without paternity & maternity confirmed) PM 6 # De novo (without paternity & maternity confirmed) PM 6_Strong For AR disorders, detected in trans with a path variant PM 3 # For AR disorders, detected in trans with a path variant PM 3_Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1
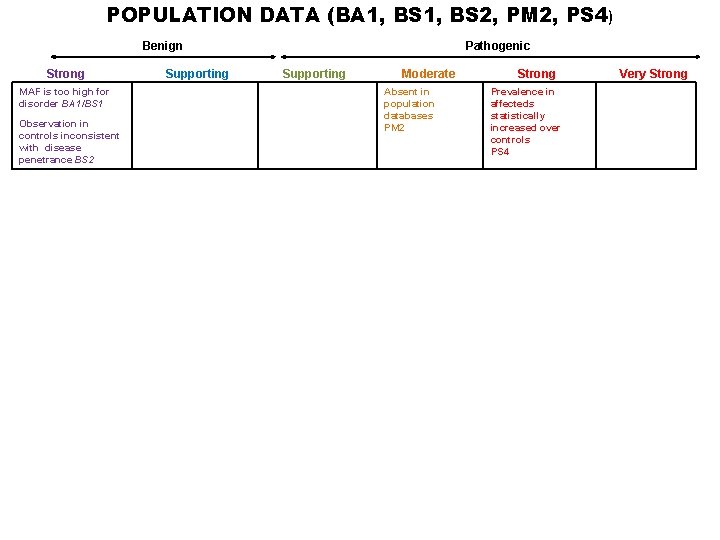
POPULATION DATA (BA 1, BS 2, PM 2, PS 4) Benign Strong MAF is too high for disorder BA 1/BS 1 Observation in controls inconsistent with disease penetrance BS 2 Supporting Pathogenic Supporting Moderate Absent in population databases PM 2 Strong Prevalence in affecteds statistically increased over controls PS 4 Very Strong
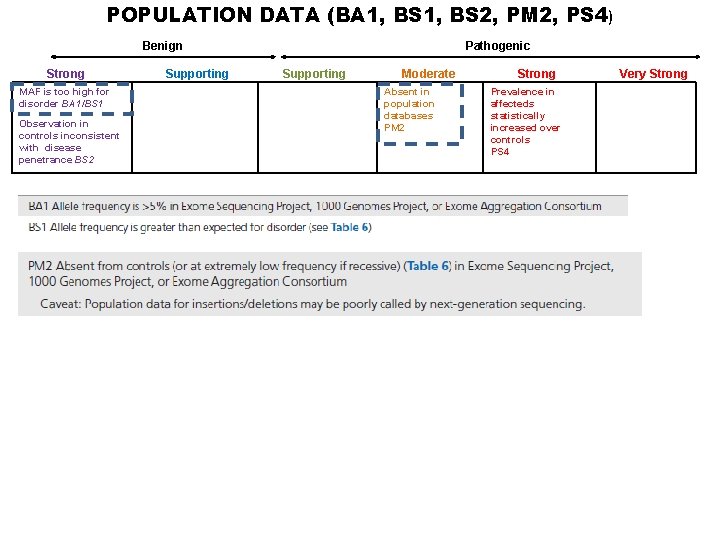
POPULATION DATA (BA 1, BS 2, PM 2, PS 4) Benign Strong MAF is too high for disorder BA 1/BS 1 Observation in controls inconsistent with disease penetrance BS 2 Supporting Pathogenic Supporting Moderate Absent in population databases PM 2 Strong Prevalence in affecteds statistically increased over controls PS 4 Very Strong
Proposed BA 1: Allele frequency is >5% in any general continental population dataset of at least 2, 000 alleles for a gene without a gene- or variant-specific recommendation
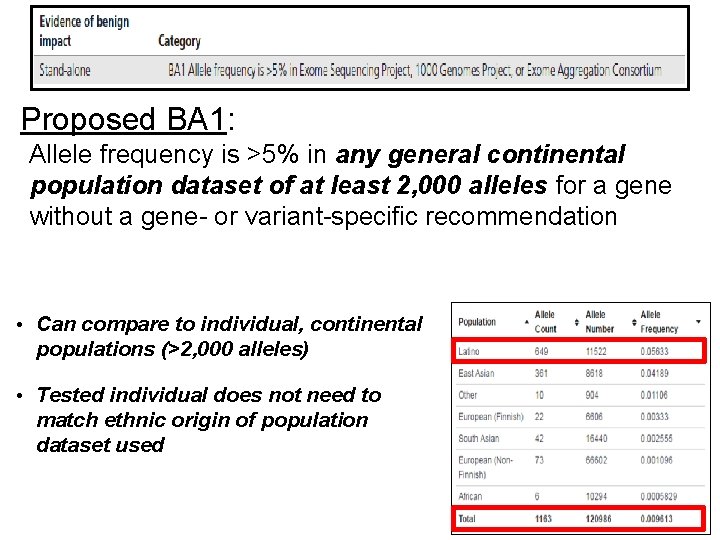
Proposed BA 1: Allele frequency is >5% in any general continental population dataset of at least 2, 000 alleles for a gene without a gene- or variant-specific recommendation • Can compare to individual, continental populations (>2, 000 alleles) • Tested individual does not need to match ethnic origin of population dataset used

Proposed BA 1: Allele frequency is >5% in any general continental population dataset of at least 2, 000 alleles for a gene without a gene- or variant-specific recommendation
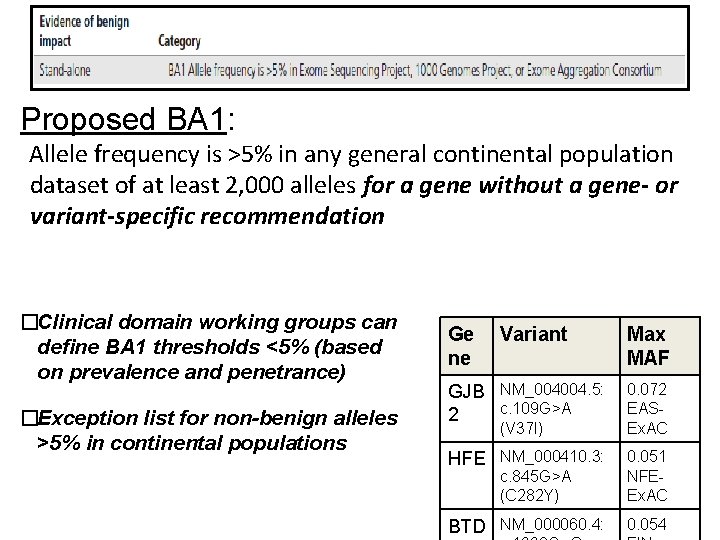
Proposed BA 1: Allele frequency is >5% in any general continental population dataset of at least 2, 000 alleles for a gene without a gene- or variant-specific recommendation �Clinical domain working groups can define BA 1 thresholds <5% (based on prevalence and penetrance) �Exception list for non-benign alleles >5% in continental populations Ge ne Variant Max MAF GJB NM_004004. 5: c. 109 G>A 2 0. 072 EASEx. AC HFE NM_000410. 3: 0. 051 NFEEx. AC BTD NM_000060. 4: 0. 054 (V 37 I) c. 845 G>A (C 282 Y)

Determine Max Credible Population AF Prevalence X Heterogeneity Penetrance BA 1 (stand alone criteria for Benign) • Use MOST conservative estimates BS 1 (Benign strong criteria) • Use less conservative estimates

Determine Max Credible Population AF BA 1 Max Credible AF: 0. 00487 (~0. 5%) • Assuming… • 1/40, 000 prevalence • 95% of Pompe due to GAA • All of GAA-Pompe caused by single variant • Penetrance 100%

Determine Max Credible Population AF BS 1 Max Credible AF: 0. 00096 (~0. 1%) • Assuming… • 1/300, 000 prevalence • 95% of Pompe due to GAA • One GAA variant account for 50% • Penetrance 85%

Great tool to help determine AF thresholds: https: //cardiodb. org/allelefrequencyapp/
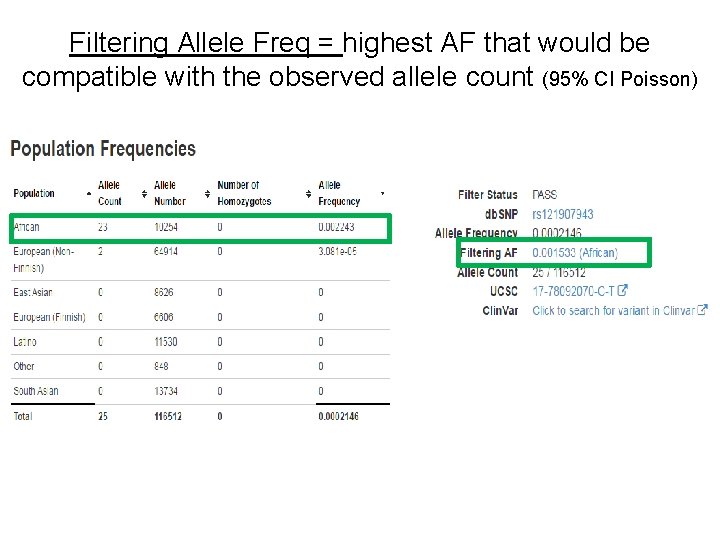
Filtering Allele Freq = highest AF that would be compatible with the observed allele count (95% CI Poisson)

PM 2 – “Absent” • Dominant versus Recessive • However, dominant does not need to be 100% absent, depending on penetrance / age of onset – Hypertrophic cardiomyopathy: <0. 00004 (~5 alleles max) – RASopathy: strictly absent

PM 2 – “Absent” • PM 2 threshold could be inverse of BS 1 – BA 1: ≥ 0. 1% – BS 1: ≥ 0. 05% – PM 2: <0. 05% • IF disorder is recessive or reduced penetrant dominant

POPULATION DATA (BA 1, BS 2, PM 2, PS 4) Benign Strong MAF is too high for disorder BA 1/BS 1 Observation in controls inconsistent with disease penetrance BS 2 Supporting Pathogenic Supporting Moderate Absent in population databases PM 2 Strong Very Strong Prevalence in affecteds statistically increased over controls PS 4 • Generally, should not be used based on occurrences in Ex. AC/gnom. AD as phenotypes are unknown • If disorder IS NOT penetrant at early age, consider removing this criteria • Can also add occurrence thresholds – RASopathy: variant identified in ≥ 3 well phenotyped individuals
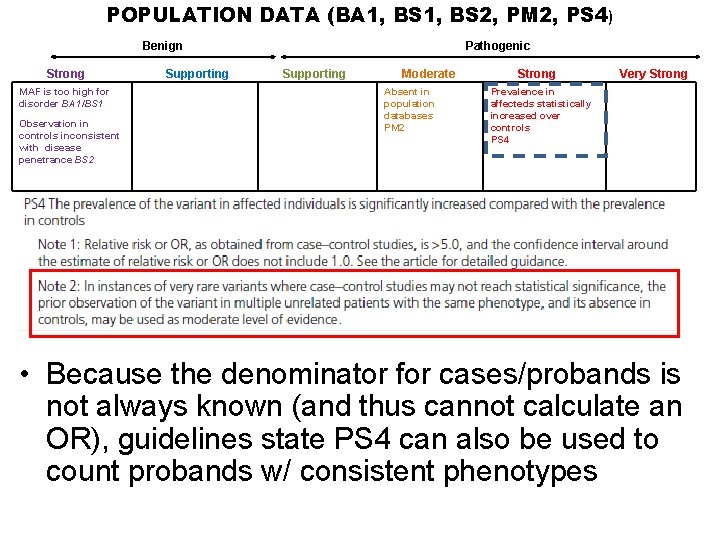
POPULATION DATA (BA 1, BS 2, PM 2, PS 4) Benign Strong MAF is too high for disorder BA 1/BS 1 Observation in controls inconsistent with disease penetrance BS 2 Supporting Pathogenic Supporting Moderate Absent in population databases PM 2 Strong Very Strong Prevalence in affecteds statistically increased over controls PS 4 • Because the denominator for cases/probands is not always known (and thus cannot calculate an OR), guidelines state PS 4 can also be used to count probands w/ consistent phenotypes

POPULATION DATA (BA 1, BS 2, PM 2, PS 4) Benign Strong Supporting Pathogenic Supporting MAF is too high for disorder BA 1/BS 1 Observation in controls inconsistent with disease penetrance BS 2 Variant identified in # probands with consistent phenotypes PS 4_Supporting Moderate Strong Absent in population databases PM 2 Prevalence in affecteds statistically increased over controls (OR method) Variant identified in ## probands with consistent phenotypes PS 4_Moderate Variant identified in ### probands with consistent phenotypes PS 4 Very Strong • PS 4 could be used for typical case-control studies with an OR OR PS 4 could be used for multiple unrelated probands with consistent phenotypes
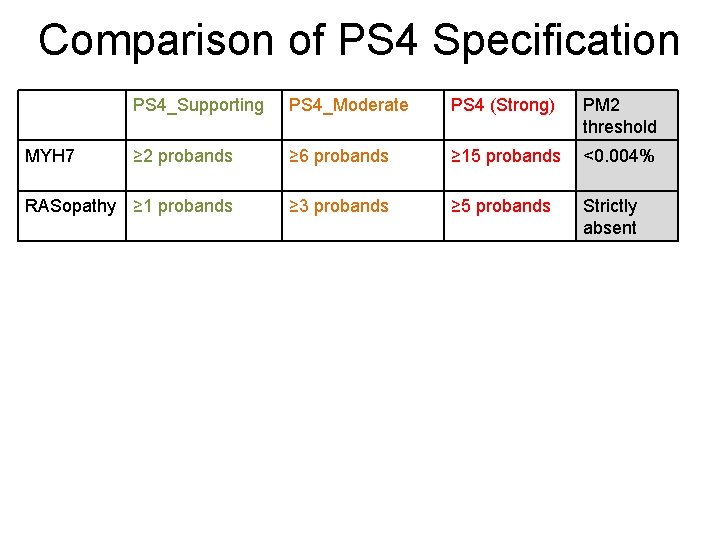
Comparison of PS 4 Specification PS 4_Supporting PS 4_Moderate PS 4 (Strong) PM 2 threshold ≥ 2 probands ≥ 6 probands ≥ 15 probands <0. 004% RASopathy ≥ 1 probands ≥ 3 probands ≥ 5 probands Strictly absent MYH 7

COMPUTATIONAL AND PREDICTIVE DATA Benign Strong Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Pathogenic Supporting Moderate Strong Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Novel missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 Same AA change as an established path variant PS 1 Silent variant w/ non predicted splice impact BP 7 In-frame indels in repeat w/out known function BP 3 Protein length changing variant PM 4 Very Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1

COMPUTATIONAL AND PREDICTIVE DATA Benign Strong Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Pathogenic Supporting Moderate Strong Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Novel missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 Same AA change as an established path variant PS 1 Very Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1 Silent variant w/ non predicted splice impact BP 7 In-frame indels in repeat w/out known function BP 3 Protein length changing variant PM 4 BP 1 could also be used for: • LOF variants in a gene where the disease: – is caused by GOF variants – is caused by dominant/negative LOF variants

COMPUTATIONAL AND PREDICTIVE DATA Benign Strong Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Pathogenic Supporting Moderate Strong Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Novel missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 Same AA change as an established path variant PS 1 Silent variant w/ non predicted splice impact BP 7 In-frame indels in repeat w/out known function BP 3 Protein length changing variant PM 4 Very Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1

COMPUTATIONAL AND PREDICTIVE DATA Benign Strong Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Silent variant w/ non predicted splice impact BP 7 In-frame indels in repeat w/out known function BP 3 Pathogenic Supporting Moderate Strong Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 Same AA change as an established path variant PS 1 Protein length changing variant PM 4 Very Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1

COMPUTATIONAL AND PREDICTIVE DATA Benign Strong Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Silent variant w/ non predicted splice impact BP 7 In-frame indels in repeat w/out known function BP 3 Pathogenic Supporting Moderate Strong Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 Same AA change as an established path variant PS 1 Very Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1 Protein length changing variant PM 4 General modifications for PM 5 and PS 1: • Allow criteria to be used for analogous genes – [HRAS, NRAS, KRAS] ; [MAP 2 K 1, MAP 2 K 2] ; [HBA 1, HBA 2] • Assessing variant c. 37 G>A (p. Gly 13 Ser) in HRAS; variant c. 38 G>A (p. Gly 13 Asp) in KRAS is established Pathogenic • Important to define mapping between genes! – Does Gly 13 in HRAS = Gly 13 in KRAS?

COMPUTATIONAL AND PREDICTIVE DATA Benign Strong Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Pathogenic Supporting Moderate Strong Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 ≥ 2 different missense changes at AA residue have been seen before PM 5_Strong Silent variant w/ non predicted splice impact BP 7 Very Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1 Same AA change as an established path variant PS 1 In-frame indels in repeat w/out known function BP 3 Protein length changing variant PM 4 General modifications for PM 5: • Allow PM 5 to be upgraded to Strong (PM 5_Strong) if ≥ 2 different pathogenic missense changes at the residue have been seen before

COMPUTATIONAL AND PREDICTIVE DATA Benign Strong Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Pathogenic Supporting Moderate Strong Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 ≥ 2 different missense changes at AA residue have been seen before PM 5_Strong Silent variant w/ non predicted splice impact BP 7 Same AA change as an established path variant PS 1 In-frame indels in repeat w/out known function BP 3 Protein length changing variant PM 4 General modifications for PS 1: • Can also be used for splice site variants Very Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1

PS 1: Splice Site Variants • Example: You are assessing IVS 3+5 G>T and IVS 3+5 G>A meets criteria for established pathogenic and has been showed to impact splicing in a functional assay. IVS 3+5 G>T has not been studied functionally but in silico splicing predictions show IVS 3+5 G>T is equally or more damaging than IVS 3+5 G>A = PS 1 met • If in the same scenario (with IVS 3+5 G>A meeting criteria for established pathogenic and has been showed to impact splicing in a functional assay) silico splicing predictions suggest IVS 3+5 G>T is not as damaging as IVS 3+5 G>A = PS 1 not met If PS 1 is met in these instances, PP 3 (in silico prediction) would NOT be applied
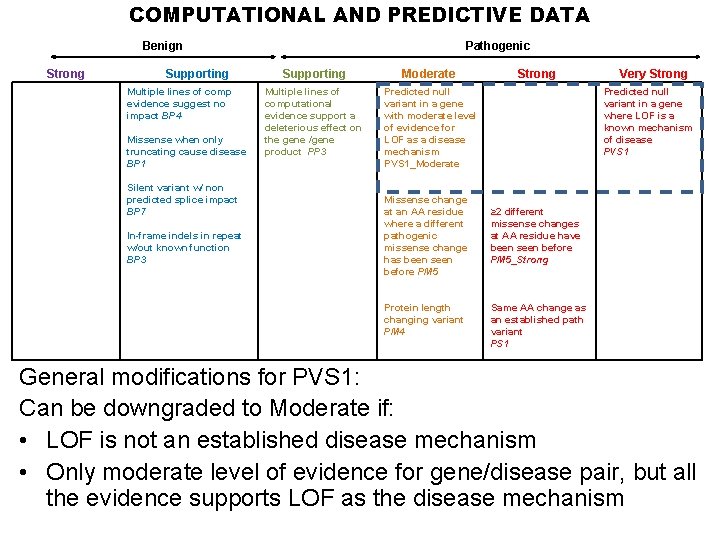
COMPUTATIONAL AND PREDICTIVE DATA Benign Strong Supporting Multiple lines of comp evidence suggest no impact BP 4 Missense when only truncating cause disease BP 1 Silent variant w/ non predicted splice impact BP 7 In-frame indels in repeat w/out known function BP 3 Pathogenic Supporting Moderate Multiple lines of computational evidence support a deleterious effect on the gene /gene product PP 3 Predicted null variant in a gene with moderate level of evidence for LOF as a disease mechanism PVS 1_Moderate Missense change at an AA residue where a different pathogenic missense change has been seen before PM 5 Protein length changing variant PM 4 Strong Very Strong Predicted null variant in a gene where LOF is a known mechanism of disease PVS 1 ≥ 2 different missense changes at AA residue have been seen before PM 5_Strong Same AA change as an established path variant PS 1 General modifications for PVS 1: Can be downgraded to Moderate if: • LOF is not an established disease mechanism • Only moderate level of evidence for gene/disease pair, but all the evidence supports LOF as the disease mechanism

DE NOVO DATA Benign Strong Supporting Pathogenic Supporting Moderate Strong De novo (paternity & maternity confirmed) PS 2 De novo (without paternity & maternity confirmed) PM 6 Very Strong

DE NOVO DATA Example: Patient had a de novo missense variant in MECP 2 but only presented with ID – no Rett phenotypes

DE NOVO DATA Benign Strong Supporting Pathogenic Supporting Moderate Strong De novo (paternity & maternity confirmed) PS 2 De novo (without paternity & maternity confirmed) PM 6 Very Strong ≥ 2 De novo occurrences (paternity & maternity confirmed) PS 2 ≥ 2 De novo occurrences (without paternity & maternity confirmed) PM 6_Strong For PS 2 and PM 6, can increase strength with increasing independent de novo occurrences
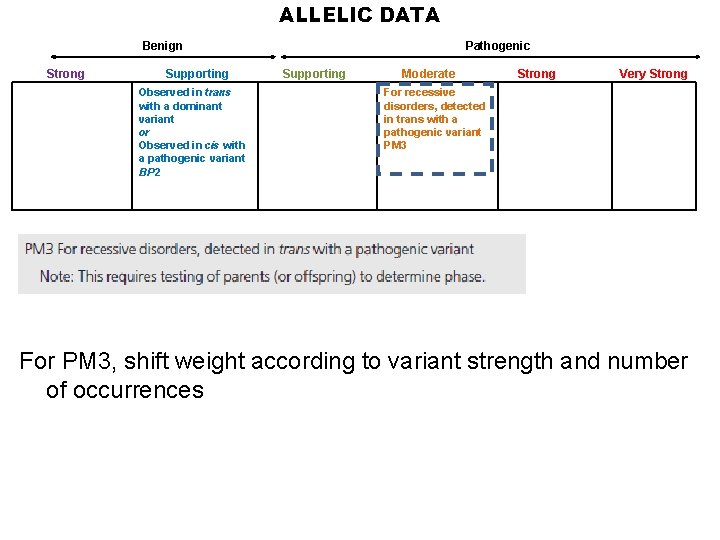
ALLELIC DATA Benign Strong Supporting Observed in trans with a dominant variant or Observed in cis with a pathogenic variant BP 2 Pathogenic Supporting Moderate Strong Very Strong For recessive disorders, detected in trans with a pathogenic variant PM 3 For PM 3, shift weight according to variant strength and number of occurrences
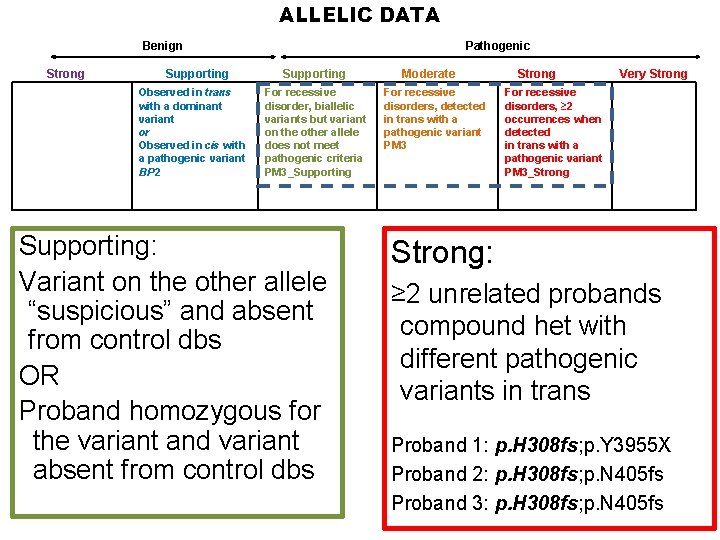
ALLELIC DATA Benign Strong Supporting Observed in trans with a dominant variant or Observed in cis with a pathogenic variant BP 2 Pathogenic Supporting For recessive disorder, biallelic variants but variant on the other allele does not meet pathogenic criteria PM 3_Supporting: Variant on the other allele “suspicious” and absent from control dbs OR Proband homozygous for the variant and variant absent from control dbs Moderate For recessive disorders, detected in trans with a pathogenic variant PM 3 Strong Very Strong For recessive disorders, ≥ 2 occurrences when detected in trans with a pathogenic variant PM 3_Strong: ≥ 2 unrelated probands compound het with different pathogenic variants in trans Proband 1: p. H 308 fs; p. Y 3955 X Proband 2: p. H 308 fs; p. N 405 fs Proband 3: p. H 308 fs; p. N 405 fs
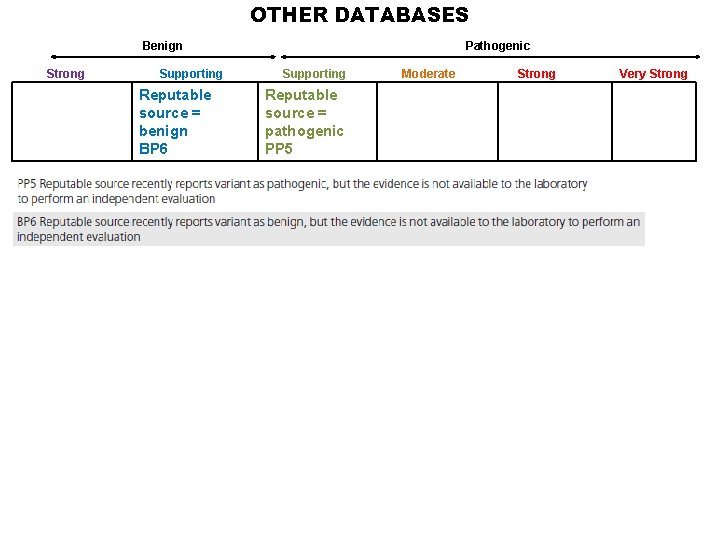
OTHER DATABASES Benign Strong Supporting Reputable source = benign BP 6 Pathogenic Supporting Reputable source = pathogenic PP 5 Moderate Strong Very Strong
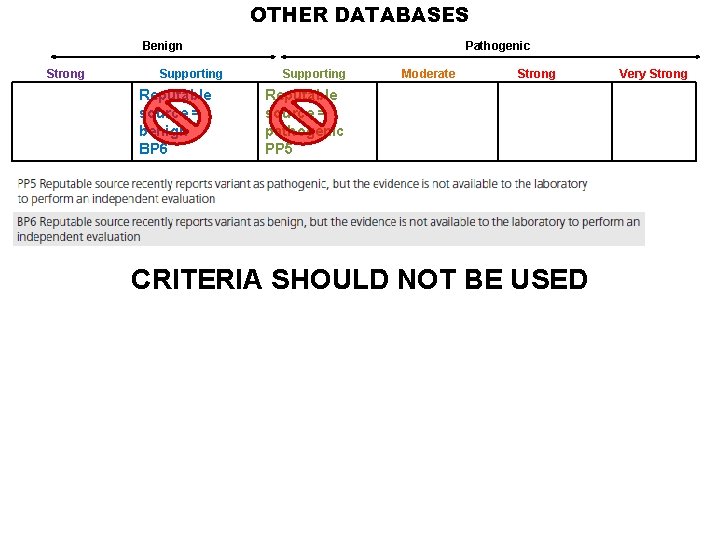
OTHER DATABASES Benign Strong Supporting Reputable source = benign BP 6 Pathogenic Supporting Moderate Strong Reputable source = pathogenic PP 5 CRITERIA SHOULD NOT BE USED Very Strong

Move to Bayes Approach ACMG/AMP criteria are compatible with this quantitative approach

Move to Bayes Approach Allow us to make better definitions for VUS and when Pathogenic and Benign data conflict (PVS# + PM#) + BP# = Pathogenic (PS# + PS#) + BS# = Uncertain significance
- Slides: 42