Bio 260 genetics topic meeting Chapter 7 Some

Bio 260 genetics topic meeting Chapter 7

Some questions first What is DNA? How does DNA protein? How does DNA change? How do organisms evolve? How do pathogens acquire antibiotic resistance? What is the difference between a bacterial species and a sub-species/strain? • How do individual cells adapt to changes in their environment? • • •
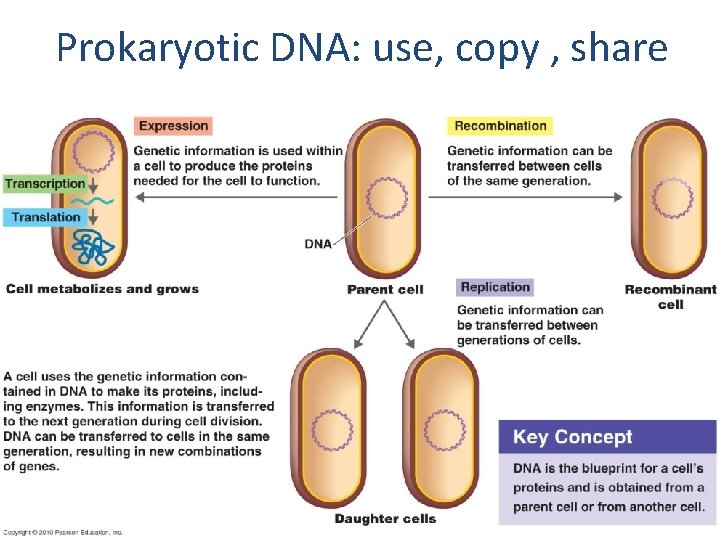
Prokaryotic DNA: use, copy , share

Define your terms • • • DNA replication Transcription Translation Gene expression Gene regulation Operons Recombination Transformation Transduction Conjugation
The importance of genetics • E. coli is found? • What is E. coli O 157: H 7?

The importance of genetics • E. coli is naturally found in the colon, where it is beneficial. • However, the pathogenic strain E. coli O 157: H 7 produces Shiga toxin. This is a strain difference. • How did E. coli acquire this gene from Shigella?

Terminology • What is a gene? • What is the genome? • Define/distinguish between genotype /phenotype? How the genes are expressed detectable physical trait Genotype Phenotype

Why we should care: the practical consequence of phenotype • Genes make proteins for example? • Microbial features – structure (eg. cell wall structure, pili) and function (fermentation; nitrogen catabolism) determined by proteins which are determined by genes • Microbial diversity and identity can be determined by sequencing the microbial genome • Genetic change (change to the genotype) is the basis of acquired drug resistance and increased or decreased pathogenesis (phenotype) • Genetic engineering is used in industry to give microbes the ability to make useful proteins

The importance of protein for cells Objects (tools, machines) made by cell for specific jobs
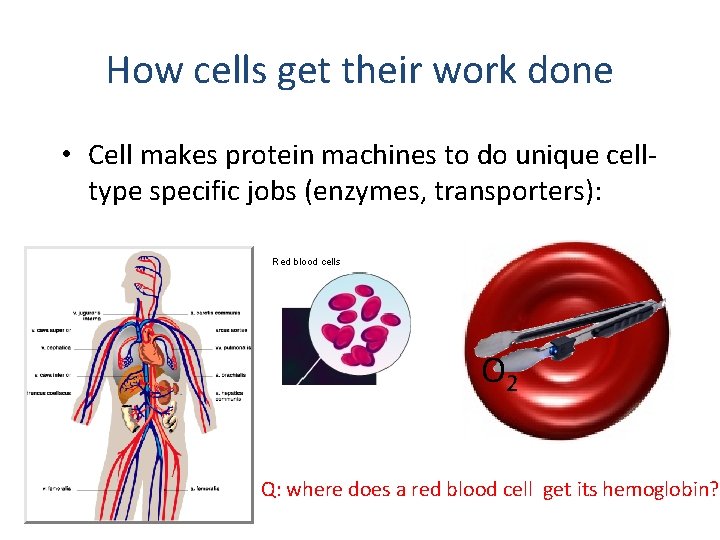
How cells get their work done • Cell makes protein machines to do unique celltype specific jobs (enzymes, transporters): Red blood cells O 2 Q: where does a red blood cell get its hemoglobin?
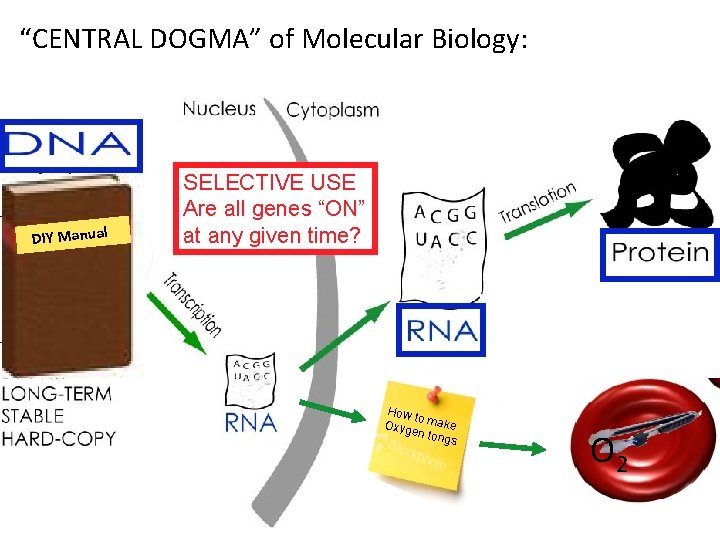
“CENTRAL DOGMA” of Molecular Biology: DIY Manual SELECTIVE USE Are all genes “ON” at any given time? How to Oxyg make en to ngs O 2
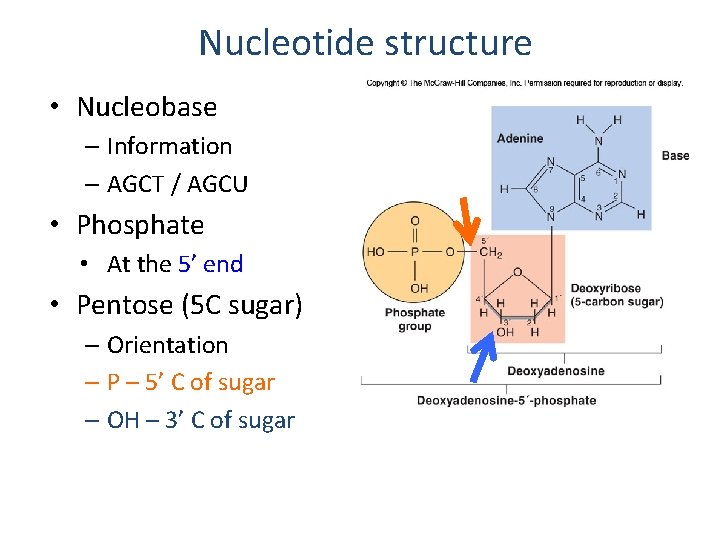
Nucleotide structure • Nucleobase – Information – AGCT / AGCU • Phosphate • At the 5’ end • Pentose (5 C sugar) – Orientation – P – 5’ C of sugar – OH – 3’ C of sugar
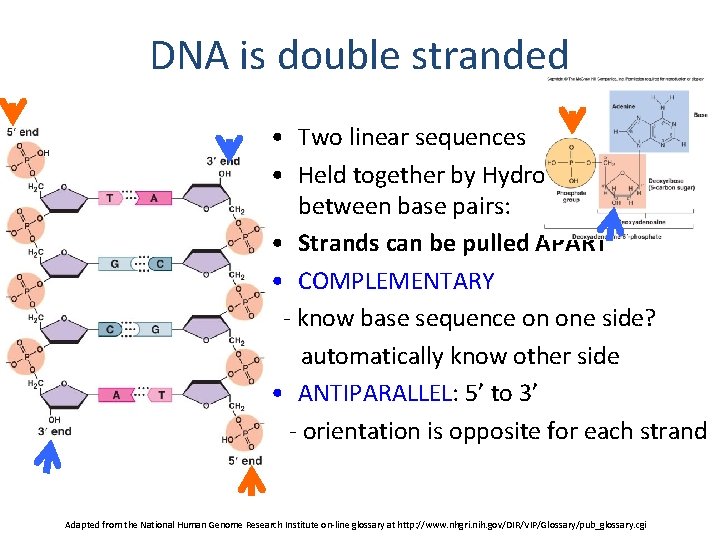
DNA is double stranded • Two linear sequences • Held together by Hydrogen bonds between base pairs: • Strands can be pulled APART • COMPLEMENTARY - know base sequence on one side? automatically know other side • ANTIPARALLEL: 5’ to 3’ - orientation is opposite for each strand Adapted from the National Human Genome Research Institute on-line glossary at http: //www. nhgri. nih. gov/DIR/VIP/Glossary/pub_glossary. cgi

Short group exercise • Draw sugar-phosphate-base structure of double stranded DNA with the following sequence: • 5’ – ATGCCCTGA – 3’ • NOTES: show it is antiparallel and complementary
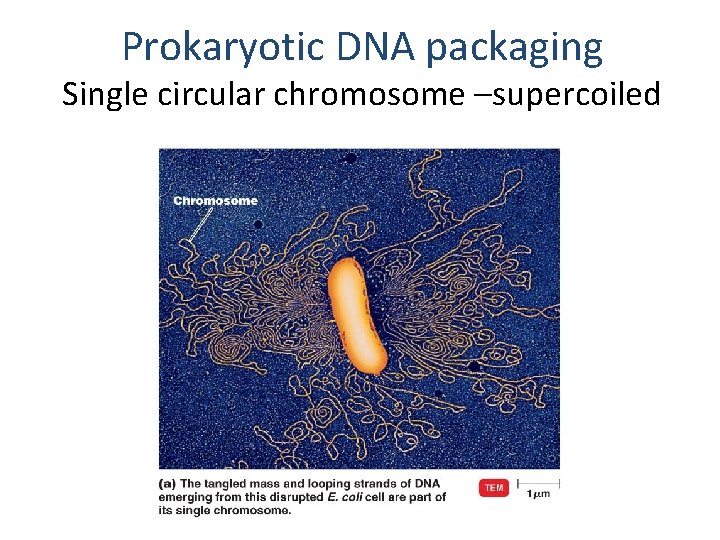
Prokaryotic DNA packaging Single circular chromosome –supercoiled
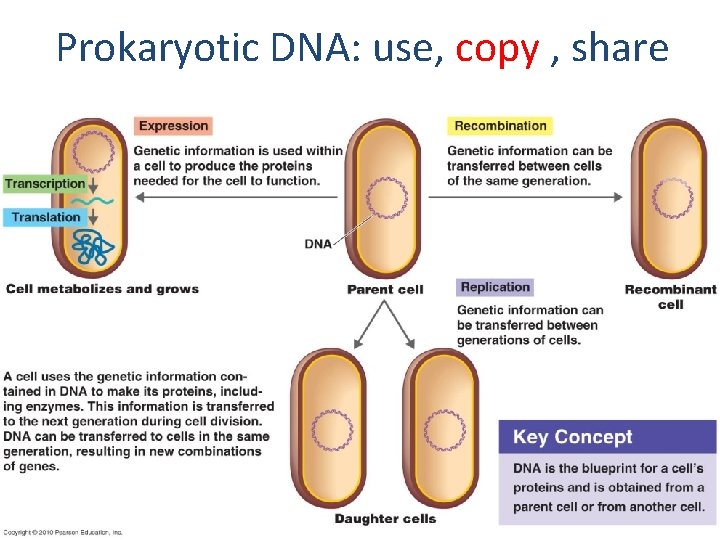
Prokaryotic DNA: use, copy , share
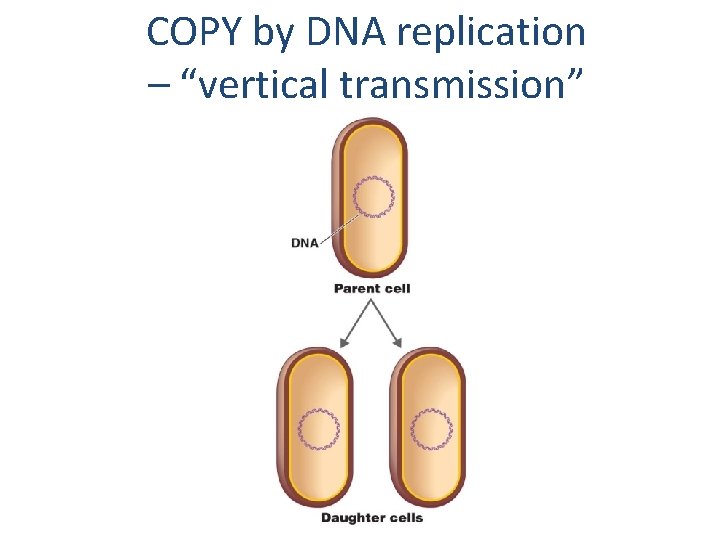
COPY by DNA replication – “vertical transmission”

DNA replication • http: //www. youtube. com/watch? v=4 jtm. OZa. I v. S 0
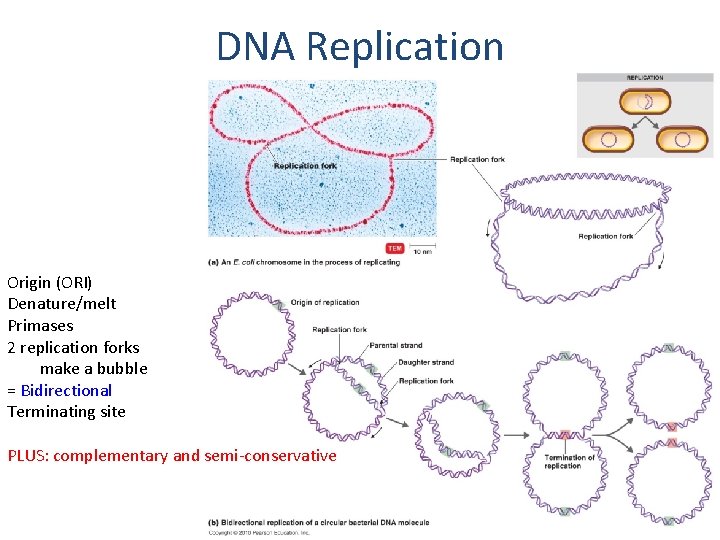
DNA Replication Origin (ORI) Denature/melt Primases 2 replication forks make a bubble = Bidirectional Terminating site PLUS: complementary and semi-conservative

DNA Replication • Replication begins at ________ – Proteins recognize and bind to site – _____ melts the double-stranded DNA – _____ synthesizes short stretches of complementary RNA called _____ to start the reaction – _____ replicates, _____ replaces RNA primers – _____ seals the breaks in the sugar-phosphate backbone • Two replication ____ create a replication _____ • Replication continues simultaneously in both directions making the process _____ • Because the DNA is antiparallel, replication creates a continuously synthesized _____ strand a discontinuously synthesized _____ strand – DNA pol III works 5’ to 3’ – one strand is easy – Other one done in short pieces “_____ fragments” by DNA pol III and pol I and primase all working together


DNA Replication • Replication begins at origin of replication – Proteins recognize and bind to site – Helicase melts the double-stranded DNA – RNA Primase synthesizes short stretches of complementary RNA called primers to start the reaction – DNA pol III replicates, DNA pol I replaces RNA primers – DNA ligase seals the breaks in the sugar-phosphate backbone • Creates two replication forks (enlarging “replication bubble”) – Leave origin going in opposite directions; bidirectional – Ultimately meet at terminating site when process complete • Leading and lagging strands (because it’s anti-parallel) – DNA pol III works 5’ to 3’ – one strand is easy – Other one done in short pieces “Okazaki fragments” by DNA pol III and pol I and primase all working together
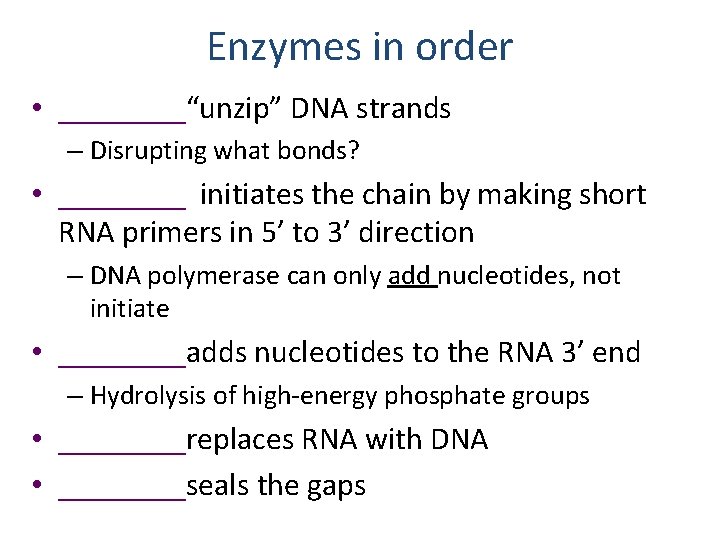
Enzymes in order • ____“unzip” DNA strands – Disrupting what bonds? • ____ initiates the chain by making short RNA primers in 5’ to 3’ direction – DNA polymerase can only add nucleotides, not initiate • ____adds nucleotides to the RNA 3’ end – Hydrolysis of high-energy phosphate groups • ____replaces RNA with DNA • ____seals the gaps

Enzymes in order • Helicase unzip” DNA strands – Disrupting what bonds? • RNA primase (a dedicated RNA polymerase) initiates the chain by making short RNA primers in 5’ to 3’ direction – DNA polymerase can only add nucleotides, not initiate • DNA polymerase I adds nucleotides to the RNA 3’ end – Hydrolysis of high-energy phosphate groups • DNA pol III replaces RNA with DNA • DNA ligase seals the gaps
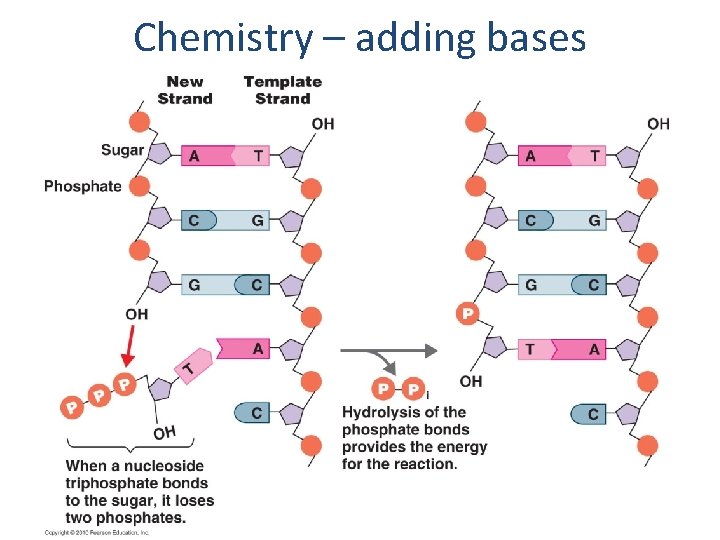
Chemistry – adding bases
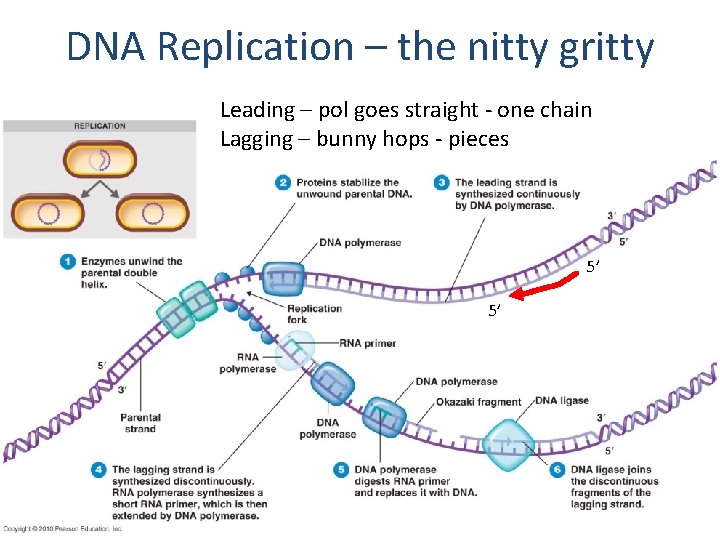
DNA Replication – the nitty gritty Leading – pol goes straight - one chain Lagging – bunny hops - pieces 5’ 5’

Test yourself
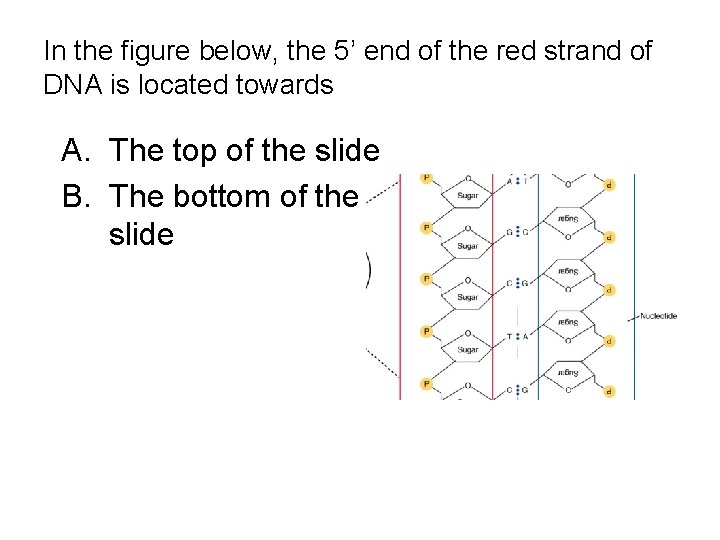
In the figure below, the 5’ end of the red strand of DNA is located towards A. The top of the slide B. The bottom of the slide

Which enzyme copies DNA during DNA replication? A. B. C. D. E. DNA ligase DNA polymerase DNA helicase DNA gyrase None of the above
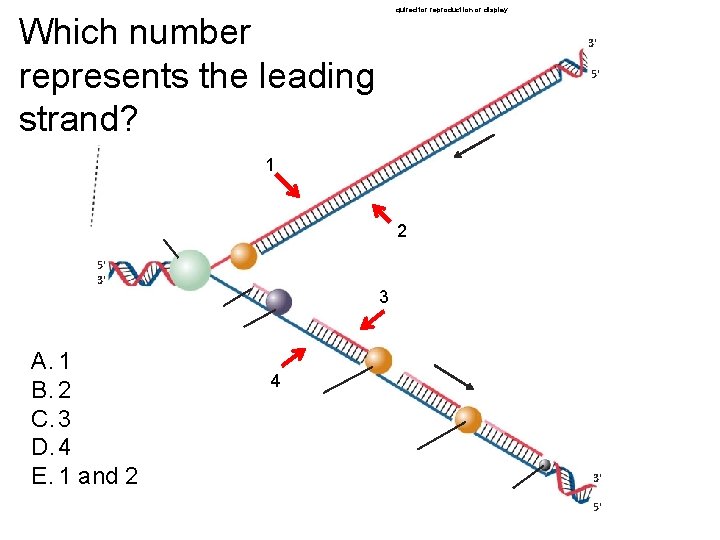
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Which number represents the leading strand? 3' 5' 1 2 5' 3' A. 1 B. 2 C. 3 D. 4 E. 1 and 2 3 4 3' 5'
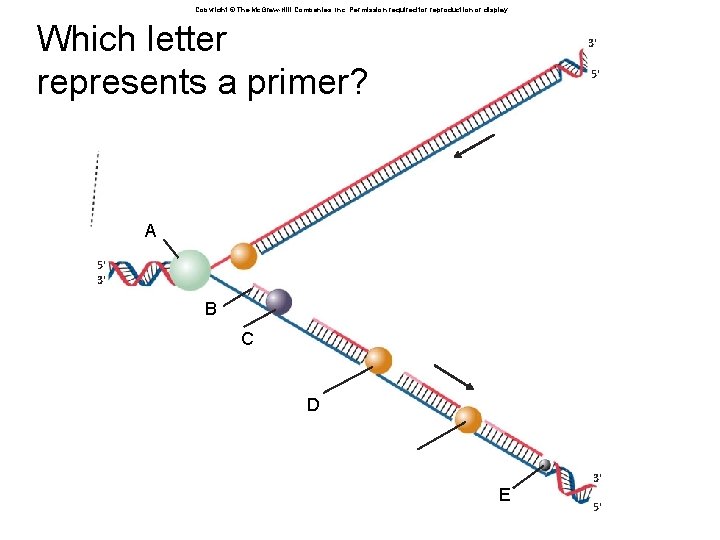
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Which letter represents a primer? 3' 5' A 5' 3' B C D E 3' 5'
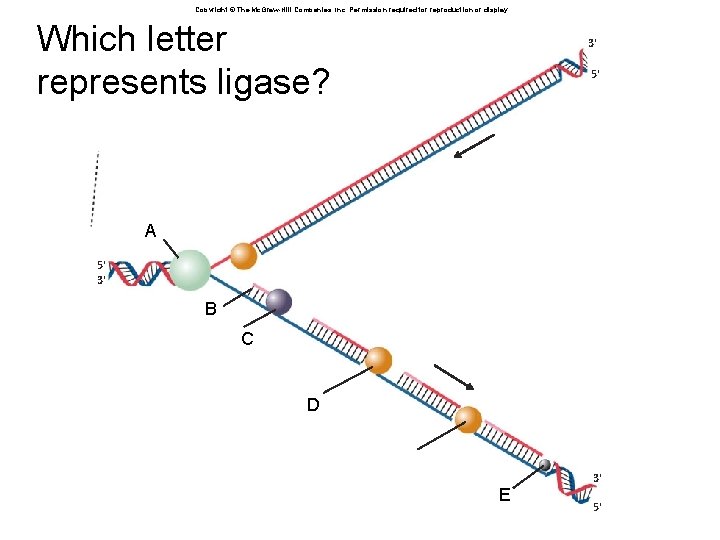
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Which letter represents ligase? 3' 5' A 5' 3' B C D E 3' 5'

A bacterial cell that had a mutation in the gene for Primase (RNA polymerase) such that the enzyme did not function would NOT be able to A. Synthesize the leading strand only B. Synthesize the lagging strand only C. Synthesize either the leading or lagging strand
Continuous replication in prokaryotes • Replication produces two copies of DNA – Each daughter cell receives one – Replication of E. coli chromosome takes ~40 minutes • Remember optimal generation time is ? • Cell can initiate new replication before previous round is complete • Each daughter cell inherits one complete chromosome already undergoing replication
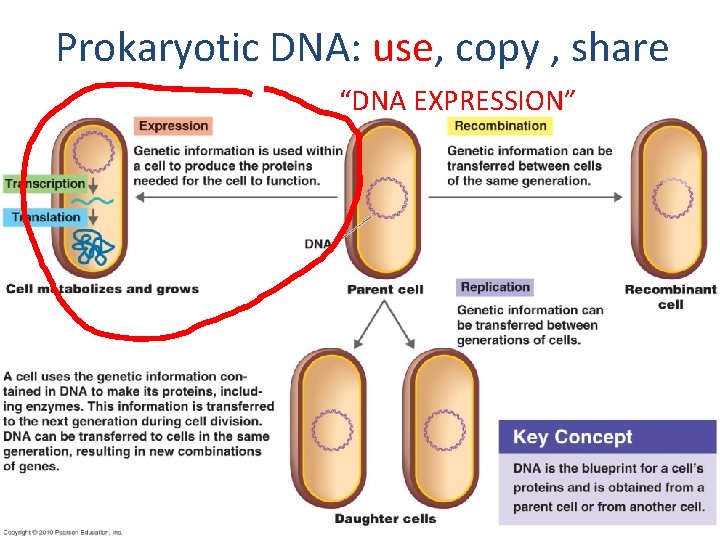
Prokaryotic DNA: use, copy , share “DNA EXPRESSION”

RNA types • 3 main types – m. RNA • Carries information from DNA to ribosome – r. RNA • Part of structure and function of ribosome – t. RNA • Carries amino acids to ribosome • Decodes m. RNA
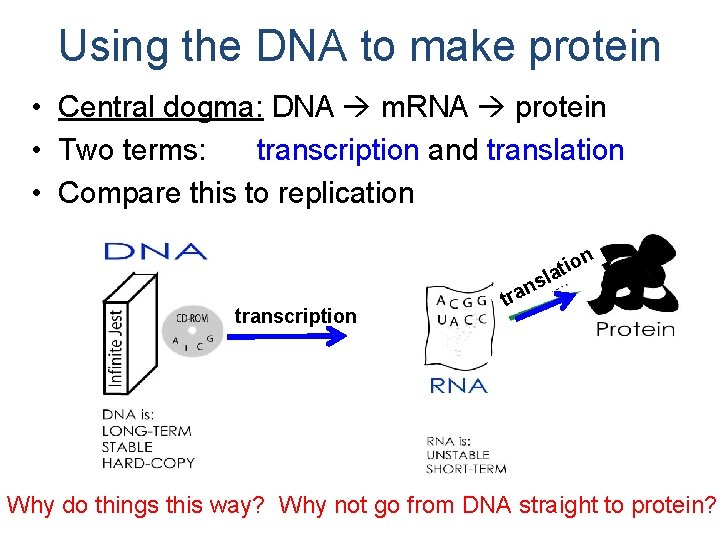
Using the DNA to make protein • Central dogma: DNA m. RNA protein • Two terms: transcription and translation • Compare this to replication io lat n s transcription n transcription Why do things this way? Why not go from DNA straight to protein?

Gene Expression: DNA m. RNA protein • http: //www. youtube. com/watch? v=41_Ne 5 m. S 2 ls&feature=related
Transcription Terminology • Promoter – Sequence in DNA – Marks beginning of genes • Bacteria may have multiple genes after one promoter • RNA polymerase – Able to open the DNA on its own (no need for _____) – Enzyme that synthesizes RNA from DNA template – Makes new RNA in 5’ to 3’ direction without needing primrs • Transcription Terminator – Sequence in DNA – Marks ends of genes
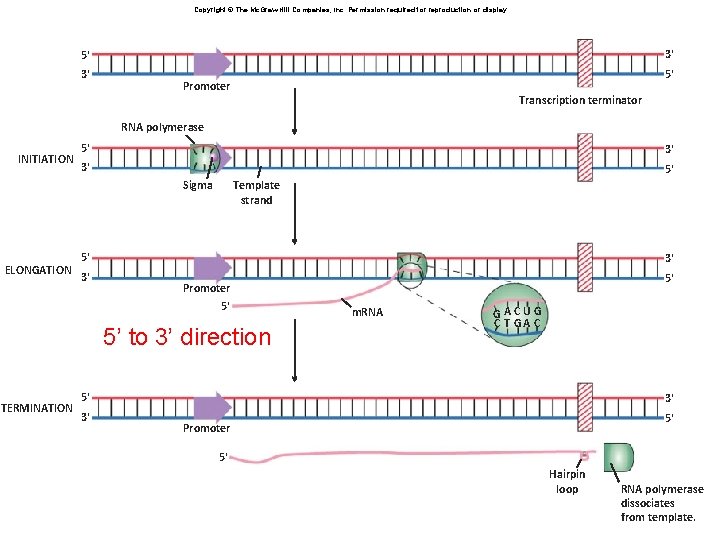
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. 3' 5' Promoter Transcription terminator RNA polymerase INITIATION 5' 3' 3' 5' Sigma ELONGATION Template strand 5' 3' 3' Promoter 5' 5’ to 3’ direction TERMINATION 5' m. RNA GACUG C T GA C 5' 3' 3' 5' Promoter 5' Hairpin loop RNA polymerase dissociates from template.


Transcription – in real life You need to understand what is happening in this picture Where is – the DNA template? Promoter? Terminator? Elongation happening? Why >1 m. RNA?

Transcription vs replication • RNA polymerase synthesizes RNA in 5’ to 3’ direction – Compare this to DNA polymerase – similar, different • Opens the DNA on its own – What enzyme did this in DNA replication? • Can initiate without primer – Where in replication do you see a similar enzyme used? • Binds to promoter (upstream of genes) – Where does replication machinery bind and start? • Stops at terminator (transcription ends) – Same as replication termination site? Why not?

True or false: All the information in the DNA of a cell is organized into genes that code for proteins. A. True B. False

Which of the following is the best description of the role of RNA polymerase? A. B. C. D. To translate m. RNA into protein To replicate the DNA To copy the code in the DNA into m. RNA To synthesize several types of RNA using the code in the DNA E. To degrade polymers of RNA

True or false: during the process of transcription, DNA and RNA nucleotides bind to each other by hydrogen bonds A. True B. False
Translation – reagent, product, enzyme? Figure 8. 2

Translation – what is the m. RNA language? • • • Carries DNA information translated in codons (3 nucleotides) begins at start codon: AUG ends at nonsense codons: UAA, UAG, UGA 64 sense codons encode 20 amino acids The genetic code is degenerate
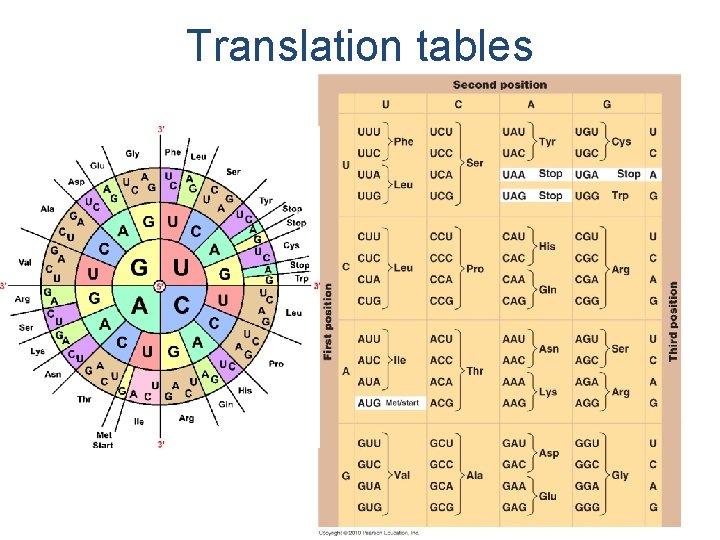
Translation tables

Codons • Start codon – AUG nearest 5’ end – Codes for Met – Marks beginning – Sets reading frame • Stop codon – UAA, UAG, UGA – Marks end – No amino acid


Genetics activity

The process of transcription begins at A. B. C. D. E. Start codons Promoters Origins of replication Transcription initiators None of the above

During translation, ribosomes function to A. Bind the m. RNA B. Bind the t. RNA C. Catalyze bond formation between amino acids D. Both A and B are correct E. All (A-C) are correct

In the table shown below, the three letter codes such as “AUG” and “ACA” represent A. Codons in m. RNA B. Anticodons in t. RNA C. Both A and B

When m. RNA is translated, the “reading frame” is directly determined by A. The location of the start codon closest to the 5’ end of the m. RNA B. The location of the ribosome binding sequence in the m. RNA C. The reading frame initiator sequence D. All of the above are correct E. None of the above are correct

A cell that has a mutation such that the tryptophan-t. RNA cannot bind tryptophan would A. Be unable to initiate translation B. Be unable to make any proteins C. Be unable to complete any proteins that contain tryptophan D. Complete the synthesis of all proteins E. Complete the synthesis of all proteins, but would be missing tryptophan from these proteins

Exons vs introns
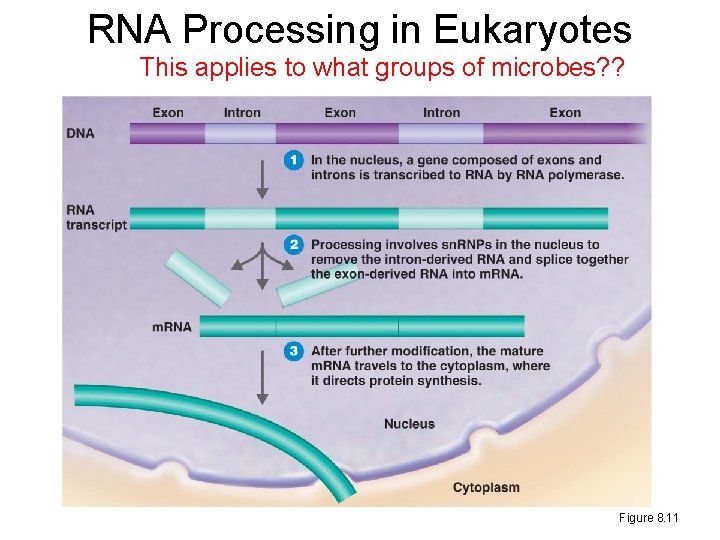
RNA Processing in Eukaryotes This applies to what groups of microbes? ? Figure 8. 11

eukaryotes v prokaryotes –differences Eukaryote • What is the difference by definition? • Replication – Linear chromosomes – In nucleus; must wait for nucleus to divide completely • Transcription – In nucleus – m. RNA processing • Translation – In cytoplasm – Must wait for transcript to exit nucleus – 80 s ribosome Prokaryote • Replication – Single circular chromosome* – In cytoplasm; can replicate again before first round is finished • Transcription – In cytoplasm – Always available • Translation – In cytoplasm – Can simultaneously translate as transcript is being made!!! – 70 s ribosome

Major differences eukaryotes v prokaryotes Eukaryote • Nucleus • Replication Prokaryote • No nucleus • Replication • Transcription • Translation – Linear chromosomes – In nucleus; must wait for nucleus to divide completely – In nucleus – m. RNA processing – In cytoplasm – Must wait for transcript to exit nucleus – 80 s ribosome – Single circular chromosome* – In cytoplasm; can replicate again before first round is finished – In cytoplasm – Always available – In cytoplasm – Can simultaneously translate as transcript is being made!!! – 70 s ribosome

Major differences eukaryotes v prokaryotes Eukaryote • Nucleus • Replication Prokaryote • No nucleus • Replication • Transcription • Translation – Linear chromosomes – In nucleus; must wait for nucleus to divide completely – In nucleus – m. RNA processing – In cytoplasm – Must wait for transcript to exit nucleus – 80 s ribosome – Single circular chromosome* – In cytoplasm; can replicate again before first round is finished – In cytoplasm – Always available – In cytoplasm – Can simultaneously translate as transcript is being made!!! – 70 s ribosome *there are exceptions

Major differences eukaryotes vs prokaryotes Eukaryote • Nucleus • Replication Prokaryote • No nucleus • Replication • Transcription • Translation – Linear chromosomes – In nucleus; must wait for nucleus to divide completely – In nucleus – m. RNA processing – In cytoplasm – Must wait for transcript to exit nucleus – 80 s ribosome This means what in terms of speed? ? – Single circular chromosome* – In cytoplasm; can replicate again before first round is finished – In cytoplasm – Always available – In cytoplasm – Can simultaneously translate as transcript is being made!!! – 70 s ribosome
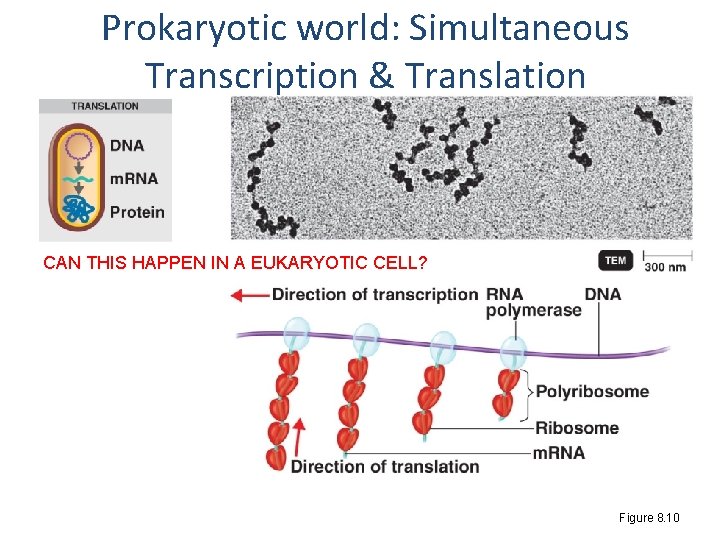
Prokaryotic world: Simultaneous Transcription & Translation CAN THIS HAPPEN IN A EUKARYOTIC CELL? Figure 8. 10

One last thing about “using” the DNA • Are all genes always made into protein all the time? • How is gene expression regulated (in bacteria)?
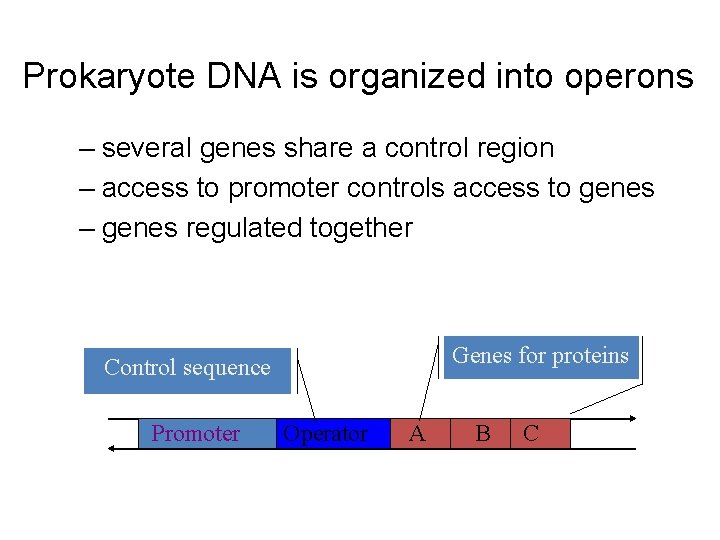
Prokaryote DNA is organized into operons – several genes share a control region – access to promoter controls access to genes – genes regulated together Genes for proteins Control sequence Promoter Operator A B C

Regulation terminology • Constitutive genes are “ON” all the time • Regulated genes are expressed only as needed – Repressible genes – Inducible genes – Catabolite repression
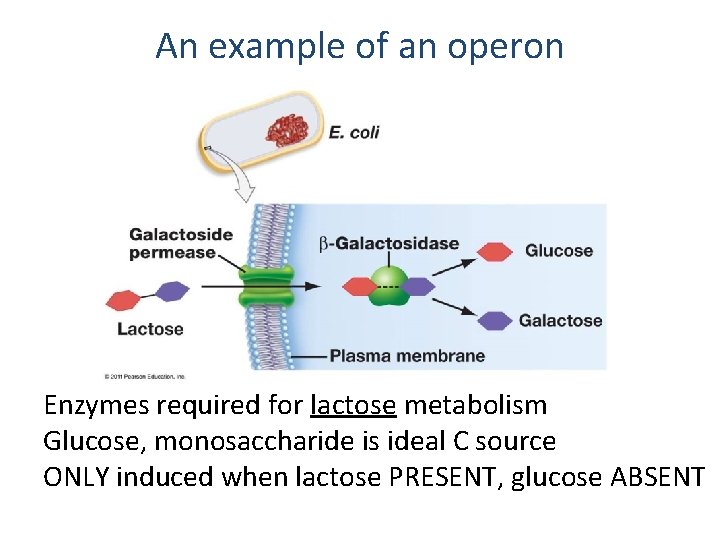
An example of an operon Enzymes required for lactose metabolism Glucose, monosaccharide is ideal C source ONLY induced when lactose PRESENT, glucose ABSENT
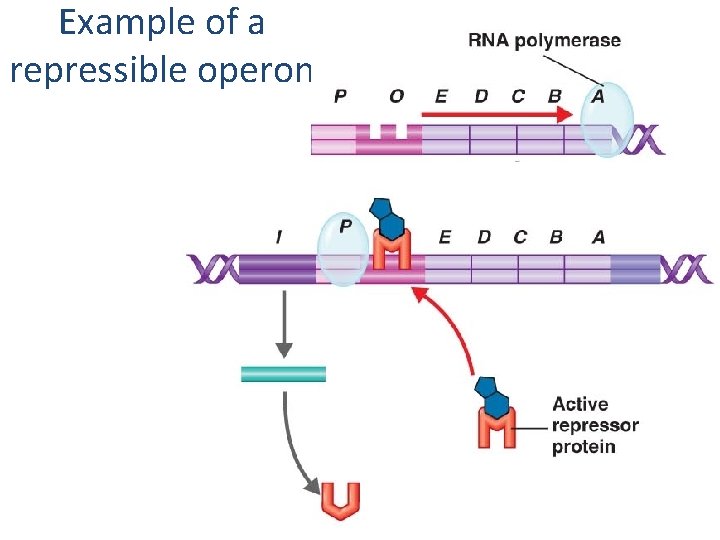
Example of a repressible operon Figure 8. 13
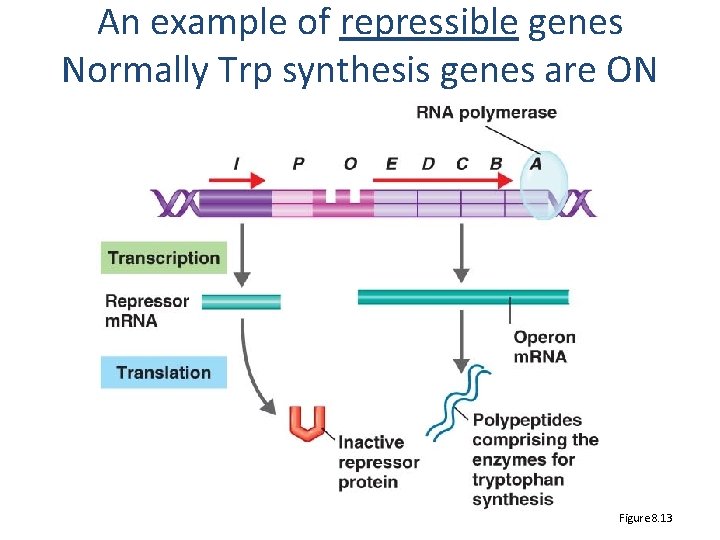
An example of repressible genes Normally Trp synthesis genes are ON Figure 8. 13

Repressible, inducible • Repressible – want these on by default – turn OFF when not needed – Eg. Synthesizing amino acids (Tryptophan) • Inducible – only need them sometimes – turn them ON only when needed – Eg. Metabolizing unusual carbon sources (lactose)
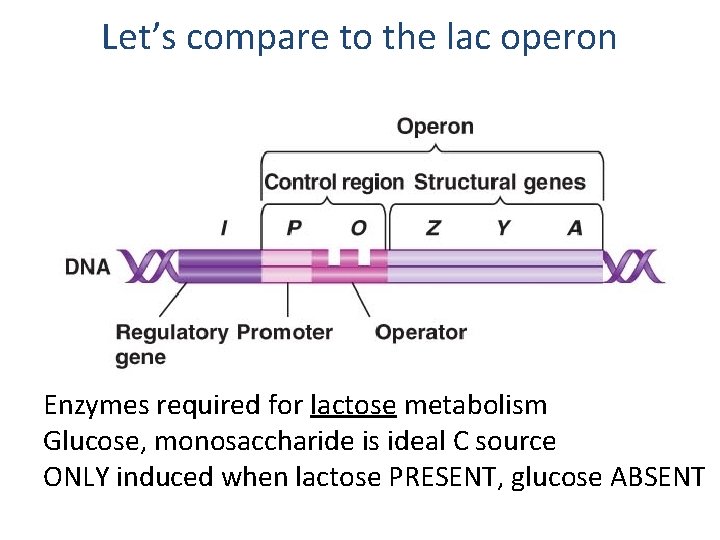
Let’s compare to the lac operon Enzymes required for lactose metabolism Glucose, monosaccharide is ideal C source ONLY induced when lactose PRESENT, glucose ABSENT
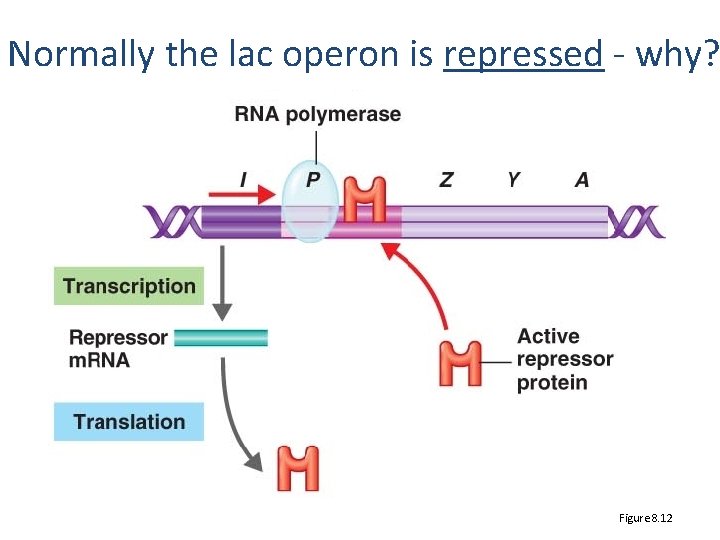
Normally the lac operon is repressed - why? Figure 8. 12
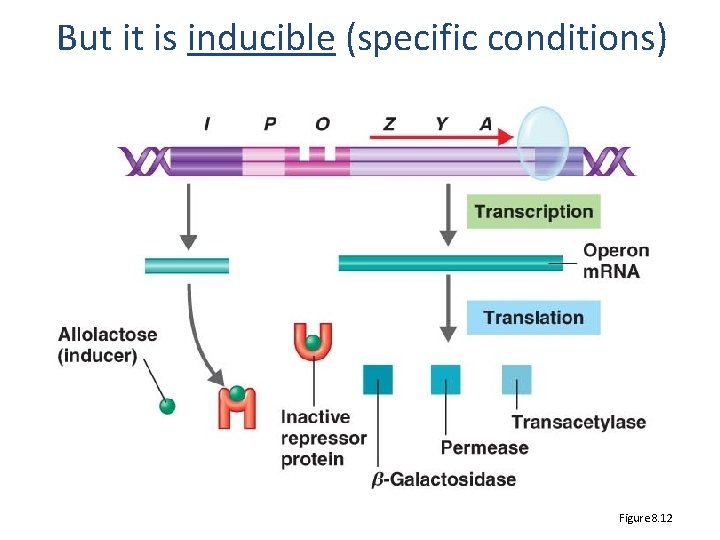
But it is inducible (specific conditions) Figure 8. 12

Is it really that simple • Of COURSE NOT • Remember: lactose operon ON only when lactose present AND glucose absent
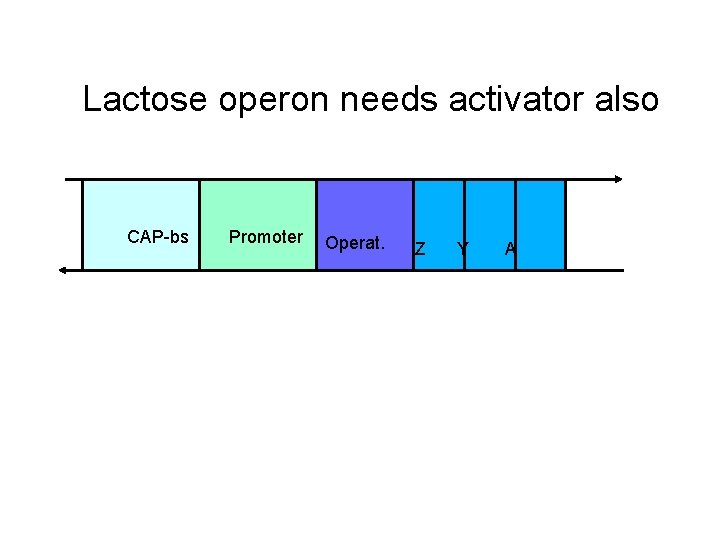
Lactose operon needs activator also CAP-bs Promoter Operat. Z Y A

CAP is an activator • Catabolite activator protein (CAP) is an activator of many catabolic operons – E. coli preferentially uses glucose – when glucose is low, need enzymes to use other carbon sources

How CAP works • CAP is controlled by cyclic AMP (c. AMP) – binds and activates CAP – When glucose is high, c. AMP is low • So CAP will be ? • Will it be available to bind activate lac operon? – When glucose is low, c. AMP is high • CAP will be ? Consequence to lac operon?
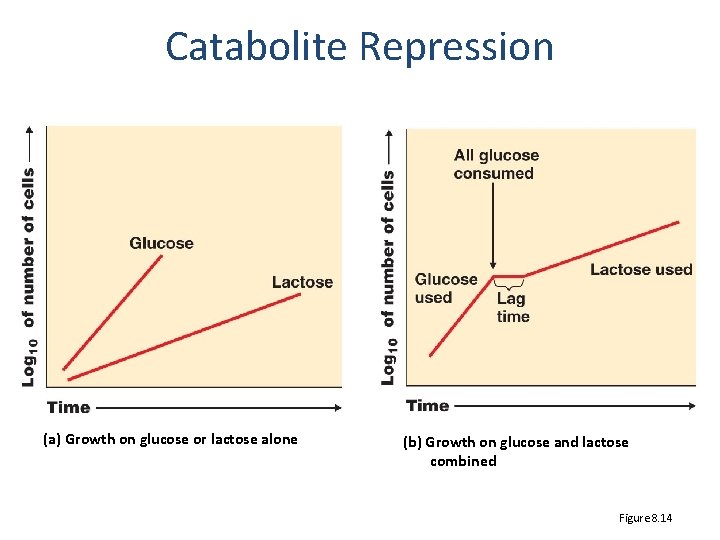
Catabolite Repression (a) Growth on glucose or lactose alone (b) Growth on glucose and lactose combined Figure 8. 14
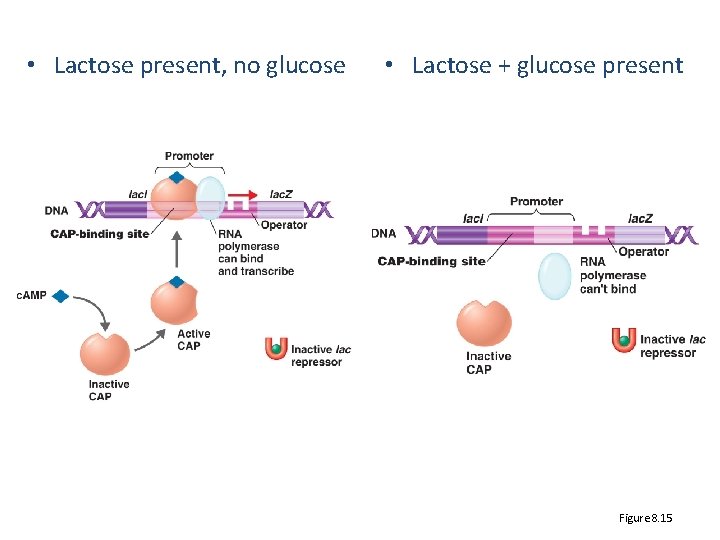
• Lactose present, no glucose • Lactose + glucose present Figure 8. 15
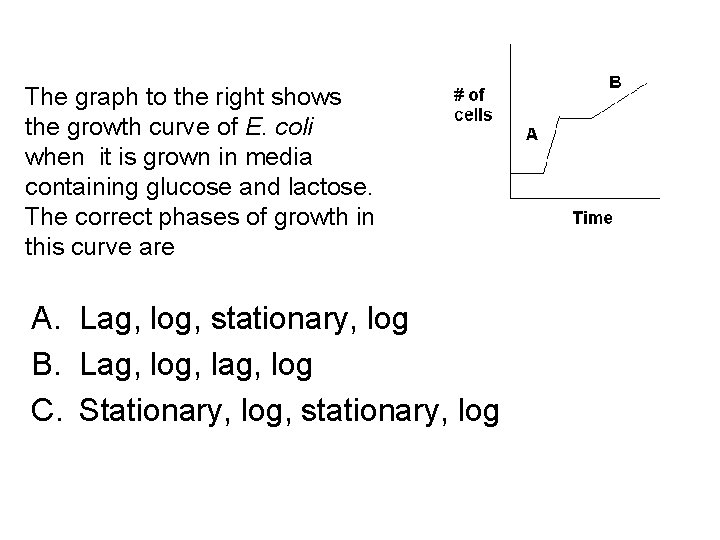
The graph to the right shows the growth curve of E. coli when it is grown in media containing glucose and lactose. The correct phases of growth in this curve are A. Lag, log, stationary, log B. Lag, log, lag, log C. Stationary, log, stationary, log

Which of the following is correct for the time marked “A” on the graph? A. Lactose is bound to the repressor, c. AMP-CAP is not bound to the CAP binding site, B-galactosidase is not being made B. Lactose is bound to the repressor, c. AMP-CAP is bound to the CAP binding site, B-galactosidase is being made C. Lactose is not bound to the repressor, c. AMP-CAP is not bound to the CAP binding site, B-galactosidase is not being made D. Lactose is not bound to the repressor, c. AMP-CAP is bound to the CAP binding site, B-galactosidase is being made

Which of the following is correct for the time marked “B” on the graph? A. Lactose is bound to the repressor, c. AMP-CAP is not bound to the CAP binding site, B-galactosidase is not being made B. Lactose is bound to the repressor, c. AMP-CAP is bound to the CAP binding site, B-galactosidase is being made C. Lactose is not bound to the repressor, c. AMP-CAP is not bound to the CAP binding site, B-galactosidase is not being made D. Lactose is not bound to the repressor, c. AMP-CAP is bound to the CAP binding site, B-galactosidase is being made

If an E. coli cell had a mutation in the Lac repressor protein such that it could not bind lactose, then A. The mutant cells would make Bgalactosidase when grown in lactose only, but not when grown in glucose only B. The mutant cells would make Bgalactosidase when grown in glucose only, but not when grown in lactose only C. The mutant cells would always make Bgalactosidase D. The mutant cells would never make Bgalactosidase

Lac vs. Trp Lac operon Trp operon • ______operon (breaks down food) • The environmental signal (lactose) acts as_____ • All by itself, the repressor is _______ • Example of an inducible operon • ______ operon (makes a building block) • The environmental signal (tryptophan) acts as _____ • All by itself, the repressor is ______ • Example of a repressible operon

Now that we know how to use the DNA • Changing the DNA – Mutation – Transduction – Conjugation – Transformation

Change in DNA can change protein Are they all visible? What is a Mutation? Are they all bad?

Mutation • • A change in the genetic material Mutations may be neutral, beneficial, or harmful Mutagen: Agent that causes mutations Spontaneous mutations: Occur without mutagen

Mutation may be beneficial (especially for microbes) • Environment changes microbes adapt • Natural selection favors those with greater fitness • How to adapt • Change when genes turned ON and OFF (mutate regulatory DNA or proteins) • Change the available genes (mutate or acquire change or add proteins) • Change in organism’s DNA alters genotype • Sequence of nucleotides in DNA • Bacteria are haploid, only one copy • May change observable characteristics, or phenotype • Also influenced by environmental conditions

Change in phenotype • Genetic change can affect organism’s phenotype – Deletion of gene for tryptophan biosynthesis yields mutant that only grows if tryptophan supplied • Growth factor required – Changing target of antibiotic or adding gene for antibiotic resistance (and others) • can survive antimicrobial treatment

Changing the DNA - in prokaryotes this includes “sharing” • Changing the DNA – Mutation – Transformation – Conjugation – Transduction

Getting us to THIS question • E. coli is found naturally in human colon, and there it is beneficial. However, the strain designated E. coli O 157: H 7 produces Shiga toxin. How did E. coli acquire this gene from Shigella?
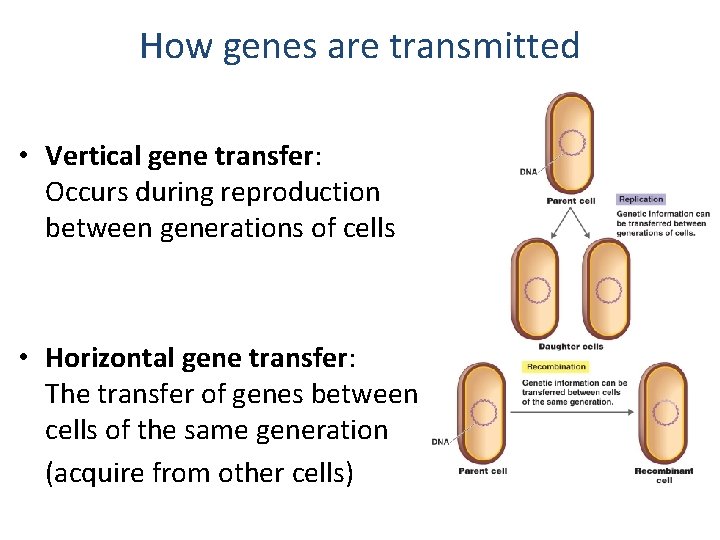
How genes are transmitted • Vertical gene transfer: Occurs during reproduction between generations of cells • Horizontal gene transfer: The transfer of genes between cells of the same generation (acquire from other cells)

Horizontal gene transfer occurs naturally by 3 mechanisms • Transformation: naked DNA uptake by bacteria • Conjugation: DNA transfer between bacterial cells • Transduction: bacterial DNA transfer by viruses
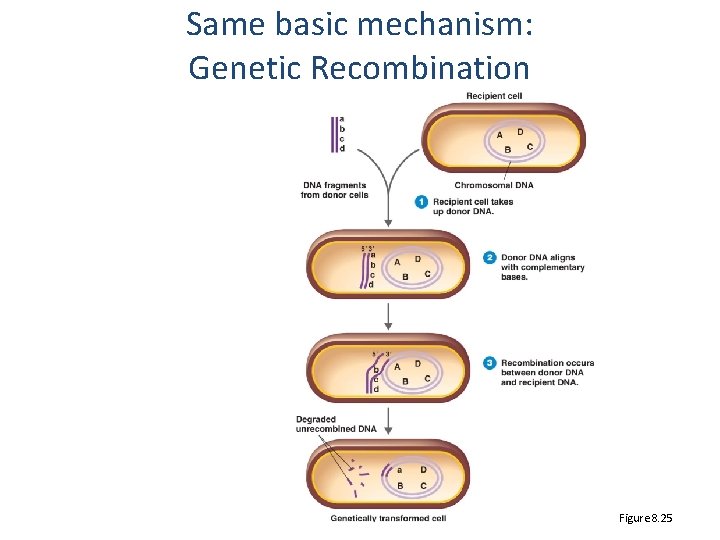
Same basic mechanism: Genetic Recombination Figure 8. 25
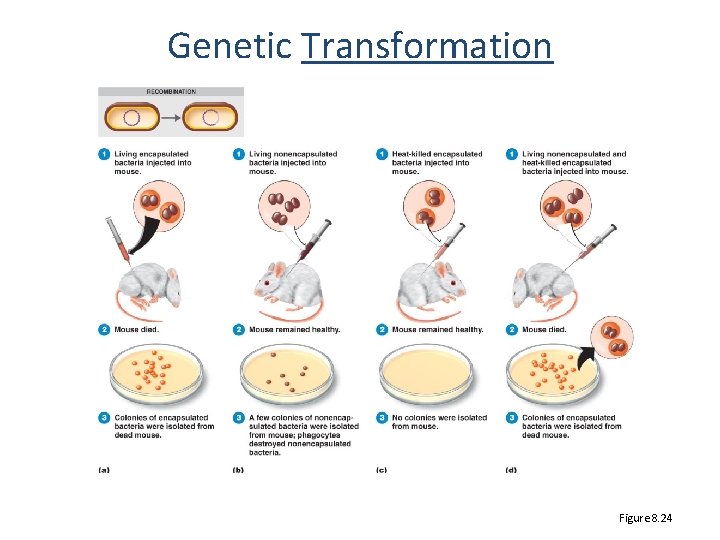
Genetic Transformation Figure 8. 24
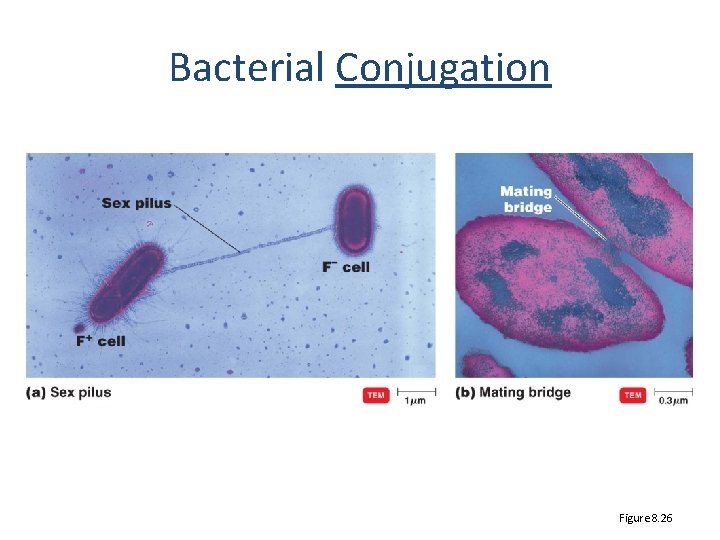
Bacterial Conjugation Figure 8. 26
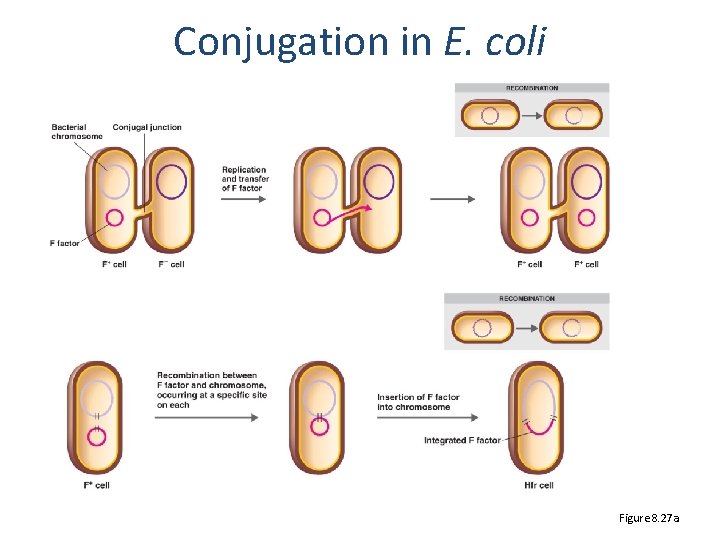
Conjugation in E. coli Figure 8. 27 a
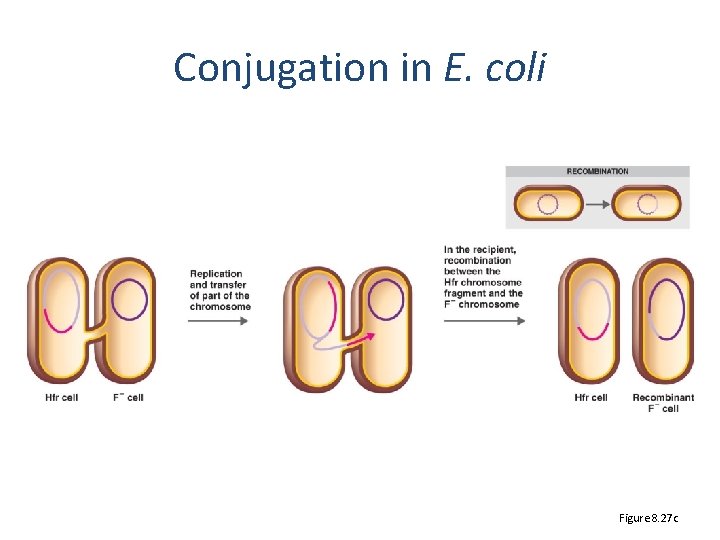
Conjugation in E. coli Figure 8. 27 c

Plasmids • Conjugative plasmid: – Carries genes for sex pili and transfer of the plasmid • Dissimilation plasmids: – Encode enzymes for catabolism of unusual compounds • R factors: – Encode antibiotic resistance
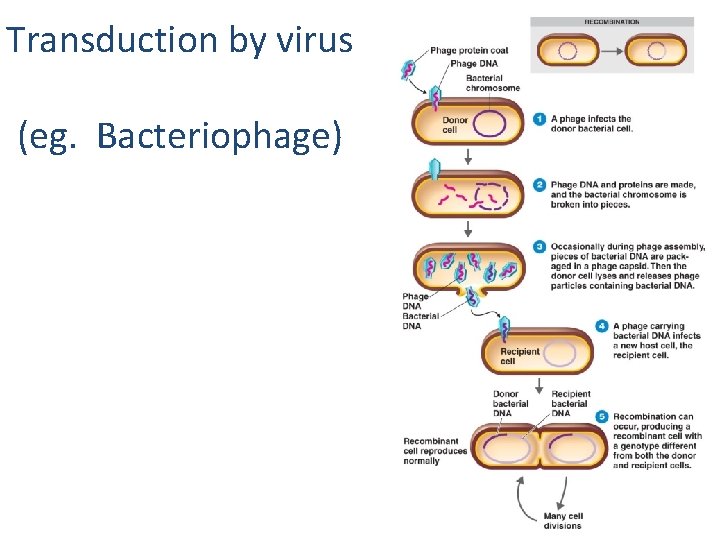
Transduction by virus (eg. Bacteriophage) Figure 8. 28
Clinical application • E. coli is found naturally in human colon, and there it is beneficial. However, the strain designated E. coli O 157: H 7 produces Shiga toxin. How did E. coli acquire this gene from Shigella? Naked DNA uptake (transformation)? Conjugation? Transduction?

Clinical application • Shiga toxin is one of the most potent toxins known to man; CDC lists it as a potential bioterrorist agent It seems likely that DNA from Shiga toxin-producing Shigella bacteria was transferred by a bacteriophage to otherwise harmless E. coli bacteria, thereby providing them with the genetic material to produce Shiga toxin act to inhibit protein synthesis within target cells by a mechanism similar to that of ricin. After entering a cell, the protein cleaves a specific adenine from the 30 S RNA of the 60 S subunit of the ribosome, thereby halting protein synthesis.

If a cell of one organism gets a piece of DNA from the cell of a different organism, what could happen? A. The cell that receives the DNA might make a new protein. B. The cell that receives the DNA might behave differently. C. The phenotype of the cell that receives the DNA might change. D. All of the above are possible. E. Nothing. The DNA will be too different.
- Slides: 104